Abstract
Since 2009, the European Association for Predictive, Preventive and Personalised Medicine (EPMA, Brussels) promotes the paradigm change from reactive approach to predictive, preventive, and personalized medicine (PPPM/3PM) to protect individuals in sub-optimal health conditions from the health-to-disease transition, to increase life-quality of the affected patient cohorts improving, therefore, ethical standards and cost-efficacy of healthcare to great benefits of the society at large. The gene-editing technology utilizing CRISPR/Cas gene-editing approach has demonstrated its enormous value as a powerful tool in a broad spectrum of bio/medical research areas. Further, CRISPR/Cas gene-editing system is considered applicable to primary and secondary healthcare, in order to prevent disease spread and to treat clinically manifested disorders, involving diagnostics of SARS-Cov-2 infection and experimental treatment of COVID-19. Although the principle of the proposed gene editing is simple and elegant, there are a lot of technological challenges and ethical considerations to be solved prior to its broadly scaled clinical implementation. This article highlights technological innovation beyond the state of the art, exemplifies current achievements, discusses unsolved technological and ethical problems, and provides clinically relevant outlook in the framework of 3PM.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Preamble
Reactive medicine as the currently implemented approach to treat clinically manifested disorders, has reached its limits from both ethical and economical points of view [1]. This can be illustrated by epidemics of non-communicable disorders demonstrating over a half of billion patients diagnosed with the diabetes mellitus and co-morbidities [2], breast cancer as the leading malignancy in female subpopulations [3], prostate cancer with the disease management costs annually increasing more rapidly than for any other cancer type [4] and alarming statistics of neurodegenerative disorders worldwide [5]. Since 2009, European Association for Predictive, Preventive and Personalised Medicine (EPMA, Brussels, www.epmanet.eu) promotes the paradigm change from reactive approach to predictive, preventive, and personalized medicine (PPPM/3PM) to reverse currently observed unfavourable trends, to protect individuals in sub-optimal health conditions from the health-to-disease transition, to increase life-quality of the affected patient cohorts improving, therefore, ethical standards and cost-efficacy of healthcare to great benefits of the society at large [6,7,8].
CRISPR/Cas gene-editing approach is applicable to primary and secondary healthcare, in order to prevent spread of diseases and to treat clinically manifested disorders. Although the principle of the proposed gene editing is simple and elegant, there are a lot of technological challenges and ethical considerations to be solved prior to its broadly scaled clinical implementation.
This article highlights technological innovation beyond the state of the art, exemplifies current achievements, discusses unsolved technological and ethical problems, and provides clinically relevant outlook in the framework of 3PM.
Technological innovation by the CRISPR/Cas gene editing
CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats)/Cas9 (CRISPR associated protein 9) system originated from bacteria and archaea, providing an adaptive immune response against viruses and plasmids. This defense system uses small RNA sequences of pathogen origin for sequence-specific detection and silencing of foreign, invading nucleic acids [9]. Although the Cas9 molecule is a part of prokaryotic defense mechanisms [10], it is fully functional in a broad spectrum of both prokaryotic and eukaryotic organisms.
A list of suitable target organisms and derived cell lines, that can be modified using the CRISPR/Cas9 system, covers the whole spectrum of life forms. Naturally, the most common are laboratory mammals such as mice [11,12,13], rats [14, 15], and rabbits [16], but also pigs [17, 18], dogs [19], and particularly human cell lines [20, 21]. Human gene editing is regulated by strict ethical norms and currently undergoes scientific, philosophical, and political discussion [22, 23]; however, in principle, it is executable. Besides the mammals, also genome of reptiles [24], amphibians (Xenopus laevis and X. tropicalis) [25, 26], fish (Danio rerio, Oryzias latipes) [26,27,28], insects (Drosophila melanogaster, mosquitoes) [29, 30], worms (Caenorhabditis elegans) [31, 32], plants [33,34,35], fungi [36, 37], bacteria [38], and viruses [39] can be edited by CRISPR/Cas9. Paradoxically, using the CRISPR/Cas9 technology is not as common in bacteria as in other organisms, likely because other methods based on homologous recombination were already available for efficient manipulation of their genomes [40].
The Cas molecules used for the gene editing originate from different organisms; thus, it is not surprising that these molecules differ in sequences as well as other properties and requirements (e.g., size of a gene, efficacy, target molecule type, or PAM sequence); however, more or less identical mechanism of action is shared by all [41]. Most frequently and first utilized is spCas9, originally isolated from Staphylococcus pyogenes [42,43,44]; later, other isolated Cas molecules from the other microorganisms started to be used as well. A list of examples contains Staphylococcus aureus [45, 46], Francisella novicida [47, 48], Neisseria meningitides [49], Campylobacter jejuni [50, 51], Streptococcus thermophilus [52, 53], Acidaminococcus spp. and Lachnospiraceae bacterium [54].
As mentioned above, the CRISPR/Cas system is commonly used in research studies to better understand biological processes. The development of precise treatment strategies should be preceded by the monitoring of genome editing experiments by NGS technology [55, 56] to eliminate unintended gene modifications[57]. This review focuses on CRISPR/Cas systems that are potentially utilized in personalized medicine.
The brief history of CRISPR/Cas9 discovery
The discovery of the CRIPSR/Cas9 mechanism started in 1987, when an unusual repetitive DNA sequence with an unknown function, which was subsequently defined as a CRISPR, was described in the Escherichia coli genome [58]. Later, similar sequence patterns were reported in a range of other bacteria and archaea, suggesting an important role for such evolutionarily conserved clusters of repeated sequences. At the beginning of this century, several conserved genes regularly present adjacent to the CRISPR region were also discovered. These genes were named CRISPR-associated sequences (Cas). To date, six CRISPR–Cas types have been described, which feature highly diverse cas gene content and operon organization. However, the function of these sequences remained unclear at that time [39, 40]. In 2005, several groups described observation that CRISPR sequences share homology with phages and plasmids; moreover, these phages and plasmids do not infect host strains harboring the homologous spacer sequences in the CRISPR. These observations concluded that these sequences are involved in the framework of a biological defense system—an adaptive immunity evolved by bacteria to destroy viral pathogens by cutting specifically their DNA with Cas proteases [40, 59,60,61]. Finally, in 2012 first publication described genome-editing technology based on CRISPR/Cas9 [9]. Recently, in 2020, the first patient legally received gene-editing therapy with CRISPR/Cas9 directly administered into the body. This treatment aims to remove mutations that cause a rare genetic condition called Leber’s congenital amaurosis, causing blindness [62]. Such an approach can be used as a paradigm for the personalized treatment of other diseases with known genetic origin.
General mechanism of CRISPR/Cas9 function
The CRISPR/Cas9 complex is an RNA-guided DNA endonuclease that recognizes target sites by RNA–DNA complementarity and produces sequence-specific double-stranded DNA break [63].
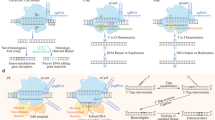
The CRISPR/Cas9 system consists of three key components: the CRISPR-associated DNA cleaving endonuclease Cas9 protein, a target DNA sequence-recognizing RNA transcribed from short DNA sequences known as protospacers (crRNA), and a trans-activating RNA (tracrRNA) [64]. The Cas9 nuclease contains two domains with endonuclease activity: RuvC and HNH, which together produces blunt double-stranded breaks. The DNA-binding crRNA recognized a 20-nucleotide-long DNA target sequence. The trans-activating crRNA (tracrRNA) anchors the crRNA to the Cas9 protein. Recognition and cleavage of a target site by Cas9 are conditioned and strictly depend on the presence of a so-called protospacer adjacent motif (PAM) sequence NGG immediately downstream of the crRNA target sequence. Cleavage occurs 3 bp upstream of the PAM [65]. The presence of a PAM sequence is the critical limitation of CRISPR/Cas9 gene editing. Currently, there are known various PAM sequences for different Cas molecules of different origins. Also, artificial Cas9 variants with expanded PAM compatibility were prepared [66]. Nowadays, the most common implementation of CRISPR genome editing in eukaryotic cells relies on a 2-component system consisting of Cas9 nuclease and a single chimeric guide RNA (sgRNA) that consist of crRNA and tracrRNA and provides both the recognition and structural function [65]. For the schematic visualization of the general mechanism of CRISPR/Cas9, see Fig. 1.
CRISPR/Cas9 and other Cas molecule-based technologies are instrumental for 3P medicine
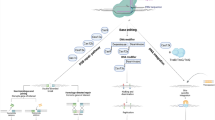
The CRISPR/cas9 system can improve human health by correcting genomes and searching for tumor manifestation, and capturing potential pathways of metastasis in organs. Similarly, in hereditary diseases, nucleases could be used to treat various diseases, including hematological and neurological disorders, autosomal dominant and X-linked disorders, and many others [67]. The basic principles of CRISPR/Cas technology, potentially useful in clinical practice, we discuss below. Generally, three main areas for CRISPR/Cas9 application in PPP are screening for inherited disease predispositions, diagnostic of molecular-based diseases, and therapy of such conditions.
CRISPR/Cas9 gene knock-out of defective genes
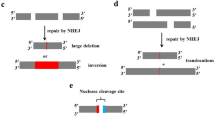
The CRISPR/Cas9 gene knock-out is a molecular genetics technique in which a target gene is inactivated. In principle, CRISPR/Cas9 gene knock-out is a combination of 2 different events. First, the CRISPR/Cas9 complex cleaves the target DNA sequence with a resulting blunt double-strand break that is subsequently repaired by the error-prone non-homologous end joining pathway (NHEJ), resulting in gene inactivation by the creation of frameshift alleles by random insertions or deletions (indels) [68, 69]. The creation of gene knockouts is one of the first and most widely used applications of the CRISPR–Cas9 system. Potentially therapeutic knockout of some genetic variants of genes involved in autoimmune disorders, such as multiple sclerosis, type 1 diabetes mellitus, psoriasis, rheumatoid arthritis, inflammatory bowel disease, systemic lupus erythematosus, and type 1 coeliac disease [70], shall be considered. For the schematic visualization of CRISPR/Cas9 knock-out, see Fig. 2.
CRISPR/Cas9 gene knock-in for restoration of physiologic gene expression
Unlike the gene knock-out, CRISPR/Cas9 gene knock-in is an insertion of DNA sequence into the genome without disrupting the ORF. It starts (just like knock-out) with CRISPR/Cas9 targeted cleavage of target DNA sequence with subsequent targeted integration of a sequence via the homology-directed repair pathway, which uses a provided DNA template to repair the cleavage [71]. A donor DNA containing a sequence surrounding the cleavage site with an insert in the frame has to be provided. For the schematic visualization of CRISPR/Cas9 knock-in, see Fig. 2.
The scale of modifications used in basic research includes the introduction of a single nucleotide–point mutation, small tags (Flag-tag, His-tag), up to larger protein sequences, very often fluorescent proteins [72]. However, this approach was used in experiments in cell lines in vitro, as well as in humanized centrosomal protein 290 (CEP290) mice in vivo carrying the mutation in the CEP290 gene, responsible for Leber congenital amaurosis type 10. The results showed the normal expression of the CEP290 gene and demonstrated the ability of CRISPR/Cas9 to edit somatic primate cells in vivo at levels that met the target therapeutic threshold. The authors suggest the application of the method for other inherited retinal disorders [73].
Nickases—a safer approach for gene editing
Off-targeting by CRISPR/Cas9 endonucleases is a significant concern. To address this problem, a paired nickase technology has been developed. To improve the specificity of CRISPR/Cas9 gene editing and to decrease the number of off-target mutagenesis events, CRISPR/Cas9 system was modified in the manner in which two single-strand nicks in proximity are created instead of a double-strand break. For this, a modified Cas9 protein called Cas9 nickase is used. Nickase is a catalytic mutant of Cas9 protein with one inactivated endonuclease domain; thus, it cleaves only one strand of the target dsDNA. When two different gRNAs targeting two neighboring sequences, however, on the opposite strands are used, then two separate single-strand breaks occur. This results in the formation of 5′ and 3′ overhangs at the cleaved target site. Induced break with overhang is either repaired by non-homologous end joining or homologous recombination-based repair mechanisms [74]. Using two close target sites on opposite strands decreases the possibility of random cleavage caused by off-target activity [75]. For the schematic visualization of nickases mechanism of function, see Fig. 3.
The catalytically dead dCas9 fused with the FokI domain works in an identical manner. The formation of DSB occurs only upon dimerization of FokI; thus, two unique gRNAs, binding to opposite strands approximately 15–25 bp apart, are necessary to bring two dCas9-FokI fusion proteins together to generate an active DNA-cleaving complex. This technology reduces the unwanted off-target activity of CRISPR, likewise the nickase approach [76, 77].
Also, nickases have been practically used in the development of sensitive and specific isothermal amplification of target genomic DNA adaptable in routine assays [78] and can potentially replace a traditional polymerase chain reaction (PCR) in non-laboratory on-site diagnostic applications [79].
CRISPR/Cas9-mediated up- and downregulation of gene expression and epigenetic modifications
A direct downregulation of gene expression can be mediated by catalytically deactivated Cas9 (dCas9) with disturbed endonuclease activity. When co-expressed with a guide RNA, it generates a DNA recognition complex, targeting specific sequences, which can interfere with transcriptional elongation, RNA polymerase binding, or transcription factor binding. This system, called CRISPR interference (CRISPRi), can be used to repress multiple target genes simultaneously, and its effects are reversible [80].
The site-specific binding of dCas9 can be utilized also for specific emplacement of various activator or repressor domains fused with dCas9, which alters the gene expression, mostly via modification of methylation [81] and/or acetylation [82]. These effector domains can write or erase histone modifications or modulate DNA methylation. Epigenome editing can be used to transiently or stably activate or repress specific genes. It was demonstrated that a catalytic domain of the human demethylase TET1cd targeted to specific methylated regions can cause highly efficient demethylation, thus up-regulation of a particular gene and a heritable change of the phenotype. In combination with dCas9, it provides tools for the targeted removal of 5mC at specific loci in the genome with high specificity and minimal off-target effects [83].
On the other hand, the use of the dCas9 fused with DNA methyltransferase catalytic domain led to a reduced expression of the gene of interest and to change in the phenotype [84]. Indeed, DNA methylation plays a critical role in regulating gene expression. Dysregulation of DNA methylation is involved in the pathogenesis of numerous diseases. Thus, the potential of technologies designed to manipulate DNA methylation at specific genomic loci is very high [85].
In parallel, if dCas9 is linked to the histone-modifying domain, it edits the chromatin histone, resulting in either activation or deactivation of the gene of choice. For example, p300 histone acetyltransferase increases H3K27 acetylation, which leads to gene activation; SID4X decreases H3K27 acetylation, thus decreasing gene expression, KRAB domain increases methylation at H3K9me3 and subsequently silences the gene expression, and paradoxically, LSD1 decreases methylation at H3K4me and also repress the gene expression [82, 86].
Epigenome editing tools have a place in precise medicine because they can be utilized in the treatment strategies of diseases, such as cancer, when some technological challenges of the method will be overcome [87]. Precise identification of epigenetically edited areas by NGS and elucidation of the interaction of edited genomes in epigenetic recombination should be taken into account [88]. For the schematic visualization of CRISPR/Cas9 utilization for up- and downregulation of gene expression and epigenetic modifications, see Fig. 4.
Targeted base editing can be useful in personalized medicine of monogenic disease
A targeted base editing could represent a promising contribution to personalized medicine, especially since many diseases are caused by small nucleotide substitutions or deletions. The CRISPR/Cas9 base-editing system directly edits single nucleotides, which avoids the dependence on homology-dependent repair [89]. Either dCas9 or Cas9 nickase (nCas9) can be fused to deaminases to induce substitutions or base edits without inducing double-strand breaks; thus, this approach minimizes the indels. Deaminases combined with Cas9 nickase and uracil DNA glycosylase inhibitors can make precise edits, changing cytidine to thymidine with high efficiency [90].
Until now, two kinds of base editors have been developed. The cytidine editor mediates the conversion of C/T (or G/A), whereas the adenosine editor affects an A/G (or T/C) substitution. The base editors are typically composed of a defective Cas9 protein (Cas9D10A or Cas9D10AH840A) and a deaminase fused to the Cas9 protein. Guided by the Cas9/sgRNA complex, the deaminase can be directed to any genomic locus to perform base editing in the single-stranded DNA (ssDNA) generated upon Cas9/sgRNA binding. By catalyzing the conversion of CAA, CAG, CGA, or TGG to TAA, TAG, or TGA codons, the cytidine base editor is capable of inactivating a target gene by generating a premature stop codon [91]. The base editors are applicable in the treatment of monogenic diseases with known pathogenic mutations and are intensively studied in experiments in vitro and in vivo. Actually, in vivo experiments showed that a single administration of base editors can have a longtime effect on many organs or tissues and improve the patient’s life that is complicated by symptomatic treatment. The rapid development of base editing tools is directly fused with the improvement in bioinformatic approaches that predict base editing outcomes. Once these shortcomings are eliminated, they can be used in personalized medicine [92]. For the schematic visualization of CRISPR/Cas9 targeted base editing, see Fig. 5.
Imaging tools based on CRISPR/Cas9
Besides the utilization of standard CRISPR/Cas9 knock-in strategy to fuse DNA sequence coding for fluorescent proteins with the gene of interest, enabling the imaging of produced protein [93, 94], CRISPR/Cas9 technology can be utilized for direct labeling of DNA and/or RNA. The dCas9/gRNA complex tagged with one or several fluorescent molecules can label specific DNA loci in living cells [95] (see Fig. 6). This approach has been proven to be effective in monitoring the localization of any genomic sequence. In this direction, another advantageous method for enriching the signal in live cell imaging is the use of the dCas9-SunTag system [96]. In this case, dCas9 is fused to an epitope tail—a protein platform loaded with multiple epitopes, each of which is recognized by a single chain variable fragment (scFv) of antibody. Simultaneously, fluorescent molecules are fused to a single chain variable fragment (scFv). While the possible number of scFv epitopes is much higher than any possible number of fluorescent proteins that can be directly fused to dCas9, this approach outperforms previous ones when the intensity of the signal is taken into consideration [97]. As an example of an imaging method that is not based on the utilization of various fluorescent proteins, a Cas9-mediated fluorescence in situ hybridization (CASFISH) should be mentioned. In this case, Halo Tag conjugated to fluorescent dye was used as a shorter replacement for fluorescent proteins [98]. Another recently developed strategy is labeling viral DNA with streptavidin-conjugated Quantum Dots (QD) connected to dCas9-Bio/gRNA complex via streptavidin/biotin binding [99]. Imaging of target sequences based on CRISPR/Cas9 technology can be used in live cells to study native chromatin organization. The method is rapid, cost-effective, comfortable, and applicable to multiple targets, which gives the prerequisite for the inclusion of the method into the diagnostics of human diseases [98].
RNA targeting by CRISPR/Cas systems
Although CRISPR/Cas9 system in natural conditions recognizes and cleaves exclusively DNA molecules, it is also possible to target RNA molecules. Repurposing dCas9 of S. pyogenes to target RNA requires, besides the usual components: dCas9 and gRNA, also PAMmers—short PAM-presenting DNA oligonucleotides. Once used, it allows the Cas9 to bind with high affinity to the single-stranded RNA and subsequently perform its enzymatic activity [63]. Other options to target RNA are the usage of Cas9 molecules of Francisella novicida (FnCas9) and/or Cas13, which are known to recognize the RNA naturally [48, 100,101,102]. In parallel with Cas9-mediated technologies focused on DNA, there are similar alternatives for RNA, for example, imaging [103, 104], base editing of full-length transcripts to repair pathogenic mutations on mRNA level [101], potential experimental therapeutic applications against RNA viruses [48], the introduction of antiviral resistance in plants [105] or oncology research—via targeting of lncRNAs [106]. A recent pandemic of the SARS-CoV-2 RNA virus allowed the development of CRISPR/Cas12 and/Cas13 molecular diagnostic kits focused on viral RNA, running in isothermal conditions and with no requirement of special equipment or training people. Colorimetric and visual detection of the pathogen in samples allows massive use of this test type anywhere and is also adjustable for other viral agents [107] (see Fig. 7).
Usage of CRISPR/Cas9 for the disease predisposition screening
Screening the disease as a part of preventive medicine represents a strategy to search for the risk markers or conditions that could turn into a disease in the future. Screening testing should be applied to individuals by cost-effective point-of-care diagnostics that advance the CRISPR/Cas system to candidate methods. The high-throughput genetic perturbation technologies are a useful tool for uncovering the involvement of particular genes in a broad spectrum of biological processes. Standard CRISPR/Cas9 knock-out combined with utilization of whole-genome or more specific gRNA libraries and subsequent identification of genes of interest represents a very effective approach for such screening [108]. Development of the CRISPR/Cas9-based loss-of-function screening assays enabled efficient identification of essential genes in mammalian cells involved in cancer therapy resistance [109], genes involved in the immune response to cancer [110], genes enabling the viral infection [111], genes associated with sepsis [112], biosynthesis pathways [113] and very probably many more.
In addition to screening based on a knock-out approach, catalytically inactive dCas9 fused with various transcriptional activator, repressor, and recruitment domains have been used to modulate gene expression without introducing irreversible mutations to the genome. In this case, gRNA libraries are focused on the upstream, regulating regions of investigated genes [114].
Purification of specific target DNA with CRISPR/Cas9-based technology
The capability of dCas9 to bind selectively a specific DNA sequence is also employed in engineered DNA-binding molecule-mediated chromatin immunoprecipitation enChIP technology. The dCas9 molecule fused to an epitope tag is being used to isolate a given genomic locus. In addition, this technology allows for retaining the intracellular molecular interactions; thus, in vitro enChIP technology is of potential use not only for sequence-specific isolation of DNA but as well as for the identification of molecules interacting with genomic regions of interest [115, 116].
The delivery strategy of the CRISPR/Cas9 system into target cells affects the outcome
Efficient and successful delivery of the CRISPR/Cas system into the target cells in vivo remains a challenge. Current delivery methods include viral and non-viral vectors and physical delivery. An effective delivery system must overcome the barriers of tissues and cell membranes; then, it should be transported to the nucleus, where it operates. The ineffective carriers may cause problems such as immune response, gene mutations, and triggering carcinogenesis. Another problem is associated with the size of packing molecules. However, nano-delivery systems seem to be efficient in experiments in vivo and in vitro [117].
As we mentioned, the selection of a suitable delivery method for Cas9/sgRNA expressing cassettes into target cells is an important step. For mammalian cells, plasmid transfection is the most common way to deliver the Cas9/sgRNA-expressing cassette. However, transfection can cause overexpression of both Cas9 protein and sgRNA, and then induce off-target mutagenesis. In addition, many cell types are hard-to-transfect cells (such as hematopoietic cells, immune cells, stem cells, etc.). A microinjection is an option but with a low efficiency of only one cell per injection. Several groups used adenovirus to deliver Cas9/sgRNA expressing cassettes, but the construction and amplification of recombinant adenovirus were laborious and time-consuming. A single lentiviral vector was designed to simultaneously deliver Cas9 and sgRNA-expressing cassettes into the target cells. This system will enable genome editing in most cell types of interest [68]. Very promising is the so-called "nanoblade" technology [118], which avoids off-target mutation induced by the application of a standard CRIPSR/Cas9 system. This has been especially a significant concern when using this technique to generate a stable knock-out cell line [68]. To overcome this problem, engineered Murine leukemia virus-like particles loaded with Cas9-sgRNA ribonucleoproteins (Nanoblades) can be used for transduction and subsequent editing without integrating a virus genome, thus reducing editing to the one-time event [118]. The goal in the nanoparticle delivery will be a tissue-specific surface of nanostructure covered with specific ligands that will be suitable for clinical disease treatment [117]. Figure 8 represents nanoblade technology for CRISPR/Cas9 delivery.
CRISPR/Cas-based diagnostics
The ability of gRNA-guided Cas molecules to recognize a specific sequence has its logical application in the diagnostics of various diseases. Indeed, several approaches to detect both specific DNA and RNA in samples have been developed recently. For this purpose, Cas12a, Cas13a, and Cas13b are employed most often. The action of these members of the Cas family differs from that of the Cas9. Unlike Cas9, upon the activation of these CRISPR/Cas complexes with a specific target, an indiscriminate cleavage of substrate nucleic acids occurs. In the diagnostic assays, substrate sequences are coupled to a fluorescent reporter. The fluorescence signal is activated only when the substrate is cleaved. The presence of a limited amount of target sequence can elicit intense fluorescence; thus, the methods based on these principles are very sensitive [119]. As an example, the DETECTR (DNA endonuclease-targeted CRISPR trans reporter) method can be mentioned. Isothermal Recombinase Polymerase Amplification (RPA) is coupled with subsequent Cas12a target DNA recognition, unleashing robust, indiscriminate cleavage of a quenched-fluorescent single-stranded DNA substrate reporter. This rapid, one-pot detection provides a straightforward platform for molecular diagnostics [120]. Another technique, named SHERLOCK (Specific High-sensitivity Enzymatic Reporter un-Locking), utilizes a combination of the isothermal RPA or reverse transcription (RT)-RPA with nuclease activity of Cas13a, followed by unspecific collateral cleavage of labeled reporter RNA resulting in signal release [121, 122]. In All-In-One Dual CRISPR/Cas12a (AIOD-CRISPR) assay, a pair of gRNAs was introduced to initiate dual CRISPR/Cas12a detection, which resulted in improved detection sensitivity. The nucleic acids (DNA and RNA) were successfully detected with a sensitivity of few copies, which is comparable with the real-time RT-PCR method [123]. Also, Cas9-based assays can be utilized for diagnostic purposes. In this case, for the identification of different strains. The combination of the isothermal amplification technique NASBA (Nucleic acid sequence-based amplification) and subsequent application of specific gRNA/Cas9 complex is used to differentiate strains in single-base discrimination. The result is either truncated or full-length DNA fragments, and only non-truncated fragments can trigger the toehold switch, leading to a color reaction [124].
After the emergence of the COVID-19 pandemic, various SARS-CoV-2 diagnostic techniques based on Cas12a and Cas13a have been developed by numerous research groups very rapidly, demonstrating the flexibility, adaptability, and versatility of this approach and its potential for future challenges [125,126,127,128,129].
In general, the methods mentioned above and many similar methods have a great potential for developing next-generation point-of-care molecular diagnostics necessary for the development of personalized medicine. For examples of CRISPR/Cas-based detection methods, see Fig. 9.
Fundamentals of the future CRISPR/Cas9-based therapy
The ability of CRISPR/Cas9-based methodology to modulate the expression of selected genes or precisely re-write the specific DNA sequence gives this technique tremendous potential in targeted therapy of a wide spectrum of both infectious and non-infectious diseases. It can find its place in the management of any condition where the removal or cleavage of specific genetic material might be beneficial. However, barely a few, if any, of the therapeutic procedures using CRISPR/Cas9 reached the state of the experimental application in human medicine. Thus, most of the discussed data shall be seen as a perspective and potential approach for future treatments.
Significant progress in this field has been made by taking advantage of cell line and animal models. Therapeutic applications for various infectious, predominantly viral diseases, monogenic disorders, or anti-cancer gene therapies have been investigated [130]. While the original role of the CRISPR/Cas system in bacteria was an antiviral defense, the attempts to utilize it as an antiviral agent are not surprising. The possibility of direct targeting of viral genomes predisposes CRISPR/Cas9 system to be a promising treatment against a wide spectrum of viruses. It has been shown in cultured cells that it can restrict the viral life cycle by introducing sequence-specific breaks in essential parts of the viral genome [131,132,133]. Based on the specific virus biology, different life stages can be targeted. For example, covalently closed circular DNA (cccDNA) of HBV is the preferred target while it is not reduced by standard treatment with nuclear analogs [134]; on the other hand, CRISPR/Cas9 treatment of HIV aims at integrated provirus removal or inactivation [135]. Similarly, in the case of HSV-1, replication of the virus can be effectively impaired by using nucleoside analogs; however, the latent virus remains unaffected. Targeting key viral genes by CRISPR/Cas9 leads to indels resulting in reduced infectivity or even completely abrogated infection. Resistant strains of escaped mutants were fully prevented by simultaneously targeting two essential viral genes by co-delivery of two gRNAs [136, 137]. When multiplexed CRISPR/Cas9 was applied to target the latent EBV genomes in a Burkitt’s lymphoma patient-derived B cell line, an eradication or significant reduction of viral load in the majority of infected cells resulted in a dramatic cell proliferation arrest and induction of apoptosis [138, 139]. In the case of HPV, already established HPV-driven tumors have to be targeted. The CRISPR/Cas9 system was employed to inactivate the viral oncogenes E6 and/or E7, thereby restoring p53/Rb levels and induction of apoptosis of HPV16-infected cells [140]. Direct targeting of RNA viruses (e.g., HCV) is also possible [48]. It is not surprising, that after the outbreak of COVID-19, several research groups attempted to adjust the CRISPR/Cas technology as a potential anti-SARS-CoV-2 therapeutics [141,142,143].
Another group of diseases that can be treated with the utilization of CRISPR/Cas are monogenic disorders caused by single gene defects. Therapies for this group of diseases are generally limited to managing symptoms regardless of the pathogenic mutation causing the disease in the patient. CRISPR/Cas system represents a promising personalized gene-editing approach in the treatment of monogenic disorders. The results of preclinical studies carried out on human stem cells, patient-derived induced pluripotent stem cells (iPSCs), or animal models showed successful correction of various types of mutations in defective genes causing monogenic diseases such as cystic fibrosis [144, 145], Duchenne muscular dystrophy [146, 147], hemophilia [148, 149], Huntington’s disease [150,151,152], and sickle cell anemia [153,154,155].
CRISPR/Cas system represents a powerful perspective tool with a wide spectrum of use in PPP medicine. The greatest possibilities of using a CRISPR-based therapeutic approach are in personalized cancer treatment. The potential of CRISPR-based screening combined with high-throughput next-generation sequencing brings great possibilities for discovering new potential therapeutic targets, biomarkers for early diagnosis of disease, monitoring of disease progress, therapy efficiency, and prognostic biomarkers [156]. Drug and immunotherapy resistance is a big problem in long-term treatment, predominantly in oncology. CRISPR/Cas system can also be utilized to identify genes and mutations responsible for drug resistance and thereby help with individualized, effective therapy [157].
CRISPR/Cas mechanism can be utilized in personalized therapy not only for gene editing and regulation but also for editing of epigenetic modification, which is one of the gene expression activation or silencing mechanisms. In tumors, oncogenes and tumor suppressor genes are often regulated via epigenetic modifications, which are also possible therapeutic targets for personalized medicine. For epigenetic editing, the most promising technology was developed based on the fusion of deactivated Cas9 protein (dCas9) without endonuclease activity serving as DNA-binding protein with an epigenetic effector (epi-effector) [158] as described above. There are several studies bringing promising results of epigenetic modulation of tumor suppressor genes or oncogenes through dCas9/effector complexes in the most common cancers, for example, activation of PTEN in breast cancer [159], BRAF1 gene in cervical cancer [160] and other genes in colon cancer [161], liver cancer [162], prostate cancer [163] or lung cancer [164] and others.
CRISPR/Cas-based technology also offers opportunities for advances in immunotherapy based on artificial CAR- (chimeric antigen receptor) T cells recognizing specific tumor-associated antigens. This method brings innovative, precise engineering for reprogramming patients’ own T cells to target tumor cells. Next-generation CAR T cells engineered with CRISPR/Cas9 system help overcome CAR-T cell therapy’s limitations such as stability, low persistence, T cell depletion, or toxic tumor microenvironment and open the possibilities for personalized immunotherapy not only for hematopoietic malignancies but also for solid tumors [165]. CRISPR/Cas9 is an effective tool for the knock-out of genes responsible for producing negative regulators of T cells like PD-1 or CTLA-4, Fas signaling, and toxic cytokines, which are factors reducing the effectiveness of CAR T cell therapy. These targets were successfully disrupted with a multiplex CRISPR/Cas9 gene-editing approach to increase the success of therapy [166,167,168]. CRISPR technology improves also the targeting of CAR encoding cassette to the specific location under the control of the appropriate promotor to ensure stable expression and prevents unspecific insertion observed in the case of viral vectors [165, 169, 170].
Outlook in the framework of 3P medicine
Predictive, preventive, and personalized approach in primary care—ethical aspects
Primary care considers pre-pregnancy check-up as being crucial for the cost-effective diagnostics and preventive strategies. To this end, an advanced screening of suboptimal health status to predict and prevent individual health risks prior to planned pregnancies meets needs of young populations and of the society at large and falls into the framework of 3P medicine. Contextually, pre-pregnancy check-up has been demonstrated as being pivotal for advanced health policy [7].
CRISPR/Cas is a feasible genome-editing technology to prevent transmission of parental life-threatening chrDNA mutations to offspring. To this end, human germline gene editing (HGGE) is the tool to introduce permanent genetic modification to the embryo by both—eliminating and introducing DNA information. Due to severe technological challenges and ethical considerations, HGGE is the subject to a detailed analysis and careful consideration by researchers, relevant scientific groups, and governmental organizations. There is a consensus amongst the majority of them that, due to a number of unanswered scientific, ethical, and policy questions, currently, it is inappropriate to perform HGGE culminating in human pregnancy. The major disagreement is about the types of research which should be allowed. For example, both—the European Group on Ethics in Science and New Technologies, EGE, and NIH in the USA (which does not fund any use of gene-editing technologies in human embryos) suggested a partial or full moratorium on HGGE research, whereas the US National Academy of Sciences and National Academy of Medicine supports basic and preclinical HGGE research [171]. Due to a limited access to the human germline material and severe ethical concerns, stem cells are suggested as the reasonable alternative model for the HGGE approaches to promote the field [172].
Technological challenges of gene editing in the mitochondrial disease
The “gatekeeper” role of mitochondria is demonstrated for the wide range of molecular, cellular, organ, and organismal functions. Compromised mitochondrial health is linked to systemic effects in multi-organ functionality. Consequently, mitochondrial health sustainability is pivotal for primary, secondary, and tertiary care [173, 174].
Mitochondrial disease (MD) is a devastating inborn pathology originating from defects either directly in mtDNA or/and in chrDNA encoding for proteins localized to mitochondria. The most severe forms of MD are most frequently linked to OXPHOS function impairments, which are fatal causing death of affected newborn infants shortly after their birth. Although many mothers are at risk of transmitting MD to their offspring, corresponding screening programs are still underdeveloped and there is no cure of MD. To this end, mitochondrial replacement therapy, and CRISPR/Cas are considered promising technological solutions to prevent MD transmission from the affected mother to offspring [175].
Being proven as a highly effective system for the nuclear genome editing, CRISPR/Cas feasibility for mitochondrial genome editing is, however, controversial [176]. Specifically, the delivery of this gene-editing system into mitochondria remains debatable. RNA import process into mitochondria is insufficiently understood and the mitochondrial DNA repair machinery differs substantially from this of the chrDNA. To this end, the basis of the CRISPR/Cas-associated editing system, namely DNA double-strand break repair, is lacking in mitochondria, in contrast to the homologous recombination-mediated repair mechanisms that are detectable there.
Consequently, CRISPR/Cas editing system application in regard to the MD is focused mainly on studying mitochondrial biology and genome-related disorders being currently restricted to distinctive tasks:
-
Ex vivo MD modeling
-
Creation of isogenic populations of diseased cells with homogenous phenotype, e.g., with clearly defined mutations causing co-enzyme Q10 deficiency to distinguish individual phenotypes in MD
-
A genome-wide library screening to investigate the relevance of individual genes to the co-enzyme Q10 biosynthesis machinery, ATP-production versus deficiency as well as to the electron-chain transfer efficacy.
These models utilizing CRISPR/Cas editing system are considered highly efficient to investigate potential therapeutic treatments tailored to the individualized profile of the MD-affected patients discriminating between moderate and severe medical conditions [176].
Data availability
Not applicable.
Code availability
Not applicable.
Abbreviations
- AIOD-CRISPR:
-
All-In-One dual CRISPR/Cas12a
- CASFISH:
-
Cas9-mediated fluorescence in situ hybridization
- CEP290:
-
Centrosomal protein 290
- CRISPR/Cas:
-
Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-associated sequences
- CRISPRi:
-
CRISPR interference
- crRNA:
-
CRISPR RNA
- dCas9:
-
Dead or deactivated Cas9
- DETECTR:
-
DNA endonuclease- targeted CRISPR trans reporter
- enChIP:
-
Engineered DNA-binding molecule-mediated chromatin immunoprecipitation
- EPMA:
-
European Association for Predictive, Preventive and Personalised Medicine
- FnCas9:
-
Cas9 molecules of Francisella novicida
- FokI:
-
Restriction endonuclease found in Flavobacterium okeanokoites
- gRNA:
-
Guide RNA
- HDR:
-
Homology-directed repair
- HGGE:
-
Human germline gene editing
- KRAB:
-
Krüppel associated box
- lncRNAs:
-
Long non-coding RNAs
- NASBA:
-
Nucleic acid sequence-based amplification
- NHEJ:
-
Non-homologous end joining pathway
- PAM:
-
Protospacer adjacent motif
- PPPM or 3PM:
-
Predictive preventive personalized medicine
- p300:
-
E1A-binding protein p300
- RPA:
-
Recombinase polymerase amplification
- scFv:
-
Single-chain variable fragment
- sgRNA:
-
Single chimeric guide RNA
- SHERLOCK:
-
Specific High-sensitivity Enzymatic Reporter un-Locking
- tracrRNA:
-
Trans-activating CRISPR RNA
References
Golubnitschaja O, Baban B, Boniolo G, Wang W, Bubnov R, Kapalla M, et al. Medicine in the early twenty-first century: paradigm and anticipation - EPMA position paper 2016. EPMA J. 2016;7:23.
Kropp M, Golubnitschaja O, Mazurakova A, Koklesova L, Sargheini N, Vo T-TKS, et al. Diabetic retinopathy as the leading cause of blindness and early predictor of cascading complications-risks and mitigation. EPMA J. 2023;14:21–42.
Mazurakova A, Koklesova L, Samec M, Kudela E, Kajo K, Skuciova V, et al. Anti-breast cancer effects of phytochemicals: primary, secondary, and tertiary care. EPMA J. 2022;13:315–34.
Ellinger J, Alajati A, Kubatka P, Giordano FA, Ritter M, Costigliola V, et al. Prostate cancer treatment costs increase more rapidly than for any other cancer-how to reverse the trend? EPMA J. 2022;13:1–7.
Golubnitschaja O, Potuznik P, Polivka J, Pesta M, Kaverina O, Pieper CC, et al. Ischemic stroke of unclear aetiology: a case-by-case analysis and call for a multi-professional predictive, preventive and personalised approach. EPMA J. 2022;13:535–45.
Wang W, Yan Y, Guo Z, Hou H, Garcia M, Tan X, et al. All around suboptimal health - a joint position paper of the Suboptimal Health Study Consortium and European Association for Predictive, Preventive and Personalised Medicine. EPMA J. 2021;12:403–33.
Evsevieva M, Sergeeva O, Mazurakova A, Koklesova L, Prokhorenko-Kolomoytseva I, Shchetinin E, et al. Pre-pregnancy check-up of maternal vascular status and associated phenotype is crucial for the health of mother and offspring. EPMA J. 2022;13:351–66.
Jain N, Nagaich U, Pandey M, Chellappan DK, Dua K. Predictive genomic tools in disease stratification and targeted prevention: a recent update in personalized therapy advancements. EPMA J. 2022;13:561–80.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 2012;337:816–21.
Barrangou R. The roles of CRISPR-Cas systems in adaptive immunity and beyond. Curr Opin Immunol. 2015;32:36–41.
Richards DY, Winn SR, Dudley S, Nygaard S, Mighell TL, Grompe M, et al. AAV-mediated CRISPR/Cas9 gene editing in murine phenylketonuria. Mol Ther Methods Clin Dev. 2020;17:234–45.
Garrett AM, Bosch PJ, Steffen DM, Fuller LC, Marcucci CG, Koch AA, et al. CRISPR/Cas9 interrogation of the mouse Pcdhg gene cluster reveals a crucial isoform-specific role for Pcdhgc4. PLoS Genet. 2019;15:e1008554.
Cho K, Ro SW, Seo SH, Jeon Y, Moon H, Kim DY, et al. Genetically engineered mouse models for liver cancer. Cancers (Basel). 2019;12:14.
Sun H, Fu S, Cui S, Yin X, Sun X, Qi X, et al. Development of a CRISPR-SaCas9 system for projection- and function-specific gene editing in the rat brain. Sci Adv. 2020;6:eaay6687.
Yu L, Wang L, Yi H, Wu X. LRP6-CRISPR prevents activation of hepatic stellate cells and liver fibrogenesis in rats. Am J Transl Res. 2020;12:397–408.
Xu Y, Liu H, Pan H, Wang X, Zhang Y, Yao B, et al. CRISPR/Cas9-mediated disruption of fibroblast growth factor 5 in rabbits results in a systemic long hair phenotype by prolonging anagen. Genes (Basel). 2020;11:297.
Tanihara F, Hirata M, Thi Nguyen N, Le Anh Q, Hirano T, Otoi T. Generation of viable PDX1 gene-edited founder pigs as providers of nonmosaics. Mol Reprod Dev. 2020;87:471–81.
Yang H, Wu Z. Genome editing of pigs for agriculture and biomedicine. Front Genet. 2018;9:360.
Mata López S, Balog-Alvarez C, Vitha S, Bettis AK, Canessa EH, Kornegay JN, et al. Challenges associated with homologous directed repair using CRISPR-Cas9 and TALEN to edit the DMD genetic mutation in canine Duchenne muscular dystrophy. PLoS One. 2020;15:e0228072.
Ginn SL, Amaya AK, Liao SHY, Zhu E, Cunningham SC, Lee M, et al. Efficient in vivo editing of OTC-deficient patient-derived primary human hepatocytes. JHEP Rep. 2020;2:100065.
Bressan RB, Pollard SM. Genome editing in human neural stem and progenitor cells. Results Probl Cell Differ. 2018;66:163–82.
Archard D, Dabrock P, Delfraissy J-F. Human-genome editing: ethics councils call to governments worldwide. Nature. 2020;579:29.
Eissenberg JC. In our image: the ethics of CRISPR genome editing. Biomol Concepts. 2021;12:1–7.
Rasys AM, Park S, Ball RE, Alcala AJ, Lauderdale JD, Menke DB. CRISPR-Cas9 gene editing in lizards through microinjection of unfertilized oocytes. Cell Rep. 2019;28:2288-2292.e3.
Dimitrakopoulou D, Tulkens D, Van Vlierberghe P, Vleminckx K. Xenopus tropicalis: joining the armada in the fight against blood cancer. Front Physiol. 2019;10:48.
Naert T, Vleminckx K. CRISPR/Cas9 disease models in zebrafish and Xenopus: the genetic renaissance of fish and frogs. Drug Discov Today Technol. 2018;28:41–52.
Shi J, Bi P, Pei J, Li H, Grishin NV, Bassel-Duby R, et al. Requirement of the fusogenic micropeptide myomixer for muscle formation in zebrafish. Proc Natl Acad Sci U S A. 2017;114:11950–5.
Beppu K, Tsutsumi R, Ansai S, Ochiai N, Terakawa M, Mori M, et al. Development of a screening system for agents that modulate taste receptor expression with the CRISPR-Cas9 system in medaka. Biochem Biophys Res Commun. 2022;601:65–72.
Hafezi Y, Sruba SR, Tarrash SR, Wolfner MF, Clark AG. Dissecting fertility functions of Drosophila Y chromosome genes with CRISPR. Genetics. 2020;214:977–90.
Anderson MAE, Purcell J, Verkuijl SAN, Norman VC, Leftwich PT, Harvey-Samuel T, et al. Expanding the CRISPR toolbox in culicine mosquitoes: in vitro validation of Pol III promoters. ACS Synth Biol. 2020;9:678–81.
Hastie E, Sellers R, Valan B, Sherwood DR. A Scalable CURE Using a CRISPR/Cas9 fluorescent protein knock-in strategy in Caenorhabditis elegans. J Microbiol Biol Educ. 2019;20:20.3.60.
Serrat X, Kukhtar D, Cornes E, Esteve-Codina A, Benlloch H, Cecere G, et al. CRISPR editing of sftb-1/SF3B1 in Caenorhabditis elegans allows the identification of synthetic interactions with cancer-related mutations and the chemical inhibition of splicing. PLoS Genet. 2019;15:e1008464.
LeBlanc C, Zhang F, Mendez J, Lozano Y, Chatpar K, Irish VF, et al. Increased efficiency of targeted mutagenesis by CRISPR/Cas9 in plants using heat stress. Plant J. 2018;93:377–86.
Moradpour M, Abdulah SNA. CRISPR/dCas9 platforms in plants: strategies and applications beyond genome editing. Plant Biotechnol J. 2020;18:32–44.
Bastet A, Zafirov D, Giovinazzo N, Guyon-Debast A, Nogué F, Robaglia C, et al. Mimicking natural polymorphism in eIF4E by CRISPR-Cas9 base editing is associated with resistance to potyviruses. Plant Biotechnol J. 2019;17:1736–50.
Stovicek V, Holkenbrink C, Borodina I. CRISPR/Cas system for yeast genome engineering: advances and applications. FEMS Yeast Res. 2017;17:fox030.
Schuster M, Kahmann R. CRISPR-Cas9 genome editing approaches in filamentous fungi and oomycetes. Fungal Genet Biol. 2019;130:43–53.
Altenbuchner J. Editing of the Bacillus subtilis Genome by the CRISPR-Cas9 System. Appl Environ Microbiol. 2016;82:5421–7.
Hupfeld M, Trasanidou D, Ramazzini L, Klumpp J, Loessner MJ, Kilcher S. A functional type II-A CRISPR–Cas system from Listeria enables efficient genome editing of large non-integrating bacteriophage. Nucleic Acids Res. 2018;46:6920–33.
Ishino Y, Krupovic M, Forterre P. History of CRISPR-Cas from encounter with a mysterious repeated sequence to genome editing technology. J Bacteriol. 2018;200:e00580-e617.
Xu CL, Park KS, Tsang SH. CRISPR/Cas9 genome surgery for retinal diseases. Drug Discov Today Technol. 2018;28:23–32.
Kulcsár PI, Tálas A, Tóth E, Nyeste A, Ligeti Z, Welker Z, et al. Blackjack mutations improve the on-target activities of increased fidelity variants of SpCas9 with 5’G-extended sgRNAs. Nat Commun. 2020;11:1223.
Yang Y-C, Chen Y-H, Kao J-H, Ching C, Liu I-J, Wang C-C, et al. Permanent inactivation of HBV genomes by CRISPR/Cas9-mediated non-cleavage base editing. Mol Ther Nucleic Acids. 2020;20:480–90.
Fujii W, Ito H, Kanke T, Ikeda A, Sugiura K, Naito K. Generation of genetically modified mice using SpCas9-NG engineered nuclease. Sci Rep. 2019;9:12878.
Lu B, Javidi-Parsijani P, Makani V, Mehraein-Ghomi F, Sarhan WM, Sun D, et al. Delivering SaCas9 mRNA by lentivirus-like bionanoparticles for transient expression and efficient genome editing. Nucleic Acids Res. 2019;47:e44.
Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature. 2015;520:186–91.
Acharya S, Mishra A, Paul D, Ansari AH, Azhar M, Kumar M, et al. Francisella novicida Cas9 interrogates genomic DNA with very high specificity and can be used for mammalian genome editing. Proc Natl Acad Sci U S A. 2019;116:20959–68.
Price AA, Sampson TR, Ratner HK, Grakoui A, Weiss DS. Cas9-mediated targeting of viral RNA in eukaryotic cells. Proc Natl Acad Sci U S A. 2015;112:6164–9.
Ibraheim R, Song C-Q, Mir A, Amrani N, Xue W, Sontheimer EJ. All-in-one adeno-associated virus delivery and genome editing by Neisseria meningitidis Cas9 in vivo. Genome Biol. 2018;19:137.
Koo T, Lu-Nguyen NB, Malerba A, Kim E, Kim D, Cappellari O, et al. Functional rescue of dystrophin deficiency in mice caused by frameshift mutations using Campylobacter jejuni Cas9. Mol Ther. 2018;26:1529–38.
Jo DH, Koo T, Cho CS, Kim JH, Kim J-S, Kim JH. Long-term effects of in vivo genome editing in the mouse retina using Campylobacter jejuni Cas9 expressed via adeno-associated virus. Mol Ther. 2019;27:130–6.
Gasiunas G, Barrangou R, Horvath P, Siksnys V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci U S A. 2012;109:E2579-2586.
Sapranauskas R, Gasiunas G, Fremaux C, Barrangou R, Horvath P, Siksnys V. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 2011;39:9275–82.
Verwaal R, Buiting-Wiessenhaan N, Dalhuijsen S, Roubos JA. CRISPR/Cpf1 enables fast and simple genome editing of Saccharomyces cerevisiae. Yeast. 2018;35:201–11.
Boel A, Steyaert W, De Rocker N, Menten B, Callewaert B, De Paepe A, et al. BATCH-GE: batch analysis of next-generation sequencing data for genome editing assessment. Sci Rep. 2016;6:30330 (Nature Publishing Group).
Martinez-Lage M, Puig-Serra P, Menendez P, Torres-Ruiz R, Rodriguez-Perales S. CRISPR/Cas9 for cancer therapy: hopes and challenges. Biomed Multidiscip Digit Publ Inst. 2018;6:105.
Park SH, Cao M, Pan Y, Davis TH, Saxena L, Deshmukh H, et al. Comprehensive analysis and accurate quantification of unintended large gene modifications induced by CRISPR-Cas9 gene editing. Sci Adv Am Assoc Adv Sci. 2022;8:eabo7676.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A. Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol. 1987;169:5429–33.
Pourcel C, Salvignol G, Vergnaud G. CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiol (Reading). 2005;151:653–63.
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiol (Reading). 2005;151:2551–61.
Hussain W, Mahmood T, Hussain J, Ali N, Shah T, Qayyum S, et al. CRISPR/Cas system: a game changing genome editing technology, to treat human genetic diseases. Gene. 2019;685:70–5.
Ledford H. CRISPR treatment inserted directly into the body for first time. Nature. 2020;579:185.
O’Connell MR, Oakes BL, Sternberg SH, East-Seletsky A, Kaplan M, Doudna JA. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature. 2014;516:263–6.
Sürün D, von Melchner H, Schnütgen F. CRISPR/Cas9 genome engineering in hematopoietic cells. Drug Discov Today Technol. 2018;28:33–9.
Hoban MD, Bauer DE. A genome editing primer for the hematologist. Blood. 2016;127:2525–35.
Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N, et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature. 2018;556:57–63.
Mani I. Chapter Eight - CRISPR-Cas9 for treating hereditary diseases. In: Singh V, editor. Progress in molecular biology and translational science [Internet]. Academic Press; 2021 [cited 2023 Mar 20]. p. 165–83. Available from: https://www.sciencedirect.com/science/article/pii/S1877117321000284
Li Y, Zhang W, Zhao J, Li S, Shan L, Zhu J, et al. Establishing a dual knock-out cell line by lentivirus based combined CRISPR/Cas9 and Loxp/Cre system. Cytotechnology. 2018;70:1595–605.
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J. RNA-programmed genome editing in human cells. Elife. 2013;2:e00471.
Lee MH, Shin JI, Yang JW, Lee KH, Cha DH, Hong JB, et al. Genome editing using CRISPR-Cas9 and autoimmune diseases: a comprehensive review. Int J Mol Sci. 2022;23:1337.
Wilson LOW, O’Brien AR, Bauer DC. The current state and future of CRISPR-Cas9 gRNA design tools. Front Pharmacol. 2018;9:749.
Pickar-Oliver A, Gersbach CA. The next generation of CRISPR-Cas technologies and applications. Nat Rev Mol Cell Biol. 2019;20:490–507.
Maeder ML, Stefanidakis M, Wilson CJ, Baral R, Barrera LA, Bounoutas GS, et al. Development of a gene-editing approach to restore vision loss in Leber congenital amaurosis type 10. Nat Med. 2019;25:229–33.
Ulaganathan K, Goud BS, Reddy MM, Kumar VP, Balsingh J, Radhakrishna S. Proteins for breaking barriers in lignocellulosic bioethanol production. Curr Protein Pept Sci. 2015;16:100–34.
Jo Y-I, Suresh B, Kim H, Ramakrishna S. CRISPR/Cas9 system as an innovative genetic engineering tool: Enhancements in sequence specificity and delivery methods. Biochim Biophys Acta. 2015;1856:234–43.
Tsai SQ, Wyvekens N, Khayter C, Foden JA, Thapar V, Reyon D, et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat Biotechnol. 2014;32:569–76.
Guilinger JP, Thompson DB, Liu DR. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat Biotechnol. 2014;32:577–82.
Wang T, Liu Y, Sun H-H, Yin B-C, Ye B-C. An RNA-guided Cas9 nickase-based method for universal isothermal DNA amplification. Angew Chem Int Ed. 2019;58:5382–6.
Zhou W, Hu L, Ying L, Zhao Z, Chu PK, Yu X-F. A CRISPR-Cas9-triggered strand displacement amplification method for ultrasensitive DNA detection. Nat Commun. 2018;9:5012.
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell. 2013;152:1173–83.
Devesa-Guerra I, Morales-Ruiz T, Pérez-Roldán J, Parrilla-Doblas JT, Dorado-León M, García-Ortiz MV, et al. DNA methylation editing by CRISPR-guided excision of 5-methylcytosine. J Mol Biol. 2020;432:2204–16.
Okada M, Kanamori M, Someya K, Nakatsukasa H, Yoshimura A. Stabilization of Foxp3 expression by CRISPR-dCas9-based epigenome editing in mouse primary T cells. Epigenetics Chromatin. 2017;10:24.
Gallego-Bartolomé J, Gardiner J, Liu W, Papikian A, Ghoshal B, Kuo HY, et al. Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain. Proc Natl Acad Sci U S A. 2018;115:E2125–34.
Lu Z, Liu Z, Mao W, Wang X, Zheng X, Chen S, et al. Locus-specific DNA methylation of Mecp2 promoter leads to autism-like phenotypes in mice. Cell Death Dis. 2020;11:85.
Urbano A, Smith J, Weeks RJ, Chatterjee A. Gene-specific targeting of DNA methylation in the mammalian genome. Cancers (Basel). 2019;11:1515.
Lau C-H, Ho JW-T, Lo PK, Tin C. Targeted transgene activation in the brain tissue by systemic delivery of engineered AAV1 Expressing CRISPRa. Mol Ther Nucleic Acids. 2019;16:637–49.
Maroufi F, Maali A, Abdollahpour-Alitappeh M, Ahmadi MH, Azad M. CRISPR-mediated modification of DNA methylation pattern in the new era of cancer therapy. Epigenomics Future Med. 2020;12:1845–59.
Wang J, Li D, Yang J, Chang L, Zhang R, Li J. CRISPR/Cas9-mediated epigenetic editing tool: An optimized strategy for targeting de novo DNA methylation with stable status via homology directed repair pathway. Biochimie. 2022;202:190–205.
Zheng K, Wang Y, Li N, Jiang F-F, Wu C-X, Liu F, et al. Highly efficient base editing in bacteria using a Cas9-cytidine deaminase fusion. Commun Biol. 2018;1:32.
Eid A, Alshareef S, Mahfouz MM. CRISPR base editors: genome editing without double-stranded breaks. Biochem J. 2018;475:1955–64.
Chen W, Zhang Y, Zhang Y, Pi Y, Gu T, Song L, et al. CRISPR/Cas9-based genome editing in Pseudomonas aeruginosa and cytidine deaminase-mediated base editing in Pseudomonas species. iScience. 2018;6:222–31.
Reshetnikov VV, Chirinskaite AV, Sopova JV, Ivanov RA, Leonova EI. Translational potential of base-editing tools for gene therapy of monogenic diseases. Front Bioeng Biotechnol. 2022;10:942440.
Deffieu MS, Cesonyte I, Delalande F, Boncompain G, Dorobantu C, Song E, et al. Rab7-harboring vesicles are carriers of the transferrin receptor through the biosynthetic secretory pathway. Sci Adv. 2021;7:eaba7803.
Matsuda T, Oinuma I. Imaging endogenous synaptic proteins in primary neurons at single-cell resolution using CRISPR/Cas9. Mol Biol Cell. 2019;30:2838–55.
Chen B, Gilbert LA, Cimini BA, Schnitzbauer J, Zhang W, Li G-W, et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell. 2013;155:1479–91.
Tanenbaum ME, Gilbert LA, Qi LS, Weissman JS, Vale RD. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell. 2014;159:635–46.
Baliou S, Adamaki M, Kyriakopoulos AM, Spandidos DA, Panayiotidis M, Christodoulou I, et al. Role of the CRISPR system in controlling gene transcription and monitoring cell fate. Mol Med Rep. 2018;17:1421–7.
Deng W, Shi X, Tjian R, Lionnet T, Singer RH. CASFISH: CRISPR/Cas9-mediated in situ labeling of genomic loci in fixed cells. Proceedings of the National Academy of Sciences. Proc Natl Acad Sci. 2015;112:11870–5.
Yang Y-B, Tang Y-D, Hu Y, Yu F, Xiong J-Y, Sun M-X, et al. Single virus tracking with quantum dots packaged into enveloped viruses using CRISPR. Nano Lett. 2020;20:1417–27.
Sampson TR, Saroj SD, Llewellyn AC, Tzeng Y-L, Weiss DS. A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature. 2013;497:254–7.
Cox DBT, Gootenberg JS, Abudayyeh OO, Franklin B, Kellner MJ, Joung J, et al. RNA editing with CRISPR-Cas13. Science. 2017;358:1019–27.
Abudayyeh OO, Gootenberg JS, Essletzbichler P, Han S, Joung J, Belanto JJ, et al. RNA targeting with CRISPR-Cas13. Nature. 2017;550:280–4.
Yang L-Z, Wang Y, Li S-Q, Yao R-W, Luan P-F, Wu H, et al. Dynamic imaging of RNA in living cells by CRISPR-Cas13 systems. Mol Cell. 2019;76:981-997.e7.
Nelles DA, Fang MY, O’Connell MR, Xu JL, Markmiller SJ, Doudna JA, et al. Programmable RNA tracking in live cells with CRISPR/Cas9. Cell. 2016;165:488–96.
Zhang T, Zheng Q, Yi X, An H, Zhao Y, Ma S, et al. Establishing RNA virus resistance in plants by harnessing CRISPR immune system. Plant Biotechnol J. 2018;16:1415–23.
Wong NK, Huang C-L, Islam R, Yip SP. Long non-coding RNAs in hematological malignancies: translating basic techniques into diagnostic and therapeutic strategies. J Hematol Oncol. 2018;11:131.
Kostyusheva A, Brezgin S, Babin Y, Vasilyeva I, Glebe D, Kostyushev D, et al. CRISPR-Cas systems for diagnosing infectious diseases. Methods. 2022;203:431–46.
Joung J, Konermann S, Gootenberg JS, Abudayyeh OO, Platt RJ, Brigham MD, et al. Genome-scale CRISPR-Cas9 knockout and transcriptional activation screening. Nat Protoc. 2017;12:828–63.
Anderson GR, Winter PS, Lin KH, Nussbaum DP, Cakir M, Stein EM, et al. A Landscape of therapeutic cooperativity in KRAS mutant cancers reveals principles for controlling tumor evolution. Cell Rep. 2017;20:999–1015.
Crowther MD, Dolton G, Legut M, Caillaud ME, Lloyd A, Attaf M, et al. Genome-wide CRISPR-Cas9 screening reveals ubiquitous T cell cancer targeting via the monomorphic MHC class I-related protein MR1. Nat Immunol. 2020;21:178–85.
Ren Q, Li C, Yuan P, Cai C, Zhang L, Luo GG, et al. A dual-reporter system for real-time monitoring and high-throughput CRISPR/Cas9 library screening of the hepatitis C virus. Sci Rep. 2015;5:8865.
Cai M, Li S, Shuai Y, Li J, Tan J, Zeng Q. Genome-wide CRISPR-Cas9 viability screen reveals genes involved in TNF-α-induced apoptosis of human umbilical vein endothelial cells. J Cell Physiol. 2019;234:9184–93.
Zhu Y, Groth T, Kelkar A, Zhou Y, Neelamegham S. A GlycoGene CRISPR-Cas9 lentiviral library to study lectin binding and human glycan biosynthesis pathways. Glycobiology. 2021;31:173–80.
Shalem O, Sanjana NE, Zhang F. High-throughput functional genomics using CRISPR-Cas9. Nat Rev Genet. 2015;16:299–311.
Fujita T, Yuno M, Fujii H. Efficient sequence-specific isolation of DNA fragments and chromatin by in vitro enChIP technology using recombinant CRISPR ribonucleoproteins. Genes Cells. 2016;21:370–7.
Fujita T, Yuno M, Fujii H. An enChIP system for the analysis of bacterial genome functions. BMC Res Notes. 2018;11:387.
Duan L, Ouyang K, Xu X, Xu L, Wen C, Zhou X, et al. Nanoparticle delivery of CRISPR/Cas9 for genome editing. frontiers in genetics [Internet]. 2021 [cited 2023 Mar 21];12. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fgene.2021.673286
Mangeot PE, Risson V, Fusil F, Marnef A, Laurent E, Blin J, et al. Genome editing in primary cells and in vivo using viral-derived Nanoblades loaded with Cas9-sgRNA ribonucleoproteins. Nat Commun. 2019;10:45.
Xiang X, Qian K, Zhang Z, Lin F, Xie Y, Liu Y, et al. CRISPR-cas systems based molecular diagnostic tool for infectious diseases and emerging 2019 novel coronavirus (COVID-19) pneumonia. J Drug Target. 2020;28:727–31.
Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, Palefsky JM, et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science. 2018;360:436–9.
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science. 2017;356:438–42.
Khodavirdipour A, Piri M, Jabbari S, Khalaj-Kondori M. Potential of CRISPR/Cas13 system in treatment and diagnosis of COVID-19. Glob Med Genet. 2021;8:7–10.
Ding X, Yin K, Li Z, Lalla RV, Ballesteros E, Sfeir MM, et al. Ultrasensitive and visual detection of SARS-CoV-2 using all-in-one dual CRISPR-Cas12a assay. Nat Commun. 2020;11:4711.
Pardee K, Green AA, Takahashi MK, Braff D, Lambert G, Lee JW, et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell. 2016;165:1255–66.
Zhu Z, Guo Y, Wang C, Yang Z, Li R, Zeng Z, et al. An ultra-sensitive one-pot RNA-templated DNA ligation rolling circle amplification-assisted CRISPR/Cas12a detector assay for rapid detection of SARS-CoV-2. Biosens Bioelectron. 2023;228:115179.
Kashefi-Kheyrabadi L, Nguyen HV, Go A, Lee M-H. Ultrasensitive and amplification-free detection of SARS-CoV-2 RNA using an electrochemical biosensor powered by CRISPR/Cas13a. Bioelectrochemistry. 2023;150:108364.
Wei J, Song Z, Cui J, Gong Y, Tang Q, Zhang K, et al. Entropy-driven assisted T7 RNA polymerase amplification-activated CRISPR/Cas13a activity for SARS-CoV-2 detection in human pharyngeal swabs and environment by an electrochemiluminescence biosensor. J Hazard Mater. 2023;452:131268.
Morales-Moreno MD, Valdés-Galindo EG, Reza MM, Fiordelisio T, Peon J, Hernandez-Garcia A. Multiplex gRNAs synergically enhance detection of SARS-CoV-2 by CRISPR-Cas12a. CRISPR J. 2023;
Wang Y, Chen H, Gao H, Wei H, Wang Y, Mu K, et al. CESSAT: A chemical additive-enhanced single-step accurate CRISPR/Cas13 testing system for field-deployable ultrasensitive detection and genotyping of SARS-CoV-2 variants of concern. Biosens Bioelectron. 2023;229:115238.
Xiao-Jie L, Hui-Ying X, Zun-Ping K, Jin-Lian C, Li-Juan J. CRISPR-Cas9: a new and promising player in gene therapy. J Med Genet. 2015;52:289–96.
Chou Y-Y, Krupp A, Kaynor C, Gaudin R, Ma M, Cahir-McFarland E, et al. Inhibition of JCPyV infection mediated by targeted viral genome editing using CRISPR/Cas9. Sci Rep. 2016;6:36921.
Kayesh MEH, Amako Y, Hashem MA, Murakami S, Ogawa S, Yamamoto N, et al. Development of an in vivo delivery system for CRISPR/Cas9-mediated targeting of hepatitis B virus cccDNA. Virus Res. 2020;290:198191.
Song J, Zhang X, Ge Q, Yuan C, Chu L, Liang H-F, et al. CRISPR/Cas9-mediated knockout of HBsAg inhibits proliferation and tumorigenicity of HBV-positive hepatocellular carcinoma cells. J Cell Biochem. 2018;119:8419–31.
Seeger C, Sohn JA. Targeting hepatitis B virus with CRISPR/Cas9. Mol Ther Nucleic Acids. 2014;3:e216.
Xu Y, Peng X, Zheng Y, Jin C, Lu X, Han D, et al. Inactivation of latent HIV-1 proviral DNA using clustered regularly interspaced short palindromic repeats/Cas9 treatment and the assessment of off-target effects. Front Microbiol. 2021;12:629153.
Roehm PC, Shekarabi M, Wollebo HS, Bellizzi A, He L, Salkind J, et al. Inhibition of HSV-1 replication by gene editing strategy. Sci Rep. 2016;6:23146.
Zhang I, Hsiao Z, Liu F. Development of genome editing approaches against herpes simplex virus infections. Viruses. 2021;13:338.
van Diemen FR, Kruse EM, Hooykaas MJG, Bruggeling CE, Schürch AC, van Ham PM, et al. CRISPR/Cas9-mediated genome editing of herpesviruses limits productive and latent infections. PLoS Pathog. 2016;12:e1005701.
Wang J, Quake SR. RNA-guided endonuclease provides a therapeutic strategy to cure latent herpesviridae infection. Proc Natl Acad Sci U S A. 2014;111:13157–62.
Kennedy EM, Kornepati AVR, Goldstein M, Bogerd HP, Poling BC, Whisnant AW, et al. Inactivation of the human papillomavirus E6 or E7 gene in cervical carcinoma cells by using a bacterial CRISPR/Cas RNA-guided endonuclease. J Virol. 2014;88:11965–72.
Abbott TR, Dhamdhere G, Liu Y, Lin X, Goudy L, Zeng L, et al. Development of CRISPR as an antiviral strategy to combat SARS-CoV-2 and Influenza. Cell. 2020;181:865-876.e12.
Nguyen TM, Zhang Y, Pandolfi PP. Virus against virus: a potential treatment for 2019-nCov (SARS-CoV-2) and other RNA viruses. Cell Res. 2020;30:189–90.
Hawsawi YM, Shams A, Theyab A, Siddiqui J, Barnawee M, Abdali WA, et al. The state-of-the-art of gene editing and its application to viral infections and diseases including COVID-19. Front Cell Infect Microbiol. 2022;12:869889.
Schwank G, Koo B-K, Sasselli V, Dekkers JF, Heo I, Demircan T, et al. Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell. 2013;13:653–8.
Firth AL, Menon T, Parker GS, Qualls SJ, Lewis BM, Ke E, et al. Functional gene correction for cystic fibrosis in lung epithelial cells generated from patient iPSCs. Cell Rep. 2015;12:1385–90.
Ousterout DG, Kabadi AM, Thakore PI, Majoros WH, Reddy TE, Gersbach CA. Multiplex CRISPR/Cas9-based genome editing for correction of dystrophin mutations that cause Duchenne muscular dystrophy. Nat Commun. 2015;6:6244.
Li HL, Fujimoto N, Sasakawa N, Shirai S, Ohkame T, Sakuma T, et al. Precise correction of the dystrophin gene in duchenne muscular dystrophy patient induced pluripotent stem cells by TALEN and CRISPR-Cas9. Stem Cell Rep. 2015;4:143–54.
Park C-Y, Kim DH, Son JS, Sung JJ, Lee J, Bae S, et al. Functional correction of large factor VIII gene chromosomal inversions in hemophilia A patient-derived iPSCs using CRISPR-Cas9. Cell Stem Cell. 2015;17:213–20.
Guan Y, Ma Y, Li Q, Sun Z, Ma L, Wu L, et al. CRISPR/Cas9-mediated somatic correction of a novel coagulator factor IX gene mutation ameliorates hemophilia in mouse. EMBO Mol Med. 2016;8:477–88.
Shin JW, Kim K-H, Chao MJ, Atwal RS, Gillis T, MacDonald ME, et al. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum Mol Genet. 2016;25:4566–76.
Monteys AM, Ebanks SA, Keiser MS, Davidson BL. CRISPR/Cas9 editing of the mutant huntingtin allele in vitro and in vivo. Mol Ther. 2017;25:12–23.
Yang S, Chang R, Yang H, Zhao T, Hong Y, Kong HE, et al. CRISPR/Cas9-mediated gene editing ameliorates neurotoxicity in mouse model of Huntington’s disease. J Clin Invest. 2017;127:2719–24.
Hoban MD, Lumaquin D, Kuo CY, Romero Z, Long J, Ho M, et al. CRISPR/Cas9-mediated correction of the sickle mutation in human CD34+ cells. Mol Ther. 2016;24:1561–9.
Park SH, Lee CM, Deshmukh H, Bao G. Therapeutic Crispr/Cas9 genome editing for treating sickle cell disease. Blood. 2016;128:4703.
Dever DP, Bak RO, Reinisch A, Camarena J, Washington G, Nicolas CE, et al. CRISPR/Cas9 β-globin gene targeting in human haematopoietic stem cells. Nature. 2016;539:384–9.
Xing H, Meng L. CRISPR-cas9: a powerful tool towards precision medicine in cancer treatment. Acta Pharmacol Sin. 2020;41:583–7.
Liu Z, Shi M, Ren Y, Xu H, Weng S, Ning W, et al. Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy. Mol Cancer. 2023;22:35.
Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 2013;154:442–51.
Moses C, Nugent F, Waryah CB, Garcia-Bloj B, Harvey AR, Blancafort P. Activating PTEN tumor suppressor expression with the CRISPR/dCas9 system. Mol Ther - Nucleic Acids. 2019;14:287–300.
Choudhury SR, Cui Y, Lubecka K, Stefanska B, Irudayaraj J. CRISPR-dCas9 mediated TET1 targeting for selective DNA demethylation at BRCA1 promoter. Oncotarget. 2016;7:46545–56.
Wang Q, Dai L, Wang Y, Deng J, Lin Y, Wang Q, et al. Targeted demethylation of the SARI promotor impairs colon tumour growth. Cancer Lett. 2019;448:132–43.
Wang H, Guo R, Du Z, Bai L, Li L, Cui J, et al. Epigenetic targeting of granulin in hepatoma cells by synthetic CRISPR dCas9 Epi-suppressors. Mol Ther - Nucleic Acids. 2018;11:23–33.
Kardooni H, Gonzalez-Gualda E, Stylianakis E, Saffaran S, Waxman J, Kypta RM. CRISPR-mediated reactivation of DKK3 expression attenuates TGF-β signaling in prostate cancer. Cancers. Multidisciplinary Digital Publishing Institute; 2018;10:165.
Morita S, Noguchi H, Horii T, Nakabayashi K, Kimura M, Okamura K, et al. Targeted DNA demethylation in vivo using dCas9-peptide repeat and scFv-TET1 catalytic domain fusions. Nat Biotechnol. 2016;34:1060–5.
Dimitri A, Herbst F, Fraietta JA. Engineering the next-generation of CAR T-cells with CRISPR-Cas9 gene editing. Mol Cancer. 2022;21:78.
Liu X, Zhang Y, Cheng C, Cheng AW, Zhang X, Li N, et al. CRISPR-Cas9-mediated multiplex gene editing in CAR-T cells. Cell Res. 2017;27:154–7.
Ren J, Liu X, Fang C, Jiang S, June CH, Zhao Y. Multiplex genome editing to generate universal CAR T cells resistant to PD1 inhibition. Clin Cancer Res. 2017;23:2255–66.
Ren J, Zhang X, Liu X, Fang C, Jiang S, June CH, et al. A versatile system for rapid multiplex genome-edited CAR T cell generation. Oncotarget. 2017;8:17002–11.
Eyquem J, Mansilla-Soto J, Giavridis T, van der Stegen SJC, Hamieh M, Cunanan KM, et al. Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection. Nature. 2017;543:113–7.
Odé Z, Condori J, Peterson N, Zhou S, Krenciute G. CRISPR-mediated non-viral site-specific gene integration and expression in T cells: protocol and application for T-cell therapy. Cancers. Multidisciplinary Digital Publishing Institute 2020;12:1704.
Ormond KE, Mortlock DP, Scholes DT, Bombard Y, Brody LC, Faucett WA, et al. Human germline genome editing. Am J Hum Genet. 2017;101:167–76.
Bekaert B, Boel A, Cosemans G, De Witte L, Menten B, Heindryckx B. CRISPR/Cas gene editing in the human germline. Semin Cell Dev Biol. 2022;131:93–107.
Koklesova L, Mazurakova A, Samec M, Kudela E, Biringer K, Kubatka P, et al. Mitochondrial health quality control: measurements and interpretation in the framework of predictive, preventive, and personalized medicine. EPMA J. 2022;13:177–93.
Koklesova L, Samec M, Liskova A, Zhai K, Büsselberg D, Giordano FA, et al. Mitochondrial impairments in aetiopathology of multifactorial diseases: common origin but individual outcomes in context of 3P medicine. EPMA J. 2021;12:27–40.
Fogleman S, Santana C, Bishop C, Miller A, Capco DG. CRISPR/Cas9 and mitochondrial gene replacement therapy: promising techniques and ethical considerations. Am J Stem Cells. 2016;5:39–52.
Tang J-X, Pyle A, Taylor RW, Oláhová M. Interrogating mitochondrial biology and disease using CRISPR/Cas9 gene editing. Genes (Basel). 2021;12:1604.
Funding
Open Access funding enabled and organized by Projekt DEAL. This publication has been produced with the support of the Integrated Infrastructure Operational Program for the project New possibilities for laboratory diagnostics and massive screening of SARS-Cov-2 and identification of mechanisms of virus behavior in the human body, ITMS: 313011AUA4, co-financed by the European Regional Development Fund.
Author information
Authors and Affiliations
Contributions
V.L. created study concepts and drafted the manuscript. V.H. and Z.K. drafted the manuscript. E.H. acquired funding and critically revised the manuscript. M.S. created the figures and edited the manuscript. O.G. contributed with PPPM-related concepts, clinical relevance, and data interpretation. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lučanský, V., Holubeková, V., Kolková, Z. et al. Multi-faceted CRISPR/Cas technological innovation aspects in the framework of 3P medicine. EPMA Journal 14, 201–217 (2023). https://doi.org/10.1007/s13167-023-00324-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-023-00324-6
Keywords
- Predictive preventive personalized medicine (PPPM / 3PM)
- Primary & secondary care
- CRISPR/Cas gene-editing technology
- Defective genes
- Restoration
- Physiologic gene expression
- Diagnostics
- Therapy
- COVID-19
- Innovation
- Health-to-disease transition
- Human germline gene editing
- Ethics
- Mitochondrial disease
- Disease severity
- Health policy