Abstract
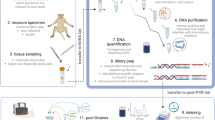
In this study, a non-invasive method for DNA sampling of terrestrial molluscs using mucus stored on FTA cards is described. DNA was isolated from FTA cards impregnated with terrestrial mollusc mucus, which were collected from four mollusc families. DNA was stored for approximately 2 months at room temperature. For two mollusc families, DNA was extracted from foot tissue as a cross-check against the DNA obtained from mucus. Amplification of mitochondrial and nuclear DNA was successful for all mucus samples and in all cases the sequences matched those obtained from tissue samples, but also sequences from the same species, genus or family (depending on their availability) from GenBank. These results show that isolating mucus on FTA cards is a very efficient alternative to sampling tissue for genetic studies of terrestrial molluscs, and particularly for conservation genetic studies. This non-invasive method is rapid, reliable and very easy to put in practice in field situations.


Similar content being viewed by others
References
Armbruster GFJ, Korte A (2006) Genomic nucleotide variation in the ITS1 rDNA spacer of land snails. J Molluscan Stud 72:211–213
Armbruster GFJ, Koller B, Baur B (2005) Foot mucus and periostracum fraction as non-destructive source of DNA in the land snail Arianta arbustorum, and the development of new microsatellite loci. Conserv Genet 6:313–316
Coote T, Clarke D, Hickman C, Murray J, Pearce-Kelly P (2004) Experimental release of endemic Partula species, extinct in the wild, into a protected area of natural habitat on Moorea. Pac Sci 58:429–434
Depraz A, Hausser J, Pfenninger M (2009) A species delimitation approach in the Trochulus sericeus/hispidus complex reveals two cryptic species within a sharp contact zone. BMC Evol Biol 9:1–10
Durnin ME, Palsboll PJ, Ryder OA, McCullough DR (2007) A reliable genetic technique for sex determination of giant panda (Ailuropoda melanoleuca) from non-invasively collected hair samples. Conserv Genet 8:715–720
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3:294–299
Geist J, Wunderlich H, Kuehn R (2008) Use of mollusc shells for DNA-based analyses. J Molluscan Stud 74:337–343
Hadfield MG, Holland BS, Olival KJ (2004) Contributions of ex situ propagation and molecular genetics to conservation of Hawaiian tree snails. In: Gordon MS, Bartol SM (eds) Experimental approaches to conservation biology. University of California Press, Berkeley, pp 16–34
Halbert ND, Raudsepp T, Chowdhary BP, Derr JN (2004) Conservation genetic analysis of the Texas State Bison Herd. J Mammal 85:924–931
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Holland BS, Hadfield MG (2002) Islands within an island: phylogeography and conservation genetics of the endangered Hawaiian tree snail Achatinella mustelina. Mol Ecol 11:365–375
Huelsenbeck JP, Ronquist F, Hall B (2001) MrBayes: bayesian inference of phylogeny. Bioinformatics 17:754–755
Jacob G, Debrunner R, Gugerli F, Schmid B, Bollmann K (2010) Field surveys of capercaillie (Tetrao urogallus) in the Swiss Alps underestimated local abundance of the species as revealed by genetic analyses of non-invasive samples. Conserv Genet 11:33–44
Johanson HC, Hyland V, Wicking C, Sturm RA (2009) DNA elution from buccal cells stored on Whatman FTA classic cards using a modified methanol fixation method. Biotechniques 46:309–311
Kawai K, Shimizu M, Hughes RN, Takenaka O (2004) A non-invasive technique for obtaining DNA from marine intertidal snails. J Mar Biol Assoc U K 84:773–774
Lee T, Burch JB, Coote T, Pearce-Kelly P, Hickman C, Meyer JY, Foighil DO (2009) Moorean tree snail survival revisited: a multi-island genealogical perspective. BMC Evol Biol 9:1–16
Luchtel DJ, Deyrup-Olson I (2001) Body wall and function. In: Gary GM (ed) The Biology of terrestrial molluscs. CABI Publishing, Oxon, pp 147–178
Oliverio M, Cervelli M, Mariottini P (2002) ITS2 rRNA evolution and its congruence with the phylogeny of muricid neogastropods (Caenogastropoda, Muricoidea). Mol Phylogenet Evol 25:63–69
Palmer ANS, Styan CA, Shearman DCA (2008) Foot mucus is a good source for non-destructive genetic sampling in Polyplacophora. Conserv Genet 9:229–231
Petersen SD, Mason T, Akber S, West R, White B, Wilson P (2007) Species identification of tarantulas using exuviae for international wildlife law enforcement. Conserv Genet 8:497–502
Seeber J, Rief A, Seeber GUH, Meyer E, Traugott M (2010) Molecular identification of detritivorous soil invertebrates from their faecal pellets. Soil Biol Biochem 42:1263–1267
Signorovitch AY, Dellaporta SL, Buss LW (2005) Molecular signatures for sex in the Placozoa. Proc Natl Acad Sci USA 102:15518–15522
Taberlet P, Luikart G (1999) Non-invasive genetic sampling and individual identification. Biol J Linn Soc 68:41–55
Uit de Weerd DR (2008) Delimitation and phylogenetics of the diverse land-snail family Urocoptidae (Gastropoda: Pulmonata) based on 28S rRNA sequence data: a reunion with Cerion. J Molluscan Stud 74:317–329
Wade CM, Mordan PB, Naggs F (2006) Evolutionary relationships among the Pulmonate land snails and slugs (Pulmonata, Stylommatophora). Biol J Linn Soc 87:593–610
Acknowledgments
These investigations would not have been possible without sampling permissions, funding and local support. We would like to thank Bernard Recorbet from the ‘Direction régionale de l’environnement, de l’aménagement et du logement de Corse’, the ‘Gordon and Betty Moore Foundation’ as part of the Moorea Biocode project and the ‘French National Research Agency’ as part of the ‘Losers’ project. This work was supported by the “Consortium National de Recherche en Génomique” and the “Service de Systématique Moléculaire” (UMS 2700 CNRS-MNHN). It is part of the agreement 2005/67 between the Genoscope and the Muséum National d’Histoire Naturelle on the project “Macrophylogeny of life” directed by Guillaume Lecointre. We thank Thierry Backeljau and Benoît Fontaine for helpful comments and fruitful discussion. We are very grateful to Neil Davis, Reo Terai, Vetea Liao and Valentine Brotherson for their help in the field in Moorea.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Régnier, C., Gargominy, O., Falkner, G. et al. Foot mucus stored on FTA® cards is a reliable and non-invasive source of DNA for genetics studies in molluscs. Conservation Genet Resour 3, 377–382 (2011). https://doi.org/10.1007/s12686-010-9345-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-010-9345-8