Abstract
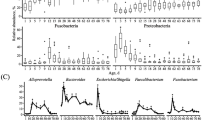
Diarrhea is a fatal disease to neonatal calves, and rotavirus is the main pathogen associated with neonatal calf diarrhea. Although previous studies have reported that the gut microbiota is changed in calves during diarrhea, less is known about whether rotavirus infection alters the structure of the gut microbiota. Here, we characterized fecal microbial communities and identified possible relationships between the gut microbiota profiles and physiological parameters. Five fecal specimens of rotavirus-infected calves from 1 to 30 days after birth and five fecal specimens of age-matched healthy calves were used for the microbial community analysis using the Illumina MiSeq sequencer. Rotavirus infection was associated with reduced rotavirus infection significantly reduced the richness and diversity of the bacterial community. Weighted unique fraction metric analysis exhibited significant differences in community membership and structure between healthy and rotavirus-infected calves. Based on relative abundance analysis and linear discriminant analysis effect size, we found that the representative genera from Lactobacillus, Subdoligranulum, Blautia, and Bacteroides were closely related to healthy calves, while the genera Escherichia and Clostridium were closely affiliated to rotavirus-infected calves. Furthermore, canonical correlation analysis and Pearson correlation coefficient results revealed that the increased relative abundances of Lactobacillus, Subdoligranulum, and Bacteroides were correlated with normal levels of physiological characteristics such as white blood cells, blood urea nitrogen, serum amyloid protein A, and glucose concentration in serum. These results suggest that rotavirus infection alters the structure of the gut microbiota, correlating changes in physiological parameters. This study provides new information on the relationship between gut microbiota and the physiological parameters of rotavirus-mediated diarrheic calves.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ammar, S.S., Mokhtaria, K., Tahar, B.B., Amar, A.A., Redha, B.A., Yuva, B., Mohamed, H.S., Abdellatif, N., and Laid, B. 2014. Prevalence of rotavirus (GARV) and coronavirus (BCoV) associated with neonatal diarrhea in calves in western Algeria. Asian Pac. J. Trop. Biomed. 4, S318–S322.
Cho, Y.I., Kim, W.I., Liu, S., Kinyon, J.M., and Yoon, K.J. 2010. Development of a panel of multiplex real-time polymerase chain reaction assays for simultaneous detection of major agents causing calf diarrhea in feces. J. Vet. Diagn. Invest. 22, 509–517.
Cho, Y.I. and Yoon, K.J. 2014. An overview of calf diarrhea - infectious etiology, diagnosis, and intervention. J. Vet. Sci. 15, 1–17.
Edrington, T.S., Dowd, S.E., Farrow, R.F., Hagevoort, G.R., Callaway, T.R., Anderson, R.C., and Nisbet, D.J. 2012. Development of colonic microflora as assessed by pyrosequencing in dairy calves fed waste milk. J. Dairy Sci. 95, 4519–4525.
Gomez, D.E., Arroyo, L.G., Costa, M.C., Viel, L., and Weese, J.S. 2017. Characterization of the fecal bacterial microbiota of healthy and diarrheic dairy calves. J. Vet. Intern. Med. 31, 928–939.
Gulliksen, S.M., Jor, E., Lie, K.I., Hamnes, I.S., Loken, T., Akerstedt, J., and Osteras, O. 2009. Enteropathogens and risk factors for diarrhea in Norwegian dairy calves. J. Dairy Sci. 92, 5057–5066.
Hur, T.Y., Jung, Y.H., Choe, C.Y., Cho, Y.I., Kang, S.J., Lee, H.J., Ki, K.S., Baek, K.S., and Suh, G.H. 2013. The dairy calf mortality: the causes of calf death during ten years at a large dairy farm in Korea. Korean J. Vet. Res. 53, 103–108.
Izzo, M.M., Kirkland, P.D., Mohler, V.L., Perkins, N.R., Gunn, A.A., and House, J.K. 2011. Prevalence of major enteric pathogens in Australian dairy calves with diarrhoea. Aust. Vet. J. 89, 167–173.
Kapikian, A.Z. and Chanock, R.M. 1996. Rotaviruses, pp. 1657–1708. In Straus, S.E. (ed.), Fields Virology Vol. 2. Lippincott-Raven, Philadelphia, USA.
Klein-Jobstl, D., Schornsteiner, E., Mann, E., Wagner, M., Drillich, M., and Schmitz-Esser, S. 2014. Pyrosequencing reveals diverse fecal microbiota in simmental calves during early development. Front. Microbiol. 5, 622.
Kostic, A.D., Xavier, R.J., and Gevers, D. 2014. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology 146, 1489–1499.
Li, R.W., Connor, E.E., Li, C., Baldwin Vi, R.L., and Sparks, M.E. 2012. Characterization of the rumen microbiota of pre-ruminant calves using metagenomic tools. Environ. Microbiol. 14, 129–139.
Lukas, F., Koppova, I., Kudrna, V., and Kopecny, J. 2007. Postnatal development of bacterial population in the gastrointestinal tract of calves. Folia Microbiol. 52, 99–104.
Margreiter, M., Ludl, K., Phleps, W., and Kaehler, S.T. 2006. Therapeutic value of a Lactobacillus gasseri and Bifidobacterium longum fixed bacterium combination in acute diarrhea: a randomized, double-blind, controlled clinical trial. Int. J. Clin. Pharmacol. Ther. 44, 207–215.
Millar, H.R., Simpson, J.G., and Stalker, A.L. 1971. An evaluation of the heat precipitation method for plasma fibrinogen estimation. J. Clin. Pathol. 24, 827–830.
Oikonomou, G., Teixeira, A.G., Foditsch, C., Bicalho, M.L., Machado, V.S., and Bicalho, R.C. 2013. Fecal microbial diversity in preweaned dairy calves as described by pyrosequencing of metagenomic 16S rDNA. Associations of Faecalibacterium species with health and growth. PLoS One 8, e63157.
Penders, J., Thijs, C., Vink, C., Stelma, F.F., Snijders, B., Kummeling, I., van den Brandt, P.A., and Stobberingh, E.E. 2006. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 118, 511–521.
Rolhion, N. and Chassaing, B. 2016. When pathogenic bacteria meet the intestinal microbiota. Philos. Trans. R. Soc. Lond. B Biol. Sci. 371, 20150504.
Singh, P., Teal, T.K., Marsh, T.L., Tiedje, J.M., Mosci, R., Jernigan, K., Zell, A., Newton, D.W., Salimnia, H., Lephart, P., et al. 2015. Intestinal microbial communities associated with acute enteric infections and disease recovery. Microbiome 3, 45.
Uetake, K. 2013. Newborn calf welfare: a review focusing on mortality rates. Animal Sci. J. 84, 101–105.
USDA. 2008. Dairy 2007 Part II: changes in the US dairy cattle industry, 1991–2007. Fort Collins: USDA-APHIS-VS, CEAH, 57–61.
Uyeno, Y., Sekiguchi, Y., and Kamagata, Y. 2010. rRNA-based analysis to monitor succession of faecal bacterial communities in Holstein calves. Lett. Appl. Microbiol. 51, 570–577.
Wotzka, S.Y., Nguyen, B.D., and Hardt, W.D. 2017. Salmonella Typhimurium diarrhea reveals basic principles of enteropathogen infection and disease-promoted DNA exchange. Cell Host Microbe 21, 443–454.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Jang, JY., Kim, S., Kwon, MS. et al. Rotavirus-mediated alteration of gut microbiota and its correlation with physiological characteristics in neonatal calves. J Microbiol. 57, 113–121 (2019). https://doi.org/10.1007/s12275-019-8549-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-019-8549-1