Abstract
Purpose
The aim of the study was to apply and optimize the process of bioconversion of pig bristle waste using keratinolytic enzymes of Bacillus cereus PCM 2849, and to evaluate the amino acid composition of the resultant hydrolysate.
Methods
Hydrolysis with concentrated culture fluid of B. cereus was applied for bioconversion of pig bristles, after thermo-chemical pretreatment with sulfite. The effect of substrate concentration, sulfite concentration during pretreatment and reaction temperature on the release of amino acids was determined using Box-Behnken design. Amino acid composition of the obtained hydrolysate was determined by HPLC. Structural condition and substructural changes of the residual substrate were evaluated with SEM microscopy and FTIR spectroscopy.
Results
The applied enzymatic preparation for bristle biodegradation was verified to contain multiple proteases of a wide molecular weight range. A regression model was developed, in which influential parameters were: linear effect of substrate concentration, followed by quadratic effects of reaction temperature, substrate concentration and pretreatment. Optimum reaction conditions were also determined. The resultant hydrolysate was rich in branched-chain amino acids. Residual substrate was detriorated and sulfitolytic cleavage of disulfides and alteration of protein secondary structures was confirmed.
Conclusions
Application of B. cereus crude keratinase allowed for partial hydrolysis of pig bristles, preceded by sulfitolytic pretreatment. A regression model was built to describe the process of hydrolysis to release free amino acids, at constant enzyme load. Hydrolysis in given conditions allowed to obtain hydrolysate rich in branched chain amino acids. The presented process poses an alternate way of management over pig bristles, a hard-to-degrade keratinous waste.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Keratinous proteins are most recalcitrant constituents of by-products generated by meat and poultry processing industries. The management over this waste remains a challenging issue, due to the unique structure and composition of keratin-based skin appendages, resulting in their exceptional sturdiness. In the face of constantly rising production levels of swine and poultry an increasing inflow of these side-products is expected. As these keratinous materials contain at least 80 % protein, processes aimed at recovery of valuable proteins, peptides or amino acids are highly desirable [1].
Since application of keratin meals in livestock feeding has been lately subject to severe legal restrictions in most European countries, in response to outbreaks of prion diseases in the past, development of novel conversion methods has become critical. Although utilization of keratinous waste through combustion or co-combustion is widely used and offers simplicity and efficiency, it requires highly specialized facilities. Moreover, the concept of destroying proteinaceous materials instead of their exploitation is questionable.
According to the EU Waste Framework Directive (Directive 2008/98/EC) two types of organic waste management have been defined, comprising aerobic and anaerobic processes. The former include composting and the latter involve biogas production. Composting of keratinous waste appears as a cost-effective and ecologically safe method, especially when closed bioreactors for dynamic composting are used. Nevertheless, to achieve beneficial efficiency of keratin biodegradation it is essential to incorporate keratin-degrading bacteria, fungi or actinomycetes [1–3]. Biomethanation, anaerobic conversion of organic waste by methanogenic bacteria, is also applicable for feather waste processing. Nevertheless, the acceptable efficiency of the process is achieved in a two-step procedure, where chemical or microbiological degradation of keratin precedes the anaerobic fermentation [4, 5].
Conventional methods used for the bioconversion of keratinous waste, generated during processing of animal raw materials, are based mainly on the thermal and chemical treatment. Steam cooking, a widely used technique for production of feather meal, although energy-consuming, does not employ chemical compounds that become troublesome after use [6]. Application of lime or other alkalies, as well as mineral acids, combined with heat treatment, is a known practice to convert keratinous materials into hydrolysates applicable as feed components, soil fertilizer, etc. and offers high process throughput [7, 8]. Both approaches allow for obtaining by-products of moderately enhanced digestibility, yet with usually deepened imbalance in the essential amino acid content. The implementation of enzymatic or microbial digestion, significantly improves the biological and technological value of keratin meals or hydrolysates by providing more advantageous amino acid balance and high digestibility [9–12]. Therefore, process designs based on the application of keratinolytic microorganisms or their proteolytic enzymes represent promising alternatives [13].
Hydrothermal pretreatment was proved to be advantageous as a method for preparation of feathers or hooves and horn prior to hydrolysis with bacterial or fungal keratinases [14, 15]. An alternative for keratinous substrate preparation is based on sulfitolytic cleavage of disulfide bonds provided by sulfite. Sulfite enhances keratin feather digestion when present in the reaction environment, but it is especially effective when applied in the thermal treatment of the keratin substrate. Significant improvement of feather and bristle digestion with microbial keratinases, was achieved at relatively low sulfite concentrations of 10 and 100 mM, respectively [16, 17].
Enzymatic digestion of keratins with purified or crude keratinases appears to be one of the preferred approaches to process keratinous waste into valuable hydrolysates. It allows for production of feather meal of improved dietary value, amino acid balance and digestibility [14, 18]. Occasionally, combined chemical and enzymatic hydrolysis also proves to be advantageous [10, 19]. Bacteria-mediated keratin breakdown is known to be a cumulative effect of proteolytic cleavage as well as auxiliary reducing factors, including released reduced thiols or indigenous disulfide reductase enzymes [20, 21]. Nevertheless, enzymatic keratinolysis in vitro, conducted in the absence of red-ox potential of living microbial cells usually requires additional reducing agents to support disulfide bonds cleavage. This could be attained either by introducing chemical reducers into reaction environment or through initial substrate pretreatment [22, 23].
Numerous keratinolytic bacteria, filamentous fungi and actinomycetes have been isolated from various environments and characterized, nevertheless bacteria of the genus Bacillus represent the dominating group. The biotechnological interest in keratinolytic bacilli results mainly from their capability to biosynthesize a diversity of extracellular keratinases, but also from low nutrient requirements that allow growth in simple media where keratin serves as a sole nutrient source, spore forming capabilities that secure culture continuity especially in composting process where disadvantageous conditions may occur, mesophilic growth conditions of most of the species ensuring low energy requirements of cultivation and also occurrence of extremophilic species producing thermostable keratinases. Applicatory potential of keratinolytic proteases or keratin hydrolysates obtained with Bacillus bacteria has been recently explored with special regard to certain branches of industry. Grazziotin et al. [18] evaluated the nutritional quality of chicken feather hydrolysates obtained in culture of B. licheniformis. Also, purified keratinase from B. licheniformis ER-15 was applied for bioconversion of feathers into feather meal of improved dietary value [14]. Likewise, keratinase from B. subtilis RSE163 proved to be applicable for production of feather hydrolysate for feed supplementation [24]. Keratin hydrolysates may serve as a source of bioactive compounds. Fakhfakh et al. [25] applied B. pumilus A1 for biodegradation of wool waste, resultant in a product of advantageous amino acid and peptidic composition, exhibiting exceptional antioxidative properties. Similarly, feathers hydrolyzed by fermentation with B. subtilis S1-4 was a source of a specific antioxidative peptide [26]. Applicability of feather hydrolysate as soil fertilizer was confirmed by Bose et al. [27]. Fermentation of feather waste with B. amyloliquefaciens 6B allowed for obtaining a product effective as agricultural nitrogen input. It also exhibited antifungal properties due to the presence of surfactin, that allowed for biocontrol of phytopathogenic fungi. Keratinolytic bacteria found multiple potential applications in leather and textile industry. Proteases from B. subtilis P13 and Bacillus sp. SB12 were effectively used in the process of dehairing, which in contrast to conventional lime-sulfide dehairing, not only allowed for the reduction of effluents, but also positively affected treated leathers [28, 29]. Hoof and horn hydrolysate produced with B. subtilis was also applied as an agent to reduce consumption of chromium salts in the process of tanning, contributing to a significantly lowered discharge of chromium [30]. Finally, production of proteolytic enzymes on keratinous waste materials constantly remains the most frequent application of keratinolytic bacteria. Due to a range of vital technological properties, e.g. thermostability, alkali and solvent stability, detergent compatibility and unique specificity, the enzymes present high predisposition for commercialization [31].
Enzymatic capabilities of Bacillus cereus species involving its immense proteolytic potential, frequently becomes a subject of applicatory research. Recently, the ability of keratin biodegradation related to biosynthesis of specific keratinases frequently undergoes detailed investigation. Several keratinolytic B. cereus strains have been described as exceptionally efficient in decomposition of chicken feather keratin, within culture conditions. The strains produced mainly serine proteinases, to utilize keratin as a nutrient source, enabling cell growth. Fermentation process allowed to obtain hydrolysates of improved lysine, methionine and threonine balance, as compared to the crude substrate [32–34]. In addition, B. cereus strains exhibiting specificity towards biodegradation of other hard keratins, like wool or ram horn, were also described. Their applicatory potential was confirmed through the analysis of obtained fermentation products [35, 36]. Highly proteolytic B. cereus strains were also proved to be applicable for the recovery of proteinaceous fractions form other agro-industrial by-products, e.g. brewer’s spent grain [37].
Bacillus cereus PCM 2849 was described previously to effectively decompose keratins, mainly feathers, in liquid culture conditions. Likewise, its crude keratinase was readily capable of efficient digestion of chicken feathers and the yield of the process could be further enhanced by substrate pretreatment [16]. The objective of the study was to optimize enzymatic digestion of pretreated pig bristle with concentrated keratinase of B. cereus PCM 2849. A Box-Behnken experimental design was applied to estimate the effect of three influential parameters: substrate content, sulfite concentration during substrate pretreatment and reaction temperature, on the release of amino acids.
Materials and Methods
Bacterial Strain and Culture Conditions
Keratinolytic bacteria B. cereus PCM 2849 applied in the study, formerly referred to as B. cereus B5esz, have been described elsewhere [38]. The strain was deposited in the Polish Collection of Microorganisms at the Institute of Immunology and Experimental Therapy in Wroclaw, Poland.
Keratinase production was carried out in 500 cm3 flasks, at 30 °C, 180 rpm, in 100 cm3 of medium consisting of (%): MgSO4 0.1, KH2PO4 0.01, FeSO4·7H2O 0.001, CaCl2 0.01, yeast extract 0.05 and degreased chicken feathers 1.0. Medium was set to pH 7.1 prior to autoclaving (121 °C, 20 min). Nutrient broth culture (glucose 1.0 % and nutrient broth 0.8 %) of approximately 1.2 × 108 cfu cm−3 served as inoculum (1 cm3 per flask).
Crude Enzyme Preparation
The bacterial culture was terminated on the 3rd day, feather debris was removed on the Whatman no. 2 filter paper and biomass was separated by centrifugation (5000 g, 20 min, 4 °C). The culture fluid was concentrated on the Labscale TFF System (Millipore) with a Pellicon XL 50 casette, Ultracel-10 PLCGC membrane (10 kDa cutoff). The obtained preparation was stored in portions at −24 °C.
Zymography
Zymographic analysis of the concentrated culture fluid was performed. The sample was mixed at the ratio 6:4 with the sample buffer (Tris–HCl 0.32 M; pH 6.8; glycerol 48 %; SDS 8 %; bromophenol blue 0.06 %). Samples were loaded onto 8 % polyacrylamide gel (5 % staking gel) containing copolymerized casein 0.1 %. Electrophoresis was performed at constant 20 mA, at 2 °C. Subsequently, the gel was washed twice with Triton-X 2.5 %, once with the incubation buffer (Tris–HCl 0.05 M, pH 7.5, containing CaCl2 2 mM and sodium azide 0.02 %) and incubated for 24 h at 30 °C in the same buffer. Staining with Coomassie Blue and decolorization with methanol: acetic acid: water (50:10:40) was applied to visualize the proteolytic activity bands.
Pretreatment of Pig Bristles
Pig bristle was subject to a two-step pretreatment, prior to enzymatic digestion. The first pretreatment step was developed in order to remove tissue remnants, non-keratinous proteins and lipids from the crude pig bristle bulk. It was performed by conducting a short-term culture of B. cereus PCM 2849 in a mixture consisting of pig bristle 10 g, distilled water 250 cm3 and bacterial inoculum in nutrient broth 50 cm3. Afterwards, the solids were separated, washed with tap water and dried. The secondary, thermo-chemical pretreatment was performed by autoclaving the substrate (121 °C, 20 min) in a solution of sodium sulfite (1 g bristle per 100 cm3 of solution), according to Łaba and Szczekała [16]. The pretreated substrate was subsequently washed with tap and distilled water to remove the pretreatment agent and finally dried at room temperature.
Enzymatic Digestion of Pig Bristle
The enzymatic digestion experiment of uncut, pretreated bristle was conducted using the concentrated culture fluid of B. cereus, in a 24-hour reaction in a mixture containing: bristle 0.2–1.0 g, enzyme solution 25 PU, buffer (Tris–HCl 0.05 M, pH 7.5) to 10 cm3, CaCl2 2 mM [16]. After digestion the reaction mixture was subsequently cooled, filtered through glass-fibre filter Marcherey-Nagel GF-1 and centrifuged (12,000 g, 10 min). Concentration of free amino groups assayed according to Snyder and Sobociński [39] served as a measure of the digestion level.
Optimization of Enzymatic Hydrolysis
Optimization of hydrolysis conditions was conducted in a 15-run Box-Behnken design, involving three independent variables, at three levels: pig bristle content (2; 6; 10 % wv−1), sulfite concentration during initial substrate pretreatment (10; 110; 210 mM) and reaction temperature (35; 50; 65 °C). Constant enzyme load and reaction time were maintained. Concentration of free amino groups released into reaction mixtures served as the response (dependent variable). The relationship between the independent variables and the response was formulated according to the second-order polynomial equation:
where Y was the predicted response, β0 was the intercept and regression coefficients were designated as follows: β1, β2, β3 (linear), β11, β22, β33 (square) and β12, β13, β23 (interaction). The experiment design, polynomial equation fit, regression and ANOVA statistics, as well as response desirability analysis were performed with Statistica 10 (StatSoft Inc.). The Profiler tool of Statistica 10 software was applied for obtaining the level of predictor variables to produce the optimum outcome.
Amino Acid Composition of Bristle Hydrolysate
Amino acid composition of bristle hydrolysate, obtained at optimized conditions, was determined using HPLC, according to Henderson et al. [40]. Initial derivatization with O-phthalaldehyde was performed. The analysis made on a HPLC 1100 Series system (Agilent Technologies) equipped with the ZORBAX Eclipse-AAA column, 4.6 × 150 mm, 3.5 µm (Agilent Technologies).
Microscopic Examination of Bristle Degradation
Visual analysis of bristles following enzymatic hydrolysis was performed using scanning electron microscopy (SEM) on a Hitachi S3400 microscope.
FTIR Measurements
The structural changes in bristles after pretreatment and enzymatic hydrolysis were analyzed by Fourier transform infrared spectroscopy (FTIR). Measurements were performed on the VERTEX 70 spectrometer with dedicated program for Fourier transformation. Specimens were prepared as KBr pellets. Measurements were recorded within the 4000–400 cm−1 range with 2 cm−1 resolution.
Results and Discussion
Enzymatic digestion of keratinous waste proteins is considered as one of the feasible methods to obtain valuable by-products. The B. cereus PCM 2849 strain is a potent keratinase producer. Application of concentrated keratinase extract for keratin hydrolysis and evaluation of crucial factors influencing the process were described previously [16, 38]. In the present study, the concentrated culture fluid of B. cereus PCM 2849 was applied for the hydrolysis of pig bristles. Zymographic analysis revealed that this enzymatic preparation contained at least 7 active proteolytic fractions. Three dominating activity bands were denoted above the molecular weight of 110 kDa and four bands within a range of 25–80 kDa, where the 60 kDa fraction prevailed (Fig. 1). Bacillus cereus 1268 inquired by Mazotto et al. [33], when cultured in feather medium, produced predominantly high molecular weight proteases of caseinolytic and keratinolytic activity.
An experimental design was employed to determine the empirical relationship between the capability of the concentrated culture fluid of B. cereus PCM 2849 to release amino acid content and variables comprising substrate concentration, the extent of pretreatment and reaction temperature, according to the layout presented in the Table 1. The Box-Cox transformation statistics revealed that, conferring to the significant χ2 value, residual sum of squares was reduced with the Box-Cox transformation with lambda of 0.1027 (Table 2). Therefore, a natural logarithm transformation of the dependent variable was applied. Consecutively, insignificant χ2 was obtained, which suggested no need for further data transformation. Transformation of dependent variables is a widely used practice in factorial design analysis, leading to improvement of residual variance and distribution, frequently required in biochemical trials [41, 42].
A regression model was developed for the process of hydrolysis, considering the release of amino acids as an outcome. A high suitability of the model was obtained, as indicated by the coefficient of determination R2 = 0.9876 and adjusted R2 = 0.9653. According to the model, significance of linear and quadratic effects describing all three variables a was confirmed, as verified by p value below 0.05. The further analysis of standardized effects revealed following order of independent variables, according to their influence on the dependent variable: X1 > X3X3 > X1X1 > X2X2. The strongest linear effect of substrate concentration was denoted, followed by quadratic effects of incubation temperature, substrate concentration and substrate pretreatment. No significant interaction between the variables was observed (Table 3).
The obtained results allowed to define the polynomial Eq. (2) to describe the regression model (significant terms underlined):
ANOVA results of the model implied their significance, according to the F value of 42.93, further confirmed by a “lack of fit” tests of insignificant rank (Table 4).
Analysis of the predicted response surface plots allowed to assess variation of the dependent variables as functions of selected independent variables. The contour plot in the Fig. 2 confirmed highest influence of the bristle content on the response variable. In the present study, enzyme to substrate ratio was modified by changing substrate content, at constant enzyme load. It was demonstrated that the maximum applicable substrate concentration was close to the border of the experimental layout. Nevertheless, it appeared that further increase of substrate concentration should not produce positive effect.
The plot of bristle load versus temperature confirmed the dominating effect of the former and the convex optimum area of the latter (Fig. 2). Excessive load of proteinaceous substrate could result in low hydrolysis yield due to enzyme inhibition by reaction products. Also keratin substrate reactivity declination might occur during prolonged incubation period [41].
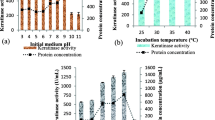
During industrial application of enzymatic catalysis it is essential to determine process temperature balanced between optimum value for enzyme activity and providing optimum enzyme stability in the specific reaction conditions. This could be attained exclusively through properly adapted experimental design. The predicted response surface plots of the response as a function of temperature and pretreatment revealed an optimum area for these parameters (Fig. 3). The most advantageous temperature for the hydrolysis was lower than the overall optimum for B. cereus PCM 2849 keratinases [38], while further increment of the pretreatment would not be legitimate. Chen et al. [43] in the inquiries on a 44.8 kDa keratinase from B. cereus H10 established comparable optima for reaction on asocasein, pH 7.57 and temperature 59 °C, using an experimental design. The enzyme also demonstrated high activity on native and milled porcine hair.
It is significant, that an optimum value of sulfite concentration during pretreatment was determined for the release of amino acids from the keratinous substrate, as indicated by a peak areas in the Figs. 3 and 4. Typically, the extent of the initial substrate pretreatment should not exhibit an optimum value, as it gradually leads to substrate degradation. Hence, excessive pretreatment would cause advanced substrate decomposition [10, 16]. In the present study an optimum for the dependent variable was denoted, therefore the desired value of keratin pretreatment should result rather from its sulfitolytic cleavage, rather than protein and amino acid release and loss during the pretreatment stage. This implied functions that produced maximum values.
Analysis of the plotted profiles for the built model allowed to obtain independent variable values that maximize the overall outcome of the response (Fig. 5). The defined values of the variables, X1 = 7.6, X2 = 122.3 and X3 = 47.7, allowed to attain maximum outcome corresponding to a predicted ln amino acid concentration of 1.68 (5.38 mg cm−3 as glycine).
The applied conditions of the enzymatic hydrolysis of pig bristles did not allow for the complete degradation of the substrate, nevertheless, symptoms of advanced decomposition were denoted. Visual analysis with SEM microscopy (Fig. 6) revealed damage and peeling of the cuticular layer (a), disorganized cuticle scales (b), breakings in the hair cortex (c) and partial exhibition of cortical cells (d).
The obtained bristle hydrolysate was especially rich in leucine and other branched-chain amino acids (BCAAs), valine and isoleucine (Table 5). It is significant that collective action of proteases in the tested culture fluid of B. cereus PCM 2849 allowed for the release of amino acids other than the most abundant ones, except for the dominating leucine. Most abundant amino acids in native pig bristles comprise glutamic acid, cysteine, leucine, serine and aspartic acid, as determined in a chemically hydrolysed substrate [44]. The different amino acid profile obtained in our study might be a result either of the crude enzyme’s specificity or the decomposition process occurring mainly in the bristles outer layers, since the substrate was purposefully undisrupted prior to hydrolysis. Elevated levels of leucine, tyrosine and methionine could result from chymotrypsin-like specificity, but not trypsin or elastase specificity [18]. However, thermal treatment might also affect the composition of bristle substrate, since Wang and Parsons [45] presented slightly elevated concentration of leucine and valine in steam-cooked hog hair meals. High amounts of BCAAs could be obtained as well by alkaline hydrolysis of animal hair [7, 46, 47]. Nevertheless, this type of treatment, in contrast to enzymatic hydrolysis, affects the initial amino acid composition of proteins by destroying some amino acids and formation of others like lysinoalanine and lanthionine [48].
Residual bristles after hydrolysis were subject to FTIR analysis in order to determine substructural changes in the substrate (Fig. 7). The appearance of a small peak at 1040 cm−1 after substrate pretreatment allowed to confirm a sulfitolytic effect to take place. It represents S–O stretching vibrations in Bunte salt residues [49]. In the sample after the hydrolysis process this signal partially remained, which, in accordance with the lack of stretching vibration from S–H bonds at 2560 cm−1, demonstrated no reduced thiols formation during hydrolysis, possibly resulting from sulfitolytic cleavage of most accessible disulfide bonds [50]. The signal between 1600 and 1700 cm−1 is usually referred to as Amide I region and it is known to be sensitive to secondary structure of proteins, it was therefore resolved into components [51]. In the initial bristle substrate exhibited nearly equal content of α-helix and β-sheet, 45 and 55 %, respectively (Fig. 8). The disordered and other structures were not included in the components resolution, since their share was below 3 % in all tested samples, similarly to characteristics of wool keratin presented by Wojciechowska et al. [52]. After thermo-chemical pretreatment the proportion was shifted towards α-helix domination (65 %), further increased through substrate hydrolysis (82 %). Similar correspondence of secondary structures is typically observed between unhydrolysed and hydrolysed keratins of hair-type appendages, however unlike in the case of feather keratin, where β-sheet and other structures prevail [49, 52].
Conclusions
Crude keratinase of B. cereus PCM 2849 was applied in biodegradation of pig bristles, an exceptionally hard to degrade keratinous waste material. As a result, partial hydrolysis of pig bristles was obtained, preceded by sulfitolytic pretreatment of the substrate. A regression model was built to optimize the process of hydrolysis to release free amino acids, at constant enzyme load. The produced amino acid cocktail was especially rich in branched chain amino acids, while residual substrate was characterized by substantial structural deterioration. The presented process of bristles bioconversion into hydrolysate could serve as a way of management of this hardly degradable biomaterial, especially for non-feed applications.
References
Korniłłowicz-Kowalska, T., Bohacz, J.: Biodegradation of keratin waste: theory and practical aspects. Waste Manage. 31, 1689–1701 (2011)
Rodziewicz, A., Łaba, W., Sobolczyk, J., Grzelak, A., Drozd, J.: Composting of keratinic waste with application of bacterial inoculum in a rotary bioreactor. Chem. Eng. Equip. 3, 95–97 (2009). [in polish]
Ichida, J.M., Krizovaa, L., LeFevreb, C.A., Keenerc, H.M., Elwellc, D.L., Burtt Jr., E.H.: Bacterial inoculum enhances keratin degradation and biofilm formation in poultry compost. J. Microbiol. Methods 47(2), 199–208 (2001)
Forgács, G., Niklasson, C., Horváth, I.S., Taherzadeh, M.J.: Methane production from feather waste pretreated with Ca(OH)2: process development and economical analysis. Waste Biomass Valorization 5(1), 65–73 (2014)
Mézes, L., Tamás, J.: Feather Waste Recycling for Biogas Production. Waste Biomass Valorization 6(5), 899–911 (2015)
Staroń, P., Banach, M., Kowalski, Z., Wzorek, Z.: Utilization of selected slaughterhouse waste by hydrolysis. Tech. Trans. Chem. 10(107), 333–341 (2010). (In Polish)
Coward-Kelly, G., Agbogbo, F.K., Holtzapple, M.T.: Lime treatment of keratinous materials for the generation of highly digestible animal feed: 2. Animal hair. Bioresour. Technol. 97, 1344–1352 (2006)
Nustorova, M., Braikova, D., Gousterova, A., Vasileva-Tonkova, E., Nedkov, P.: Chemical, microbiological and plant analysis of soil fertilized with alkaline hydrolysate of sheep’s wool waste. World J. Microbiol. Biotechnol. 22(4), 383–390 (2006)
Karthikeyan, R., Balaji, S., Sehgal, P.K.: Industrial applications of keratins-a review. J. Sci. Ind. Res. 66, 710–715 (2007)
Mokrejs, P., Krejci, O., Svoboda, P., Vasek, V.: Modeling technological conditions for breakdown of waste sheep wool. Rasayan J. Chem. 4(4), 728–735 (2011)
Costa, J.C., Barbosa, S.G., Sousa, D.Z.: Effects of pretreatment and bioaugmentation strategies on the anaerobic digestion of chicken feathers. Bioresour. Technol. 120, 114 (2012)
Kumar, A.G., Swarnalatha, S., Gayathri, S., Nagesh, N., Sekaran, G.: Characterization of an alkaline active–thiol forming extracellular serine keratinase by the newly isolated Bacillus pumilus. J. Appl. Microbiol. 104, 411–419 (2008)
Lasekan, A., Bakar, F.A., Hashim, D.: Potential of chicken by-products as sources of useful biological resources. Waste Manage. 33, 552–565 (2013)
Tiwary, E., Gupta, R.: Rapid conversion of chicken feather to feather meal using dimeric keratinase from Bacillus licheniformis ER-15. J. Bioprocess. Biotech. 2(4), 123 (2012)
Veselá, M., Friedrich, J.: Amino acid and soluble protein cocktail from waste keratin hydrolysed by a fungal keratinase of Paecilomyces marquandii. Biotechnol. Bioprocess Eng. 14, 84–90 (2009)
Łaba, W., Szczekała, K.B.: Keratinolytic proteases in biodegradation of pretreated feathers. Pol. J. Environ. Stud. 22(4), 1101–1109 (2013)
Łaba, W., Kopeć, W., Chorążyk, D., Kancelista, A., Piegza, M., Malik, K.: Biodegradation of pretreated pig bristles by Bacillus cereus B5esz. Int. Biodeter. Biodegrad. 100, 116–123 (2015)
Grazziotin, A., Pimentel, F.A., De Jong, E.V., Brandelli, A.: Nutritional improvement of feather protein by treatment with microbial keratinase. Anim. Feed Technol. 126, 135–144 (2006)
Kim, W.K., Lorenz, E.S., Patterson, P.H.: Effect of enzymatic and chemical treatments on feather solubility and digestibility. Poult. Sci. 81, 95–98 (2002)
Cedrola, S.M.L., De Melo, A.C.N., Mazotto, A.M., Lins, U., Zingali, R.B., Rosado, A.S., Peixoto, R.S., Vermelho, A.B.: Keratinases and sulfide from Bacillus subtilis SLC to recycle feather waste. World J. Microbiol. Biotechnol. 28, 1259–1269 (2012)
Rahayu, S., Syah, D., Suhartono, M.T.: Degradation of keratin by keratinase and disulfide reductase from Bacillus sp. MTS of Indonesian origin. Biocatal. Agric. Biotechnol. 1, 152–158 (2012)
Ramnani, P., Gupta, R.: Keratinases vis-à-vis conventional proteases and feather degradation. World J. Microbiol. Biotechnol. 23, 1537–1540 (2007)
Cai, C., Chen, J., Qi, J., Yin, Y., Zheng, X.: Purification and characterization of keratinase from a new Bacillus subtilis strain. J. Zhejiand Univ. Sci. B. 9(9), 713–720 (2008)
Gupta, S., Nigam, A., Singh, R.: Purification and characterization of a Bacillus subtilis keratinase and its prospective application in feed industry. Acta Biol. Szeged. 59(2), 197–204 (2015)
Fakhfakh, N., Ktari, N., Siala, R., Nasri, M.: Wool-waste valorization: production of protein hydrolysate with high antioxidative potential by fermentation with a new keratinolytic bacterium, Bacillus pumilus A1. J. Appl. Microbiol. 115, 424–433 (2013)
Wan, M.Y., Dong, G., Yang, B.Q., Feng, H.: Identification and characterization of a novel antioxidant peptide from feather keratin hydrolysate. Biotechnol. Lett. 38, 643–649 (2016)
Bose, A., Pathan, S., Pathak, K., Keharia, H.: Keratinolytic protease production by Bacillus amyloliquefaciens 6B using feather meal as substrate and application of feather hydrolysate as organic nitrogen input for agricultural soil. Waste Biomass Valorization 5(4), 595–605 (2014)
Pillai, P., Mandge, S., Archana, G.: Statistical optimization of production and tannery applications of a keratinolytic serine protease from Bacillus subtilis P13. Process Biochem. 46, 1110–1117 (2011)
Briki, S., Hamdi, O., Landoulsi, A.: Enzymatic dehairing of goat skins using alkaline protease from Bacillus sp SB12. Protein Expr. Purif. 121, 9–16 (2016)
Balaji, S., Karthikeyan, R., Senthil Kumar, M., Chandra Babu, N.K., Sehgal, P.K.: Microbial degradation of horn meal with Bacillus subtilis and its application in leather processing: a two fold approach. J. Am. Leather Chem. As. 103(3), 89–93 (2008)
Brandelli, A., Sala, L., Kalil, S.J.: Microbial enzymes for bioconversion of poultry waste into added-value products. Food Res. Int. 73, 3–12 (2015)
Ghosh, A., Chakrabarti, K., Chattopadhyay, D.: Degradation of raw feather by a novel high molecular weight extracellular protease from newly isolated Bacillus cereus DCUW. J. Ind. Microbiol. Biotechnol. 35(8), 825–834 (2008)
Mazotto, A.M., De Melo, A.C.N., Macrae, A., Rosado, A.S., Peixoto, R., Cedrola, S.M.L., Couri, S., Zingali, R.B., Villa, A.L.V., Rabinovitch, L., Chaves, J.Q., Vermelho, A.B.: Biodegradation of feather waste by extracellular keratinases and gelatinases from Bacillus spp. World J. Microbiol. Biotechnol. 27, 1355–1365 (2011)
Lo, W.H., Too, J.R., Wu, J.Y.: Production of keratinolytic enzyme by an indigenous feather-degrading strain Bacillus cereus Wu2. J. Biosci. Bioeng. 114(6), 640–647 (2012)
Kurbanoglu, E.B., Algur, O.F.: Single-cell protein production from ram horn hydrolysate by bacteria. Bioresour. Technol. 85, 125–129 (2002)
Adiguzel, A.C., Bitlisli, B.O., Yasa, I., Eriksen, N.T.: Sequential secretion of collagenolytic, elastolytic, and keratinolytic proteases in peptide-limited cultures of two Bacillus cereus strains isolated from wool. J. Appl. Microbiol. 107, 226–234 (2009)
Kotlar, C.E., Ponce, A.G., Roura, S.I.: Characterization of a novel protease from Bacillus cereus and evaluation of an eco-friendly hydrolysis of a brewery byproduct. J. Inst. Brew. 121, 558–565 (2015)
Łaba, W., Rodziewicz, A.: Keratinolytic potential of feather-degrading Bacillus polymyxa and Bacillus cereus. Pol. J. Environ. Stud. 19(2), 371–378 (2010)
Snyder, S.L., Sobociński, P.Z.: An improved 2,4,6-trinitrobenzenesulfonic acid method for the determination of amines. Anal. Biochem. 64, 284–288 (1975)
Henderson, J.W., Ricker, R.D., Bidlingmeyer, B.A., Woodward, C.: Rapid, accurate, sensitive, and reproducible HPLC analysis of amino acids. http://www.chem.agilent.com/Library/chromatograms/59801193.pdf (2000). Accessed Dec 2015
Eslahi, N., Dadashian, F., Nejad, N.H.: Optimization of enzymatic hydrolysis of wool fibers for nanoparticles production using response surface methodology. Adv. Powder Technol. 24, 416–426 (2013)
Sharma, P., Singh, L., Dilbaghi, N.: Optimization of process variables for decolorization of Disperse Yellow 211 by Bacillus subtilis using Box-Behnken design. J. Hazard. Mater. 164, 1024–1029 (2009)
Chen, K.N., Huang, J.C., Chung, C.I., Kuo, W.Y., Chen, M.J.: Identification and characterization of H10 enzymes isolated from Bacillus cereus H10 with keratinolytic and proteolytic activities. World J. Microbiol. Biotechnol. 27, 349–358 (2011)
Mahan, D.C., Shields, R.G.: Essential and nonessential amino acid composition of pigs from birth to 145 kilograms of body weight, and comparison to other studies. J. Anim. Sci. 76(2), 513–521 (1998)
Wang, X., Parsons, C.M.: Effect of processing systems on protein quality of feather meals and hog hair meals. Poult. Sci. 76, 491–496 (1997)
Przybył, A., Madziar, M., Koligot, A.: Suitability of modified pig bristles for extruded feed mixtures for carp fry. Arch. Pol. Fish. 7, 113–122 (1999)
Hertrampf, J.W., Piedad-Pascual, F.: Handbook on ingredients for aquaculture feeds. Kluwer Academic Publishers, Dordrecht (2000)
Garcia, R.A., Pyle, D.J., Piazza, G.J., Wen, Z.: Hydrolysis of animal protein meals for improved utility in non-feed applications. Appl. Eng. Agric. 27(2), 269–275 (2011)
Eslahi, N., Dadashian, F., Nejad, N.H.: An investigation on keratin extraction from wool and feather waste by enzymatic hydrolysis. Prep. Biochem. Biotechnol. 43(7), 624–648 (2013)
Paul, T., Das, A., Mandal, A., Halder, S.K., DasMohapatra, P.K., Pati, B.R., Mondal, K.C.: Valorization of chicken feather waste for concomitant production of keratinase, oligopeptides and essential amino acids under submerged fermentation by Paenibacillus woosongensis TKB2. Waste Biomass Valorization 5, 575–584 (2014)
Aluigi, A., Zoccola, M., Vineis, C., Tonin, C., Ferrero, F., Canetti, M.: Study on the structure and properties of wool keratin regenerated from formic acid. Int. J. Biol. Macromol. 41, 266–273 (2007)
Wojciechowska, E., Włochowicz, A., Wesełucha-Birczyńska, A.: Application of Fourier-transform infrared and Raman spectroscopy to study degradation of the wool fiber keratin. J. Mol. Struct. 511–512, 307–318 (1999)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Łaba, W., Chorążyk, D., Pudło, A. et al. Enzymatic Degradation of Pretreated Pig Bristles with Crude Keratinase of Bacillus cereus PCM 2849. Waste Biomass Valor 8, 527–537 (2017). https://doi.org/10.1007/s12649-016-9603-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12649-016-9603-4