Abstract
Background
MicroRNA-155 (miR-155) may function as a diagnostic biomarker of breast cancer (BC). Nevertheless, the available evidence is controversial. Therefore, we performed this study to summarize the global predicting role of miR-155 for early detection of BC and preliminarily explore the functional roles of miR-155 in BC.
Methods
We first collected published studies and applied the bivariate meta-analysis model to generate the pooled diagnostic parameters of miR-155 in diagnosing BC such as sensitivity, specificity and area under curve (AUC). Then, we applied function enrichment and protein–protein interactions (PPI) analyses to explore the potential mechanisms of miR-155.
Results
A total of 21 studies were finally included. The results indicated that miR-155 allowed for the discrimination between BC patients and healthy controls with a sensitivity of 0.87 (95% CI 0.78–0.93), specificity of 0.82 (0.72–0.89), and AUC of 0.91 (0.88–0.93). In addition, the overall sensitivity, specificity and AUC for circulating miR-155 were 0.88 (0.76–0.95), 0.83 (0.72–0.90), and 0.92 (0.89–0.94), respectively. Function enrichment analysis revealed several vital ontologies terms and pathways associated with BC occurrence and development. Furthermore, in the PPI network, ten hub genes and two significant modules were identified to be involved in some important pathways associated with the pathogenesis of BC.
Conclusions
We demonstrated that miR-155 has great potential to facilitate accurate BC detection and may serve as a promising diagnostic biomarker for BC. However, well-designed cohort studies and biological experiments should be implemented to confirm the diagnostic value of miR-155 before it can be applied to routine clinical procedures.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Breast cancer (BC) remains the most prevalent cause of cancer mortality in females worldwide [1]. Like all other malignant tumors, the prognosis of BC is highly associated with the extent how early this disease is diagnosed [2]. Currently, several conventional methods, including imaging examination and biopsy, and some tumor markers such as CEA and CA15-3 have been confirmed to be beneficial for detection of BC [3]. Nevertheless, there are some deficiencies as they are invasive and harmful procedure or not sensitive enough for accurate BC diagnosis. Therefore, the development of efficient diagnostic methods or tumor biomarkers for BC is urgently needed.
MicroRNAs are a class of 18–25 nucleotide, non-coding RNAs that regulate gene expression through targeting messenger RNAs and triggering either translational repression or RNA degradation [4]. Increasing studies have demonstrated that microRNAs were highly involved in various physiological and pathological processes such as cell proliferation, differentiation, and apoptosis [5]. Moreover, a large number of articles have proved that the abnormal expressions of microRNAs have a direct association with the development and progression of various cancers including BC. Subsequent evidence also indicated that circulating microRNAs were highly stabile and could be easily extracted and measured, revealing that circulating microRNAs might serve as promising biomarkers for early detection of BC [6].
Among a wide spectrum of microRNAs, microRNA-155 (miR-155) has gained great attention and numerous studies have revealed the critical role of miR-155 in the development of some cancers including colorectal cancer, oesophageal squamous cell carcinoma, lung cancer and BC [7]. Recently, several studies have revealed that aberrant expression of miR-155 may become a potential diagnostic marker of several types of cancers including BC [8]. Nevertheless, the diagnostic accuracy of miR-155 in BC was inconsistent or even contradictory among different studies. In addition, the biomarker roles of miR-155 during the initiation and progression of BC are still poorly illustrated by previous studies.
Thus, we first performed a meta-analysis, to synthesize available evidence on miR-155 as a diagnostic biomarker in patients with BC. Furthermore, we designed a bioinformatics study to uncover the potential biomarker roles of miR-155.
Methods
Literature search strategy
To identify the relevant studies, we searched through the online PubMed, EMBASE and Web of Science databases until May 1, 2020. The key words used for literature retrieval were as follows: (“microRNA-155” OR “miR-155” OR “miRNA-155”) and (“breast cancer” OR “breast tumor” OR “breast carcinoma” OR “breast neoplasm”). The references of all the relevant publications were manually searched to obtain additional eligible studies.
Inclusion and exclusion criteria
Studies were considered eligible if they met the following criteria: (1) all the patients with BC must have been confirmed by pathological examination; (2) the levels of miR-155 were measured; (3) they presented sufficient data to allow the establishment of two-by-two tables, including true positives (TP), false positives (FP), true negatives (TN), and false negatives (FN).
Studies were excluded if they were: (1) reviews, meta-analysis, letters, commentaries, or abstracts presented in conferences; (2) lacking sufficient data for calculation of specificity and sensitivity with 95% confidence intervals (CIs); (3) duplication of previous publications.
Data extraction and quality assessment
Two investigators (Xuemin Liu and Qingyu Chang) independently extracted all the data from eligible studies. A third investigator (Haiqiang Wang) was responsible for reconciling disagreements when the results were controversial. The extracted data elements of this study for diagnosis included: the first author’s name, country of research, year of publication, age of patients, ethnicity, sample size, sample source, detection method, sensitivity, specificity, TP, FP, FN, and TN. We estimated the quality of each study using the revised Quality Assessment of Diagnostic Accuracy Studies (QUADAS-2), which has been confirmed to be an effective tool for assessing the quality of diagnostic tests [9].
Statistical analysis
The data analysis to assess the diagnostic role of miR-155 in BC was performed by STATA 14.0. The bivariate meta-analysis model was applied to obtain the pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), and diagnostic odds ratio (DOR) [10]. Furthermore, the summary receiver operator characteristic (SROC) curve was generated based on pooled sensitivity and specificity of included studies and the corresponding area under the SROC curve (AUC) was calculated to estimate the accuracy of miR-155 in BC detection [11]. Statistical heterogeneity was evaluated using Cochran-Q test and I2 test. P value < 0.05 for Cochran’s Q test, or I2 > 50%, suggested an existence of significant heterogeneity [12]. The Spearman correlation coefficient of logarithm sensitivity and 1-specificity was calculated for measuring the threshold effect. In addition, subgroup, meta-regression and sensitivity analyses were employed to investigate the potential sources of heterogeneity among included studies. Finally, publication bias was checked with the Deeks’ funnel plot [13]. A P value less than 0.05 for two-tailed was considered statistically significant.
Target genes prediction of miR-155
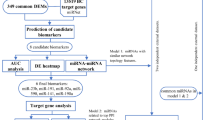
We then performed a bioinformatics analysis to explore the potential function of miR-155. First, the microRNA prediction data were downloaded from the microRNA target gene prediction website miRTarBase, which has been one of the most comprehensive databases with experimentally validated microRNA–target interactions [14]. We only selected the human microRNA–target information for further analysis.
GO enrichment and pathway analysis
Gene ontology (GO) analysis was carried out for annotating the parental genes according to the biological processes (BP), cellular components (CC) and molecular functions (MF) [15]. Besides, we provided pathway enrichment analyses for miR-155 targets using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [16]. To enable better recognition of the biological functions of miR-155, we used the online tool Database for Annotation, Visualization and Integrated Discovery (DAVID) to perform GO enrichment and KEGG analyses [17]. P value < 0.05 was considered statistically significant.
Establishing the PPI network, selection of hub genes and module analysis
To identify the hub regulatory genes and to investigate the interactions among the target genes of miR-155, a protein–protein interaction (PPI) network analysis was conducted. The Search Tool for the Retrieval of Interacting Genes (STRING) is a biological database constructed for collecting PPI information, and then analyzing the functional interactions between proteins [18]. All the targets of miR-155 were uploaded onto STRING to retrieve their PPI data and only the interactions with a combined score > 0.4 validated by experiments or collected by text mining were set to be the cutoff criterion. Afterwards, the PPI network was visualized with Cytoscape software [19]. The CytoNCA plug-in of Cytoscape was used to identify the hub genes from PPI network, and the hub genes were selected according to three different centrality measures, containing betweenness centrality, closeness centrality and degree centrality [20]. Furthermore, the plug-in of Molecular Complex Detection (MCODE) in Cytoscape software was employed to search the significant modules in PPI network [21]. Finally, KEGG pathway enrichment analyses were performed with the hub genes and genes involved in the selected modules. The threshold P value < 0.05 was considered statistically significant.
Results
Descriptive assessment and study characteristics
In total, 657 potentially eligible studies were obtained from online database searching. After manually scanning the titles, abstracts and key data, 600 records were excluded because of meta-analysis, letters, reviews, duplicate studies, or their irrelevance to the present analysis. After screening full texts of the remaining 57 articles, 39 publications were further excluded because they failed to satisfy our inclusion criteria. Finally, 18 publications containing 21 studies were considered in the meta-analysis [22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39]. Our flow diagram of each stage for inclusion and exclusion was presented in Fig. 1.
The main characteristics of each enrolled publication were summarized in Table 1. The publication years of the included studies were from 2012 to 2020. All the studies performed reverse transcription quantitative PCR (RT-qPCR) for miR-155 detection, and the specimen type included serum, plasma, blood, urine, and tissue. The quality of the included articles was evaluated by querying the QUADAS-2 scores. As a result, the overall methodological qualities of the study designs of selected studies were generally good.
Diagnostic performance of miR-155 in BC
The forest plots of sensitivity and specificity of miR-155 in the diagnosis of BC are presented at Fig. 2. The overall analysis indicated the pooled sensitivity and specificity were 0.87 (95% CI 0.78–0.93) and 0.82 (95% CI 0.72–0.89), respectively. We also noted that the pooled PLR was 4.8 (95% CI 3.0–7.7), NLR was 0.16 (95% CI 0.09–0.29), and DOR was 30 (95% CI 11–80). The SROC curve for the enrolled diagnostic tests is shown at Fig. 3. The corresponding AUC was 0.91 (95% CI 0.88–0.93), revealing that miR-155 has relatively high diagnostic accuracy for BC.
Test of heterogeneity and subgroup analysis
The Spearman correlation coefficient was 0.35 with a P value of 0.12, indicating no obvious heterogeneity from threshold effect in the present data pooling.
The I2 values of sensitivity and specificity were 93.58% (95% CI 91.74–95.43%; P < 0.01) and 91.03% (95% CI 88.17–93.88%; P < 0.01), respectively, indicating significant heterogeneity from non-threshold effect in our study. To investigate the between-study heterogeneity, subgroup analyses based on ethnicity (Asian or non-Asian), sample size (N > 100 or N < 100) and sample source (serum, plasma and tissue) were conducted (Table 2). A comparison of sample size in large sample size (N > 100) and small sample size (N < 100) indicated that the specificity (0.85 versus 0.77), DOR (36 versus 24), PLR (5.7 versus 3.8), and AUC (0.92 versus 0.89) were higher in studies with large sample size than studies with small sample size, suggesting that large cohort studies are warranted and worthwhile. For studies on Asian populations, the pooled sensitivity was 0.90, specificity was 0.73, PLR was 3.3, NLR was 0.14, DOR was 23, and AUC was 0.84. For studies on non-Asian populations, the pooled sensitivity was 0.83, specificity was 0.87, PLR was 6.7, NLR was 0.19, DOR was 35, and AUC was 0.92. The sensitivity of studies using plasma was 0.84 and the specificity was 0.75, with a pooled DOR of 15 and AUC of 0.80. Nevertheless, for serum-based studies, the sensitivity of studies was 0.89 and the specificity was 0.86, with a pooled DOR of 52 and AUC of 0.94, indicating that the use of serum miR-155 as relatively reliable matrix in diagnosing BC. Since most sample sources concentrated on circulating samples, we performed subgroup analysis based on this point. The results indicated that the pooled sensitivity, specificity, PLR, NLR, DOR, and AUC (Fig. 3) for circulating miR-155 in the detection of BC were 0.88 (95% CI 0.76–0.95), 0.83 (95% CI 0.72–0.90), 5.2 (95% CI 2.9–9.3), 0.14 (95% CI 0.06–0.31), 37 (95% CI 11–123) and 0.92 (95% CI 0.89–0.94), respectively.
Subsequently, we attempted to explain the heterogeneity by exploring some study characteristics including sample size, sample source, and ethnicity, through meta-regression analyses. However, no significant results were identified.
Sensitivity analysis and publication bias
Afterwards, influence analysis was employed to evaluate the influence of single studies on overall estimates (Fig. 4). The random-effect bivariate model was demonstrated to be robust for the calculation of the pooled parameters by conducting the goodness of fit and bivariate normality analyses. From the illustrated results, a study from Zaleski et al. [31] was identified as an outlier. After excluding the outlier, the overall pooled sensitivity for miR-155 increased from 0.87 to 0.89, specificity increased from 0.82 to 0.83, DOR increased from 30 to 39, PLR increased from 4.8 to 5.2, NLR decreased from 0.16 to 0.13, and AUC increased from 0.91 to 0.92. Moreover, the I2 values for sensitivity and specificity were not significantly influenced by the outlier study. In general, the overall results did not show significant changes, indicating that the overall pooled estimates could not be affected by a single study.
We then used the Deeks’ funnel plot to check for publication bias. The slope coefficient presented a P value of 0.71 (Fig. 5), revealing a low likelihood of publication bias.
Functional and pathway enrichment analyses
GO functional annotation and KEGG signaling pathway enrichment analysis were performed with the genes regulated by miR-155 to obtain an in-depth and comprehensive understanding of the biological activities of miR-155. The target genes of miR-155 were retrieved from miRTarBase and then conducted with functional GO enrichment analysis. GO analysis results indicated that BP terms of the PPI network were significantly enriched in cell cycle arrest, regulation of pri-miRNA transcription, peptidyl-serine phosphorylation, small GTPase-mediated signal transduction and negative regulation of protein ubiquitination, CC terms in nucleoplasm, cytosol, nuclear chromatin, focal adhesion and extracellular exosome, and MF terms in RNA polymerase II core promoter proximal region sequence-specific DNA binding, chromatin binding, ATP binding and GTP binding (Fig. 6).
KEGG pathway enrichment analysis indicated that the genes regulated by miR-155 were significantly enriched in pathways in cancer, microRNAs in cancer, TNF signaling pathway, FoxO signaling pathway, prolactin signaling pathway, T cell receptor signaling pathway, signaling pathways regulating pluripotency of stem cells, HIF-1 signaling pathway, and PI3K-Akt signaling pathway (Fig. 7). According to previous studies and KEGG results, it could be speculated that the prolactin signaling pathway may be one of the most important pathways. The chart arising from the KEGG pathway analysis is presented at Fig. 8.
PPI network integration and selection of hub genes
We applied the STRING online database to analyze the 965 identified target genes of miR-155 and to set up a PPI network, which consisted of 907 nodes interacting with the 11.742 average numbers of neighbors. The results were downloaded and visualized by Cytoscape software for further analysis. The hub genes were selected by CytoNCA plug-in, and the top significant hub genes were obtained by betweenness centrality, closeness centrality and degree centrality methods, respectively (Fig. 9). At last, 10 hub genes were identified, including TP53, AKT1, EGFR, MYC, CTNNB1, IL6, JUN, STAT3, CASP3, and CCND1. To clarify the signal pathways associated with the hub genes during the initiation and progression of BC, we performed KEGG pathway enrichment analysis using DAVID software. The results in Fig. 7 disclosed the most vital KEGG pathways of the hub genes regulated by miR-155 associated with BC including pathways in cancer, proteoglycans in cancer, Jak-STAT signaling pathway, PI3K-Akt signaling pathway, microRNAs in cancer, MAPK signaling pathway, and FoxO signaling pathway.
Module analysis
Based on the MCODE plug-in, the top two significant modules of the above PPI network were identified and reconstructed with Cytoscape (Fig. 10). Then, KEGG pathway analysis was conducted to predicate the potential module function and cellular pathways. A list of pathways that were notably enriched by the module nodes were pathways in cancer, FoxO signaling pathway, proteoglycans in cancer, PI3K-Akt signaling pathway, prolactin signaling pathway, cell cycle, microRNAs in cancer, HIF-1 signaling pathway, focal adhesion, T cell receptor signaling pathway, TNF signaling pathway, and sphingolipid signaling pathway (Fig. 10).
Discussion
Sensitive and specific tumor biomarkers are vital for early cancer detection and diagnosis and for adopting novel therapeutic trials and prevention strategies in clinic. Increasing evidence has demonstrated that microRNAs can take part in various critical processes containing tumor cell mutation, proliferation, invasion, progression and metastasis. Abnormal microRNA expression has been identified in extensive researches, and microRNAs could function as tumor suppressor or promoter in a variety of cancers. Among all studied microRNAs, miR-155 is no doubt one of the most attractive members, which has been explored to be a potential valuable biomarker for cancer detection and survival prediction. Nevertheless, the sample sizes in most studies are relatively small. Moreover, it is still inconclusive regarding the diagnostic roles of miR-155 expression in BC. In the present study, we systematically reviewed clinical studies on the application of miR-155 for diagnosis of BC in recent years and performed this study to clarify the clinical significance of miR-155 expression in BC.
In the comprehensive analysis of diagnostic accuracy of miR-155 and BC, the pooled sensitivity and specificity of miR-155 in the detection of BC was 0.87 and 0.82, respectively. The AUC, which is recommended for evaluating the accuracy of a diagnostic test, was calculated to be 0.91, suggesting that miR-155 has relatively high clinical significance for diagnosis of BC. The value of a DOR is also an indicator of test performance; it ranges from 0 to infinity, with higher values suggesting better test identification. The pooled DOR was 30 in our study, revealing that miR-155 assays had excellent test performance in the diagnosis of BC. The overall PLR value was 4.8, revealing that the probability of having BC in a person with a positive test result was approximately fivefold higher compared to non-cancer patients. The pooled NLR was 0.16, meaning that the probability of a patient having BC is 16% if the miR-155 assay exhibits a negative result. Taken together, these results suggest a higher level of accuracy and better diagnostic characteristics for miR-155 in the diagnosis of BC than traditional detection methods.
To explore the potential sources of the heterogeneity, we carried out subgroup analysis and meta-regression analysis. Our subgroup analysis based on the sample size indicated that miR-155 with large sample size had a higher accuracy than miR-155 with small sample size, suggesting a large samples of clinical study is needed. Meanwhile, the subgroup analyses based on specimen types indicated that serum had relatively higher diagnostic value compared to plasma, indicating serum may be applied as more suitable sources of clinical specimens than others for BC detection. However, this should be interpreted with caution as levels could differ significantly if the source was serum versus plasma, because of coagulation processes. More studies should be conducted to further explore the most suitable sources of specimens. Due to the genetic heterogeneity, a difference in diagnostic accuracy has been displayed among different ethnicities. The meta-regression result revealed that these factors may not be the potential sources contributing to heterogeneity in this study. In addition, the sensitivity analysis indicated that the overall pooled estimates could not be affected by a single study.
Currently, some efforts have been devoted to exploring the functional roles of miR-155 in the pathogenic mechanism and malignant progression of cancer. However, the underlying molecular mechanisms involved in cancer progression are still largely unclear. Thus, we used some bioinformatics analysis methods to explore the functional activities of miR-155 targets involved in the occurrence and development of BC. GO analysis indicated that miR-155 was associated with some vital biological processes, basic cell components and the binding function of some key molecules. Moreover, KEGG functional enrichment analysis identified a series of important pathways, which are closely associated with the initiation and progression of BC. For example, pathways in cancer and microRNAs in cancer directly reflected the associations between miR-155 and some vital pathways involved in BC. TNF signaling has been one of the most significant cellular pathways with important functions in homeostasis and disease pathogenesis [40]. Studies have convinced the roles of FoxO signaling pathway in regulating genes essential for cell proliferation, cell death, senescence, angiogenesis, cell migration and metastasis [41]. The prolactin pathway, known as a hormone involved in normal breast development and lactation, has been long demonstrated to be involved in the progression of human breast cancer [42]. It has been confirmed that T cell receptor signaling pathway has important roles in the host adaptive immune system and T cell-based adoptive immunotherapy has been shown to be a promising treatment for various types of cancers. The abnormal expression of T cell receptor may lead to carcinogenesis [43]. HIF-1 has been recognized as an essential component in changing the transcriptional response of tumors under hypoxia and plays important roles in crucial aspects of cancer biology, including angiogenesis, cell survival, glucose metabolism and invasion [44]. PI3K-Akt signaling pathway was demonstrated to be hyperactivated in most of the breast carcinomas and could regulate various biological processes such as cell growth, differentiation, migration, and survival, as well as angiogenesis and metabolism [45]. Moreover, previous evidence has reported that microRNAs have critical roles in cellular activities implicated in BC cell growth, migration and metastasis by targeting the PI3K/AKT oncogenic signaling pathway [46]. These functional enrichment results may help us understand the mechanisms of miR-155 during the initiation and progression of BC.
PPI analysis has been recognized as an entry point for better interpretation of associations between different proteins on a genome-wide scale, and might be helpful to provide novel insights into protein function explanation. We constructed the PPI network with the target genes of miR-155 and identified the top ten hub genes. These hub genes were predominantly associated with some important pathways, most of which have been confirmed to be related to BC. In addition, there is growing evidence that proteoglycans can adjust extensive normal and pathological activities, such as morphogenesis, tissue repair, inflammation, vascularization and cancer metastasis [47]. Rapidly emerging ground-breaking discoveries have revealed that Jak-STAT signaling pathway regulates almost all immune regulatory processes, containing those that are involved in tumor cell recognition and tumor-driven immune escape [48]. Alterations in this pathway may contribute to the development of BC and metastatic spread and targeting of JAK-STAT signaling in BC may provide potential therapeutic strategies to overcome drug resistance [49]. The MAPK signaling pathway plays an important part in organizing great constitution network that mediates a variety of physiological processes, such as cell growth, differentiation, as well as apoptotic cell death and dysregulation of the MAPK signaling cascades has been confirmed to be associated with the pathogenesis of various human cancer types including BC [50, 51]. The hub genes may provide potential predictor and therapeutic targets for BC patients.
The module analysis of the constructed PPI network was then carried out and the top two significant modules were selected for conducting KEGG pathway enrichment analysis of the genes included. The enriched results indicated that the nodes involved in the modules were highly related to some vital pathways. In addition to the most pathways which were already discussed above, cell cycle has been identified as a hallmark of cancer involved in cellular proliferation and cancer development. The dysregulation of cell cycle control may lead to cancer progression [52]. Moreover, focal adhesion kinase (FAK) is a cytoplasmic non-receptor tyrosine kinase involved in almost every aspect of cancer, from invasion to metastasis to epithelial–mesenchymal transition (EMT) and maintenance of cancer stem cells [53]. Sphingolipids have been the largest family of bioactive lipids that participate in a wide variety of biological mechanisms, including cell death and proliferation and are linked with numerous aspects of tumorigenesis [54]. The PPI analysis further uncovered the potential mechanisms of miR-155 involved in BC occurrence and development. However, further studies are still urgently demanded to validate the hub genes and pathways, and deeper mechanisms would be uncovered.
We should point out that there are some limitations in our study. First, the cutoff values of miR-155 in the enrolled diagnostic tests were not uniform, which might contribute to the observed heterogeneity. More studies should be conducted to reach an appropriate cutoff value to be accurate enough. Second, the sample sizes for plasma, whole blood, tissue and urine are relatively small. It is difficult to make a definitive conclusion about the accuracy of miR-155 in diagnosing BC from these sample sources. Thus, further evaluations of the utility of miR-155 from different sample sources as biomarkers for BC detection should be performed. Third, the lack of studies for different ethnicities evaluating the miR-155 diagnostic performance complicated the exploration of its clinical application. More studies should be conducted to evaluate the diagnostic accuracy of miR-155 for different ethnicities. In addition, due to the fact that all studies that are used for the data pooling have patients of various TNM stage (from stage I to stage IV), there is considerable heterogeneity in detection of BC. Without patient-level data, it would be very difficult to tell whether miR-155 is actually useful as a marker in early detection of BC. Besides, BRCA1-deficient tumors have been shown to have increased levels of miR-155, which could be relevant for identifying possible respondents to PARP-1 inhibitors [55, 56]. Most of the mechanistic studies on miR-155 and BC have been done on these patients, which was ignored in this study. Last, due to lack of experiment verification, the function and mechanisms of miR-155 need to be further studied in vivo and in vitro.
Despite all this, we revealed that miR-155 had great potential in early diagnosis for BC and further studies to explore its clinical application are warranted and worthwhile. Some interesting findings will stimulate further research with rigid criteria and large study populations to resolve any remaining controversy of the diagnostic value of miR-155 in BC patients. The bioinformatics analysis results definitely point out the molecular basis to better understand the pathogenesis of miR-155 involved in BC and provide valuable novel markers or targets for the diagnosis and treatment of BC.
Conclusion
In conclusion, the present study according to published articles demonstrated that miR-155 has strong potential to serve as a novel noninvasive biomarker for detection of BC. In addition, integrated bioinformatics analysis identified some key hub genes and pathways, which may help understand why miR-155 could possess excellent biomarker characteristics and provide some new clues to explore the exact etiology and mechanism underlying the initiation and progression of BC. To strengthen our findings, the diagnostic value of miR-155 in BC should be further confirmed by large-scale and standard investigations. Moreover, further studies are urgently demanded to validate the hub genes and pathways, and further mechanisms would be uncovered.
Data availability
The data supporting the conclusions of this article are within the article.
Abbreviations
- BC:
-
Breast cancer
- AUC:
-
Area under the curve
- TP:
-
True positive
- FP:
-
False positive
- FN:
-
False negative
- TN:
-
True negative
- PLR:
-
Positive likelihood ratios
- NLR:
-
Negative likelihood ratios
- DOR:
-
Diagnostic odds ratio
- CI:
-
Confidence interval
- SROC:
-
Summary receiver operator characteristic
- QUADAS-2:
-
Quality Assessment of Diagnostic Accuracy Studies 2
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PPI:
-
Protein–protein interaction
- DAVID:
-
Database for Annotation, Visualization, and Integrated Discovery
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30.
DeSantis CE, Ma J, Gaudet MM, Newman LA, Miller KD, Goding Sauer A, Jemal A, Siegel RL. Breast cancer statistics, 2019. CA Cancer J Clin. 2019;69(6):438–51.
Schunemann HJ, Lerda D, Quinn C, Follmann M, Alonso-Coello P, Rossi PG, Lebeau A, Nystrom L, Broeders M, Ioannidou-Mouzaka L, Duffy SW, Borisch B, Fitzpatrick P, Hofvind S, Castells X, Giordano L, Canelo-Aybar C, Warman S, Mansel R, Sardanelli F, Parmelli E, Grawingholt A, Saz-Parkinson Z, European Commission Initiative on Breast Cancer Contributor G. Breast cancer screening and diagnosis: a synopsis of the European breast guidelines. Ann Intern Med. 2020;172(1):46–56.
McGeary SE, Lin KS, Shi CY, Pham TM, Bisaria N, Kelley GM, Bartel DP. The biochemical basis of microRNA targeting efficacy. Science. 2019;366:6472.
Treiber T, Treiber N, Meister G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat Rev Mol Cell Biol. 2019;20(1):5–20.
He Y, Deng F, Yang S, Wang D, Chen X, Zhong S, Zhao J, Tang J. Exosomal microRNA: a novel biomarker for breast cancer. Biomark Med. 2018;12(2):177–88.
Wu C, Liu Q, Liu B. MicroRNA-155 hallmarks promising accuracy for the diagnosis of various carcinomas: results from a meta-analysis. Dis Markers. 2015;2015:327287.
Hou Y, Wang J, Wang X, Shi S, Wang W, Chen Z. Appraising MicroRNA-155 as a Noninvasive Diagnostic Biomarker for Cancer Detection: A Meta-Analysis. Medicine (Baltimore). 2016;95(2):e2450.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, Leeflang MM, Sterne JA, Bossuyt PM, Group Q-. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155(8):529–536.
Liu Y, Chen Y, Chu H. A unification of models for meta-analysis of diagnostic accuracy studies without a gold standard. Biometrics. 2015;71(2):538–47.
Rosman AS, Korsten MA. Application of summary receiver operating characteristics (sROC) analysis to diagnostic clinical testing. Adv Med Sci. 2007;52:76–82.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327(7414):557–60.
Deeks JJ, Macaskill P, Irwig L. The performance of tests of publication bias and other sample size effects in systematic reviews of diagnostic test accuracy was assessed. J Clin Epidemiol. 2005;58(9):882–93.
Huang HY, Lin YC, Li J, Huang KY, Shrestha S, Hong HC, Tang Y, Chen YG, Jin CN, Yu Y, Xu JT, Li YM, Cai XX, Zhou ZY, Chen XH, Pei YY, Hu L, Su JJ, Cui SD, Wang F, Xie YY, Ding SY, Luo MF, Chou CH, Chang NW, Chen KW, Cheng YH, Wan XH, Hsu WL, Lee TY, Wei FX, Huang HD. miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res. 2020;48(D1):D148–54.
Hinderer EW 3rd, Flight RM, Dubey R, MacLeod JN, Moseley HNB. Advances in gene ontology utilization improve statistical power of annotation enrichment. PLoS ONE. 2019;14(8):e0220728.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30.
Dennis G Jr, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4(5):P3.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, Jensen LJ, Mering CV. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13.
Otasek D, Morris JH, Boucas J, Pico AR, Demchak B. Cytoscape automation: empowering workflow-based network analysis. Genome Biol. 2019;20(1):185.
Tang Y, Li M, Wang J, Pan Y, Wu FX. CytoNCA: a cytoscape plugin for centrality analysis and evaluation of protein interaction networks. Biosystems. 2015;127:67–72.
Bader GD, Hogue CW. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics. 2003;4:2.
Sun Y, Wang M, Lin G, Sun S, Li X, Qi J, Li J. Serum microRNA-155 as a potential biomarker to track disease in breast cancer. PLoS ONE. 2012;7(10):e47003.
Eichelser C, Flesch-Janys D, Chang-Claude J, Pantel K, Schwarzenbach H. Deregulated serum concentrations of circulating cell-free microRNAs miR-17, miR-34a, miR-155, and miR-373 in human breast cancer development and progression. Clin Chem. 2013;59(10):1489–96.
Mar-Aguilar F, Mendoza-Ramirez JA, Malagon-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, Rodriguez-Padilla C, Resendez-Perez D. Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers. 2013;34(3):163–9.
Erbes T, Hirschfeld M, Rucker G, Jaeger M, Boas J, Iborra S, Mayer S, Gitsch G, Stickeler E. Feasibility of urinary microRNA detection in breast cancer patients and its potential as an innovative non-invasive biomarker. BMC Cancer. 2015;15:193.
Shaker O, Maher M, Nassar Y, Morcos G, Gad Z. Role of microRNAs -29b-2, -155, -197 and -205 as diagnostic biomarkers in serum of breast cancer females. Gene. 2015;560(1):77–82.
Gao S, Wang Y, Wang M, Li Z, Zhao Z, Wang RX, Wu R, Yuan Z, Cui R, Jiao K, Wang L, Ouyang L, Liu R. MicroRNA-155, induced by FOXP3 through transcriptional repression of BRCA1, is associated with tumor initiation in human breast cancer. Oncotarget. 2017;8(25):41451–64.
Han JG, Jiang YD, Zhang CH, Yang YM, Pang D, Song YN, Zhang GQ. A novel panel of serum miR-21/miR-155/miR-365 as a potential diagnostic biomarker for breast cancer. Ann Surg Treat Res. 2017;92(2):55–66.
Fan T, Mao Y, Sun Q, Liu F, Lin JS, Liu Y, Cui J, Jiang Y. Branched rolling circle amplification method for measuring serum circulating microRNA levels for early breast cancer detection. Cancer Sci. 2018;109(9):2897–906.
Huang SK, Luo Q, Peng H, Li J, Zhao M, Wang J, Gu YY, Li Y, Yuan P, Zhao GH, Huang CZ. A panel of serum noncoding RNAs for the diagnosis and monitoring of response to therapy in patients with breast cancer. Med Sci Monit. 2018;24:2476–88.
Zaleski M, Kobilay M, Schroeder L, Debald M, Semaan A, Hettwer K, Uhlig S, Kuhn W, Hartmann G, Holdenrieder S. Improved sensitivity for detection of breast cancer by combination of miR-34a and tumor markers CA 15–3 or CEA. Oncotarget. 2018;9(32):22523–36.
Arabkari V, Clancy E, Dwyer RM, Kerin MJ, Kalinina O, Holian E, Newell J, Smith TJ. Relative and absolute expression analysis of MicroRNAs associated with luminal a breast cancer- a comparison. Pathol Oncol Res. 2020;26(2):833–44.
Soleimanpour E, Babaei E, Hosseinpour-Feizi MA, Montazeri V. Circulating miR-21 and miR-155 as potential noninvasive biomarkers in Iranian Azeri patients with breast carcinoma. J Cancer Res Ther. 2019;15(5):1092–7.
Song X, Liu Z, Yu Z. LncRNA NEF is downregulated in triple negative breast cancer and correlated with poor prognosis. Acta Biochim Biophys Sin (Shanghai). 2019;51(4):386–92.
Swellam M, Ramadan A, El-Hussieny EA, Bakr NM, Hassan NM, Sobeih ME, EzzElArab LR. Clinical significance of blood-based miRNAs as diagnostic and prognostic nucleic acid markers in breast cancer: Comparative to conventional tumor markers. J Cell Biochem. 2019;120(8):12321–30.
Swellam M, Zahran RFK, Abo El-Sadat Taha H, El-Khazragy N, Abdel-Malak C. Role of some circulating MiRNAs on breast cancer diagnosis. Arch Physiol Biochem. 2019;125(5):456–64.
Bitaraf A, Babashah S, Garshasbi M. Aberrant expression of a five-microRNA signature in breast carcinoma as a promising biomarker for diagnosis. J Clin Lab Anal. 2020;34(2):e23063.
Raeisi F, Mahmoudi E, Dehghani-Samani M, Hosseini SSE, Ghahfarrokhi AM, Arshi A, Forghanparast K, Ghazanfari S. Differential expression profile of miR-27b, miR-29a, and miR-155 in chronic lymphocytic leukemia and breast cancer patients. Mol Ther Oncolytics. 2020;16:230–7.
Peng Q, Zhao P, Shen Y, Cheng M, Wu Y, Zhu Y. Prognostic implication and functional exploration for microRNA-20a as a molecular biomarker of gastrointestinal cancer. BMC Cancer. 2020;20(1):420.
Dostert C, Grusdat M, Letellier E, Brenner D. The TNF Family of Ligands and Receptors: Communication Modules in the Immune System and Beyond. Physiol Rev. 2019;99(1):115–60.
Jiramongkol Y, Lam EW. FOXO transcription factor family in cancer and metastasis. Cancer Metastasis Rev. 2020;39(3):681–709.
Urbaniak A, Jablonska K, Podhorska-Okolow M, Ugorski M, Dziegiel P. Prolactin-induced protein (PIP)-characterization and role in breast cancer progression. Am J Cancer Res. 2018;8(11):2150–64.
Chandran SS, Klebanoff CA. T cell receptor-based cancer immunotherapy: emerging efficacy and pathways of resistance. Immunol Rev. 2019;290(1):127–47.
Kachamakova-Trojanowska N, Podkalicka P, Bogacz T, Barwacz S, Jozkowicz A, Dulak J, Loboda A. HIF-1 stabilization exerts anticancer effects in breast cancer cells in vitro and in vivo. Biochem Pharmacol. 2020;175:113922.
Hoxhaj G, Manning BD. The PI3K-AKT network at the interface of oncogenic signalling and cancer metabolism. Nat Rev Cancer. 2020;20(2):74–88.
Rahmani F, Ferns GA, Talebian S, Nourbakhsh M, Avan A, Shahidsales S. Role of regulatory miRNAs of the PI3K/AKT signaling pathway in the pathogenesis of breast cancer. Gene. 2020;737:144459.
Espinoza-Sanchez NA, Gotte M. Role of cell surface proteoglycans in cancer immunotherapy. Semin Cancer Biol. 2020;62:48–67.
Owen KL, Brockwell NK, Parker BS. JAK-STAT Signaling: A Double-Edged Sword of Immune Regulation and Cancer Progression. Cancers (Basel). 2019;11:12.
Tabassum S, Abbasi R, Ahmad N, Farooqi AA. Targeting of JAK-STAT signaling in breast cancer: therapeutic strategies to overcome drug resistance. Adv Exp Med Biol. 2019;1152:271–81.
Dwyer AR, Truong TH, Ostrander JH, Lange CA. Steroid receptors as MAPK signaling sensors in breast cancer: let the fates decide. J Mol Endocrinol. 2020;65(1):T35–48.
Lee S, Rauch J, Kolch W. Targeting MAPK signaling in cancer: mechanisms of drug resistance and sensitivity. Int J Mol Sci. 2020;21:3.
Ding L, Cao J, Lin W, Chen H, Xiong X, Ao H, Yu M, Lin J, Cui Q. The roles of cyclin-dependent kinases in cell-cycle progression and therapeutic strategies in human breast cancer. Int J Mol Sci. 2020;21:6.
Zhou J, Yi Q, Tang L. The roles of nuclear focal adhesion kinase (FAK) on cancer: a focused review. J Exp Clin Cancer Res. 2019;38(1):250.
Samaha D, Hamdo HH, Wilde M, Prause K, Arenz C. Sphingolipid-transporting proteins as cancer therapeutic targets. Int J Mol Sci. 2019;20:14.
Chang S, Wang RH, Akagi K, Kim KA, Martin BK, Cavallone L, Kathleen Cuningham Foundation Consortium for Research into Familial Breast C, Haines DC, Basik M, Mai P, Poggi E, Isaacs C, Looi LM, Mun KS, Greene MH, Byers SW, Teo SH, Deng CX, Sharan SK. Tumor suppressor BRCA1 epigenetically controls oncogenic microRNA-155. Nat Med. 2011;17(10):1275–82.
Pasculli B, Barbano R, Fontana A, Biagini T, Di Viesti MP, Rendina M, Valori VM, Morritti M, Bravaccini S, Ravaioli S, Maiello E, Graziano P, Murgo R, Copetti M, Mazza T, Fazio VM, Esteller M, Parrella P. Hsa-miR-155-5p up-regulation in breast cancer and its relevance for treatment with poly[ADP-Ribose] polymerase 1 (PARP-1) inhibitors. Front Oncol. 2020;10:1415.
Acknowledgements
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
LX collected and processed data, and drafted the manuscript. CQ took part in the statistical analysis, while WH and QH were responsible for the bioinformatics analysis. JY drafted and revised the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no competing interest.
Consent for publication
All the authors approve the publication of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Liu, X., Chang, Q., Wang, H. et al. Discovery and function exploration of microRNA-155 as a molecular biomarker for early detection of breast cancer. Breast Cancer 28, 806–821 (2021). https://doi.org/10.1007/s12282-021-01215-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-021-01215-2