Abstract
The hormone receptor-positive (estrogen and/or progesterone receptor (PR)-positive) and HER2-negative breast cancer (BC) subtype is a biologically heterogeneous entity that includes luminal A-like (LumA-like) and luminal B-like (LumB-like) subtypes. Decreased PR levels is a distinctive biological feature of LumB-like tumors. These tumors also show reduced sensitivity to endocrine therapies and poorer prognosis than LumA-like tumors. Identification of biomarkers to accurately predict disease relapse in these subtypes is crucial in order to select effective therapies. We identified the tumor suppressor PDCD4 (programmed cell death 4), located in the nucleus (NPDCD4), as an independent prognostic factor of good clinical outcome in LumA-like and LumB-like subtypes. NPDCD4-positive LumB-like tumors presented overall and disease-free survival rates comparable to those of NPDCD4-positive LumA-like tumors, indicating that NPDCD4 improves the outcome of LumB-like patients. In contrast, NPDCD4 loss increased the risk of disease recurrence and death in LumB-like compared with LumA-like tumors. This, along with our results showing that LumB-like tumors present lower NPDCD4 positivity than LumA-like tumors, suggests that NPDCD4 loss contributes to endocrine therapy resistance in LumB-like BCs. We also revealed that PR induces PDCD4 transcription in LumB-like BC, providing a mechanistic explanation to the low PDCD4 levels in LumB-like BCs lacking PR. Finally, PDCD4 silencing enhanced BC cell survival in a patient-derived explant model of LumB-like disease. Our discoveries highlight NPDCD4 as a novel biomarker in LumA- and LumB-like subtypes, which could be included in the panel of immunohistochemical markers used in the clinic to accurately predict the prognosis of LumB-like tumors.
Similar content being viewed by others
Introduction
The hormone receptor (HR)-positive (estrogen receptor (ER) and/or progesterone receptor (PR)-positive) and HER2-negative BC subtype comprises up to 70% of all breast cancers (BCs) [1]. Treatment options for these patients include endocrine therapies directed to block ER action, either by antagonizing ligand binding to ER (e.g., tamoxifen), downregulating ER levels (e.g., fulvestrant), or blocking estrogen synthesis (e.g., aromatase inhibitors) (reviewed in [2]). Despite most women with HR-positive and HER2-negative (HR+/HER2−) disease respond to endocrine therapy, 30% of patients will relapse due to de novo or acquired resistance to these agents [1]. HR+/HER2− BC is a biologically heterogeneous entity that includes luminal A-like (LumA-like) and luminal B-like (LumB-like) subtypes [3]. LumB-like tumors show reduced sensitivity to endocrine therapies and poorer prognosis than LumA-like tumors [2, 4]. The St. Gallen Consensus guidelines have endorsed the use of immunohistochemical markers to define LumA-like and LumB-like BC subtypes [3]. Lower PR levels in LumB-like tumors than in LumA-like tumors is one of the distinctive biological features of these subtypes [3, 5]. However, accurate differentiation between LumA-like and LumB-like subsets as well as the prediction of their outcome remain a clinical challenge. Therefore, developing biomarkers guiding therapeutics for these subtypes is urgently required.
Tumor suppressor PDCD4 (programmed cell death 4), originally discovered as a gene upregulated during apoptosis [6], efficiently inhibits tumor development, growth, invasion, and metastasis [7,8,9]. Loss of PDCD4 expression is associated with poor prognosis in solid tumors including BC [10,11,12,13,14,15]. The best-acknowledged PDCD4 function is as inhibitor of protein translation, which occurs by PDCD4 interaction with the eukaryotic translation initiation factor 4A (eIF4A) or by its binding to specific mRNAs [16, 17]. Accumulating evidence also demonstrated that PDCD4 regulates gene transcription [18,19,20]. Downregulation of PDCD4 is associated with increased expression of the oncogenic microRNA-21 (miR-21), which targets PDCD4 mRNA [21]. The ErbB-2/HER2 pathway also modulates PDCD4 via its canonical and non-canonical mechanisms of actions [22, 23]. PDCD4 expression was detected in both the nucleus and the cytoplasm in normal breast and BC tissues [24,25,26]. The shuttle of PDCD4 between these subcellular compartments would have important effects on function regulation and seems to be cell-type specific [27]. To our knowledge, there are no studies exploring the clinical relevance of either nuclear or cytoplasmic PDCD4 in BC.
Here, we identify nuclear PDCD4 (NPDCD4) as an independent prognostic factor of good clinical outcome in LumA-like and LumB-like BCs, which can be included in the panel of immunohistochemical markers routinely used in the clinic to accurately predict the prognosis of LumB-like tumors. Notably, we clearly demonstrate that PR plays a key role in PDCD4 regulation in BC cells. Most interestingly, we show that PDCD4 silencing enhances tumor cell survival in a patient-derived explant (PDE) model of LumB-like disease.
Materials and Methods
Patients
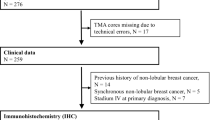
Paraffin-embedded tissue sections from 619 archived invasive BCs were selected from the Histopathology Department of Temuco Hospital (Chile), Hospital General de Agudos “Juan A. Fernández” (Argentina), Instituto de Oncología Henry Moore (Argentina), and Hospital Aeronáutico Central (Argentina) from 2001 to 2014. The median follow-up time was of 73 months (range, 2.4–144 months). This study was conducted according to the provisions of the Declaration of Helsinki and informed written consents were obtained from all patients. Study protocols were approved by the Review Boards on Human Research of the participating institutions. Patients included were women aged 18–85 (alive or deceased), were diagnosed with stage I–III BC as primary tumor, and whose data on baseline clinical features and outcomes were complete. Patients underwent surgical treatment, received adjuvant chemotherapy with anthracyclins, taxanes, and/or platinum compounds; and/or endocrine therapy with tamoxifen or anastrazole and/or trastuzumab; and/or radiation therapy according to standard protocols. The primary endpoints were disease-free survival (DFS), distant metastasis-free survival (DMFS), and local relapse-free survival (LRFS). DFS was defined as the time from BC diagnosis to the first recording of a recurrence or death, whichever came first. DMFS and LRFS were defined as the time from BC diagnosis to the first recording of a distant metastasis or a local recurrence, respectively. Local relapse was defined as recurrences of BC occurring in the ipsilateral breast, regional lymph nodes, and skin from the breast. Distant relapse was defined as recurrences of BC occurring beyond the confines of the ipsilateral breast, chest wall, or regional lymph nodes. Sites of distant relapse included the brain (and central nervous system), liver, lung, bone, pleural/peritoneal, and supraclavicular nodes. The secondary endpoint was the overall survival (OS). Pre-treatment patient staging was classified according to the American Joint Committee on Cancer (AJCC) system [28] through the Elston and Ellis histological grading system [29]. Clinicopathological data of the cohort is shown in Table 1.
Immunohistochemistry (IHC) and Histopathological Analysis
Antigen retrieval was performed in 10 mM sodium citrate buffer pH 6 for 20 min at 92 °C. Slides were incubated with a rabbit monoclonal PDCD4 antibody (D29C6, dilution 1:100 overnight at 4 °C; Cell Signaling Technology, USA), or rabbit IgG. Sections were incubated with a biotinylated anti-rabbit secondary antibody, dilution 1:400, for 30 min and then incubated with the VECTASTAIN® Elite® ABC-HRP Kit (Vector, USA) and developed in 3,3-diaminobenzidine (Cell Marque, USA). Slides were counterstained with Mayer’s hematoxylin solution (Sigma-Aldrich, USA). PDCD4 was independently evaluated by three board-certified clinical pathologists (MGP, PG, and AD) who ignored sample identity. Score discrepancies were re-evaluated and reconciled on a multiple-headed microscope. PDCD4 was scored as reported [11, 23]. ER, PR, and HER2 were evaluated by IHC as reported [30]. Guidelines for Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK) were followed in this work [31].
Publicly Available BC Databases Assayed for Validation
In order to validate the clinical impact of PDCD4 expression, the online Kaplan–Meier plotter database was used as previously described [32]. In addition, we used the METABRIC cohort. The METABRIC dataset [33], which includes BC-specific survival (BCSS), clinical data, and mRNA gene expression of primary breast tumors, was downloaded from cBioportal on 2019-03-01. This dataset counts with samples classified according to their gene expression profiles in GENIUS (Gene Expression progNostic Index Using Subtypes) subgroups (HER2+, ER+/HER2− high proliferation, ER+/HER2− low proliferation, ER−/HER2−, [34]). We used the data of 1222 patients classified as ER+/HER2− and displaying either low or high proliferation gene expression profiles (representing the LumA-like and LumB-like BC subtypes, respectively) and used the data file containing log2-transformed data, as provided by the original authors. Patients were segregated into high and low expression groups based on PDCD4 mRNA median values.
BC Cell Lines Data Mining
For analysis of PDCD4 expression in BC cell lines, we assessed the Cancer Cell Line Encyclopedia (CCLE) dataset [35] for transcriptomic characterization (RNA-seq, n = 50). To classify BC cell lines, literature mining was performed for ER, PR, and HER2 status as well as their intrinsic BC subtype. Processed mRNA expression data from the CCLE was downloaded from the CCLE portal (https://portals.broadinstitute.org/ccle, release version dated 2019-02-01). Two-sided Pearson’s correlation test was used to evaluate statistical significance in all these analyses. For multiple linear regression analysis among PDCD4, ER-alpha and PR mRNA levels, the R scatterplot3d package was used. PDCD4 mRNA levels in BC cell lines expressing or not ER and PR were retrieved from a publicly available dataset (GSE55350 [36]) using the GEO2R interactive web tool (http://www.ncbi.nlm.nih.gov/geo/info/geo2r.html).
Cell Lines and Treatments
T47D, MCF-7, BT-474, SK-BR-3, and HCC1569 cell lines were obtained from the American Type Culture Collection and JIMT-1 cells from the German Resource Center for Biological Material. T47D-Y cells were a gift from K. Horwitz (University of Colorado, Denver, CO). All lines were authenticated by short tandem repeat DNA profiling and were regularly screened for mycoplasma infection. Experiments in Figs. 3e, f and 4a–c were performed in complete media. T47D and MCF-7 cells were starved in 0.1% charcoalized fetal calf serum for 48 h before stimulation with medroxyprogesterone acetate (MPA).
Western Blot (WB)
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis and immunoblots were performed as reported [37]. Proteins were electroblotted onto nitrocellulose, and membranes were immunoblotted with the following antibodies: anti-PDCD4 (D29C6) from Cell Signaling Technology (Beverly, MA, USA); anti-ErbB-2/HER2 (C-18, sc-284), anti-ER-alpha (MC-20, sc-542), anti-GAPDH (0411, sc-47724), anti-Histone H3 (C16R, sc-8654-R) and anti-lamin B1 (B10, sc-374015) from Santa Cruz Biotechnology (Santa Cruz, CA, USA); anti-β tubulin (T0198) from Sigma-Aldrich (Saint Louis, MO, USA); and anti-PR (hPRa7) from Thermo Fisher Scientific (Waltham, MA; USA). Proteins were visualized with HRP-conjugated secondary antibodies (Vector Laboratories, Burlingame, CA, USA), using Amersham ECL Detection Reagent (GE Healthcare). Nuclear soluble, chromatin-bound and cytoplasmic protein fractions were isolated using the Subcellular Protein Fractionation Kit (78840, ThermoFisher) according to the manufacturer’s instructions. β Tubulin, lamin B1, and histone H3 were used to control cellular fractionation efficiency. Experiments in which PDCD4, ErbB-2/HER2, PR and ER-alpha protein levels were explored were repeated two to three times. Signal intensities of proteins were analyzed by densitometry using the ImageJ software (National Institutes of Health) and normalized to β tubulin or GAPDH bands. Densitometry was performed at different exposures to assure quantification within the linear detection range.
Plasmid Transient Transfections
Plasmids used in this work are detailed in [37, 38]. T47D-Y cells were transiently transfected with plasmids encoding human wild-type PR-B (kindly provided by K. Horwitz) or human wild-type PR-A (a gift from D. Edwards, Baylor College of Medicine, Houston TX). The plasmid encoding PR-B engineered to contain a point mutation in a conserved cysteine in the first zinc finger of the DNA-binding domain (C587A-PR), which lacks the ability to bind to DNA, was also a gift of K. Horwitz. Cells were transfected for 48 h with 2 μg of expression vectors using X-tremeGENE HP (Roche) following manufacturer’s instructions [23, 38].
siRNA Transfections
Small interfering RNAs (siRNAs) targeting human PR and PDCD4 were synthesized by Dharmacon. siRNA sequences were as follows: PR siRNA #1, 5′-CUAACAAAUCAUCGACUUA-3′; PR siRNA #2, 5′-GUAGUCAAGUGGUCUAAAU-3′ [39]; and PDCD4 siRNA, 5′-GGUGGCUGGAACAUCUAUU-3′ [40]. Experiments were performed with two different siRNA sequences for PR. We here present the results obtained with only one of them. A control siRNA from Dharmacon that does not target any known mammalian gene was used as negative control. Transfection of siRNA duplexes was performed using DharmaFECT #1 transfection reagent (Dharmacon, Inc.) following manufacturer’s instructions, using 100 nM of siRNAs, for 48 h. In all experiments, the knockdown effects produced by specific siRNAs were assessed at protein levels by western blot.
RNA Preparation and Real-Time RT-PCR
RNA was isolated and reverse transcription was performed as previously described [41]. The following primers were used for human PDCD4 cDNA: 5′-TATGATGTGGAGGAGGTGGATGTGA-3′; 5′-CCTTTCATCCAAAGGCAAAACTACA-3′. The primers used to amplify glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as a normalization control were 5′-CAGTCAGCCGCATCTTCTTTTG-3′; 5′-ACCAGAGTTAAAAGCAGCCCT-3′. The fold change of mRNA expression was calculated by normalizing the absolute PDCD4 mRNA amounts to GAPDH mRNA levels, used as an internal control.
ChIP Assays and Real-Time qPCR
ChIP assays were performed as described [37]. The antibodies used for ChIP assays were anti-PR (hPRa7) from Sigma-Aldrich; anti-p300 (clone RW128) and anti-acetyl histone H4 (06–866) from Millipore (Billerica, MA, USA); and IgG (PP64), from Sigma-Aldrich, which was used as negative control. Primers used for qPCR spanning the region containing PREs at positions -3699 and -3568, relative to human PDCD4 transcription start site (TSS), were 5′-GCAGCCCTAGGCTTCAGATATT-3′ and 5′-TGCCTACCAAATCAATCCACA-3′.
Analysis of Publicly Available ChIP-Seq Data
ChIP-seq and associated input datasets were downloaded from the publicly available resource Cistrome Database [42]. Curated data from Mohammed et al. [43] (CistromeDB ID: 53982, 53983, 53984, 54058, 54062, and 54066) was loaded and visualized with WashU EpiGenome Browser [42], allowing binding pattern comparison in the region comprising the human PDCD4 coding region and promoter.
PDE Culture of a LumB-Like BC Tumor
The BC tissue sample was obtained after written informed consent from the patient and approval of the Ethics Committees of IBYME and Sanatorio Sagrado Corazón. PDE culture was performed as reported [44]. Briefly, BC tissue was obtained and transported to the laboratory in DMEM-F12 (Dulbecco’s modified Eagle’s medium/nutrient mixture F12) supplemented with 10% fetal calf serum (FCS) within 1 h of surgery. Under sterile conditions, tissue was placed onto a 10-cm plate in fresh DMEM-F12, rinsed, and using a surgical blade, a 1-mm3-thick longitudinal section of the tissue sample was cut and placed into neutral-buffered formalin for paraffin embedding (day 0 sample). This section was used to determine the cancer content of the tissue following staining with hematoxylin and eosin (H&E) and IHC. The remaining tissue was dissected into 1-mm3 pieces (PDEs) and placed in quadruplicate on top of a pre-soaked gelatin sponge with 500 μL DMEM-F12 supplemented with 10% FCS inside 24-well plates. PDEs were transfected with siRNAs targeting PDCD4 or control siRNAs for 48 h. A PDE cultured for 48 h in complete media was used as the control condition. At the end of the experiment, PDEs were formalin-fixed and paraffin embedded for IHC and H&E or harvested for WB. Cell cytotoxicity was evaluated in PDE culture medium by a lactate dehydrogenase (LDH) release assay.
Statistics
Analyses were performed using the STATA version 15 software (Stata Inc., College Station, TX). Correlations between categorical variables were performed using the χ2 test or Fisher’s exact test when the number of observations obtained was under five. Cumulative survival probabilities were calculated according to the Kaplan–Meier method, and statistical significance was analyzed by log-rank or Wilcoxon test. Univariate and multivariate analyses were performed using Cox proportional hazards regression models. Adjustment for significant confounders was done to avoid increased bias and variability, unreliable confidence interval coverage, and problems with the model associated to the small size of our sample [45]. Variables that were statistically significant (P < 0.05) in the univariate model were included in the multivariate analysis. Given that clinical stage grading includes both tumor size and lymph nodes status information, when the three confounders were statistically significant, only the clinical stage was included in the multivariate model. The hazard ratio (HR) and its 95% confidence interval (CI) were calculated. Statistical differences were determined by two-tailed Student’s t test, when two groups were compared, or one-way ANOVA followed by Tukey’s test, when three or more groups were compared, using Prism 6 (GraphPad Software). All tests of statistical significance were two-sided. P values < 0.05 were regarded as statistically significant.
Results
NPDCD4 Expression Is Associated with Good Clinical Outcome in BC
To explore the clinical relevance of nuclear and cytoplasmic PDCD4 expression in BC, we conducted a retrospective study in a cohort of 619 primary invasive breast carcinomas from Latin American women. Clinical and pathological characteristics of specimens are shown in Table 1. IHC was used to study both NPDCD4 and cytoplasmic PDCD4 (CPDCD4) protein expression. NPDCD4 was scored, as previously reported, considering both the percentage of positive cells and staining intensity [11, 23]. A score of 0 represents no staining, 1+ weak nuclear staining in 1–30% of cells, 2+ intermediate staining in 31–70%, and 3+ strong staining in > 70% of cells. CPDCD4 levels were graded according to staining intensities (score 0, negative; score 1+, weak; score 2+, intermediate; score 3+, strong) [11]. Representative images of BC tissues corresponding to the different scores of NPDCD4 and CPDCD4 are shown in Fig. 1a and Supplementary Fig. 1. Positive and negative controls were run in parallel (Fig. 1b). Scores of 2+ and 3+ were considered positive for NPDCD4 and CPDCD4 expression. The majority of invasive BCs in our cohort (367 out of 619, 59.3%) lacked both NPDCD4 and CPDCD4 (Supplementary Table 1). Among the 619 tumors analyzed, 215 were NPDCD4-positive (34.7%) and 132 CPDCD4-positive (21.3%) (Supplementary Table 1). Only 44.2% of NPDCD4-positive patients (95 out of 215) also expressed CPDCD4, indicating that NPDCD4 is not necessarily accompanied by CPDCD4 (Supplementary Table 1). NPDCD4 positivity was associated with a lower clinical stage (P = 0.00026), more differentiated tumors (P = 6.43 × 10−6), and ER and PR expression (P = 9.15 × 10−9 and P = 2.76 × 10−9, respectively) (Table 2). Loss of NPDCD4 was associated with higher tumor size (P = 1.80 × 10−6). On the other hand, CPDCD4 was only associated with ER and PR expression (P = 0.033 and P = 0.001, respectively) (Table 2). Then, we explored the clinical relevance of either NPDCD4 or CPDCD4 in BC. Kaplan–Meier (KM) analysis revealed that BC patients bearing NPDCD4-positive tumors showed significantly higher overall survival (OS) and disease-free survival (DFS) as compared to patients whose tumors lacked NPDCD4 (Fig. 1c). Distant metastasis-free survival (DMFS), but not local relapse-free survival (LRFS), was significantly longer in NPDCD4-positive tumors than in NPDCD4-negative tumors (Fig. 1c). On the other hand, KM analysis showed that CPDCD4 only correlated with a higher OS in BC (Fig. 1c). Univariate analysis revealed that while NPDCD4 was associated with longer OS and DFS, CPDCD4 was only associated with longer OS (Fig. 1d). Higher tumor size, lymph node stage, clinical stage, tumor grade, and absence of ER and PR expression were associated with poor outcome in the univariate analysis (Fig. 1d). Multivariate analysis adjusted for tumor grade, clinical stage, and ER/PR status showed that only NPDCD4 and clinical stage were independent predictors of longer OS and DFS in BC (Fig. 1e). Our findings reveal that NPDCD4, but not CPDCD4, is associated with a less aggressive phenotype and good clinical outcomes in BC.
NPDCD4 expression is associated with good clinical outcome in BC. a NPDCD4 and CPDCD4 expression levels in primary invasive BC samples were evaluated by IHC and scored considering the percentage of positive cells and/or staining intensity, as previously reported [10,11,12,13,14,15, 22, 23]. Shown are representative images from tumor samples displaying scores 0 to 3+. Scores of 2+ and 3+ were considered positive for both NPDCD4 and CPDCD4 expression. b Positive and negative controls of PDCD4 IHC staining. Representative images from normal colorectal tissue used as a positive control for PDCD4 staining as previously reported [10,11,12,13,14,15, 22, 23]. Negative controls were obtained by using an isotype-nonspecific IgG antibody during staining procedure. c KM survival analysis and log-rank test were performed to correlate NPDCD4 or CPDCD4 expression with overall survival (OS), disease-free survival (DFS), distant metastasis-free survival (DMFS), or local relapse-free survival (LRFS) in BC patients. d, e Forest plots showing the hazard ratios (HR, squares) and 95% confidence intervals (CI, horizontal lines) of Cox univariate (d) and multivariate (e) analysis. Statistically significant P values are shown in bold. Scale bars = 20 μm
NPDCD4 Expression Is an Independent Biomarker of Good Clinical Outcome in LumA-Like and LumB-Like BC
Next, we investigated the prognostic value of NPDCD4 in surrogate BC subtypes. Patients were classified into clinical, IHC-derived BC subtypes according to the St. Gallen Consensus [3] as follows: HR-positive and HER2-negative (HR+/HER2−), HR-positive and HER2-positive (HR+/HER2+), HR-negative and HER2-positive (HR−/HER2+), and triple negative (TN, HR−/HER2−). A total of 565 out of 619 (91.3%) patients in our cohort presented complete histopathological and clinical data and were classified into clinical BC subtypes (Table 1). The incidence of HR+/HER2−, HR+/HER2+, HR−/HER2+, and TN subtypes in this cohort was comparable to that observed in Caucasian cohorts [4, 46, 47]. Besides, HR+/HER2− BC patients were classified as LumA-like or LumB-like based on the criteria of Perou and coworkers, where tumors were defined as LumA-like if they were ER-positive and presented a positive PR status ≥ 20%, or as LumB-like if they were ER-positive with low (< 20%) or negative PR expression [5]. As well acknowledged [2, 4], LumA-like tumors showed better prognoses than LumB-like tumors. LumA-like BCs presented a 5-year OS estimate of 90.3% (95%CI 86.1–94.5) and a 5-year DFS estimate of 82.2% (95%CI 76.7–87.6), while LumB-like BCs presented 5-year OS and DFS estimates of 77.7% (95%CI 69.9–85.6) and 73.3% (95%CI 64.9–81.7), respectively (Supplementary Fig. 2). The OS and DFS estimates of LumA- and LumB-like BCs from our cohort were comparable to those observed in other previously reported cohorts [48,49,50,51]. We found higher prevalence of NPDCD4 positivity in LumA-like (44.7%), LumB-like (33.0%), and HR+/HER2+ (46.2%) tumors as compared to HR−/HER2+ (10.8%) and TN (20.7%) ones (P = 0.030; Fig. 2a). Interestingly, NPDCD4 positivity was significantly lower in LumB-like tumors than in LumA-like tumors (33.0%, 95%CI 24.3–41.8 vs 44.7%, 95%CI 38.3–51.5, P = 0.0425). We then evaluated the association between NPDCD4 and clinicopathological characteristics in BC subtypes. NPDCD4 inversely correlated with clinical stage and tumor grade in LumA-like tumors, and with tumor size in LumB-like and TN BCs (Table 3). No association between clinicopathological characteristics and NPDCD4 was observed in HER2+ BCs, either HR+/HER2+ or HR−/HER2+ (Table 3). KM analysis revealed that NPDCD4 was associated with longer OS in LumA-like, LumB-like, and in HR+/HER2+ BCs (Fig. 2b). Our results also revealed that NPDCD4 was associated with longer DFS and DMFS in LumA-like and in LumB-like BCs, but not in HR+/HER2+ BCs (Fig. 2b). No significant association between NPDCD4 and clinical outcome was observed in either HR−/HER2+ or TN BCs (Fig. 2b). Interestingly, LumB-like patients displaying NPDCD4-positive tumors presented estimated OS, DFS, and DMFS rates similar to those observed in LumA-like tumors also expressing NPDCD4 (Fig. 2c), indicating that NPDCD4 improves the clinical outcome in LumB-like BC. In contrast, loss of NPDCD4 resulted in lower OS, DFS, and DMFS in LumB-like patients compared to LumA-like patients (Fig. 2c). Moreover, loss of NPDCD4 increased the risk of disease recurrence, distant metastasis recurrence and death by 50%, 61%, and 51%, respectively, in LumB-like tumors compared to LumA-like tumors (Fig. 2c). It is worth noting that no association between NPDCD4 and the treatment regimens (endocrine therapy and chemotherapy) was observed in patients bearing LumA-like and LumB-like tumors (Supplementary Table 2). Univariate analysis revealed that NPDCD4 was associated with longer OS in LumA-like and LumB-like BCs (Fig. 2d). Other clinicopathological parameters, such as tumor size, lymph nodes stage, and clinical stage were also associated with the clinical outcome (Fig. 2d). Multivariate analysis adjusted for clinical stage, identified NPDCD4 positivity as a significant and independent predictor of longer OS in LumA-like BC and of longer OS and DFS in LumB-like BC (Fig. 2d). To validate our findings, we took advantage of publicly available human BC datasets as surrogates to investigate the clinical impact of PDCD4. Using the online Kaplan–Meier plotter [32], we found that high PDCD4 mRNA levels are associated with good clinical outcome in luminal A (n = 1933) and luminal B (n = 1149) BC patients (Fig. 2e). In addition, the METABRIC dataset was used as a second cohort for validation. Our analysis considered METABRIC samples classified as ER+/HER2−, displaying either low or high proliferation gene expression profiles (representing the LumA-like and LumB-like BCs, respectively) [33]. KM analysis revealed that high PDCD4 mRNA levels were associated with longer BC-specific survival (BCSS) in both LumA-like (n = 619) and LumB-like (n = 603) BCs (Fig. 2f). No correlation between PDCD4 and the treatment regimens was observed in patients with LumA-like and LumB-like tumors from METABRIC cohort (Supplementary Table 2). Univariate analysis showed that PDCD4 mRNA was associated with longer BCSS in LumA-like and LumB-like BCs (Fig. 2g). Other clinicopathological parameters, such as tumor size, lymph nodes status, clinical stage and tumor grade were also associated with clinical outcome in LumA-like and/or LumB-like BCs. Multivariate analysis adjusted for tumor size, lymph nodes status, and tumor grade identified PDCD4 mRNA expression as an independent predictor of longer BCSS in both LumA-like and LumB-like BCs from the METABRIC cohort (Fig. 2g). Altogether, these findings demonstrate that NPDCD4 is an independent biomarker of good clinical outcome in LumA-like and LumB-like BC subtypes. Most importantly, our results might help to identify a subset of LumB-like patients with low levels or even absence of NPDCD4 expression and decreased OS, and DFS rates, which may underlie their reduced sensitivity to endocrine therapies.
NPDCD4 expression is an independent biomarker of good clinical outcome in LumA-like and LumB-like BC. a Graphical representation of NPDCD4-positive or NPDCD4-negative expression in tumor specimens corresponding to distinct surrogate BC subtypes. Patients were classified into the surrogate BC subtypes following St. Gallen International Expert Consensus, and NPDCD4 expression was detected by IHC and scored as detailed in Fig. 1. The number of positive or negative cases is shown as the percentage of the total number of cases analyzed by subtype. b KM survival analysis and log-rank test were performed to correlate NPDCD4 expression with overall survival (OS), disease-free survival (DFS), or distant metastasis-free survival (DMFS) in patients classified according to their BC subtype. c KM survival analysis and log-rank test performed in LumA-like and LumB-like BC tumors which express NPDCD4 (NPDCD4-positive) or not (NPDCD4-negative). Hazard ratios (HR) and 95% confidence intervals (CI) were also calculated using log-rank analysis. d Forest plots showing the HR (squares) and 95% CI (horizontal lines) of Cox univariate and multivariate analysis for LumA-like (left panel) and LumB-like (right panel) BC subtypes. e KM estimates of relapse-free survival (RFS) for patients with luminal A or luminal B BCs in relation to PDCD4 mRNA levels from KM plotter analysis [32]. KM plots of patient survival stratified by gene expression levels (high/low) using the median mRNA expression value of each group as cutoff point (Probe ID 202731_at). The analysis was performed in luminal A and luminal B tumors classified based on their intrinsic subtype as indicated by the online KM plotter. Red line represents low expression and blue line high expression. The hazard ratio (HR) including 95% confidence interval (CI) is shown. f KM survival analysis and log-rank test were performed to correlate PDCD4 mRNA levels with BC-specific survival (BCSS) in METABRIC BC patients from LumA-like (left panel) and LumB-like (right panel) BC subtypes. Patients were stratified by PDCD4 gene expression levels (high/low) using the median mRNA expression value of each group as cutoff point. g Forest plots showing the HR (squares) and 95% CI (horizontal lines) of Cox univariate and multivariate analysis for LumA-like (left panel) and LumB-like (right panel) BCs from the METABRIC cohort
PDCD4 Expression Depends on ER and PR in BC Cells
Since higher PDCD4 expression was observed in patients with HR-positive BCs compared to those bearing HR-negative tumors, we hypothesized that HRs (ER and/or PR) modulate PDCD4 expression. To test our theory, we explored the association between PDCD4 and ER and PR expression in BC cells. We interrogated the CCLE database [35] and found a positive correlation between PDCD4 and ER mRNA levels and between PDCD4 and PR mRNA levels (Fig. 3a). Moreover, multiple linear regression showed a stronger correlation among all three PDCD4, ER, and PR messages (Fig. 3b). To further explore the relationship between PDCD4 and HR, CCLE BC cell lines were classified into the different BC subtypes based on previous studies reporting ER, PR, and HER2 expression in said lines (Supplementary Table 3). Among the BC cells from the CCLE, 11 were classified as HR+/HER2−: LumB-like, 9 as HR+/HER2+, 6 as HR−/HER2+, and 24 as TN. As observed for NPDCD4 protein in our BC cohort, we found that PDCD4 mRNA levels were higher in cells belonging to LumB-like and HR+/HER2+ BC subtypes than in cells from the HR−/HER2+ and TN BC subtypes (Fig. 3c). We also studied PDCD4 mRNA levels in a dataset of pure ER+/PR+ luminal and ER−/PR− luminobasal lines generated from the same parental T47D BC cells by Horwitz and coworkers [36]. Consistently, we found that the pure ER+/PR+ cell line showed significantly higher PDCD4 levels than the pure ER−/PR− one (Fig. 3d). Next, we examined PDCD4 expression in a selected panel of BC cell lines: T47D and MCF-7 cells representative of the luminal subtype; BT-474 cells representative of the HR+/HER2+ BC subtype; and SK-BR-3, JIMT-1, and HCC1569 cells from the HR−/HER2+ BC subtype. Supporting the fact that expression of HRs is associated with higher levels of PDCD4 (Fig. 3c), we found higher levels of PDCD4 mRNA and protein in cell lines which express ER and PR (Fig. 3e, f). The ErbB-2/HER2 pathway has already been found to decrease PDCD4 levels via upregulation of microRNA-21 (miR-21), which targets PDCD4 [22, 23]. Consistently, among BC cells with comparable levels of HRs, we found that HER2-overexpressing BT-474 cells presented lower PDCD4 mRNA and protein levels than T47D cells, which lack HER2 overexpression (Fig. 3e, f). We found higher levels of PDCD4 mRNA and protein in T47D cells than in MCF-7 (Fig. 3e, f). Subcellular fractionation studies showed that T47D cells expressed PDCD4 both in the cytoplasmic and in the nuclear compartment, and, within the nucleus, PDCD4 was bound to the chromatin fraction (Fig. 3g). In contrast, PDCD4 was mainly located in the cytoplasm in MCF-7 cells (Fig. 3g). Although both T47D and MCF-7 cell lines display ER, MCF-7 cells show lower levels of constitutive PR expression (Fig. 3f). These findings, along with the fact that NPDCD4 positivity was significantly lower in LumB-like tumors (Fig. 2a), which either lack PR or display lower PR expression, than in LumA-like tumors, raise the exciting possibility that PR plays a key role in the regulation of PDCD4 levels.
PDCD4 expression depends on ER and PR in BC cells. a Correlation of PDCD4 mRNA levels with ER-alpha (left panel) or PR (right panel) in BC cell lines from the CCLE database (n = 50). Each dot represents a cell line. P value determined by linear regression analysis. b Multivariate linear regression analysis of ER-alpha, PR, and PDCD4 mRNA levels within the CCLE BC cells. For visualization of multivariate data relationship and a better comparison of ER-alpha, PR, and PDCD4 mRNA expression association, the regression plane is shown. c Differential PDCD4 mRNA levels in BC cells within the CCLE dataset grouped according to their BC subtype. Cell lines were classified into different BC subtypes as detailed in Supplementary Table 3. Whiskers represent the 5–95% distribution, boxes represent the interquartile range (25th and 75th percentiles), and the horizontal line in the box represents the median value. For c vs a: P < 0.01; for d vs a and b: P < 0.001; and for c vs b: P < 0.05. d PDCD4 mRNA levels were analyzed in a publicly available dataset of pure ER+/PR+ luminal versus pure ER−/PR− luminobasal cells generated by Horwitz and coworkers [36]. PDCD4 mRNA values are expressed as log2 transformed median-centered intensity. e PDCD4 mRNA expression levels were determined by RT-qPCR in a panel of BC cell lines. The relative expression levels of mRNA was calculated by normalizing the absolute levels of PDCD4 mRNA to GAPDH mRNA levels, which were used as an internal control, and setting the value of PDCD4 mRNA expression in T47D cells to 1. Data are presented as mean ± SEM from three independent experiments. For b and c vs a: P < 0.0001; for c vs b: P < 0.05. f PDCD4 protein expression was analyzed by WB in total cell lysates from representative BC cell lines with an anti-PDCD4 antibody. GAPDH was used as a loading control. The bar plot on the lower panel represents the values obtained after densitometry of PDCD4 bands and normalization to GAPDH bands from three independent experiments. Data are presented after setting the value of PDCD4 protein expression of T47D cells as 1. For b, c, d, e and f vs a: P < 0.001; for d, e, and f vs b, and e and f vs c: P < 0.01; for d vs c: P < 0.05. The control of HER2, PR, and ER expressions in the BC cell lines is shown in the right panel. HER2, PR, and ER protein expression was analyzed by WB. It is to note that in order to reveal the significantly lower PR expression levels in MCF-7 cells as compared to T47D cells, longer exposure times of the PR WB were required. β Tubulin was used as the loading control. g Cytosolic (C), nuclear (N), and chromatin (Chr) protein lysates were analyzed by WB. Total (T) lysates were blotted in parallel. β Tubulin, lamin B, and histone H3 were used to control cellular fractionation efficiency. The experiments shown in f to g are representative of three independent ones
PR Is a Key Regulator of PDCD4 Expression in BC Cells
We took advantage of the exquisite experimental model developed by Horwitz and coworkers comprising the T47D cell line which has become the major model to study the actions of progestins in human cells due to the constitutively high levels of PR. This model includes sublines derived from the T47D cell line, which enable the study of specific functions of PR [52]. PDCD4 protein levels were significantly higher in T47D cells than in the PR-null T47D-Y cells, and reconstitution of either PR isoform B (PR-B) or isoform A (PR-A) in T47D-Y cells restored PDCD4 expression (Fig. 4a). As another experimental strategy to demonstrate that PR participates in the regulation of PDCD4, we silenced PR expression in T47D cells and, as expected, we obtained the same results than those when using the T47D and the T47D-Y cells (Fig. 4b). Moreover, analysis of PDCD4 mRNA levels in T47D and T47D-Y cells also confirmed that PR knockdown decreased PDCD4 levels (Fig. 4c). To explore the molecular mechanisms involved in PDCD4 modulation by PR we performed bioinformatics studies to identify the presence of progesterone response elements (PREs) in PDCD4 promoter [53]. We identified PREs at positions − 4292, − 3599, and − 3568 bp relative to the human PDCD4 transcription start site (TSS) (Fig. 4d and Supplementary Fig. 3). Chromatin immunoprecipitation (ChIP)-sequencing data of PR in T47D cells treated with a synthetic progestin [43] obtained from the Cistrome database, revealed a PR binding peak located at − 3740 to − 3484 bp from the TSS of PDCD4 gene, which encompasses the − 3599 bp and − 3568 bp PREs identified by us (Supplementary Fig. 4). We performed ChIP assays to validate said PREs in our experimental conditions, i.e., in cells starved in charcoalized fetal calf serum (chFCS) and treated or not with MPA for 1 h. We found that the synthetic progestin MPA induced PR binding to the region containing these sites (Fig. 4d). Notably, PR was not significantly recruited to the PREs in the absence of MPA when the receptor is transcriptionally inactive. p300, a coactivator with histone acetyltransferase activity and marker of active gene transcription, was also recruited to these sites (Fig. 4d). Analysis of the local chromatin architecture revealed high levels of histone H4 acetylation (AcH4), a marker of chromatin activation (Fig. 4d). To study whether direct binding of PR to DNA is required for PDCD4 transcriptional regulation, T47D-Y cells were transfected with the C587A-PR mutant, which expresses a PR harboring a substitution of the cysteine 587 for alanine that renders the receptor unable to bind to DNA or to tether to other transcription factors bound to DNA [54]. Inability of PR capacity of binding to the DNA successfully reduced MPA-induced PR loading to the PDCD4 promoter, inhibited p300 recruitment, and significantly decreased AcH4 levels (Fig. 4d), further demonstrating requirement of PR binding function for PDCD4 transcriptional regulation. Progestins were found to induce T47D cells to progress through one round of cell division, followed by growth arrest at the G1/S phase of the second cycle [55, 56]. MPA treatment of T47D cells for 30 h, which is compatible with a condition where cells are arrested, showed an increase in PDCD4 protein levels (Fig. 4e and Supplementary Fig. 5). Similar results were obtained in MCF-7 cells, which, although at lower levels, still express functional PR (as evidenced by the observed PR downregulation after 36 h of MPA treatment) (Supplementary Fig. 6). Induction of PDCD4 mRNA levels in T47D cells was also observed in the presence of MPA at 18 h treatment (Fig. 4e). MPA failed to induce PDCD4 upregulation in the PR-null T47D-Y cells and in T47D-Y-C587-PR cells, the latter expressing a PR protein unable to bind to DNA, further confirming the requirement of PR DNA-binding function for progestin-mediated PDCD4 modulation (Fig. 4f). Altogether, our findings revealed that PR is a direct transcriptional regulator of PDCD4 in LumB-like BC cells.
PR is a key regulator of PDCD4 expression in BC cells. a PDCD4 expression was analyzed by WB as in Fig. 3f, in T47D-Y cells lacking PR expression or in T47D-Y cells transfected with vectors encoding PR isoform A (T47D-Y-A) or PR isoform B (T47D-Y-B). The middle panel shows the successful reconstitution of PR-A and PR-B expression. PDCD4 and PR bands underwent densitometry and values were normalized to β tubulin protein, used as a loading control, setting the value of control cells as 1. b T47D cells were transfected with PR or control siRNA and PDCD4 expression was evaluated by WB. Middle panel shows successful inhibition of PR expression by siRNA. PDCD4 and PR bands underwent densitometry and values are depicted as in panel a. Experiments shown in (a, b) were repeated twice with similar results. c PDCD4 mRNA expression levels were determined by RT-qPCR. The relative expression levels of mRNA upon PR depletion was calculated by normalizing the absolute levels of PDCD4 mRNA to GAPDH levels, which were used as an internal control, and setting the value of wild-type T47D cells to 1. Data are presented as mean ± SEM from three independent experiments. For b vs a: P < 0.05. d Recruitment of PR, p300, and H4 acetylation levels (AcH4) at the human PDCD4 promoter was analyzed by ChIP in cells stimulated with MPA (10 nM) for 1 h. Immunoprecipitated DNA was amplified by qPCR using primers (green arrows) flanking PREs (denoted in red) found by in silico analysis (MatInspector [http://www.genomatix.de]). Amounts of immunoprecipitated DNA were normalized to inputs and are reported relative to the amount obtained by IgG immunoprecipitation, used as negative control, which was set to one. Data is expressed as n-fold chromatin enrichment over IgG immunoprecipitation. Results are the mean ± SEM from three independent experiments. For b vs a: P < 0.001. e MPA induces PDCD4 protein and mRNA expression in T47D cells. Left panel: PDCD4 protein expression was analyzed by WB after stimulation with MPA (10 nM) for 30 h. PDCD4 WB bands underwent densitometry and values were normalized to GAPDH protein bands, used as a loading control, setting the value of untreated cells to 1. The experiment shown is a representative of three with similar results. Right panel: PDCD4 mRNA levels were determined by RT-qPCR as in panel c. Fold change of mRNA levels upon MPA treatment (10 nM, 18 h) was calculated by normalizing the absolute levels of PDCD4 mRNA to GAPDH levels, setting the value of untreated cells to 1. Data are presented as mean ± SEM from three independent experiments. For b vs. a: P < 0.05. f T47D-Y cells and T47D-Y cells transfected with a C587A-PR mutant (T47D-Y-C587A-PR) were analyzed for PDCD4 protein expression by WB and data was analyzed as in panel e. PDCD4 bands of T47D-Y and T47D-Y-C587A-PR cells correspond to a longer exposure time of the blots as compared with the bands of T47D cells, albeit all of them were performed in the linear range of detection. The experiments shown are representative of three, with similar results
Silencing PDCD4 Inhibits Cell Death in a PDE Model of LumB-Like BC
For preclinical assessment of PDCD4 biological effects in LumB-like BC, we developed a PDE from a freshly resected primary invasive mucinous breast carcinoma [44]. The primary tumor specimen presented a LumB-like phenotype, since it displayed positive ER expression and negative PR and HER2 expressions (Fig. 5a). As for the clinical management of this tumor, the patient was treated with therapies according to the LumB-like phenotype. Histological analysis revealed that the PDE maintained in culture for 48 h presented a tumor and stroma morphology similar to that observed in both the primary specimen and the original resected tumor tissue (i.e., day 0) (Fig. 5a). Moreover, ER, PR, and HER2 IHC staining showed that the PDE presented a LumB-like BC subtype which resembled the original resected tumor and the primary specimen (Fig. 5a). NPDCD4 expression was detected in the PDE (Fig. 5b). We found that the explant in which PDCD4 was silenced by transfection with PDCD4-specific siRNAs showed tumor morphology and histologic grade comparable to those observed in the control siRNA-transfected PDE (Fig. 5b). We also found that PDCD4 silencing increased the percentage of neoplastic tissue, supporting that PDCD4 modulates BC cell survival (Fig. 5b). Upon breakdown of the plasma membrane, the lactate dehydrogenase (LDH) enzyme is released from the cells into the culture medium, serving as a cell cytotoxicity marker [57]. Interestingly, we found that PDCD4 knockdown induced a significant decrease of LDH release (Fig. 5c), indicating inhibition of cell death. Altogether these findings demonstrate that PDCD4 abrogation favors BC cell survival in a PDE model of LumB-like BC.
Silencing PDCD4 inhibits cell death in a PDE model of LumB-like BC. a Method used for PDE tissue culture. After surgery, the pathologist resects a sample of the primary tumor specimen, which was dissected into 1 mm3 fragments. One of these fragments was immediately placed into neutral-buffered formalin for paraffin embedding (day 0) and other fragments were cultured in 24-well plates on a gelatin sponge under different treatment conditions (day 2). Lower panels show representative hematoxylin and eosin (H&E), ER, PR, and HER2 IHC staining of the primary tumor specimen, the tumor resected tissue (day 0), and the PDE (day 2). b Representative H&E and PDCD4 IHC stainings from PDEs transfected with PDCD4-specific siRNAs or control siRNAs for 48 h. The quantification of the percentage of neoplastic tissue of the entire section by H&E staining is depicted in the right panel. Lower panel shows WB control of efficacy of PDCD4 inhibition. PDCD4 bands underwent densitometry and values were normalized to β tubulin protein bands setting the value of control siRNA-transfected PDE to 1. c LDH release from PDEs transfected as in panel b. An aliquot of culture supernatant was removed for LDH assay. For b vs a, P < 0.001. Scale bars = 25 μm
Discussion
Identifying biomarkers for an accurate prediction of disease relapse in LumA-like and LumB-like BCs is crucial to select the most adequate adjuvant therapy. Here, we revealed NPDCD4 positivity as an independent biomarker of good clinical outcome in both LumA-like and LumB-like BCs. Our findings are particularly relevant for LumB-like BCs, which, among the luminal subtype, are the ones which show poor prognosis and reduced sensitivity to endocrine therapies aimed at inhibiting ER action [2, 4].
Nuclear and cytoplasmic levels of PDCD4 were found to decrease when normal breast tissue progresses to invasive ductal carcinoma [26]. The study of our BC cohort showed that NPDCD4 is an independent predictor of longer OS in LumA-like BCs and of longer OS and DFS in patients with LumB-like tumors. Consistent with PDCD4 role as inhibitor of metastasis [8, 9], we also revealed that NPDCD4 was associated with longer DMFS in said BC subtypes. Interestingly, LumB-like patients displaying NPDCD4-positive tumors presented favorable clinical outcomes, comparable to those observed in LumA-like patients also expressing NPDCD4. In contrast, NPDCD4 loss resulted in increased risk of disease recurrence and death in LumB-like tumors compared with LumA-like tumors. We also confirmed our results by in silico analysis using mRNA expression data from larger cohorts. NPDCD4 has already been associated with good clinical outcome in esophageal, colorectal, pancreatic ductal, and renal cell carcinomas [10,11,12,13]. On the other hand, CPDCD4 has been associated with good outcome in ovarian and nasopharyngeal carcinomas [14, 58]. The fact that PDCD4 is localized predominantly in the nucleus in normal breast tissues [25], along with our own findings of the clinical relevance of NPDCD4 in BC, supports the notion that NPDCD4 is functionally more relevant than CPDCD4 in BC.
Although most women with luminal-like disease respond to endocrine therapies, some of them, mainly patients bearing LumB-like BCs, develop fatal relapses [2]. LumA-like tumors express high ER and PR levels, are typically low grade, and have a low proliferative fraction (low Ki67), while LumB-like tumors present low or negative PR expression, are high grade, and have a high proliferative fraction [3, 59]. Patients within the LumB-like BC subtype receive chemotherapy in addition to endocrine treatment, whereas most patients with LumA-like disease avoid cytotoxic therapy [3]. Multigene assays, such as the 21 gene recurrence score or the PAM50 risk of relapse score (reviewed in [60]), are critical for a refined prognosis of LumA-like and LumB-like tumors. However, these assays are expensive and, therefore, less accessible than routine pathology tests [61]. Our findings showed that NPDCD4 is a prognostic factor of good clinical outcome in both LumA-like and LumB-like tumors defined according to their PR levels. Although further prospective studies would be required to validate our findings by immunohistochemistry in a larger cohort, our results were confirmed in LumA and LumB tumors from the KMplotter database and in LumA-like and LumB-like from the METABRIC cohort, in which patients were classified based on the proliferation fraction determined by their proliferation gene expression profile. Our studies demonstrate that both NPDCD4 protein and PDCD4 mRNA levels are associated with good clinical outcome in LumA-like and LumB-like BCs, independently of the criteria used to define these subtypes. Preclinical studies showed that PDCD4 expression is downregulated in BC cells resistant to tamoxifen, fulvestrant, and aromatase inhibitors [62,63,64]. Besides, PDCD4 silencing reduced the sensitivity of BC cells to tamoxifen [65], further demonstrating that PDCD4 loss is involved in resistance to endocrine therapy. Our findings identified a subset of LumB-like BCs with low levels or even absence of NPDCD4 expression and poor prognoses, indicating that NPDCD4 loss might contribute to endocrine therapy resistance in the clinic. In light of our findings, we recommend that NPDCD4 detection could be included in the panel of IHC markers routinely used in the clinic to accurately predict the prognosis of LumB-like patients and to guide addition of chemotherapy or targeted therapies for LumB-like patients bearing NPDCD4-negative tumors.
The study of the mechanisms controlling PDCD4 expression is crucial to elucidate why PDCD4 is lost during BC progression. PDCD4 regulation by ER has previously been reported. ER-alpha pathway inhibits microRNA-21 expression, leading to increased PDCD4 expression [64, 66]. Moreover, both ER-alpha and PDCD4 levels are decreased in tamoxifen- and fulvestrant-resistant BC cells as compared to those in sensitive ones [63, 64], further confirming ER involvement in PDCD4 modulation. PDCD4 regulation by PR has not been reported. A key differential biological feature between LumA-like and LumB-like tumors is that the latter either lack PR or show lower PR levels than LumA-like BCs. In this frame, one of our most exciting findings is that PR is a direct transcriptional regulator of PDCD4. This fact may underlie the inability of LumB-like tumors, showing low PR levels, to upregulate PDCD4, and also demonstrates a direct correlation between our findings in the clinic and those in BC models. PR possesses an intriguing role in BC biology. Accumulating evidence demonstrated that PR activation by progestogens, via both genomic and non-genomic mechanisms, favors BC cells proliferation and increases BC risk in patients (reviewed in [67, 68]). However, PR expression is an independent prognostic marker of good clinical outcome in ER-positive BC [69,70,71]. These seemingly contradictory effects of PR action may be explained by PR biphasic regulation of BC cell growth [55, 56]. Pioneering findings by Horwitz and coworkers demonstrated that progestins accelerate T47D BC cells through one round of cell division, followed by growth arrest at the G1 phase of the second cycle. Therefore, despite the initial proliferative burst, the effects of a single pulse of progestins are growth inhibitory [55]. Interestingly, our results demonstrated that MPA induces the expression of tumor suppressor PDCD4 in a context where growth is arrested in BC cells. This is consistent with previous findings showing that progestin-induced growth arrest is accompanied by increased levels of the cyclin-dependent protein kinase inhibitor p21, a cell-cycle inhibitor [55]. Here, we reveal a novel transcriptional mechanism by which PR induces PDCD4. Indeed, MPA induced PR binding to its PREs at the PDCD4 promoter, leading to PDCD4 expression. Our present data contribute to the understanding of the molecular mechanisms underlying PR growth inhibitory effects in BC models and of PR role as a biomarker of favorable prognosis, regardless of ER expression.
Our findings in a PDE model of LumB-like BC showed that PDCD4 silencing favors tumor cell survival, supporting the rationale for developing therapeutic strategies aimed at restoring PDCD4 expression in LumB-like BC.
In conclusion, our novel findings highlight the importance of NPDCD4 clinical assessment to accurately predict the outcomes of LumB-like BCs and to guide the addition of cytotoxic therapies. Current therapeutic strategies for HR+ advanced BC include inhibitors of cyclin-dependent kinases 4 and 6 (CDK4/6i), such as palbociclib and ribiciclib, which inhibit CDK4/6 kinase activity, thus preventing cell cycle progression and proliferation (reviewed in [72]). Since PDCD4 acts as a natural inhibitor of CDK4 [9, 24], detection of PDCD4 may serve as a biomarker to identify patients who will benefit from CDK4/6i. Moreover, combination of endocrine therapy with CDK4/6i would be a useful therapeutic strategy for NPDCD4-negative LumB-like tumors even in early BC.
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Musgrove EA, Sutherland RL (2009) Biological determinants of endocrine resistance in breast cancer. Nat Rev Cancer 9(9):631–643. https://doi.org/10.1038/nrc2713
Osborne CK, Schiff R (2011) Mechanisms of endocrine resistance in breast cancer. Annu Rev Med 62:233–247. https://doi.org/10.1146/annurev-med-070909-182917
Curigliano G, Burstein HJ, Winer P, Gnant M, Dubsky P, Loibl S, Colleoni M, Regan MM, Piccart-Gebhart M, Senn HJ, Thurlimann B, Andre F, Baselga J, Bergh J, Bonnefoi H, Brucker Y, Cardoso F, Carey L, Ciruelos E, Cuzick J, Denkert C, Di Leo A, Ejlertsen B, Francis P, Galimberti V, Garber J, Gulluoglu B, Goodwin P, Harbeck N, Hayes DF, Huang CS, Huober J, Hussein K, Jassem J, Jiang Z, Karlsson P, Morrow M, Orecchia R, Osborne KC, Pagani O, Partridge AH, Pritchard K, Ro J, Rutgers EJT, Sedlmayer F, Semiglazov V, Shao Z, Smith I, Toi M, Tutt A, Viale G, Watanabe T, Whelan TJ, Xu B (2017) De-escalating and escalating treatments for early-stage breast cancer: the St. Gallen International Expert Consensus Conference on the Primary Therapy of Early Breast Cancer 2017. Ann Oncol 28(8):1700–1712. https://doi.org/10.1093/annonc/mdx308
Kennecke H, Yerushalmi R, Woods R, Cheang MC, Voduc D, Speers CH, Nielsen TO, Gelmon K (2010) Metastatic behavior of breast cancer subtypes. J Clin Oncol 28(20):3271–3277. https://doi.org/10.1200/JCO.2009.25.9820
Prat A, Cheang MC, Martin M, Parker JS, Carrasco E, Caballero R, Tyldesley S, Gelmon K, Bernard PS, Nielsen TO, Perou CM (2013) Prognostic significance of progesterone receptor-positive tumor cells within immunohistochemically defined luminal A breast cancer. J Clin Oncol 31(2):203–209. https://doi.org/10.1200/JCO.2012.43.4134
Shibahara K, Asano M, Ishida Y, Aoki T, Koike T, Honjo T (1995) Isolation of a novel mouse gene MA-3 that is induced upon programmed cell death. Gene 166(2):297–301. https://doi.org/10.1016/0378-1119(95)00607-9
Cmarik JL, Min H, Hegamyer G, Zhan S, Kulesz-Martin M, Yoshinaga H, Matsuhashi S, Colburn NH (1999) Differentially expressed protein Pdcd4 inhibits tumor promoter-induced neoplastic transformation. Proc Natl Acad Sci U S A 96(24):14037–14042. https://doi.org/10.1073/pnas.96.24.14037
Yang HS, Matthews CP, Clair T, Wang Q, Baker AR, Li CC, Tan TH, Colburn NH (2006) Tumorigenesis suppressor Pdcd4 down-regulates mitogen-activated protein kinase kinase kinase kinase 1 expression to suppress colon carcinoma cell invasion. Mol Cell Biol 26(4):1297–1306. https://doi.org/10.1128/MCB.26.4.1297-1306.2006
Jansen AP, Camalier CE, Colburn NH (2005) Epidermal expression of the translation inhibitor programmed cell death 4 suppresses tumorigenesis. Cancer Res 65(14):6034–6041. https://doi.org/10.1158/0008-5472.CAN-04-2119
Fassan M, Cagol M, Pennelli G, Rizzetto C, Giacomelli L, Battaglia G, Zaninotto G, Ancona E, Ruol A, Rugge M (2010) Programmed cell death 4 protein in esophageal cancer. Oncol Rep 24(1):135–139. https://doi.org/10.3892/or_00000838
Mudduluru G, Medved F, Grobholz R, Jost C, Gruber A, Leupold JH, Post S, Jansen A, Colburn NH, Allgayer H (2007) Loss of programmed cell death 4 expression marks adenoma-carcinoma transition, correlates inversely with phosphorylated protein kinase B, and is an independent prognostic factor in resected colorectal cancer. Cancer 110(8):1697–1707. https://doi.org/10.1002/cncr.22983
Nagao Y, Hisaoka M, Matsuyama A, Kanemitsu S, Hamada T, Fukuyama T, Nakano R, Uchiyama A, Kawamoto M, Yamaguchi K, Hashimoto H (2012) Association of microRNA-21 expression with its targets, PDCD4 and TIMP3, in pancreatic ductal adenocarcinoma. Mod Pathol 25(1):112–121. https://doi.org/10.1038/modpathol.2011.142
Li X, Xin S, Yang D, Li X, He Z, Che X, Wang J, Chen F, Wang X, Song X (2012) Down-regulation of PDCD4 expression is an independent predictor of poor prognosis in human renal cell carcinoma patients. J Cancer Res Clin Oncol 138(3):529–535. https://doi.org/10.1007/s00432-011-1121-y
Wei NA, Liu SS, Leung TH, Tam KF, Liao XY, Cheung AN, Chan KK, Ngan HY (2009) Loss of programmed cell death 4 (Pdcd4) associates with the progression of ovarian cancer. Mol Cancer 8:70. https://doi.org/10.1186/1476-4598-8-70
Meric-Bernstam F, Chen H, Akcakanat A, Do KA, Lluch A, Hennessy BT, Hortobagyi GN, Mills GB, Gonzalez-Angulo A (2012) Aberrations in translational regulation are associated with poor prognosis in hormone receptor-positive breast cancer. Breast Cancer Res 14(5):R138. https://doi.org/10.1186/bcr3343
Yang HS, Jansen AP, Komar AA, Zheng X, Merrick WC, Costes S, Lockett SJ, Sonenberg N, Colburn NH (2003) The transformation suppressor Pdcd4 is a novel eukaryotic translation initiation factor 4A binding protein that inhibits translation. Mol Cell Biol 23(1):26–37. https://doi.org/10.1128/mcb.23.1.26-37.2003
Biyanee A, Ohnheiser J, Singh P, Klempnauer KH (2015) A novel mechanism for the control of translation of specific mRNAs by tumor suppressor protein Pdcd4: inhibition of translation elongation. Oncogene 34(11):1384–1392. https://doi.org/10.1038/onc.2014.83
Yang HS, Knies JL, Stark C, Colburn NH (2003) Pdcd4 suppresses tumor phenotype in JB6 cells by inhibiting AP-1 transactivation. Oncogene 22(24):3712–3720. https://doi.org/10.1038/sj.onc.1206433
Shiota M, Izumi H, Tanimoto A, Takahashi M, Miyamoto N, Kashiwagi E, Kidani A, Hirano G, Masubuchi D, Fukunaka Y, Yasuniwa Y, Naito S, Nishizawa S, Sasaguri Y, Kohno K (2009) Programmed cell death protein 4 down-regulates Y-box binding protein-1 expression via a direct interaction with Twist1 to suppress cancer cell growth. Cancer Res 69(7):3148–3156. https://doi.org/10.1158/0008-5472.CAN-08-2334
Hwang SK, Baker AR, Young MR, Colburn NH (2014) Tumor suppressor PDCD4 inhibits NF-kappaB-dependent transcription in human glioblastoma cells by direct interaction with p65. Carcinogenesis 35(7):1469–1480. https://doi.org/10.1093/carcin/bgu008
Frankel LB, Christoffersen NR, Jacobsen A, Lindow M, Krogh A, Lund AH (2008) Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 283(2):1026–1033. https://doi.org/10.1074/jbc.M707224200
Huang TH, Wu F, Loeb GB, Hsu R, Heidersbach A, Brincat A, Horiuchi D, Lebbink RJ, Mo YY, Goga A, McManus MT (2009) Up-regulation of miR-21 by HER2/neu signaling promotes cell invasion. J Biol Chem 284(27):18515–18524. https://doi.org/10.1074/jbc.M109.006676
Venturutti L, Romero LV, Urtreger AJ, Chervo MF, Cordo Russo RI, Mercogliano MF, Inurrigarro G, Pereyra MG, Proietti CJ, Izzo F, Diaz Flaque MC, Sundblad V, Roa JC, Guzman P, Bal de Kier Joffé ED, Charreau EH, Schillaci R, Elizalde PV (2016) Stat3 regulates ErbB-2 expression and co-opts ErbB-2 nuclear function to induce miR-21 expression, PDCD4 downregulation and breast cancer metastasis. Oncogene 35(17):2208–2222. https://doi.org/10.1038/onc.2015.281
Goke R, Barth P, Schmidt A, Samans B, Lankat-Buttgereit B (2004) Programmed cell death protein 4 suppresses CDK1/cdc2 via induction of p21(Waf1/Cip1). Am J Phys Cell Physiol 287(6):C1541–C15C6. https://doi.org/10.1152/ajpcell.00025.2004
Qi L, Bart J, Tan LP, Platteel I, Sluis T, Huitema S, Harms G, Fu L, Hollema H, Berg A (2009) Expression of miR-21 and its targets (PTEN, PDCD4, TM1) in flat epithelial atypia of the breast in relation to ductal carcinoma in situ and invasive carcinoma. BMC Cancer 9:163. https://doi.org/10.1186/1471-2407-9-163
Wen YH, Shi X, Chiriboga L, Matsahashi S, Yee H, Afonja O (2007) Alterations in the expression of PDCD4 in ductal carcinoma of the breast. Oncol Rep 18(6):1387–1393. https://doi.org/10.3892/or.18.6.1387
Lankat-Buttgereit B, Goke R (2009) The tumour suppressor Pdcd4: recent advances in the elucidation of function and regulation. Biol Cell 101(6):309–317. https://doi.org/10.1042/BC20080191
Singletary SE, Allred C, Ashley P, Bassett LW, Berry D, Bland KI, Borgen PI, Clark G, Edge SB, Hayes DF, Hughes LL, Hutter RV, Morrow M, Page DL, Recht A, Theriault RL, Thor A, Weaver DL, Wieand HS, Greene FL (2002) Revision of the American Joint Committee on Cancer staging system for breast cancer. J Clin Oncol 20(17):3628–3636. https://doi.org/10.1200/JCO.2002.02.026
Page DL, Ellis IO, Elston CW (1995) Histologic grading of breast cancer. Let’s do it. Am J Clin Pathol 103(2):123–124. https://doi.org/10.1093/ajcp/103.2.123
Schillaci R, Guzman P, Cayrol F, Beguelin W, Diaz Flaque MC, Proietti CJ, Pineda V, Palazzi J, Frahm I, Charreau EH, Maronna E, Roa JC, Elizalde PV (2012) Clinical relevance of ErbB-2/HER2 nuclear expression in breast cancer. BMC Cancer 12(1):74. https://doi.org/10.1186/1471-2407-12-74
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) Reporting recommendations for tumor marker prognostic studies. J Clin Oncol 23(36):9067–9072. https://doi.org/10.1200/JCO.2004.01.0454
Gyorffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat 123(3):725–731. https://doi.org/10.1007/s10549-009-0674-9
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Graf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Langerod A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Borresen-Dale AL, Brenton JD, Tavare S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486(7403):346–352. https://doi.org/10.1038/nature10983
Haibe-Kains B, Desmedt C, Rothe F, Piccart M, Sotiriou C, Bontempi G (2010) A fuzzy gene expression-based computational approach improves breast cancer prognostication. Genome Biol 11(2):R18. https://doi.org/10.1186/gb-2010-11-2-r18
Ghandi M, Huang FW, Jane-Valbuena J, Kryukov GV, Lo CC, McDonald ER III, Barretina J, Gelfand ET, Bielski CM, Li H, Hu K, Andreev-Drakhlin AY, Kim J, Hess JM, Haas BJ, Aguet F, Weir BA, Rothberg MV, Paolella BR, Lawrence MS, Akbani R, Lu Y, Tiv HL, Gokhale PC, de Weck A, Mansour AA, Oh C, Shih J, Hadi K, Rosen Y, Bistline J, Venkatesan K, Reddy A, Sonkin D, Liu M, Lehar J, Korn JM, Porter DA, Jones MD, Golji J, Caponigro G, Taylor JE, Dunning CM, Creech AL, Warren AC, McFarland JM, Zamanighomi M, Kauffmann A, Stransky N, Imielinski M, Maruvka YE, Cherniack AD, Tsherniak A, Vazquez F, Jaffe JD, Lane AA, Weinstock DM, Johannessen CM, Morrissey MP, Stegmeier F, Schlegel R, Hahn WC, Getz G, Mills GB, Boehm JS, Golub TR, Garraway LA, Sellers WR (2019) Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 569(7757):503–508. https://doi.org/10.1038/s41586-019-1186-3
Knox AJ, Scaling AL, Pinto MP, Bliesner BS, Haughian JM, Abdel-Hafiz HA, Horwitz KB (2014) Modeling luminal breast cancer heterogeneity: combination therapy to suppress a hormone receptor-negative, cytokeratin 5-positive subpopulation in luminal disease. Breast Cancer Res 16(4):418. https://doi.org/10.1186/s13058-014-0418-6
Beguelin W, Diaz Flaque MC, Proietti CJ, Cayrol F, Rivas MA, Tkach M, Rosemblit C, Tocci JM, Charreau EH, Schillaci R, Elizalde PV (2010) Progesterone receptor induces ErbB-2 nuclear translocation to promote breast cancer growth via a novel transcriptional effect: ErbB-2 function as a coactivator of Stat3. Mol Cell Biol 30(23):5456–5472. https://doi.org/10.1128/MCB.00012-10
Proietti CJ, Rosemblit C, Beguelin W, Rivas MA, Diaz Flaque MC, Charreau EH, Schillaci R, Elizalde PV (2009) Activation of Stat3 by heregulin/ErbB-2 through the co-option of progesterone receptor signaling drives breast cancer growth. Mol Cell Biol 29(5):1249–1265. https://doi.org/10.1128/MCB.00853-08
Izzo F, Mercogliano F, Venturutti L, Tkach M, Inurrigarro G, Schillaci R, Cerchietti L, Elizalde PV, Proietti CJ (2014) Progesterone receptor activation downregulates GATA3 by transcriptional repression and increased protein turnover promoting breast tumor growth. Breast Cancer Res 16(6):491. https://doi.org/10.1186/s13058-014-0491-x
Santhanam AN, Baker AR, Hegamyer G, Kirschmann DA, Colburn NH (2010) Pdcd4 repression of lysyl oxidase inhibits hypoxia-induced breast cancer cell invasion. Oncogene 29(27):3921–3932. https://doi.org/10.1038/onc.2010.158
Cordo Russo RI, Beguelin W, Diaz Flaque MC, Proietti C, Venturutti L, Galigniana NM, Tkach M, Guzman P, Roa JC, O'Brien N, Charreau EH, Schillaci R, Elizalde PV (2015) Targeting ErbB-2 nuclear localization and function inhibits breast cancer growth and overcomes trastuzumab resistance. Oncogene 34(26):3413–3428. https://doi.org/10.1038/onc.2014.272
Mei S, Qin Q, Wu Q, Sun H, Zheng R, Zang C, Zhu M, Wu J, Shi X, Taing L, Liu T, Brown M, Meyer CA, Liu XS (2017) Cistrome Data Browser: a data portal for ChIP-Seq and chromatin accessibility data in human and mouse. Nucleic Acids Res 45(D1):D658–DD62. https://doi.org/10.1093/nar/gkw983
Mohammed H, Russell IA, Stark R, Rueda OM, Hickey TE, Tarulli GA, Serandour AA, Birrell SN, Bruna A, Saadi A, Menon S, Hadfield J, Pugh M, Raj GV, Brown GD, D’Santos C, Robinson JL, Silva G, Launchbury R, Perou CM, Stingl J, Caldas C, Tilley WD, Carroll JS (2015) Progesterone receptor modulates ERalpha action in breast cancer. Nature 523(7560):313–317. https://doi.org/10.1038/nature14583
Centenera MM, Hickey TE, Jindal S, Ryan NK, Ravindranathan P, Mohammed H, Robinson JL, Schiewer MJ, Ma S, Kapur P, Sutherland PD, Hoffmann CE, Roehrborn CG, Gomella LG, Carroll JS, Birrell SN, Knudsen KE, Raj GV, Butler LM, Tilley WD (2018) A patient-derived explant (PDE) model of hormone-dependent cancer. Mol Oncol 12(9):1608–1622. https://doi.org/10.1002/1878-0261.12354
Vittinghoff E, McCulloch CE (2007) Relaxing the rule of ten events per variable in logistic and Cox regression. Am J Epidemiol 165(6):710–718. https://doi.org/10.1093/aje/kwk052
Ross JS, Slodkowska EA, Symmans WF, Pusztai L, Ravdin PM, Hortobagyi GN (2009) The HER-2 receptor and breast cancer: ten years of targeted anti-HER-2 therapy and personalized medicine. Oncologist 14(4):320–368. https://doi.org/10.1634/theoncologist.2008-0230
Garrido-Castro AC, Lin NU, Polyak K (2019) Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer Discov 9(2):176–198. https://doi.org/10.1158/2159-8290.CD-18-1177
Cheang MC, Chia SK, Voduc D, Gao D, Leung S, Snider J, Watson M, Davies S, Bernard PS, Parker JS, Perou CM, Ellis MJ, Nielsen TO (2009) Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst 101(10):736–750. https://doi.org/10.1093/jnci/djp082
Dawood S, Hu R, Homes MD, Collins LC, Schnitt SJ, Connolly J, Colditz GA, Tamimi RM (2011) Defining breast cancer prognosis based on molecular phenotypes: results from a large cohort study. Breast Cancer Res Treat 126(1):185–192. https://doi.org/10.1007/s10549-010-1113-7
Garcia Fernandez A, Gimenez N, Fraile M, Gonzalez S, Chabrera C, Torras M, Gonzalez C, Salas A, Barco I, Cirera L, Cambra MJ, Veloso E, Pessarrodona A (2012) Survival and clinicopathological characteristics of breast cancer patient according to different tumour subtypes as determined by hormone receptor and Her2 immunohistochemistry. A single institution survey spanning 1998 to 2010. Breast. 21(3):366–373. https://doi.org/10.1016/j.breast.2012.03.004
Hennigs A, Riedel F, Gondos A, Sinn P, Schirmacher P, Marme F, Jager D, Kauczor HU, Stieber A, Lindel K, Debus J, Golatta M, Schutz F, Sohn C, Heil J, Schneeweiss A (2016) Prognosis of breast cancer molecular subtypes in routine clinical care: a large prospective cohort study. BMC Cancer 16(1):734. https://doi.org/10.1186/2Fs12885-016-2766-3
Sartorius CA, Groshong SD, Miller LA, Powell RL, Tung L, Takimoto GS, Horwitz KB (1994) New T47D breast cancer cell lines for the independent study of progesterone B- and A-receptors: only antiprogestin-occupied B-receptors are switched to transcriptional agonists by cAMP. Cancer Res 54(14):3868–3877
Clarke CL, Graham JD (2012) Non-overlapping progesterone receptor cistromes contribute to cell-specific transcriptional outcomes. PLoS One 7(4):e35859. https://doi.org/10.1371/journal.pone.0035859
Takimoto GS, Tasset DM, Eppert AC, Horwitz KB (1992) Hormone-induced progesterone receptor phosphorylation consists of sequential DNA-independent and DNA-dependent stages: analysis with zinc finger mutants and the progesterone antagonist ZK98299. Proc Natl Acad Sci U S A 89(7):3050–3054. https://doi.org/10.1073/pnas.89.7.3050
Groshong SD, Owen GI, Grimison B, Schauer IE, Todd MC, Langan TA, Sclafani RA, Lange CA, Horwitz KB (1997) Biphasic regulation of breast cancer cell growth by progesterone: role of the cyclin-dependent kinase inhibitors, p21 and p27(Kip1). Mol Endocrinol 11(11):1593–1607. https://doi.org/10.1210/mend.11.11.0006
Lange CA, Richer JK, Horwitz KB (1999) Hypothesis: progesterone primes breast cancer cells for cross-talk with proliferative or antiproliferative signals. Mol Endocrinol 13(6):829–836. https://doi.org/10.1210/mend.13.6.0290
Goliwas KF, Richter JR, Pruitt HC, Araysi LM, Anderson NR, Samant RS, Lobo-Ruppert SM, Berry JL, Frost AR (2017) Methods to evaluate cell growth, viability, and response to treatment in a tissue engineered breast cancer model. Sci Rep 7(1):14167. https://doi.org/10.1038/s41598-017-14326-8
Zhen Y, Liu Z, Yang H, Yu X, Wu Q, Hua S, Long X, Jiang Q, Song Y, Cheng C, Wang H, Zhao M, Fu Q, Lyu X, Chen Y, Fan Y, Liu Y, Li X, Fang W (2013) Tumor suppressor PDCD4 modulates miR-184-mediated direct suppression of C-MYC and BCL2 blocking cell growth and survival in nasopharyngeal carcinoma. Cell Death Dis 4:e872. https://doi.org/10.1038/cddis.2013.376
Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thurlimann B, Senn HJ (2013) Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol 24(9):2206–2223. https://doi.org/10.1093/annonc/mdt303
Prat A, Ellis MJ, Perou CM (2011) Practical implications of gene-expression-based assays for breast oncologists. Nat Rev Clin Oncol 9(1):48–57. https://doi.org/10.1038/nrclinonc.2011.178
Burstein HJ, Curigliano G, Loibl S, Dubsky P, Gnant M, Poortmans P, Colleoni M, Denkert C, Piccart-Gebhart M, Regan M, Senn HJ, Winer EP, Thurlimann B (2019) Estimating the benefits of therapy for early-stage breast cancer: the St. Gallen International Consensus Guidelines for the primary therapy of early breast cancer 2019. Ann Oncol 30(10):1541–1557. https://doi.org/10.1093/annonc/mdz235
Chen Z, Yuan YC, Wang Y, Liu Z, Chan HJ, Chen S (2015) Down-regulation of programmed cell death 4 (PDCD4) is associated with aromatase inhibitor resistance and a poor prognosis in estrogen receptor-positive breast cancer. Breast Cancer Res Treat 152(1):29–39. https://doi.org/10.1007/s10549-015-3446-8
Fan M, Yan PS, Hartman-Frey C, Chen L, Paik H, Oyer SL, Salisbury JD, Cheng AS, Li L, Abbosh PH, Huang TH, Nephew KP (2006) Diverse gene expression and DNA methylation profiles correlate with differential adaptation of breast cancer cells to the antiestrogens tamoxifen and fulvestrant. Cancer Res 66(24):11954–11966. https://doi.org/10.1158/0008-5472.CAN-06-1666
Manavalan TT, Teng Y, Appana SN, Datta S, Kalbfleisch TS, Li Y, Klinge CM (2011) Differential expression of microRNA expression in tamoxifen-sensitive MCF-7 versus tamoxifen-resistant LY2 human breast cancer cells. Cancer Lett 313(1):26–43. https://doi.org/10.1016/j.canlet.2011.08.018
Jansen AP, Camalier CE, Stark C, Colburn NH (2004) Characterization of programmed cell death 4 in multiple human cancers reveals a novel enhancer of drug sensitivity. Mol Cancer Ther 3(2):103–110
Wickramasinghe NS, Manavalan TT, Dougherty SM, Riggs KA, Li Y, Klinge CM (2009) Estradiol downregulates miR-21 expression and increases miR-21 target gene expression in MCF-7 breast cancer cells. Nucleic Acids Res 37(8):2584–2595. https://doi.org/10.1093/nar/gkp117
Truong TH, Lange CA (2018) Deciphering steroid receptor crosstalk in hormone-driven cancers. Endocrinology 159(12):3897–3907. https://doi.org/10.1210/en.2018-00831
Proietti CJ, Cenciarini ME, Elizalde PV (2018) Revisiting progesterone receptor (PR) actions in breast cancer: insights into PR repressive functions. Steroids 133:75–81. https://doi.org/10.1016/j.steroids.2017.12.015
Costa SD, Lange S, Klinga K, Merkle E, Kaufmann M (2002) Factors influencing the prognostic role of oestrogen and progesterone receptor levels in breast cancer--results of the analysis of 670 patients with 11 years of follow-up. Eur J Cancer 38(10):1329–1334. https://doi.org/10.1016/s0959-8049(02)00067-9
Purdie CA, Quinlan P, Jordan LB, Ashfield A, Ogston S, Dewar JA, Thompson AM (2014) Progesterone receptor expression is an independent prognostic variable in early breast cancer: a population-based study. Br J Cancer 110(3):565–572. https://doi.org/10.1038/bjc.2013.756
Blows FM, Driver KE, Schmidt MK, Broeks A, van Leeuwen FE, Wesseling J, Cheang MC, Gelmon K, Nielsen TO, Blomqvist C, Heikkila P, Heikkinen T, Nevanlinna H, Akslen LA, Begin LR, Foulkes WD, Couch FJ, Wang X, Cafourek V, Olson JE, Baglietto L, Giles GG, Severi G, McLean CA, Southey MC, Rakha E, Green AR, Ellis IO, Sherman ME, Lissowska J, Anderson WF, Cox A, Cross SS, Reed MW, Provenzano E, Dawson SJ, Dunning AM, Humphreys M, Easton DF, Garcia-Closas M, Caldas C, Pharoah PD, Huntsman D (2010) Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med 7(5):e1000279. https://doi.org/10.1371/journal.pmed.1000279
O'Leary B, Finn RS, Turner NC (2016) Treating cancer with selective CDK4/6 inhibitors. Nat Rev Clin Oncol 13(7):417–430. https://doi.org/10.1038/nrclinonc.2016.26
Acknowledgments
In memory of EH Charreau, our mentor and dear colleague. We thank WD Tilley (Dame Roma Mitchell Cancer Research Laboratories, University of Adelaide, Australia) for his advice and help to perform PDE culture protocol and A Molinolo (UCSD, USA) for his constant help and support. We would also like to acknowledge H Martinetto for allowing us to use their facilities, S Muñoz for assistance with statistical studies, P Pomata for the technical assistance, and Fundación René Baron and Fundación Williams for their institutional support.
Funding
This work was supported by the National Agency of Scientific Promotion of Argentina (IDB/PICT 2012-668, PID 2012-066, PICT 2015-1587, and PICT 2017-1072, all awarded to PVE; PICT 2017-0419, awarded to RICR); the National Cancer Institute of Argentina (INC 2016 research grant, awarded to PVE); and the Foundation Alberto J. Roemmers Research Grant awarded to RCR and PVE.
Author information
Authors and Affiliations
Contributions
PVE, RICR, and SM were responsible for the conception and design of the study. SM, MFC, VAC, CJP, RS, RICR, and PVE developed methodology. SM, MFC, VAC, MGP, LV, FI, ARD, PG, AD, JCR, MEC, SB, SF, DLDV, CL, FA, TC, EC, SA, EGD, FGG, RS, RICR, and PVE acquired the data (and also acquired and managed patients, provided facilities, and so on). RICR, PVE, SM, MFC, VAC, MGP, PG, AD, CJP, EGD, and RS analyzed and interpreted the data. RICR, PVE, and SM wrote the manuscript. RICR, PVE, RS, CJP, and EGD revised the manuscript. RICR and PVE supervised the study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethics Approval and Consent to Participate
This study was performed in line with the principles of the Declaration of Helsinki and informed written consents were obtained from all patients. Study protocols were approved by the Review Boards on Human Research of the following institutions: Universidad de La Frontera-Temuco Hospital (Chile), Hospital General de Agudos Juan A. Fernández (approval number: 201217, Argentina), Instituto de Oncología Henry Moore (Argentina), Hospital Aeronáutico Central (Argentina), and Sanatorio Sagrado Corazón (Argentina).
Consent for Publication
Not applicable.
Code Availability
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Supplementary Fig. 1
(PNG 742 kb).
Supplementary Table 1
(DOCX 8 kb).
Supplementary Fig. 2
(PNG 191 kb).
Supplementary Table 2
(DOCX 10 kb).
Supplementary Table 3
(DOCX 332 kb).
Supplementary Fig. 3
(PDF 72 kb).
Supplementary Fig. 4
(PNG 217 kb).
Supplementary Fig. 5
(PNG 188 kb).
Supplementary Fig. 6
(PDF 179 kb).
Rights and permissions
About this article
Cite this article
Madera, S., Chervo, M.F., Chiauzzi, V.A. et al. Nuclear PDCD4 Expression Defines a Subset of Luminal B-Like Breast Cancers with Good Prognosis. HORM CANC 11, 218–239 (2020). https://doi.org/10.1007/s12672-020-00392-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12672-020-00392-4