Abstract
Environmental DNA (eDNA) consists of DNA fragments shed from organisms into the environment, and can be used to identify species presence and abundance. This study aimed to reveal the dispersion and degradation processes of eDNA in the sea. Caged fish were set off the end of a pier in Maizuru Bay, the Sea of Japan, and their eDNA was traced at sampling stations located at the cage and 10, 30, 100, 300, 600 and 1000 m distances from the cage along two transect lines. Sea surface water was collected at each station at 0, 2, 4, 8, 24 and 48 h after setting the cage, and again after removing the cage. Quantitative PCR analyses using a species-specific primer and probe set revealed that the target DNA was detectable while the cage was present and for up to 1 h after removing the cage, but not at 2 h or later. Among the 57 amplified samples, 45 (79%) were collected within 30 m from the cage. These results suggest that eDNA can provide a snapshot of organisms present in a coastal marine environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Efficient management of fisheries resources requires prompt and precise evaluation of the spatial distribution and abundance of target species. Traditional methods for this purpose, such as net-towing, visual census, and acoustic survey, are generally effort-demanding (Murphy and Jenkins 2010). Environmental DNA (eDNA) technology can serve to evaluate the presence and abundance of aquatic animals or plants by sampling environmental water and detecting DNA fragments released from organisms. The source of eDNA can be epidermis, mucus, feces, or urine of species (Turner et al. 2014; Klymus et al. 2015; Wilcox et al. 2015). eDNA is typically detected by quantitative PCR (qPCR), using a set of species-specific primers and probe. Typical eDNA surveys can be performed by collecting only a liter or less of water (Ficetola et al. 2008; Takahara et al. 2012; Thomsen et al. 2012a, b). Advances in such methods are expected to promote less costly and non-invasive monitoring of the ecosystem for conservation and biomass estimation. Thus, eDNA technology has a great potential for the management of aquatic resources.
eDNA was first applied to vertebrates to detect the invasive American bullfrog Rana catesbeiana in Europe (Ficetola et al. 2008). Since then, it has been used to identify habitats of rare species (Thomsen et al. 2012b; Pilliod et al. 2013; Rees et al. 2014; Fukumoto et al. 2015) and to indicate other introduced species’ invasions (Takahara et al. 2013; Piaggio et al. 2014). Recent studies have shown a positive relationship between eDNA concentration and biomass in experimental surveys, e.g., Takahara et al. (2012), and field surveys in freshwater (e.g., Thomsen et al. 2012b; Pilliod et al. 2013; Eichmiller et al. 2014; Wilcox et al. 2016; Doi et al. 2017). eDNA studies have also been conducted in the marine environment (Thomsen et al. 2012a; Valentini et al. 2016; Kelly et al. 2014). Yamamoto et al. (2016) demonstrated that eDNA concentration can be used to assess the biomass of jack mackerel Trachurus japonicus, with findings confirmed using sonar. eDNA metabarcoding using universal primers enables fish fauna to be comprehensively and simultaneously surveyed (Miya et al. 2015). Yamamoto et al. (2017) detected 128 fish species by eDNA metabarcoding of water samples collected during a 1-day cruise, including 40 species observed by underwater visual census conducted in this area during the preceding 14 years. In experimental studies, the sequence reads of fish and amphibians in eDNA metabarcoding correlated with their biomass (Kelly et al. 2014; Evans et al. 2016). In the sea, the species detected by eDNA metabarcoding matched trawling catch data, and eDNA sequence read numbers were correlated with abundance data (Thomsen et al. 2016). Quantification in eDNA metabarcoding has recently been established by simultaneously using artificial DNA internal standards with known concentration (Ushio et al. 2018). As such, eDNA is being developed as a suitable tool across many fields for estimating both biomass and species diversity.
eDNA released by marine or freshwater organisms does not remain at the site of its release, but is dispersed and degraded over time. Further, information as to the timing and source of eDNA release cannot be resolved based on the eDNA itself. Thus, a firm understanding of eDNA dynamics in nature constitutes essential information for estimating distribution and abundance of aquatic organisms. Dispersion or degradation of eDNA in the natural environment has been reported in rivers (Deiner and Altermatt 2014; Jane et al. 2015; Sansom and Sassoubre 2017), but there is limited information for the marine environment (Yamamoto et al. 2016; O’Donnell et al. 2017). Owing to the multi-directional currents present in oceanic habitats, the processes of eDNA dispersion and degradation may be more complicated than those in rivers or ponds.
The present study aimed to show the spatiotemporal scale of dispersion and degradation of eDNA in a coastal marine system, using caged live striped jack Pseudocaranx dentex as a known source of eDNA in Maizuru Bay, the Sea of Japan. This species was selected as it was presumed to be absent in this area based on results of long-term underwater visual census and eDNA metabarcoding in this area (Masuda 2008; Yamamoto et al. 2017). A cage containing live fish was set off the end of a pier, and eDNA dispersion and degradation were periodically estimated with spatiotemporal monitoring.
Materials and methods
Field experiment
Hatchery-reared striped jack juveniles were purchased from the Aquaculture Research Institute of Kindai University and transported to Maizuru Fisheries Research Station (MFRS), Kyoto University on June 5, 2015. Forty-nine fish with a cumulative biomass of 426.9 g were stocked in a covered mesh cage (1.15 × 1.15 × 1.15 m, 7 mm mesh size) in a 500-l black polyethylene tank. The tank water was recirculated using four sets of aerators equipped with filtering materials (Roka Boy M, GEX, Osaka, Japan). Fish were fed with pellets (Otohime, Marubeni Nisshin Feed Co., Ltd., Tokyo, Japan) beginning 2 days after stocking.
On June 9, 2015, a cage was set ~ 1 m from the surface at the tip of a 30-m floating pier at the research station (35°28′N, 135°22′E). The area is in a protected inner bay. The sea bottom is muddy silt with a small number of artificial structures such as discarded concrete blocks. The water depth where the cage was set is ~ 8 m. The fish were not fed during the experiment. Sampling stations were set above the cage ~ 1 m from the school of fish and at six linear distant points extending northwest and northeast (six in each direction; Fig. 1). These stations were named St. 1, W10–W1000 and E10–E1000, where “W” or “E” and the number of each station name represent northwest or northeast and distance in meters from the cage, respectively. We took a course of first going to W1000, then along the transect line to W10, E10, E1000 and then back to the port. Care was taken not to run the boat across sampling stations except during sampling. Sampling at St. 1 was conducted directly from the floating pier so as not to disturb fish when the boat was between W30 and W10. Two-liter samples of sea surface water were collected using a ladle (1.4 m in length, 3.2 l in capacity) in triplicate at each station at roughly 0, 1, 2, 4, 8, 24 and 48 h after setting the cage, as well as 40 min prior at St. 1 as a control. It took 22–34 min to visit all the stations and conduct water sampling. Therefore, the actual timing of each sampling was on average ~ 14 min after the above-mentioned ‘hours’ after setting the cage. These samples were placed in a cooling box on ice to minimize the degradation of eDNA until filtration was performed.
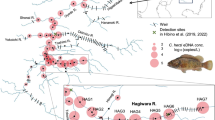
Location of caged striped jack (St. 1) and water sampling stations along the northwest and northeast transect lines. Two triangles represent where current flow meters and thermometers were deployed. A thermometer was also set at St. 1. Circle size indicates total eDNA concentration over all time points
The cage was removed from the sea after 48 h, and sampling was repeated at 0, 1, 2, 4, 8 and 24 h after this point. Sampling was also conducted at St. 1, 48 h after removing the cage. Current flow meters (COMPACT-EM, JFE Advantech, Hyogo, Japan) were set NW and NE of the cage, and conductivity-temperature meters (INFINITY-CT, JFE Advantech, Hyogo, Japan) were placed NW, NE, and at St. 1 at a depth of 2 m from the surface (Fig. 1).
Vertical dispersion of eDNA was also examined to confirm the validity of sampling surface water in studying horizontal dispersion. Triplicate water samples were collected using a van Dorn sampler at a distance of ~ 1 m from the edge of the cage, at the sea surface, midwater (4 m) and bottom (8 m) at 29, 32 and 37 h after setting the cage. The timing of these samplings did not overlap with those of horizontal dispersion.
Filtration and DNA extraction from water samples
Filtration was conducted in the order of water sampling. We started to filter samples within ~ 40 min (30 min on board and 10 min for transportation to the laboratory) after collection using 47-mm-diameter glass microfiber filters (nominal pore size 0.7 µm, GE Healthcare Life Science, Whatman) through an aspirator. It took ~ 30 min to complete filtration per one sampling time point. Filters containing trapped eDNA were folded, with the surface in contact with the sample facing inward, then wrapped with aluminum foil and preserved at − 20 °C. Samples from northwest and northeast transects were filtered in separate laboratories to avoid cross-contamination, with each transect provided to the opposite lab at each time point to negate handling bias. All instruments used for filtration were chlorinated using 0.1% hypochlorous acid to degenerate remnant DNA and were rinsed with tap water and distilled water prior to the next filtration step. The amount of water filtered was 1 litre for each sample. The same amount of artificial sea water (MARIN MERIT, Matsuda, Osaka, Japan) was filtered once per series of samples in each transect as a negative control. DNA was extracted according to the procedure detailed in Yamamoto et al. (2016).
Species-specific primers and probe for striped jack
Sequences of mitochondrial cytochrome b (cyt b) were obtained from the National Center for Biotechnology Information (NCBI) for Pseudocaranx dentex (Sequence ID: EF392607) and five other carangid fishes found in this area, including: Trachurus japonicus (AP003091.1), Decapterus maruadsi (EF512291.1), Seriola quinqueradiata (AB263290.1), Seriola dumerili (EF439584.1), and Kaiwarinus equula (KM201334.1). We searched for an appropriate primer binding site of five base pairs with a pair unique to striped jack at the 3′-end, and designed a primer targeting this region (Fig. 2). The sequences of primer pairs and probe were as follows: 5′-ACGCAGTTTTTATTCTGGC-3′ for the forward primer, 5′-CTAGGAAGTATAGGGC-3′ for the reverse primer, and 5′-FAM-CGTGGCTATCCTCACCTGAATCGGG–TAMRA-3′ for the probe. The PCR product size was 130 bp. Only striped jack DNA was amplified when qPCR was conducted using DNA extracted from tissue at levels of 10 and 100 pg per reaction with the above six species as template. Real-Time TaqMan® PCR (Life Technologies) was performed to measure the copy number of striped jack cytochrome b using a LightCycler 96 System (Roche Diagnostics, Mannheim, Germany). The total volume of qPCR reaction solution was 20 µl, including 2 µl DNA extract, 900 nM of each primer (final concentration), 125 nM TaqMan probe (final concentration), and 10 µl TaqMan master mix (TaqMan Gene Expression Master Mix, Life Technologies). PCR was performed according to the following thermal cycling: 2 min at 50 °C and 10 min at 95 °C, followed by 55 cycles of 15 s at 95 °C and 1 min at 60 °C. qPCR was performed with both eDNA samples and the artificial DNA fragments as a dilution series containing 3 × 101 to 3 × 104 copies. The artificial DNA of striped jack cytochrome b was pUC57 plasmids containing 330 bp of partial cyt b gene, or 130 bp of the PCR target and 100 bp of upstream and downstream regions. Prior to qPCR analysis, the plasmid was digested with a restriction enzyme (Hind III). qPCR was performed in triplicate for each eDNA sample as well as the artificial DNA. In all the runs, R2 values of the calibration curve were greater than 0.99, with the slope range between − 4.190 and − 3.350 and an intercept range between 39.66 and 43.36. In addition, a 2-µl pure water sample was analyzed simultaneously in triplicate as a PCR negative control. Different laboratory rooms were used for DNA extraction and PCR to prevent cross-contamination. The average of qPCR triplicates (including the template with 0 copies) was taken to represent eDNA concentration (number of copies per liter) in each sample.
To confirm the specificity of the primer and probe design, the sequence of qPCR products that showed amplification by qPCR was analyzed and BLAST searching was conducted using the NCBI nucleotide database. In the qPCR, 64 water samples showed amplification. Sequencing revealed that 61 samples that were successful in the read out of the total 64 samples matched with the striped jack cyt b gene sequence (Sequence ID: EF392607.1). Therefore, we assumed that the designed primers and probe set were specific for striped jack.
Statistical analyses
Prior to statistical analysis, eDNA concentration was log10 (x + 1) transformed to improve homogeneity of variance. Since the vertical dispersion experiment showed that most eDNA can be assumed to remain near the surface and disperse horizontally, we applied a power regression between the distance from the cage and eDNA concentration to evaluate the dispersion. A coefficient of determination was obtained for each time point. We compared eDNA concentrations among sampling stations along the northwest or northeast transect lines using a nested ANOVA, with nesting after setting or removing the cage and time points, followed by Tukey’s HSD test. eDNA concentration was compared among time points after setting or removing the cage by nested ANOVA, with nesting in sampling stations, followed by Tukey’s HSD test. Then, we calculated the detectability of eDNA concentration at each sampling station after removing the cage as percent eDNA remaining per hour, using the average of eDNA concentration at 0 and 1 h time points after removing the cage, as follows:
where A is the average eDNA concentration 1 h after removing the cage, B is the average eDNA concentration immediately after removing the cage, and Dtime is detectability (%/h). Detectability of eDNA over distance was calculated using data between St. 1 and W10 or W30, and St. 1 and E10 or E30 at all time points after setting the cage, as follows:
where A is the average eDNA concentration at each station, B is the average eDNA concentration at St. 1, and Dst is detectability in distance, presented as a percentage.
We also compared eDNA concentrations at three different depths (0, 4, and 8 m) at St. 1 using ANOVA followed by Tukey’s HSD test. R statistical software version 3.2.0 (2015.04.16; R Core Team 2015) was used for these analyses.
Results
Striped jack DNA was not detected in samples collected at St. 1 before setting the cage, implying that neither wild nor anthropogenically originated striped jack DNA was present. No striped jack DNA was detected in any negative controls during filtration or PCR, indicating that there was no contamination during the experiment.
The concentration of eDNA gradually decreased with distance from the cage (Fig. 3). Amplified samples were confirmed in 39 of the 273 (13 stations × 7 time points × 3 replicates) samples collected at 0–48 h after setting the cage (Fig. 3). Among these amplified samples, 28 (72%) were located within 30 m of the cage. Meanwhile, a substantial proportion of samples (9 out of 21, after setting the cage) at St. 1 showed no detection of DNA (0 copies). eDNA was detectable for up to 1 h after removing the cage. The detectability of eDNA concentration at each sampling station was 0–36.5%/h after removing the cage (Table 1). These results indicate that eDNA concentration at a station decreased over 63.5% in 1 h. At 0–1 h after removing the cage, target eDNA was detected in 18 out of 78 samples (13 × 2 × 3). Among these, 17 (94%) were located within 30 m of the cage. Taken together, amplified samples were confirmed in 57 out of the total 351 samples at the total of 0–48 h after setting the cage and 1–2 h after removing. Among the amplified samples, 45 (79%) were obtained within 30 m of the cage. However, after 2–48 h, eDNA was not detected in any samples. The decrease in eDNA concentration from its source was fitted to the power regression model (Figs. 3, 4), and the coefficient of determination for each time point and direction ranged from 0.00 to 0.93.
Concentration of eDNA at each sampling station at a, b: 0 h; c, d: 1 h; e, f: 2 h; g, h: 4 h; i, j: 8 h; k, l: 24 h and m, n: 48 h after setting the cage of fish. Left and right column graphs represent the west and east transect lines, respectively. Open circles represent the average data from PCR triplicates of each water sample (n = 3 in each station and time point)
Mean ± SD water temperature was 21.4 ± 0.4 °C and salinity was 31.7 ± 0.8. The flow velocity during the field experiment was 5 ± 4 (mean ± SD, range: 0–25) cm/s, with directions mainly east or southeast except for time points of 0–4 h after setting the cage (Fig. S1a, b). The direction of current for 1 h before each sampling time point was compared with the total eDNA concentration along the transects (Fig. 3); there were only two out of nine time points where more eDNA was detected along the transect in the direction of currents (24 h after setting the cage and 1 h after removing the cage), whereas an opposite trend was observed in six out of the others. The tidal range was 19 cm during the survey period according to data from the Maizuru Tidal Station of the Japan Meteorological Agency (Fig. S1c).
eDNA concentration decreased exponentially with distance from the source, and was significantly different among sampling stations to the east and west (p < 0.05, nested ANOVA; Fig. 5). In the west, the concentration of eDNA was highest at St. 1, followed by W10 and W30. eDNA concentration at W100 was significantly lower than that at St. 1 and W10. eDNA concentration was not significantly different between W100 and W1000 (Fig. 5a). In the east, the concentration of eDNA was significantly higher at St. 1 and E10 than at E100 or E1000. eDNA concentration was not significantly different between E30 and E1000 (Fig. 5b).
Spatial change of eDNA concentration for all the time points along the northwest (a) and northeast (b) transects after setting the cage and northwest (c) and northeast (d) transects after removing the cage. Open circles represent eDNA concentration at each time point 0–48 h after setting the cage, and those at 0 h and 1 h after removing the cage. Different lowercase letters represent significant differences in ANOVA (p < 0.05) followed by Tukey’s HSD test
The change in detectability over distance after removing the cage was as follows: St. 1–W10, 3.2%; St. 1–W30, 0.7%; St. 1–E10, 21.2%; St. 1–E30, 2.4%. Because eDNA was not detected at 2 h after the removal of the cage (Fig. 6), we assumed that eDNA transferred from adjacent stations was negligible for the above estimation.
Temporal change of eDNA concentration accumulated for all the sampling stations after setting (a, b) or removing the cage (c, d). Different shadings represent different sampling stations as shown in the inserted legends. Different lowercase letters represent significant differences in nested ANOVA (p < 0.05) followed by Tukey’s HSD test
eDNA concentration was significantly different among time points after setting and removing the cage (p < 0.05, nested ANOVA). The concentration of eDNA was significantly higher at 0 h after setting the cage than at other time points (Tukey’s HSD test, p < 0.05; Fig. 6a, b). eDNA concentration at 0 h after removing the cage was also significantly higher than at 1 and 2 h, in both the east and west transects (p < 0.05, Tukey’s HSD test; Fig. 6c, d).
In the comparison of depth layers, eDNA was detected in 8 of the 27 samples collected at 29 h, 32 h and 37 h after setting the cage, among which 5 samples were from surface water (Fig. 7). A significant difference in eDNA concentration was identified between depth layers at 37 h, with the highest value observed in surface water (Fig. 7c).
eDNA concentration collected from three different layers of the water column in St. 1 at a 29 h, b 32 h and c 37 h after setting the cage. Open circles represent average data from PCR triplicates of each water sample (n = 3). Different lowercase letters represent significant differences in ANOVA (p < 0.05) followed by Tukey’s HSD test
Discussion
This field experiment evaluated the spatiotemporal detection range of eDNA using qPCR. We found that eDNA was detectable at stations mostly within 30 m from the source (among 57 amplified samples, 45, or 79%, were obtained within 30 m from the cage; Figs. 3, 4). Our study also indicates that the target DNA was detectable during the time the cage was present and up to 1 h after removing the cage, but not after 2 h had elapsed (Fig. 6). Previous studies conducted in rivers reported longer dispersion distances. For example, Jane et al. (2015) found that eDNA of brook trout Salvelinus fontinalis in a cage could be detected at 240 m downstream from its source. Deiner and Altermatt (2014) reported that eDNA of daphnia Daphnia longispina could be detected at 12.3 km and DNA from the pelecypod Unio tumidus could be found at 9.1 km downstream. The limited dispersion of eDNA in our study may be partly because the marine costal study area has a more limited current speed with complicated flow direction, and involves a larger open area than relatively small rivers.
The dispersion area of eDNA in the sea demonstrated in the present study was within the range suggested by previous studies (Fig. 5). A previous study comparing eDNA concentration and echo intensity in sonar in Maizuru Bay suggested that eDNA reflects the source within a distance of 10–150 m (Yamamoto et al. 2016), although in their study, sampling stations were set at 400-m intervals and the estimation of dispersion and degradation area was based on a statistical model. Meanwhile, nearly 100% of eDNA shed by shiner perch Cymatogaster aggregate, a nearshore fish, could be detected within 100 m from the shore (O’Donnell et al. 2017).
We did not find any explainable relationship between the pattern of current direction and eDNA dispersion (Fig. 3, Fig. S1). The tidal amplitude is generally small in the Sea of Japan, and tidal currents are very weak. Therefore, the current we measured may not have been a major factor in the dispersion of eDNA. The distance between the location of current flow meters and the cage (~ 300 m from NW/NE and St. 1) may also have hindered monitoring of the currents affecting dispersion. Future studies should be conducted in an area where the current is strong to focus on the relationship between current and eDNA dispersion.
The decrease in eDNA in the sea over time could be due to not only dispersion but also degradation. The decrease in eDNA in the sea demonstrated in the present study (Table 1) was much faster than the degradation reported in previous tank experiments. Degradation rates of eDNA for freshwater fishes in tanks are reported as 10.5%/h in common carp Cyprinus carpio (Barnes et al. 2014) and 15.9%/h in bluegill sunfish Lepomis macrochirus (Maruyama et al. 2014). The degradation rates of eDNA of marine fishes in tanks are 4.6%/h in European flounder Platichthys flesus, 1.5%/h in three-spined stickleback Gasterosteus aculeatus (Thomsen et al. 2012a), and 5.5–10.1%/h in northern anchovy Engraulis mordax, Pacific sardine Sardinops sagax, and Pacific chub mackerel Scomber japonicus (Sassoubre et al. 2016). In the present study, eDNA was not detected at any stations more than 1 h after removing the cage. The detectability of eDNA between 0 and 1 h after removing the cage was 0–36.5%/h. Although previous studies of eDNA degradation in a tank used a model of power decline, the degradation rate of eDNA in the sea occurred more quickly, and did not fit this model. eDNA retention time in the sea may be shorter owing to the combination of biological decay and physical dispersion. Additionally, the degradation of eDNA could have been accelerated by high water temperature (21.4 °C) because warm temperature hastens degradation of eDNA due to microbial activity (Strickler et al. 2015; Lance et al. 2017; Tsuji et al. 2017).
It is also important to consider that the emission of DNA might not occur constantly but intermittently. Our dataset showed a number of cases where eDNA was not detected at the nearest station to the cage (Fig. 3). This may be partly due to dispersion caused by flow and the limited probability of catching the DNA fragments during water sampling. However, fish were constantly swimming and forming a group during the experiment. The detected eDNA pattern may also indicate that eDNA was emitted intermittently, with a fluctuating rate of release. A large amount of eDNA was detected immediately after setting and removing the cage, most likely because of handling (Fig. 6). Less eDNA was detected at 1–8 h after setting the cage. A large quantity of eDNA was also detected at 24 h and 48 h, suggesting that a large emission of eDNA could occur temporarily. Fish activity generally increases when prey such as crustaceans are present (Masuda et al. 2012). Feeding behavior contributes to an increased release rate of eDNA in carp, suggesting that feces and intestinal tissues are major sources of eDNA (Klymus et al. 2015). Feeding, defecation, and response to a potential predator were not temporally consistent and thus might have affected the fluctuation of DNA emission.
There were some cases where eDNA was detected at a relatively long distance from the source, with the most distant observation occurring at W1000 immediately after setting the cage (Fig. 1). The summertime current in Maizuru Bay is generally weak, with an average velocity of 5–20 cm/s at the surface (Miwa and Ikeno 2007). The measured hourly velocity immediately after setting the cage was 0.4–13.5 cm/s. Such a slow current can transport eDNA only about 14–480 m in 1 h, making it difficult to explain the rapid transport observed. None of the negative controls in either filtration or PCR (45 in total) showed any amplification, suggesting that contamination after the sampling process is unlikely. Considering that the fish seem to have emitted a substantial amount of eDNA when being introduced in the cage, the potential risk of contamination may have been high at this time point. Alternatively, striped jack is distributed as food for human consumption in this region, and so we cannot exclude the possibility that eDNA in this sample may have come from drainage discarded from houses or vessels. Jo et al. (2017) recently developed a method using two sets of primers with long and short DNA fragments to distinguish between eDNA emitted from live fish and eDNA that is relatively old and degraded. We suggest that long fragments of eDNA would likely be present only near the cage, as opposed to the short fragments we targeted in this study; this would require further investigation to resolve.
In examining eDNA at different depths, we found that eDNA was most frequently present at the surface (Fig. 7). This implies that eDNA tended to stay in the layer where it was released, or tended to be buoyant in surface water, and justifies sampling from sea surface water in our experiment. Moreover, this result coincides with a previous finding that the vertical distribution of eDNA reflects the layer in which organisms live, based on a study by Minamoto et al. (2017) wherein eDNA of the jellyfish Chrysaora pacifica was more abundant near the bottom where this species tends to accumulate. On the other hand, we did detect eDNA in the middle and bottom layers, implying that some eDNA may have sunk after emission. eDNA at the surface might be detectable relatively soon after emission, while eDNA may be detected near the bottom at distant points after transport and gradual settlement. In a lake, eDNA of bigheaded Asian carp Hypophthalmichthys spp. was detected in greater quantities in sediment than in surface water (Turner et al. 2015), suggesting that settled eDNA could be preserved in sediment. On the other hand, eDNA of African jewelfish Hemichromis letourneuxi in an experimental pond was present in greater quantities near the surface than at the bottom (Moyer et al. 2014). These findings indicate that vertical distribution of eDNA depends on both the depth of source organisms and the behavior of eDNA, with the latter dependent on its chemical condition and physical factors in the environment.
Although the use of eDNA as a tool for marine studies still has many obstacles, the present investigation implies that the area of eDNA dispersion in the sea is as narrow as 30 m and the duration of persistence could be as short as 1 h when emitted from a limited source. Collectively, these results suggest that eDNA reflects presence and abundance of organisms not in a large scale of time and space, but within a few hours and tens of meters. Further study should elucidate how biomass and other factors influence the distance of dispersion, degradation, and concentration of eDNA.
Change history
30 August 2019
A Correction to this paper has been published: https://doi.org/10.1007/s12562-019-01341-z
13 June 2022
A Correction to this paper has been published: https://doi.org/10.1007/s12562-022-01612-2
References
Barnes MA, Turner CR, Jerde CL, Renshaw MA, Chadderton WL, Lodge DM (2014) Environmental conditions influence eDNA persistence in aquatic systems. Environ Sci Technol 48:1819–1827
Deiner K, Altermatt F (2014) Transport distance of invertebrate environmental DNA in a natural river. PLoS One 9:e88786
Doi H, Inui R, Akamatsu Y, Kanno K, Yamanaka H, Takahara T, Minamoto T (2017) Environmental DNA analysis for estimating the abundance and biomass of stream fish. Freshw Biol 62:30–39
Eichmiller JJ, Bajer PG, Sorensen PW (2014) The relationship between the distribution of common carp and their environmental DNA in a small lake. PLoS One 9:e112611
Evans NT, Olds BP, Renshaw MA, Turner CR, Li Y, Jerde CL, Lodge DM (2016) Quantification of mesocosm fish and amphibian species diversity via environmental DNA metabarcoding. Mol Ecol Resour 16:29–41
Ficetola GF, Miaud C, Pompanon F, Taberlet P (2008) Species detection using environmental DNA from water samples. Biol Lett UK 4:423–425
Fukumoto S, Ushimaru A, Minamoto T (2015) A basin-scale application of environmental DNA assessment for rare endemic species and closely related exotic species in rivers: a case study of giant salamanders in Japan. J Appl Ecol 52:358–365
Jane SF, Wilcox TM, McKelvey KS, Young MK, Schwartz MK, Lowe WH, Letcher BH, Whiteley AR (2015) Distance, flow and PCR inhibition: eDNA dynamics in two headwater streams. Mol Ecol Resour 15:216–227
Jo T, Murakami H, Masuda R, Sakata MK, Yamamoto S, Minamoto T (2017) Rapid degradation of longer DNA fragments enables the improved estimation of distribution and biomass using environmental DNA. Mol Ecol Resour 27:25–33
Kelly RP, Port JA, Yamahara KM, Crowder LB (2014) Using environmental DNA to census marine fishes in a large mesocosm. PLoS One 9:e86175
Klymus KE, Richter CA, Chapman DC, Paukert C (2015) Quantification of eDNA shedding rates from invasive bighead carp Hypophthalmichthys nobilis and silver carp Hypophthalmichthys molitrix. Biol Conserv 183:77–84
Lance RF, Klymus KE, Richter CA, Guan X, Farrington HL, Carr MR, Thompson N, Chapman DC, Baerwaldt KL (2017) Experimental observations on the decay of environmental DNA from bighead and silver carps. Manag Biol Invasion 8:343–359
Maruyama A, Nakamura K, Yamanaka H, Kondoh M, Minamoto T (2014) The release rate of environmental DNA from juvenile and adult fish. PLoS One 9:e114639
Masuda R (2008) Seasonal and interannual variation of subtidal fish assemblages in Wakasa Bay with reference to the warming trend in the Sea of Japan. Environ Biol Fish 82:387–399
Masuda R, Matsuda K, Tanaka M (2012) Laboratory video recordings and underwater visual observations combined to reveal activity rhythm of red-spotted grouper and banded wrasse, and their natural assemblages. Environ Biol Fish 95:335–346
Minamoto T, Fukuda M, Katsuhara KR, Fujiwara A, Hidaka S, Yamamoto S, Takahashi K, Masuda R (2017) Environmental DNA reflects spatial and temporal jellyfish distribution. PLoS One 12:e0173073
Miwa H, Ikeno H (2007) Three dimensional analysis of flow field and water environment in Maizuru Bay with consideration of density distribution. B Maizuru Natl Coll Tec 42:47–58 (In Japanese)
Miya M, Sato Y, Fukunaga T, Fukunaga T, Sado T, Poulsen JY, Sato K, Minamoto T, Yamamoto S, Yamanaka H, Araki H, Kondoh M, Iwasaki W (2015) MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. R Soc Open Sci 2:150088
Moyer GR, Díaz-Ferguson E, Hill JE, Shea C (2014) Assessing environmental DNA detection in controlled lentic systems. PLoS One 9:e103767
Murphy HM, Jenkins GP (2010) Observational methods used in marine spatial monitoring of fishes and associated habitats: a review. Mar Freshw Res 61:236–252
O’Donnell JL, Kelly RP, Shelton AO, Samhouri JF, Lowell NC, Williams GD (2017) Spatial distribution of environmental DNA in a nearshore marine habitat. PeerJ 5:e3044
Piaggio AJ, Engeman RM, Hopken MW, Humphrey JS, Keacher KL, Bruce WE, Avery ML (2014) Detecting an elusive invasive species: a diagnostic PCR to detect burmese python in Florida waters and an assessment of persistence of environmental DNA. Mol Ecol Resour 14:374–380
Pilliod DS, Goldberg CS, Arkle RS, Waits LP, Richardson J (2013) Estimating occupancy and abundance of stream amphibians using environmental DNA from filtered water samples. Can J Fish Aquat Sci 70:1123–1130
Rees HC, Bishop K, Middleditch DJ, Patmore JRM, Maddison BC, Gough KC (2014) The application of eDNA for monitoring of the great crested newt in the UK. Ecol Evol 4:4023–4032
Sansom BJ, Sassoubre LM (2017) Environmental DNA (eDNA) shedding and decay rates to model freshwater mussel eDNA transport in a river. Environ Sci Technol 51:14244–14253
Sassoubre LM, Yamahara KM, Gardner LD, Block BA, Boehm AB (2016) Quantification of environmental DNA (eDNA) shedding and decay rates for three marine fish. Environ Sci Technol 50:10456–10464
Strickler KM, Fremier AK, Goldberg CS (2015) Quantifying effects of UV-B, temperature, and pH on eDNA degradation in aquatic microcosms. Biol Cons 183:85–92
Takahara T, Minamoto T, Yamanaka H, Doi H, Kawabata Z (2012) Estimation of fish biomass using environmental DNA. PLoS One 7:e35868
Takahara T, Minamoto T, Doi H (2013) Using environmental DNA to estimate the distribution of an invasive fish species in ponds. PLoS One 8:e56584
Thomsen PF, Kielgast J, Iversen LL, Møller PR, Rasmussen M, Willerslev E (2012a) Detection of a diverse marine fish fauna using environmental DNA from seawater samples. PLoS One 7:e41732
Thomsen PF, Kielgast J, Iversen LL, Wiuf C, Rasmussen M, Gilbert MTP, Willerslev E (2012b) Monitoring endangered freshwater biodiversity using environmental DNA. Mol Ecol 21:2565–2573
Thomsen PF, Møller PR, Sigsgaard EE, Knudsen SW, Jørgensen OA, Willerslev E (2016) Environmental DNA from seawater samples correlate with trawl catches of subarctic, deepwater fishes. PLoS One 11:e0165252
Tsuji S, Ushio M, Sakurai S, Minamoto T, Yamanaka H (2017) Water temperature-dependent degradation of environmental DNA and its relation to bacterial abundance. PLoS One 12:e0176608
Turner CR, Barnes MA, Xu CCY, Jones SE, Jerde CL, Lodge DM (2014) Particle size distribution and optimal capture of aqueous macrobial eDNA. Methods Ecol Evol 5:676–684
Turner CR, Uy KL, Everhart RC (2015) Fish environmental DNA is more concentrated in aquatic sediments than surface water. Biol Conserv 183:93–102
Ushio M, Murakami H, Masuda R, Sado T, Miya M, Sakurai S, Yamanaka H, Minamoto T, Kondoh M (2018) Quantitative monitoring of multispecies fish environmental DNA using high-throughput sequencing. MBMG 2:e23297
Valentini A, Taberlet P, Miaud C, Civade R, Herder J, Thomsen PF, Dejean T (2016) Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol Ecol 25:929–942
Wilcox TM, McKelvey KS, Young MK, Lowe WH, Schwartz MK (2015) Environmental DNA particle size distribution from brook trout (Salvelinus fontinalis). Conserv Genet Resour 7:639–641
Wilcox TM, McKelvey KS, Young MK, Sepulveda AJ, Shepard BB, Jane SF, Schwartz MK (2016) Understanding environmental DNA detection probabilities: a case study using a stream-dwelling char Salvelinus fontinalis. Biol Conserv 194:209–216
Yamamoto S, Minami K, Fukaya K, Takahashi K, Sawada H, Murakami H, Tsuji S, Hashizume H, Kubonaga S, Horiuchi T, Hongo M, Nishida J, Okugawa Y, Fujiwara A, Fukuda M, Hidaka S, Suzuki K.W, Miya M, Araki H, Yamanaka H, Maruyama A, Miyashita K, Masuda R, Minamoto T, Kondo M (2016) Environmental DNA as a ‘snapshot’ of fish distribution: a case study of Japanese jack mackerel in Maizuru Bay, Sea of Japan. PLoS One 11:e0149786
Yamamoto S, Masuda R, Sato Y, Sado T, Araki H, Kondoh M, Minamoto T, Miya M (2017) Environmental DNA metabarcoding reveals local fish communities in a species-rich coastal sea. Sci Rep 7:40368
Acknowledgements
We thank Yoshihito Ogura for navigating the boat and Masahiro Mukai and Aina Tanimoto (MFRS) for assisting with filtration procedures. This study was supported by CREST of JST (grant number: JPMJCR13A2) and the Sasakawa Scientific Research Grant from The Japan Science Society.
Author information
Authors and Affiliations
Corresponding author
Additional information
The original online version of this article was revised due to a retrospective Open Access order.
Electronic supplementary material
Below is the link to the electronic supplementary material.
12562_2018_1282_MOESM1_ESM.docx
Fig. S1 Observed flow velocities (a and b) and sea level (c) during the experiment. The length and angle of stick represent flow velocity and direction, respectively, recorded at NW (a) and NE (b) in Fig. 1. The eDNA sampling time points are represented by dotted lines
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Murakami, H., Yoon, S., Kasai, A. et al. Dispersion and degradation of environmental DNA from caged fish in a marine environment. Fish Sci 85, 327–337 (2019). https://doi.org/10.1007/s12562-018-1282-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12562-018-1282-6