Abstract
Reducing the amount of Helicobacter pylori in the stomach by selective bacterial–bacterial cell interaction was sought as an effective and novel method for combating the stomach pathogen. Lactobacillus reuteri DSM17648 was identified as a highly specific binding antagonist to H. pylori among more than 700 wild-type strains of Lactobacillus species. Applying a stringent screening procedure, the strain DSM17648 was identified as selective binder to H. pylori cells under in vivo gastric conditions. The strain DSM17648 co-aggregates the pathogen in vivo and in vitro. The specific co-aggregation occurs between Lact. reuteri DSM17648 and different H. pylori strains and serotypes, as well as H. heilmannii, but not with Campylobacter jejuni or other commensal oral and intestinal bacteria. Lact. reuteri DSM17648 was shown in a proof-of-concept single-blinded, randomized, placebo-controlled pilot study to significantly reduce the load of H. pylori in healthy yet infected adults. Reducing the amount of H. pylori in the stomach by selective bacterial–bacterial cell interaction might be an effective and novel method for combating the stomach pathogen. Lact. reuteri DSM17648 might prove useful as an adhesion blocker in antibiotic-free H. pylori therapies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Helicobacter pylori is a recognized pathogen and carcinogen causing gastritis, ulcers and gastric cancer. More than 50 % of the world population is infected with this stomach bacterium [1, 2]. Severity of the clinical manifestations of the infection is associated with bacterial load [3–5]. Currently, the only therapeutic option is eradication of the pathogen by a combination of several antibiotics and a proton-pump inhibitor (triple therapy; Maastricht IV/Florence Consensus Report) [6]. Eradication therapy is associated with severe side effects and development of antibiotic resistances [7]. Reducing the amount of H. pylori in the stomach by selective bacterial–bacterial surface interaction represents an alternative method for combating the stomach pathogen. Specific co-aggregation has been widely discussed as a means to return to homeostasis in diseased states [8–10]. While H. pylori resides in the mucus where it is present in its motile form, mucus is constantly produced by the epithelium and shed into the stomach lumen. This continuously releases planktonic H. pylori cells into the stomach. The Lactobacillus strain identified in this study specifically captures such H. pylori cells. As spray-drying or freeze-drying procedures allow the retention of binding activity, structures in the cell wall unaffected by the drying procedures are supposed to be responsible for the aggregation activity.
Previous papers describing the use of Lact. reuteri in H. pylori-related clinical studies show a reduction in H. pylori-associated urease activity by Lact. reuteri strain ATCC55730 [11]. Dore et al. [12] showed that the oral application of Lact. reuteri strain DSM17938, when used in combination with pantoprazole twice a day for 8 weeks, resulted in significant reduction in the urease breath test. Emara et al. [13] used a Lact. reuteri preparation (a mixture of strains Lact. reuteri DSM17938 and Lact. reuteri ATCC PTA6475 in combination with a triple therapy). The Lactobacillus supplementation increased the Gastrointestinal Symptom Rating Scale (GSRS) score significantly, but did not improve the eradication rate.
The aim of the present study was to characterize the binding activity of Lact. reuteri strain DSM17648 to H. pylori in vitro and to determine the impact of 14 days of oral intake of lyophilized Lact. reuteri DSM17648 cells (non-viable) on H. pylori load in a single-blinded, placebo-controlled study.
Materials and Methods
Microorganisms and Cultivation
Lactobacillus strains (strains from ORGANOBALANCE GmbH, Berlin, Germany) were grown in Lactobacillus MRS medium [14] at 37 °C, and H. pylori DSM21031 and Campylobacter jejuni DSM 4688T were grown in Brucella/FBS broth (BD, Heidelberg, Germany, with 10 % (v/v) fetal bovine serum, Biochrom, Berlin, Germany) at 37 °C in microaerobic atmosphere. Other Helicobacter species were grown in Brucella/FBS broth containing additionally 0.75 % (v/v) Vitox supplement (Oxoid, Wesel, Germany) [15].
The taxonomic identification of the Lactobacillus strains to the species level relied on 16 S-rDNA sequence analysis (sequencing done LCG Genomics, taxonomic classification done by Nadicom, Karlsruhe, Germany) using the primers 27f (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492r (5′-ACGGYTACCTTGTTACGACTT-3′) [16] and on phenotypic characterization using the API 50 CH system and apiweb™ software (bioMerieux, France). Bacterial counts were determined from calibration curves of optical density versus microscopic cell counts using a Neubauer chamber (Carl Roth, Karlsruhe, Germany).
Chemicals and Enzymes
Sugars, sugar substitutes and inorganic chemicals were reagent grade (Merck, Darmstadt; Carl Roth, Karlsruhe; Germany), and proteases (protease from Streptomyces griseus, proteinase K from Tritirachium album, trypsin from bovine pancreas and pepsin from bovine pancreas) were of the highest commercially available grade (Sigma, Taufkirchen, Germany).
Screening for Co-aggregates
Co-aggregation was performed with stationary-phase cells of lactobacilli (A600 = 4, in PBS) and H. pylori (A600 = 2, in artificial gastric juice pH 4, 0.3 % (w/v) pepsin, 0.5 % (w/v) sodium chloride [17]). Cells were mixed and immediate flocculation was observed. Co-aggregates could be observed visually as flocking structures, whereas no such structures were present in controls of the single strains (see also Fig. 1). If no aggregates were detected after 10 min, pairs were judged as non-co-aggregating.
a Microscopic analysis of co-aggregation of Lact. reuteri DSM17648 with H. pylori DSM21031 in artificial stomach juice (pH 4), A H. pylori DSM21031 stained with hexidium iodide. B Lact. reuteri DSM17648 stained with CFDA. C Co-aggregate showing clumping of both strains. Bright field fluorescence microscopy (OLYMPUS BX60 microscope, 100-W mercury lamp U-RFL-T, Olympus, Japan), magnification ×1000. b Co-aggregation of Lact. reuteri DSM17648 with H. pylori DSM21031 is macroscopically visible. A H. pylori DSM21031; B H. pylori DSM21031 and Lact. reuteri DSM17648 co-aggregates; C Lact. reuteri DSM17648
For some experiments, cells were stained separately using either hexidium iodide (HI, 10 µg/mL) or carboxyfluorescein diacetate succinimidyl ester (CFDA-SE 1 µg/mL) (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. Excess dye was removed by extensive washing with PBS. Equal amounts of cells were mixed and vortexed for 10 s prior to phase-contrast and fluorescence microscopy [10].
Flow Cytometry
Lactobacillus cells (without staining) and H. pylori cells stained with CFDA-SE were used for co-aggregation by mixing suspensions of the strain DSM17648 and H. pylori strain DSM21031T in a ratio of 1:1 (cell/cell) to a final volume of 100 µL and subsequent shaking for 15 min. The mixture was added to 990 µL 0.5 % (v/v) sodium chloride (pH 4) in FACS tubes (BD, Heidelberg, Germany). A non-co-aggregating Lactobacillus strain was used as a negative control. Samples were analyzed using a flow cytometer (FACSCalibur, BD, Heidelberg, Germany). Cell co-aggregation was quantified by determining events with a high fluorescence intensity (>5 × 102) via channel FL1-H (Ex 488 nm, Em 530/30) [18, 19].
Scanning Electron Microscopy (SEM)
Cells were prepared as described above and re-suspended in PBS. Co-aggregation was induced by mixing suspensions of the strain DSM17648 and H. pylori strain DSM21031T at a ratio of 1:1 (cell/cell). After 20-min incubation at room temperature, the resulting co-aggregates were pelleted by centrifugation (7,150×g, 1 min, Hettich Mikro 22R, Tuttlingen, Germany). The supernatant was carefully discarded, and co-aggregates were either frozen in liquid nitrogen, freeze-dried and sputtered with palladium (1,800× picture) or fixed in 4 % glutaraldehyde, dehydrated in graded ethanol solutions, dried in liquid CO2 and sputtered with palladium (11,000× picture) before SEM. SEM was done using a FEI Quanta 200 FEG Field emission scanning electron microscope. Some images were colorized according to the bacillary or spiral shape to facilitate viewing (eye of science Meckes and Ottawa GbR, Reutlingen, Germany).
Sugar and pH Effects on Co-aggregation
Co-aggregation was tested in the presence of 25 mM sucrose with known co-aggregating pairs. Analogous simultaneous incubations were done testing lactose, glucose, maltose, iso-maltose, fructose or sorbitol to detect possible interference with the ability of the strain DSM17648 to co-aggregate H. pylori. To evaluate the pH dependency of co-aggregation, DSM17648 cells were re-suspended in 0.1 M Sorensen buffer (0.1 M glycine, 0.1 M NaCl) adjusted to pH 2.0 and in McIlvaine citrate–phosphate buffer (0.1 M citrate, 0.2 M disodium phosphate) adjusted to pH values from 3.0 to 8.0, in 1.0 pH unit intervals. H. pylori cells were re-suspended in artificial gastric juice [17]. Co-aggregation was then assessed in a ratio of 1:1 (DSM17648/H. pylori). pH values of final mixtures were controlled. No pure cultures evidenced auto-aggregation within this pH range.
Protease Pre-treatment
The strain DSM17648 and H. pylori were grown separately to stationary phase, harvested by centrifugation, washed in PBS, and 1 mL aliquots adjusted to A600 = 4 (for DSM17648) or A600 = 2 (for H. pylori) in monopotassium phosphate/calcium chloride buffer (pH 7.0) containing either one of four proteases: protease Strep. griseus Type XIV (5.7 U/mg), proteinase K (51 U/mg), trypsin (40 U/mg) or pepsin (2,950 U/mg) at a final concentration of 2.0 mg/mL. After incubation for 1 h at 37 °C, cells were washed, re-suspended again in PBS (pH 7.0), and 500 µL aliquots of each preparation mixed, and co-aggregation assayed visually and microscopically.
Spray Drying and Lyophilization of Cells
Spray drying of cells was done using a Büchi B-191 spray dryer, inlet temperature 140 °C, aspiration 75 % and pump rate 5 % [20]. NaCl [75 % (w/w) final concentration] was used as carrier substance. Before lyophilization, cells were washed, re-suspended in 15 % (w/v) skimmed milk powder and frozen at −80 °C. Lyophilization was done under vacuum (0, 1 mbar) for 24 h [21].
Study Population
The original setting of the study was a placebo-controlled co-twin control design with one twin receiving the active treatment while the co-twin received a placebo. Concordance rates for H. pylori infection in monozygotic twins have been reported at 80 % [22], whereas in dizygotic twins they are 60 %. Heritability for quantitative levels of H. pylori colonization has been estimated at 0.8. Historical prevalence of infection by H. pylori for the general population was reported as 45 %, although more recent studies suggest a reduction to approximately 25 % in the Western world [23, 24]. For Germany, a prevalence of 39 % was reported in 1996 [25]. Based on those figures, the first screening phase was planned to include analysis for 64 twin pairs, expecting 29 pairs with at least one affected twin, and 23 concordant pairs, i.e., pairs with positive findings for both twins. As incidence rates found in the screening phase were lower than expected from published figures, the original design was then adapted to include singletons in a pre–post design. A second screening phase included twins as well as singletons. Subjects were included if they had reached the age of 18 and had a positive H. pylori finding in the 13C urea breath test (Helicobacter Test INFAI®, Dd ≥ 4 ‰). Informed consent was obtained from all persons for being included in the study. Additional informed consent was obtained from all patients for which identifying information is included in this article. Exclusion criteria were any medication interfering with the action of the lactobacilli, previous surgical procedures affecting stomach or small intestine with potential interference with the study, e.g., gastrectomy or gastric bypass, diabetes type 1 or 2, familiar lipid metabolism diseases, any other major disease, weight changes >3 kg over the last 3 months, pregnancy or lactation, alcohol or drug abuse, or psychiatric diseases.
Study Protocol
The study was approved by the local ethics advisory committee (Charité, Berlin, Germany) and was conducted according to the Declaration of Helsinki [26]. As the trial was not a clinical trial, the trial was not registered, as at the time of the trial in Germany it was not customary for pilot type trials to be registered.
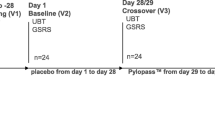
The test product (active ingredient) consisted of lyophilized dead cells of the strain DSM17648, prepared as solid tablets for oral application. Each tablet contained 5 × 109 cells (determined by counting in Neubauer chamber), and the daily dosage of four tablets translates into 2 × 1010 cells. Verum and placebo tablets were identical in weight (250 mg), size, color and flavor. Within concordant affected twin pairs, treatment was randomized in parallel for a time period of 14 days. In singletons, active treatment and placebo were given in a single-blinded non-randomized crossover design. The first period of 14 days was the placebo phase; after a second breath test, active treatment was given for another 14 days, followed by a breath test. Four to six weeks after the treatment phase, a follow-up breath test was conducted. Subjects were instructed to take two tablets after breakfast as well as after their evening meal. During the treatment phase, no lifestyle or dietary changes were to be initiated and no probiotic food products or cranberries were to be used. Subjects were asked to fill in a study-specific questionnaire to document well-being, any potential side effects, smoking, alcohol use, nutrition and medication.
Measurements
Detection of H. pylori infection in the screening phase and quantification of colonization to verify effects of the strain DSM17648 were accomplished by a breath test, as this diagnostic approach is best suited to screening as well as detection of intra-individual changes [27]. Helicobacter Test INFAI® is a breath test for direct noninvasive quantitative detection of the bacterium H. pylori [28]. The test is based on urease activity of H. pylori. Specificity (98.5 %) and sensitivity (97.9 %) of Helicobacter Test INFAI® are comparable to traditional invasive diagnostic methods (endoscopy or biopsy). As the breath test reflects the current status of colonization by H. pylori, it is well suited to detect reduction in or eradication of the bacteria [29, 30].
The test is based on the hydrolysis of 13C urea to ammonium and 13C-enriched carbon dioxide, which is detectable in the breath. Patients ingest a small amount of the 13C urea isotope. Carbon dioxide resulting from the degradation of urea contains this isotope, detectable by mass spectrometry. As there is a small amount of naturally occurring 13C even in the absence of urease activity, breath samples are taken before and after the ingestion of 13C urea. If there is no difference, the test is negative, indicating no infection with H. pylori. There is a quantitative relation between urease activity and amount of 13C in breath that indirectly relates to the level of colonization by H. pylori.
Statistics
All historical and clinical data were entered into a dedicated trial database. Statistical analysis was conducted using SPSS version 16.0.2. We computed differences in 13C urea breath test (UBT) values against initial measurements: ΔActive = 13C UBT Active − 13C UBT Initial, ΔPlacebo = 13C UBT Placebo − 13C UBT Initial, ΔWash-out = 13C UBT Wash-out − 13C UBT Initial. Additionally, the absolute test values between the various study time-points were compared: 13C UBT Initial, 13C UBT Verum (after 14-day verum treatment), 13C UBT Placebo (after 14-day placebo treatment), 13C UBT Wash-out (4–6 weeks after verum treatment). Data from twin pairs were combined and analyzed as in singletons. The co-twin control design is comparable to a crossover design, but without any potential carry over effects. There was no randomized order for verum/placebo treatment, as no continuing placebo effect was expected.
All data were tested for deviations from normal distribution by Kolmogorov–Smirnov test. Mean differences were computed by pairwise t test. Potential relations between response to treatment and initial level of colonization were explored by linear regression. An error level of 5 % was set as threshold for significance. Results are reported as mean ± standard deviation (SD); figures present the standard error of the mean (SEM).
Results
Co-aggregation Analysis of Lact. reuteri DSM17648
Lactobacilli that co-aggregate H. pylori were sought among a large Lactobacillus strain collection. The in-house, private strain collection has been assembled from wild-type strains of diverse origin, such as food sources, plants, vegetables or human skin. Strains are classified according to the physiological characteristics prior to being included in the screening process. Among 700 Lactobacillus strains tested, only eight were found to co-aggregate with spiral forms of H. pylori strain DSM21031, without exhibiting any auto-aggregation (Fig. 1). Three of the co-aggregating lactobacilli (strains DSM17648, DSM17647 and DSM17651) were identified as Lact. fermentum (API method). One of these—Lact. fermentum DSM17648 [classified as Lact. reuteri by 16 S-rDNA sequencing and sequence alignment (100 % identical to accession numbers CP000705, CP006603, CP006011 (at 99 % coverage))]—was analyzed in depth (Table 1). Numerous other Lact. fermentum and Lact. reuteri strains were tested in parallel for auto-aggregation and co-aggregation under the described conditions. None of them formed co-aggregates with H. pylori. Lactobacillus and Helicobacter strains did not auto-aggregate (Fig. 1). To confirm that both species were present in the aggregates, cells were stained separately using either hexidium iodide or carboxyfluorescein diacetate succinimidyl ester. Both the strain DSM17648 and H. pylori DSM21031 participated in the aggregation (Fig. 1). Co-aggregation occurs within seconds after mixing the strains. Quantification of co-aggregate formation between Lact. reuteri DSM17648 and H. pylori DSM21031 by flow cytometry (Fig. 2) shows that one Lactobacillus cell binds 2–3 Helicobacter cells. Interestingly, the co-aggregation activity is preserved during lyophilization or spray drying of whole cells of Lact. reuteri DSM17648 and persists in non-viable cells (Table 3). Spray-dried or lyophilized cells of strain DSM17648 induced co-aggregate formation with the same sensitivity as untreated cells. Expression of co-aggregation activity is dependent on the growth phase of Lact. reuteri DSM17648, and it is present at entry into stationary growth and during stationary phase. SEM images of co-aggregates were prepared to analyze cellular sites of the attachment. Figure 3 shows that single cells of the strain DSM17648 bind several H. pylori cells resulting in cross-linking of the co-aggregates. Binding sites on the cells of the strain DSM17648 appear evenly distributed over the cell surface, and binding sites on H. pylori cells do not seem to be present on flagellar structures.
Co-aggregation of Lact. reuteri DSM17648 with H. pylori DSM21031 was analyzed by flow cytometry (e). H. pylori cells were CFDA stained. Samples were analyzed using flow cytometry, and cell co-aggregation was quantified by determining the events with a high FL (>5 × 102, area within green frame). Co-aggregation was not observed when strains were analyzed separately (a–c) nor when a non-aggregating Lactobacillus strain was used as a control (d)
As the interaction between cells of Lact. reuteri DSM17648 and H. pylori DSM21031 involves cell surfaces, we tested for possible interference by surface modulating treatments. Co-aggregation persists in the presence of sugar (sucrose, lactose, glucose, fructose, maltose, isomaltose, and sorbitol). It occurs at comparable efficiency at room temperature and at 37 °C, and co-aggregation activity is observed over a wide pH range (pH 2.0—corresponding to empty stomach conditions—up to pH 8, including typical pH values after meals). No pure cultures evidenced auto-aggregation within this pH range. Slightly smaller co-aggregates formed at pH 2 compared with pH 8 in vitro. Thus, aggregation of H. pylori by Lact. reuteri DSM17648 occurs at pH values and conditions encountered in the human stomach.
The susceptibility to protease inactivation of the co-aggregation determinants on the surfaces of both the strain DSM17648 and H. pylori DSM21031 was tested after treatment with protease Strep. griseus Type XIV, proteinase K, trypsin or pepsin. Incubating Lact. reuteri DSM17648 with any protease before co-aggregation reduced binding to H. pylori DSM21031 by 30 %, but did not eliminate it completely. H. pylori required pretreatment with the protease pepsin (as is naturally present in gastric fluids) to be fully active in co-aggregation with the strain DSM17648.
Lactobacillus reuteri DSM17648 does not co-aggregate with common non-Helicobacter members of the human flora. Neither the major intestinal commensals nor C. jejuni detectably co-aggregate with Lact. reuteri DSM17648 (Table 2), and no auto-aggregation was observed. Co-aggregation is active with different H. pylori strains (type I and type II strains) as well as with H. heilmannii strains (type I and type II) and with H. canis of animal source. Thus, the strain DSM17648 specifically co-aggregates H. pylori without interfering with other bacteria of the commensal intestinal flora.
Pilot Study
The strain DSM17648 was used in a placebo-controlled pilot study to evaluate the effect of the strain DSM17648 in asymptomatic Helicobacter-positive test persons after a two-week application. Screening included 128 subjects, 47 twin pairs and 34 singletons; 27 subjects had a positive breath test result. Overall Helicobacter prevalence was 21 %; 6 twin pairs were concordant and 10 pairs discordant positive. Fourteen independent treatments were started with no dropouts during the trial phase. All 6 concordant twin pairs participated in the study, as well as 4 discordant twin pairs and 4 singletons. Due to the large inter-individual variability of quantitative measures of colonization (13C UBT Initial), analysis of H. pylori reduction by the strain DSM17648 was primarily based on intra-individual changes after active treatment or placebo (Δverum vs. Δplacebo). Treatment by placebo did not result in a significant change in 13C UBT (Δplacebo −0.6 ± 5.3), whereas verum treatment significantly reduced 13C UBT values (Δverum −4.9 ± 7.8, p = 0.026 vs. placebo), indicating significant reduction in H. pylori. Absolute values of 13C UBT at baseline measurement, after placebo and after verum treatment were 14.1 ± 9.9, 12.7 ± 7.2 (ns vs. initial) and 11.9 ± 5.9 (p vs. initial 0.01, p vs. placebo 0.03), respectively. To allow for a detailed evaluation of response to the strain DSM17648, individual values for 13C UBT are plotted in Fig. 4. After verum treatment, the majority of subjects showed a reduction in H. pylori colonization. Responses showed some variability, from no reduction to a delta of more than 20. In comparison, after 2 weeks of placebo, some subjects had lowered values while others had increases in the same magnitude, indicating no systematic effect.
Values of 13C UBT after wash-out (x ± y) are not significantly different from verum treatment values. The effect of reduced Helicobacter values lasts over the actual treatment. There was some dependency of treatment response on initial values (r 2 = 0.66, p = 0.01, Fig. 5). With increasing level of colonization, the lowering effect caused by the strain DSM17648 becomes stronger. For placebo treatment, the same dependency was found albeit to a lesser degree (r 2 = 0.35, p = 0.02), probably reflecting regression to the mean effects. A direct placebo effect on immune response cannot be ruled out. This potential effect is significantly lower than the specific action of the strain DSM17648. During the course of the study, there was neither any change in life style, e.g., in physical activities or diet, nor health as indicated by the questionnaire. No side effects were reported in either study group.
Discussion
The specific and fast co-aggregation of defined Lactobacillus strains of the species Lact. reuteri/fermentum, Lact. brevis and Lact. pentosus (typified Lact. reuteri DSM17648) with H. pylori under gastric conditions is novel. Co-aggregation of Lact. reuteri DSM17648 also occurs with H. heilmannii and H. canis. Major commensals in the intestinal environment are not co-aggregated by Lact. reuteri DSM17648. Numerous other lactobacilli in a very large collection of Lactobacillus wild-type strains (collected from various natural habitats) including many other Lact. fermentum, Lact. brevis and Lact. pentosus strains fail to manifest H. pylori-specific co-aggregation. Co-aggregation activity is not affected by sugars and pH (over a wide range between 2 and 8), and it requires a pepsin pretreatment of spiral form cells of H. pylori. It does not occur with far less infective coccoidal H. pylori cells [31], and it is dependent on a Lact. reuteri DSM17648 cell surface factor that is present at the end of the exponential growth phase and during stationary phase. Co-aggregation between H. pylori and the strain DSM17648 occurs within seconds. While Chen et al. [32] observed a very slow interaction between some lactobacilli from food source, the present paper is to our knowledge the first description of a rapid and efficient co-aggregation of H. pylori by a specific Lactobacillus strain under gastric conditions. It is proposed that Lact. reuteri DSM17648 interferes with mobility of H. pylori and its adherence to the gastric mucosa by entangling the cells into aggregates and masking H. pylori surface sites that are ordinarily available for binding to human epithelium. Once bound, co-aggregates will be flushed from the stomach by natural bowel movement. Interestingly, the aggregation activity was preserved when the cells were killed by freeze drying or spray drying. It can be assumed that binding is due to specific surface molecules on the Lact. reuteri DSM17648 cells which are strain specific and are resistant to such process steps. Such surface molecules might include lipoteichoic acid and carbohydrate structures.
This novel anti-H. pylori activity has not been described previously as a mode of action for probiotic treatment of H. pylori infections. This hypothesis was tested in a proof-of-concept in vivo study. Our data demonstrate the significant decrease in H. pylori load by a two-week application of Lact. reuteri DSM17648 in healthy subjects with detectable H. pylori infection in a general population sample. The principal outcome criterion was the reduction in H. pylori as measured by 13C urease breath test (Helicobacter Test INFAI®) after a 14-day supplementation period of Lact. reuteri DSM17648 at a daily dose of 2 × 1010 non-viable lyophilized cells. Data obtained in a parallel clinical study support the data reported in this paper [33]. Previous studies with non-specific probiotics require the application of live microorganisms while Lact. reuteri DSM17648 is active as non-viable cell preparation. This will greatly reduce any potential side effects and will help ensure stable activity in a potential consumer product and in pharmaceutical and medicinal formulations.
Our study reveals a novel Lact. reuteri strain (the strain DSM17648) that features unique properties as it specifically aggregates with planktonic H. pylori in the stomach. Freeze-dried (and spray-dried) preparations significantly reduce the H. pylori load (measured by urease breath test) after a 14-day oral treatment period in H. pylori-positive test persons.
Lact. reuteri strain DSM17648 can become a central part of a strategy to avoid using antibiotics and combating antibiotic resistances in H. pylori infections, in reducing H. pylori load, either as a prophylactic food additive or a medical cure to treat H. pylori-induced stomach diseases.
References
Suerbaum S, Michetti P (2002) Helicobacter pylori infection. N Engl J Med 347:1175–1186
Lopes D, Nunes C, Martins MCL, Sarmento B, Reis S (2014) Eradication of Helicobacter pylori: past, present and future. J Control Release 189:169–186
Kusters JG, van Vliet AHM, Kuipers EJ (2006) Pathogenesis of Helicobacter pylori infection. Clin Microbiol Rev 19:449–490
Varbanova M, Malfertheiner P (2011) Bacterial load and degree of gastric mucosal inflammation in Helicobacter pylori infection. Dig Dis 29:592–599
Tokunaga Y, Hata K, Ryo J, Kitaoka A, Tokuka A, Ohsumi K (1998) Density of Helicobacter pylori infection in patients with peptic ulcer perforation. J Am Coll Surg 186:659–663
Malfertheiner P, Megraud F, O’Morain CA, Atherton J, Axon AT, Bazzoli F, Gensini GF, Gisbert JP, Graham DY, Rokkas T, El-Omar EM, Kuipers EJ, Kuipers EJ (2012) Management of Helicobacter pylori infection—the Maastricht IV/Florence consensus report. Gut 61:646–664
Wu TS, Hu HM, Kuo FC, Kuo CH (2014) Eradication of Helicobacter pylori infection. Kaohsiung J Med Sci 30:167–172
Reid G, McGroarty JA, Domingue PAG, Chow AW, Bruce AW, Eisen A, Costerton JW (1990) Coaggregation of urogenital bacteria in vitro and in vivo. Curr Microbiol 20:47–52
Younes JA, van der Mei HC, van den Heuvel E, Busscher HJ, Reid G (2012) Adhesion forces and coaggregation between vaginal staphylococci and lactobacilli. PLoS One 7:e36917
Lang C, Böttner M, Holz C, Veen M, Ryser M, Reindl A, Pompejus M, Tanzer JM (2010) Specific Lactobacillus/Mutans Streptococcus co-aggregation. J Dent Res 89:175–179
Francavilla R, Lionetti E, Castellaneta SP, Magista AM, Maurogiovanni G, Bucci N, De Canio A, Indrio F, Cavallo L, Ierardi E, Miniello VL (2008) Inhibition of Helicobacter pylori infection in humans by Lactobacillus reuteri ATCC 55730 and effect on eradication therapy: a pilot study. Helicobacter 13:127–134
Dore MP, Cuccu M, Pes GM, Manca A, Graham DY (2014) Lactobacillus reuteri in the treatment of Helicobacter pylori infection. Intern Emerg Med 9:649–654
Emara MH, Mohamed SY, Abdel-Aziz HR (2014) Lactobacillus reuteri in management of Helicobacter pylori infection in dyspeptic patients: a double-blind placebo-controlled randomized clinical trial. Therap Adv Gastroenterol 7:4–13
De Man JC, Rogosa M, Sharpe ME (1960) A medium for the cultivation of lactobacilli. J Appl Bacteriol 23:130–135
Whitmire JM, Merrell DS (2012) Successful culture techniques for Helicobacter species: general culture techniques for Helicobacter pylori. Methods Mol Biol 921:17–27
Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA, Olsen GJ (2008) Critical evaluation of two primers commonly used for amplification of bacterial 16 S rRNA genes. Appl Environ Microbiol 74:2461–2470
Charteris WP, Kelly PM, Morelli L, Collins JK (1998) Development and application of an in vitro methodology to determine the transit tolerance of potentially probiotic Lactobacillus and Bifidobacterium species in the upper human gastrointestinal tract. J Appl Microbiol 84:759–768
da Silva TL, Reis A, Kent CA, Roseiro JC, Hewitt CJ (2005) The use of multi-parameter flow cytometry to study the impact of limiting substrate, agitation intensity, and dilution rate on cell aggregation during Bacillus licheniformis CCMI 1034 aerobic continuous culture fermentations. Biotechnol Bioeng 92:568–578
Geng J, Beloin C, Ghigo J-M, Henry N (2014) Bacteria hold their breath upon surface contact as shown in a strain of Escherichia coli, using dispersed surfaces and flow cytometry analysis. PLoS One 9:e102049
To BCS, Etzel MR (1997) Spray drying, freeze drying, or freezing of three different lactic acid bacteria species. J Food Sci 62:576–578
Desmond C, Stanton C, Fitzgerald GF, Collins K, Ross RP (2001) Environmental adaptation of probiotic lactobacilli towards improvement of performance during spray drying. Int Dairy J 11:801–808
Malaty HM, Engstrand L, Pedersen NL, Graham DY (1994) Helicobacter pylori infection: genetic and environmental influences. A study of twins. Ann Intern Med 120:982–986
Stettin D, Waldmann A, Wolters M, Trunz B, Schauder P, Hahn A (2007) Infection with Helicobacter pylori—outcome of a cross-sectional investigation. Dtsch Med Wochenschr 132:2677–2682
Fischbach W, Malfertheiner P, Hoffmann JC, Bolten W, Bornschein J, Götze O, Höhne W, Kist M, Koletzko S, Labenz J, Layer P, Miehlke S, Morgner A, Peitz U, Preiss JC, Prinz C, Rosien U, Schmidt WE, Schwarzer A, Suerbaum S, Timmer A, Treiber G, Vieth M (2009) S3-Leitlinie “Helicobacter pylori und gastroduodenale Ulkuskrankheit”. Z Gastroenterol 47:68–102
Breuer T, Sudhop T, Hoch J, Sauerbruch T, Malfertheiner P (1996) Prevalence of and risk factors for Helicobacter pylori infection in the western part of Germany. Eur J Gastroenterol Hepatol 8:47–52
World Medical Association (2013) Declaration of Helsinki, ethical principles for medical research involving human subjects. JAMA 310:2191–2194
Calvet X, Lehours P, Lario S, Mégraud F (2010) Diagnosis of Helicobacter pylori infection. Helicobacter 15(Suppl 1):7–13
Labenz J, Stolte M, Aygen M, Hennemann O, Bertrams J, Borsch G (1993) Qualitative und semiquantitative invasive und nichtinvasive diagnosik der Helicobacter pylori kolonization der gastralen mukosa. Z Gastroenterol 31:437–443
Peng NJ, Lai KH, Lo GH, Hsu PI (2009) Comparison of noninvasive diagnostic tests for Helicobacter pylori infection. Med Princ Pract 18:57–61
Debongnie JC, Pauwels S, Raat A, de Meeus Y, Haot J, Mainguer P (1991) Quantification of Helicobacter pylori infection in gastritis and ulcer disease using a simple and rapid carbon-14-urea breath test. J Nucl Med 32:1192–1198
Cole SP, Cirillo D, Kagnoff MF, Guiney DG, Eckmann L (1997) Coccoid and spiral Helicobacter pylori differ in their abilities to adhere to gastric epithelial cells and induce interleukin-8 secretion. Infect Immun 65:843–846
Chen X, Tian F, Liu X, Zhao J, Zhang HP, Zhang H, Chen W (2010) In vitro screening of lactobacilli with antagonistic activity against Helicobacter pylori from traditionally fermented foods. J Dairy Sci 93:5627–5634
Mehling H, Busjahn A (2013) Non-viable Lactobacillus reuteri DSMZ17648 (Pylopass™) as a new approach to Helicobacter pylori control in humans. Nutrients 5:3062–3073
Acknowledgments
The authors acknowledge the excellent technical assistance of Christina Balcke and Melanie Just.
Conflict of Interest
Caterina Holz, Andreas Busjahn, Heidrun Mehling, Stefanie Arya, Mewes Boettner, Hajar Habibi declare that they have no conflict of interest. Christine Lang owns stock in Organobalance GmbH.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Holz, C., Busjahn, A., Mehling, H. et al. Significant Reduction in Helicobacter pylori Load in Humans with Non-viable Lactobacillus reuteri DSM17648: A Pilot Study. Probiotics & Antimicro. Prot. 7, 91–100 (2015). https://doi.org/10.1007/s12602-014-9181-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-014-9181-3