Abstract
Introduction
Lung cancer is one of the most common cancer malignancies and the principal cause of cancer-associated deaths worldwide. Non-small cell lung cancers (NSCLCs) account for more than 80% of all lung cancer cases. Recent studies showed that the genes of the integrin alpha (α) (ITGA) subfamily play a fundamental role in various cancers. However, little is known about the expression and roles of distinct ITGA proteins in NSCLCs.
Methods
Gene Expression Profiling Interactive Analysis and UALCAN (University of ALabama at Birmingham CANcer) web resources and The Cancer Genome Atlas (TCGA), ONCOMINE, cBioPortal, GeneMANIA, and Tumor Immune Estimation Resource databases were used to evaluate differential expression, correlations between the expression levels of individual genes, the prognostic value of overall survival (OS) and stage, genetic alterations, protein–protein interactions, and the immune cell infiltration of ITGAs in NSCLCs. We used R (v. 4.0.3) software to conduct gene correlation, gene enrichment, and clinical correlation of RNA sequencing data of 1016 NSCLCs from TCGA. To evaluate the expression of ITGA5/8/9/L at the expression and protein levels, qRT-PCR, immunohistochemistry (IHC), and hematoxylin and eosin (H&E) were performed, respectively.
Results
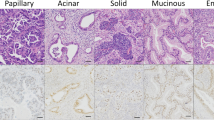
Upregulated levels of ITGA11 messenger RNA and downregulated levels of ITGA1/3/5/7/8/9/L/M/X were observed in the NSCLC tissues. Lower expression of ITGA5/6/8/9/10/D/L was discovered to be expressively associated with advanced tumor stage or poor patient prognosis in patients with NSCLC. A high mutation rate (44%) of the ITGA family was observed in the NSCLCs. Gene Ontology functional enrichment analyses results revealed that the differentially expressed ITGAs could be involved in roles related to extracellular matrix (ECM) organization, collagen-containing ECM cellular components, and ECM structural constituent molecular functions. The results of the Kyoto Encyclopedia of Genes and Genomes analysis revealed that ITGAs may be involved in focal adhesion, ECM–receptor interaction, and amoebiasis; the expression of ITGAs was significantly correlated with the infiltration of diverse immune cells in NSCLCs. ITGA5/8/9/L was also highly correlated with PD-L1 expression. The validation results for marker gene expression in NSCLC tissues by qRT-PCR, IHC, and H&E staining indicated that the expression of ITGA5/8/9/L decreased compared with that in normal tissues.
Conclusion
As potential prognostic biomarkers in NSCLCs, ITGA5/8/9/L may fulfill important roles in regulating tumor progression and immune cell infiltration.








Similar content being viewed by others
References
Araujo LH, Timmers C, Bell EH, et al. Genomic characterization of non-small-cell lung cancer in African Americans by targeted massively parallel sequencing. J Clin Oncol. 2015;33(17):1966–73.
Wang W, Zhao M, Cui L, et al. Characterization of a novel HDAC/RXR/HtrA1 signaling axis as a novel target to overcome cisplatin resistance in human non-small cell lung cancer. Mol Cancer. 2020;19(1):134.
Kim JW, Marquez CP, Kostyrko K, et al. Antitumor activity of an engineered decoy receptor targeting CLCF1-CNTFR signaling in lung adenocarcinoma. Nat Med. 2019;25(11):1783–95.
Wu S, Shen G, Mao J, Gao B. CT radiomics in predicting EGFR mutation in non-small cell lung cancer: a single institutional study. Front Oncol. 2020;10: 542957.
Liu H, Li D, Fang H, Ning J. Species-specific function of microRNA-7702 in human colorectal cancer cells via targeting TADA1. Am J Transl Res. 2018;10(8):2579–89.
Han Y, Su C, Yu D, et al. Cholecystokinin attenuates radiation-induced lung cancer cell apoptosis by modulating p53 gene transcription. Am J Transl Res. 2017;9(2):638–46.
Arnaout MA. Integrin structure: new twists and turns in dynamic cell adhesion. Immunol Rev. 2002;186:125–40.
Tadokoro S, Shattil SJ, Eto K, et al. Talin binding to integrin beta tails: a final common step in integrin activation. Science. 2003;302(5642):103–6.
Kim C, Ye F, Ginsberg MH. Regulation of integrin activation. Annu Rev Cell Dev Biol. 2011;27:321–45.
Takada Y, Ye X, Simon S. The integrins. Genome Biol. 2007;8(5):215.
Campbell ID, Humphries MJ. Integrin structure, activation, and interactions. Cold Spring Harb Perspect Biol. 2011;3(3):a004994.
Hynes RO. Integrins: bidirectional, allosteric signaling machines. Cell. 2002;110(6):673–87.
Larson RS, Corbi AL, Berman L, Springer T. Primary structure of the leukocyte function-associated molecule-1 alpha subunit: an integrin with an embedded domain defining a protein superfamily. J Cell Biol. 1989;108(2):703–12.
Humphries JD, Byron A, Humphries MJ. Integrin ligands at a glance. J Cell Sci. 2006;119(Pt 19):3901–3.
Dmitriev AA, Kashuba VI, Haraldson K, et al. Genetic and epigenetic analysis of non-small cell lung cancer with NotI-microarrays. Epigenetics. 2012;7(5):502–13.
Desgrosellier JS, Cheresh DA. Integrins in cancer: biological implications and therapeutic opportunities. Nat Rev Cancer. 2010;10(1):9–22.
Cox D, Brennan M, Moran N. Integrins as therapeutic targets: lessons and opportunities. Nat Rev Drug Discov. 2010;9(10):804–20.
Shattil SJ, Kim C, Ginsberg MH. The final steps of integrin activation: the end game. Nat Rev Mol Cell Biol. 2010;11(4):288–300.
Moser M, Nieswandt B, Ussar S, Pozgajova M, Fässler R. Kindlin-3 is essential for integrin activation and platelet aggregation. Nat Med. 2008;14(3):325–30.
Shen B, Delaney MK, Du X. Inside-out, outside-in, and inside-outside-in: G protein signaling in integrin-mediated cell adhesion, spreading, and retraction. Curr Opin Cell Biol. 2012;24(5):600–6.
Legate KR, Wickström SA, Fässler R. Genetic and cell biological analysis of integrin outside-in signaling. Genes Dev. 2009;23(4):397–418.
Schwartz MA, Ginsberg MH. Networks and crosstalk: integrin signalling spreads. Nat Cell Biol. 2002;4(4):E65–8.
Li Y, Xiao Y, Lin H-P, et al. In vivo β-catenin attenuation by the integrin α5-targeting nano-delivery strategy suppresses triple negative breast cancer stemness and metastasis. Biomaterials. 2019;188:160–72.
Ryu J, Koh Y, Park H, et al. Highly expressed integrin-α8 induces epithelial to mesenchymal transition-like features in multiple myeloma with early relapse. Mol Cells. 2016;39(12):898–908.
Guo WH, Bian JJ, Tian GF, Lyu ZX, Gui YX, Ye L. Expression of Fermintin family homologous protein 2 in non-small cell lung cancer and its clinical significance. Zhonghua Bing Li Xue Za Zhi. 2018;47(10):780–3.
Haas TL, Sciuto MR, Brunetto L, et al. Integrin α7 is a functional marker and potential therapeutic target in glioblastoma. Cell Stem Cell. 2017;21(1):35–50.e9.
Gong L, Zheng Y, Liu S, Peng Z. Fibronectin regulates the dynamic formation of ovarian cancer multicellular aggregates and the expression of integrin receptors. Asian Pac J Cancer Prev. 2018;19(9):2493–8.
Chang H-W, Yen C-Y, Chen C-H, et al. Evaluation of the mRNA expression levels of integrins α3, α5, β1 and β6 as tumor biomarkers of oral squamous cell carcinoma. Oncol Lett. 2018;16(4):4773–81.
Rhodes DR, Yu J, Shanker K, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6(1):1–6.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–102.
Nagy Á, Lánczky A, Menyhárt O, Győrffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018;8(1):9227.
The TCGA legacy. Cell. 2018;173(2):281–2.
Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13.
Warde-Farley D, Donaldson SL, Comes O, et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010;38(Web server issue):W214–20.
Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–58.
Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–10.
Caswell PT, Norman JC. Integrin trafficking and the control of cell migration. Traffic. 2006;7(1):14–21.
Xiong J, Balcioglu HE, Danen EHJ. Integrin signaling in control of tumor growth and progression. Int J Biochem Cell Biol. 2013;45(5):1012–5.
Felding-Habermann B, Mueller BM, Romerdahl CA, Cheresh DA. Involvement of integrin alpha V gene expression in human melanoma tumorigenicity. J Clin Investig. 1992;89(6):2018–22.
Chuang Y-C, Wu H-Y, Lin Y-L, et al. Blockade of ITGA2 induces apoptosis and inhibits cell migration in gastric cancer. Biol Proced Online. 2018;20:10.
Gong J, Lu X, Xu J, Xiong W, Zhang H, Yu X. Coexpression of UCA1 and ITGA2 in pancreatic cancer cells target the expression of miR-107 through focal adhesion pathway. J Cell Physiol. 2019;234(8):12884–96.
Lu Y, Li C, Chen H, Zhong W. Identification of hub genes and analysis of prognostic values in pancreatic ductal adenocarcinoma by integrated bioinformatics methods. Mol Biol Rep. 2018;45(6):1799–807.
Lemma SA, Kuusisto M, Haapasaari K-M, et al. Integrin alpha 10, CD44, PTEN, cadherin-11 and lactoferrin expressions are potential biomarkers for selecting patients in need of central nervous system prophylaxis in diffuse large B-cell lymphoma. Carcinogenesis. 2017;38(8):812–20.
Pan Y, Liu G, Yuan Y, Zhao J, Yang Y, Li Y. Analysis of differential gene expression profile identifies novel biomarkers for breast cancer. Oncotarget. 2017;8(70):114613–25.
Zhang R, Zhang T-T, Zhai G-Q, et al. Evaluation of the HOXA11 level in patients with lung squamous cancer and insights into potential molecular pathways via bioinformatics analysis. World J Surg Oncol. 2018;16(1):109.
Ando T, Kage H, Matsumoto Y, et al. Integrin α11 in non-small cell lung cancer is associated with tumor progression and postoperative recurrence. Cancer Sci. 2020;111(1):200–8. https://doi.org/10.1111/cas.14257.
Parajuli H, Teh M-T, Abrahamsen S, et al. Integrin α11 is overexpressed by tumour stroma of head and neck squamous cell carcinoma and correlates positively with alpha smooth muscle actin expression. J Oral Pathol Med. 2017;46(4):267–75.
Shang L, Ye X, Zhu G, et al. Prognostic value of integrin variants and expression in post-operative patients with HBV-related hepatocellular carcinoma. Oncotarget. 2017;8(44):76816–31.
Haider S, Wang J, Nagano A, et al. A multi-gene signature predicts outcome in patients with pancreatic ductal adenocarcinoma. Genome Med. 2014;6(12):105.
Yan P, He Y, Xie K, Kong S, Zhao W. Analyses for potential key genes associated with gastric cancer. PeerJ. 2018;6:e6092.
Zheng W, Jiang C, Li R. Integrin and gene network analysis reveals that ITGA5 and ITGB1 are prognostic in non-small-cell lung cancer. Onco Targets Ther. 2016;9:2317–27.
Dingemans AM, van den Boogaart V, Vosse BA, van Suylen RJ, Griffioen AW, Thijssen VL. Integrin expression profiling identifies integrin alpha5 and beta1 as prognostic factors in early stage non-small cell lung cancer. Mol Cancer. 2010;17(9):152. https://doi.org/10.1186/1476-4598-9-152.
Mallawaaratchy DM, Buckland ME, McDonald KL, et al. Membrane proteome analysis of glioblastoma cell invasion. J Neuropathol Exp Neurol. 2015;74(5):425–41.
Kok-Sin T, Mokhtar NM, Ali Hassan NZ, et al. Identification of diagnostic markers in colorectal cancer via integrative epigenomics and genomics data. Oncol Rep. 2015;34(1):22–32.
Yang X, Deng Y, He R-Q, et al. Upregulation of HOXA11 during the progression of lung adenocarcinoma detected via multiple approaches. Int J Mol Med. 2018;42(5):2650–64.
Wang Z, Li Y, Xiao Y, et al. Integrin α9 depletion promotes β-catenin degradation to suppress triple-negative breast cancer tumor growth and metastasis. Int J Cancer. 2019;145(10):2767–80.
Fan J, Kang X, Zhao L, Zheng Y, Yang J, Li D. Long noncoding RNA CCAT1 functions as a competing endogenous RNA to upregulate ITGA9 by sponging MiR-296-3p in melanoma. Cancer Manag Res. 2020;12:4699–714.
Ross RW, Galsky MD, Scher HI, et al. A whole-blood RNA transcript-based prognostic model in men with castration-resistant prostate cancer: a prospective study. Lancet Oncol. 2012;13(11):1105–13.
Fernández-Sevilla LM, Valencia J, Flores-Villalobos MA, et al. The choroid plexus stroma constitutes a sanctuary for paediatric B-cell precursor acute lymphoblastic leukaemia in the central nervous system. J Pathol. 2020;252(2):189–200.
Zhou M, Zhang Z, Bao S, et al. Computational recognition of lncRNA signature of tumor-infiltrating B lymphocytes with potential implications in prognosis and immunotherapy of bladder cancer. Brief Bioinform. 2021;22(3): bbaa047.
Sun J, Zhang Z, Bao S, et al. Identification of tumor immune infiltration-associated lncRNAs for improving prognosis and immunotherapy response of patients with non-small cell lung cancer. J Immunother Cancer. 2020;8(1):e000110.
Acknowledgements
We are particularly grateful to all the people who have given us help with our article.
Funding
This work was supported by the Performance incentives guide special key project of Chongqing scientific research institutes (cstc2019jxj1130005, Yuequan Jiang) and the Laboratory opening fund of Chongqing University Cancer Hospital (Fei Teng). The Rapid Service Fee was funded by the authors.
Author Contributions
Yu Huang and Dong-Ming Guo conceived the idea and conceptualized the study. Shi Bu and Wei Xu collected the data and analyzed the data. Qing-Chun Cai and Jian Xu drafted the manuscript, then Yue-Quan Jiang and Fei Teng reviewed the manuscript. All authors read and approved the final draft.
Disclosures
Yu Huang, Dong-Ming Guo, Shi Bu, Wei Xu, Qing-Chun Cai, Jian Xu, Yue-Quan Jiang and Fei Teng have nothing to disclose.
Compliance with Ethics Guidelines
The data has been taken from the ONCOMINE database, the Gene Expression Profiling Interactive Analysis web resource, The Cancer Genome Atlas (TCGA) database, the UALCAN web resource, the cBioPortal database, the GeneMANIA database, and the Tumor Immune Estimation Resource database. IRB approval was not required as all the database information in the article was publicly available without any limitation and data was deidentified.
Data Availability
The data sets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Corresponding authors
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, Y., Guo, DM., Bu, S. et al. Systematic Analysis of the Prognostic Significance and Roles of the Integrin Alpha Family in Non-Small Cell Lung Cancers. Adv Ther 40, 2186–2204 (2023). https://doi.org/10.1007/s12325-023-02469-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12325-023-02469-2