Abstract
Genetic tools, which can be used for the morphology study of specific neurons, pathway-selective connectome mapping, neuronal activity monitoring, and manipulation with a spatiotemporal resolution, have been widely applied to the understanding of complex neural circuit formation, interactions, and functions in rodents. Recently, similar genetic approaches have been tried in non-human primates (NHPs) in neuroscience studies for dissecting the neural circuits involved in sophisticated behaviors and clinical brain disorders, although they are still very preliminary. In this review, we introduce the progress made in the development and application of genetic tools for brain studies on NHPs. We also discuss the advantages and limitations of each approach and provide a perspective for using genetic tools to study the neural circuits of NHPs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thanks to recent advancements in molecular biology, cell biology, synthetic biology, gene editing, and virus studies, several powerful genetic tools have been developed for specific neuron labeling, circuit tracing, neuronal activity recording, and perturbing with high spatiotemporal resolution. The development of gene-editing methods makes it possible and efficient to generate sizable transgenic animal lines that target specific cell types. Various viral vectors with unique tropisms play crucial roles in delivering genetic material to different cell types and mediating anterograde/retrograde transport among neurons. Engineered opsins and designer receptors exclusively activated by designer drugs (DREADDs) enable the reversible control of activity in specific neuronal populations. Calcium indicators, equipped with microscopes, enable long-term and large-scale tracking of the activities of distinct cell types. The widespread application of these genetic tools in small animals such as rodents, zebrafish, and Drosophila, has allowed the precise dissection of neural circuits and has largely revolutionized the research paradigm in neuroscience.
Non-human primates (NHPs) are important animal models in neuroscience research, as they have brain network organization, cognitive circuits, brain structures (e.g., enlargement of the prefrontal cortex), and a proportion of neocortex similar to humans [1, 2]. In addition, NHPs are more similar to humans at cellular-molecular levels than rodents. For instance, a population of slow-spiking neurons expressing nitric oxide synthase (nNOS), neuropeptide Y (NPY), and GABAA receptor α1 subunit (GABAAR-α1) -"ivy cells" - are abundant in the neocortex of primates, while in mice they are predominantly located in the hippocampus [3, 4]. The structural basis for supporting such a considerable operating system is the complex brain that consists of billions of neurons, trillions of synaptic connections, and specialized neural circuits. Efforts in unraveling the neural mechanisms of brain physiology and pathology in NHPs could deepen our understanding of the human brain. However, brain studies on NHPs are still limited. Traditional electrophysiology, pharmacological intervention, functional magnetic resonance imaging (fMRI), and intrinsic imaging have been widely used in NHPs to reveal the correlations and causal relationships between specific behaviors and neuronal activity. Yet, traditional methods lack the ability to precisely identify cells and are limited in how they can modulate and record cells. Genetic tools, as more powerful neuroscience approaches, have been lagging in NHPs due to various limitations. Here, we review the genetic tools and related advances that have been applied to primates, including cell-type targeting approaches, viral vector delivery technology, and genetic tools for neuronal activity manipulation and tracking. We believe that genetic approaches have the potential to open new horizons and to reveal the wiring logic and inner mechanisms of neural circuits involved in complex physiological and pathological functions.
Cell-Type Specific Targeting in Monkey
A fundamental technical problem in neuroscience studies is the fact that the nervous system is highly heterogeneous, consisting of many intermixed cells. Distinct cell types cannot be identified by a single criterion. Common classification criteria are molecular, morphological, and neurophysiological properties. The neural underpinnings of behaviors, including the information processing of parallel, convergent, divergent, feedback, and streamlining, are maintained by the assembly and the interactions of distinct cell types in the mammalian brain. Gaining access to different cell types is necessary in order to better understand how the brain works in NHPs. Recently, the rapid development of sequencing technology has provided scientists with a resource to finely identify cell types and gene regulatory networks. Fueled by innovations in genome editing techniques, genetic tools have a great inherent potential to distinguish and study specific cell types. Here, we discuss cell-type-targeting methods in primates and look ahead to new approaches to gain insight into the complexity of mesoscopic neural circuits in primates (Fig. 1).
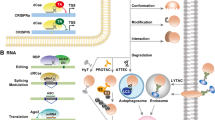
Genetic cell-type-targeting strategies via viral vectors. A Principle overview of several genetic strategies to target cell types. Promoter/enhancer-based approaches work at the gene transcription stage. Generally, the promoter (dark blue box) determines the transcription initiation sites, while the enhancer (sapphire blue box) determines the transcription efficiency of genes. Enhancers activated by specific TFs move close to their target promoter through a looping mechanism, and their interaction is stabilized by cohesion proteins (gray balls). RNA sensor/mAGNET work at the gene translation stage. In programmable RNA sensor tools, sesRNA hybridizes to target endogenous mRNA, and ADAR enzymes (orange ball) repair mismatched sites of the stop codon (red circle) switching on the translation of required proteins. For mAGNET, abundant exogenous miRNAs recruit the RISC (dark green sphere) and inhibit nonspecific protein translation by hybridizing to their complementary miRNA in off-target cells. B Sequence elements of four targeting approaches. Red scissors, repair sites; red circle, stop codon; orange circle, self-cleaving peptide T2A; TFs, transcription factors; Pol II, RNA polymerase II; mAGNET, miRNA-guided neuron tag; RISC, RNA silencing complex.
Cre Knock-in-Based Cell-Type Targeting
Conventional Cre knock-in lines, generated from embryonic stem cells by gene targeting, make it possible to target specific cell types reliably, revolutionizing the study of neural circuits in mice [5, 6]. The development of genome editing methods (zinc-finger nucleases, transcription activator-like effector proteins, and the CRISPR/Cas9 system) allows the generation of gene-modified monkeys [7, 8]. Gene-modified monkey disease models carrying indels or point mutations by CRISPR/Cas9 or a CRISPR-based base editor have been reported in many studies with high efficiency [9, 10]. Yet, the efficiency of precise large-fragment knock-in via homologous recombination in embryo gene editing is still relatively low, making it acceptable to generate knock-in mouse models, but not knock-in monkey models. To date, no Cre knock-in monkey model has been reported due to the low efficiency, high cost, and limited available resources. Alternative approaches, such as exogenous gene expression via local injection of viral vectors driven by promoters, enhancers, and other regulators have been so far considered more feasible. We recommend that efforts be made towards improving the efficiency of this approach and developing a long-term plan for the generation of Cre knock-in monkey models.
Promoter-Based Cell-Type Targeting
A promoter is a DNA sequence upstream of a gene where relevant proteins, such as RNA polymerase II (RNAPII) and general transcription factors bind to the transcription start site of that gene. The function of the promoter is generally thought to be to determine the initial transcription location and direction of a single RNA transcript. The endogenous expression pattern of a gene of interest can be mimicked by inserting exogenous genes after a known promoter of this gene. Although the expression patterns of exogenous genes are far from satisfactory in vivo, the promoter-driven cell-type-targeting approach remains the simplest and quickest way to target specific neurons in monkeys today.
The promoter most widely used in monkeys is the Ca2+/calmodulin-dependent protein kinase II alpha (CaMKIIα). This gene is mainly expressed in excitatory neurons and its promoter preferentially targets the same cell type, enabling high-level transgene expression in excitatory neurons, especially in layer 5 of the neocortex [11, 12]. The human synapsin-1 promoter (hSyn) has an expression pattern similar to the CaMKIIα promoter in monkeys, resulting in particularly robust expression in the deep layers [11–14]. Other studies have also selected fused cytomegalovirus (CMV), human thymocyte-1(hThy-1), or the CMV enhancer, chicken β-actin promoter, and rabbit β-globin splice acceptor site (CAG) as the ubiquitous promoter [14]. In addition to the general cell type, the tyrosine hydroxylase (TH) promoter, coupled with the Cre/loxP system can be used to target TH+ neurons in the locus coeruleus and substantia nigra of primates [15]. The L7/Pcp2 promoter can be used to track Purkinje cells in the cerebellum of monkeys [16]. This strategy depends on the specificity of a native promoter. However, the expression pattern of most genes is not promoter dependent, leading to unspecific labeling in the promoter-driven cell-targeting approach. Indeed, the specificity of many promoters is likely the result of random integration next to highly specific cis-regulatory elements [17]. In conclusion, the promoter-driven cell-type-targeting approach can achieve some degree of specificity, but such specificity is relatively modest and largely falls short in the finer targeting of specific cell types.
Enhancer-Based Cell-Type Targeting
Investigations of eukaryotic transcriptional regulation have revealed that the same gene can be expressed in distinct cell types via the activation and inactivation of different sets and combinations of enhancers or other cis-regulatory elements [12, 13]. Enhancers, typical genomic cis-regulatory elements, can modulate gene expression patterns in different tissues irrespective of their orientation or distance by a looping mechanism [18]. Recent progress in profiling the landscape of accessible chromatin of single cells has dramatically expanded our ability to identify enhancers [19]. Enhancer-driven gene expression (EDGE) can mimic endogenous marker gene expression patterns and transfer artificial genes into specific cell types by hijacking enhancers. Several studies have shown that EDGE can be used to generate useful genetic tools for accessing specific cell subclasses in rodents (e.g., layer 5 pyramidal tract neurons) [17, 20,21,22,23,24,25]. There are also a few reports on monkeys. A collection of parvalbumin (Pvalb) enhancers showing highly specific access to Pvalb cells in the primate neocortex has been defined by Mich et al. [26]. The mouse Dlx5/6 enhancer has provided a powerful tool for GABAergic neuron manipulation in macaques [27]. An intersectional strategy using the human Dlx5/6 enhancer and the somatostatin (Sst) promoter has confined the expression of exogenous proteins to Sst+ interneurons in the marmoset cortex and the rhesus macaque cortex [28]. Currently, the enhancer-based cell-type targeting system is the most powerful and simple genetic tool available to target specific cell types in primates. Future research towards building up a comprehensive toolkit of enhancers for precise cell-type labeling in monkeys would add another layer of sophistication to basic research on the NHP brain, and truly deepen our understanding of the assembly and organization of neural circuits and their operating principles.
Programmable RNA-based Cell-Type Targeting
Engineered RNA-specific sensors have recently been shown to be promising for directly targeting cell types at endogenous mRNA levels. Several laboratories have reported independently programmable RNA sensing systems that harness endogenous adenosine deaminases acting on RNA (ADAR) repair systems to detect intracellular transcripts and target specific cell types [29,30,31,32]. In these systems, a programmable RNA sensor encodes the exogenous protein for which transcription is interrupted by artificially inserted stop codons in the context of a sequence complementary to the target transcript of the genes of interest. Only in the presence of target transcripts in a certain type of neuron, the sense–edit–switch RNA (sesRNA) would hybridize with the target mRNA and form double-stranded RNA molecules (dsRNA). The ADAR enzyme is then recruited to convert A to I at the mismatch sequence, switching on the expression of reporter proteins. The capacity of sesRNA in operating and tracking the activities of specific cell types has been proven in cultured cell lines, behaving mice, and human brain slices. Such a system, which bypasses complex and indirect DNA transcription processes, provides a relatively flexible and straightforward approach to target cell subclasses across species, especially for monkeys and humans [33].
MicroRNA-Based Cell-Type targeting
The microRNA (miRNA)-based genetic system can also operate at endogenous mRNA levels. This approach – miRNA-guided neuron tag (mAGNET) – suppresses transgene expression in off-target cell populations to enable gene expression in specific cell types. Specific miRNA from off-target cells or tissues recruits the RNA-silencing complex (RISC) and degrades the transcripts by hybridizing to their complementary miRNA recognition sites incorporated in viral vectors. Conversely, low levels of miRNA in on-target cells have little impact on the translation of exogenous proteins. This strategy has been successfully applied to identify and label interneurons in rodents [34]. In addition, the miRNA-mediated de-targeting system is also used to down-regulate the transcripts of exogenous delivery proteins in the dorsal root ganglion (DRG) of primates with the aim of reducing toxicity and the immune response of AAV-mediated gene therapies [35]. Although not commonly used, the miRNA-mediated de-targeting approach offers another option for researchers to label general cell populations.
In summary, targeting specific cell types in NHPs remains a daunting challenge. Viral-mediated cell targeting methods can overcome the defects of traditional Cre knock-in monkey models in terms of time, cost, and resource requirements. Enhancers often outperform native promoters in their ability to target cell types with high stability and specificity. While EDGE has so far withstood many tests and is promising for cell-type targeting in NHPs, researchers often run into the issue of enhancer validation and elusive transcription regulation in vivo. Recently, engineered RNA sensors, a novel genetic tool, have made tag events occur during the mRNA-based translation process. This system is convenient and flexible in terms of probe design, yet its efficiency and stability need further verifications in NHPs. the mAGNET also provides an alternative for researchers who do not need to label fine cell subtypes while requiring safety. Scientists can pick the appropriate genetic targeting strategy according to their research needs.
Viral Tools for Neuroscience Studies in Monkeys
The application of engineered viral tools in neuroscience started a new era of exploring neural circuits at the molecular, single synapse, and neuron populations levels. As the mainstay of modern neuroscience tools, viral vectors have the advantage of rapid and high-level exogenous gene expression, cell tropism, and propagation directionality that can be harnessed for neuronal tracing, manipulation, and monitoring.
Viral tools have been widely applied in rodents, However, achieving viral-mediated robust, local, and pathway-selective exogenous gene expression in NHPs remains a challenging task, due to low expression levels, aggressive immune responses, and weak anterograde and retrograde transport efficiency. Here, we summarize recent progress in the use of viral tools for studies of primate neural circuits (Fig. 2).
Key properties of representative viral tools for neuroscience studies in primates. Green cells and lines represent cells and fibers infected by viral vectors, and yellow dots indicate the co-expression of TVA/G and RVdG. The number of dots is proportional to the strength of local infection, the number and thickness of lines are proportional to retro/anterograde traffic strength, and dotted lines indicate possible projected performances. Retrograde, viral particles are usually taken up by axon terminals and retro-transported to the presynaptic soma; Anterograde, viral particles are usually taken up by cell bodies and transported to axon terminals; VSV-G, VSV glycoprotein, protein G; RbV-G, RbV glycoprotein, protein G; FUG-B, fusion glycoprotein type B; FUG-C, fusion glycoprotein type C; FUG-E, fusion glycoprotein type E; EnVA-RVdG, avian ASLV type A envelope protein (cognate ligand for TVA)- glycoprotein (G)-deleted rabies virus.
Adeno-associated Virus (AAV)
Recombinant adeno-associated viruses (AAVs), single-stranded DNA parvovirus with a payload capacity of approximately 3.5-5 kb, are key viral tools for neuroscience research. AAVs have a number of advantages, such as minimal toxicity, low immunogenicity, and long-term stable expression across multiple species. Newly developed AAV variants provide better results in terms of precise targeting; they are minimally invasive and can maximize cargo transgene capacity [36]. Different AAV serotypes have unique tropisms and infection properties, including organs tropism, cell types tropism, and direction of spread in neurons [37]. Here, we discuss the tropisms and biodistribution of known AAV capsid variants in primates as a reference for AAV serotype selection.
A systematic evaluation of AAV serotypes in the primate brain has indicated that AAV2 has the lowest transfection efficiency compared with other serotypes (AAV1, AAV5, AAV8 and AAV9) in the macaque cortex [11], AAV5 is the most effective vector in transducing cells and axonal terminals in the substantia nigra pars compacta (SNc) and striatum, followed by AAV1 and AAV2, while the least effective is AAV3 [38]. In this study, AAV2 has shown greater tropism to neurons (98%), while AAV5 transduces neurons and glial cells with equal efficacy. AAV8 and AAV9 exhibit local axonal terminal transport and retrograde transport across primate species. Yet, retrograde efficiency depends on the injection site: for instance, retrograde efficiency in the lateral geniculate nucleus (LGN) is relatively low compared to the SNc [39]. In conclusion, the infection results depend on many complex factors. Due to the limited sample sizes and experience, it is not yet possible to conclude the transduction performance of different AAV serotypes. Most researchers would pick suitable AAV serotypes based on their experience and personal preferences.
There are also many engineered variants derived from its parent capsids to suit customized study requirements. Among those capsid variants, AAV.DJ exhibits more robust and widespread local and axon terminal transport with neuronal tropism in the putamen and motor cortex of monkeys than AAV2 and AAV5 [40, 41]. AAV2.retro mediates stronger retrograde transport in the caudate and putamen of primates than AAV8 and AAV9, but in some brain regions, such as the occipital lobe and the cerebellum, it shows insufficient retrograde ability and robust axon terminal labeling [42]. AAV2.1 exhibits low neurotoxicity, and efficient gene transduction in large regions of the macaque brain [43]. AAV2.HBKO, AAV.CAP-B10, AAV.CAP-Mac, AAV.MaCPNS1, and AAV.MaCPNS2 have shown varying degrees of CNS neuronal tropism after intracranial injections in both marmosets and macaques [44, 46, 51].
In summary, AAVs are prominent gene delivery vehicles and have the potential to be tools for research on complex neural circuits in NHPs. Two major limitations of off-the-shelf AAV capsids would need to be addressed, both of which have been addressed in rodents: firstly, the efficiency of anterograde or retrograde circuit tracing in primates; secondly, the ability to cross the blood-brain barrier (BBB) and target brain cells of newborn or adult primates by intravenous administration. In rodents. AAV engineering can be done in three main ways: discovering naturally occurring AAV, rational design, and directed evolution [47, 53]. Among these, multiplexed-Cre recombination–based AAV targeted evolution (M-CREATE) has become the mainstream approach for capsid evolution, due to the high signal-to-noise ratio, mechanistic diversity, and screening capacity with cell type specificity. All reports published so far have applied the engineering strategy of screening in mice and validation in primates. Given the differences among the species considered, this approach is likely to cause early exits of ideal variants during the early screening rounds and lead to unpredictable infection patterns in NHPs. For instance, AAV.PHP.eB shows enhanced neuron-biased transduction after intracranial injections in C57BL/6J mice, but the BBB-penetrant ability can’t be kept in macaques, partly due to the absence of Ly6a in primates [45, 48,49,50]. In addition, AAV.MaCPNS1 and AAV.MaCPNS2 are transduced in the PNS in rodents, and both the PNS and CNS in NHPs [51]. Yet it is difficult to carry out directed evolution-based screening in primates due to a number of limitations. A combination of rational design and library screening may therefore be a better capsid modification and engineering approach in monkeys. It is worth noting that AAV.CPP.16, a recently reported rational design-based AAV9 variant by inserting cell-penetrating peptides (CPPs) between amino acids 588 and 589, has exhibited enhanced transduction efficiency after systemic delivery in young and adult macaques [52]. The reality, however, is that a rational design-based engineering strategy is hard to use as the molecular mechanisms of AAV tropism are still unclear [53]. More efforts in exploring the biological mechanisms of AAV infection are required to develop the next generation of AAV variant screening approaches in NHPs.
Lentivirus Pseudotype (LV)
Lentivirus pseudotype (LVs) is a single-stranded RNA retrovirus with a capsid derived from HIV-1 and envelope proteins exchanged with those of another virus; in this way, the original viral characteristics are modified, affecting its tropism and transport direction [37]. There are two major types of lentivirus pseudotypes: VSV-G, in which the glycoprotein is the one from the vesicular stomatitis virus, and RbV-G, in which the glycoprotein is derived from the rabies virus. RbV-G has higher neurotoxicity than VSV-G. It is worth noting that RbV has outstanding retrograde traffic ability [54]. VSV-G has broad tropism and exhibits powerful local transduction in the primate CNS. NHP researchers have successfully transfected and manipulated the cortex, putamen, thalamus, and external globus pallidus (GPe) of monkeys via VSV-G [55,56,57]. Recently, new LV pseudotypes, HIV-1 lentivector fused with glycoprotein type B (FuG-B), glycoprotein type C (FuG-C), and glycoprotein type E (FuG-E), have started to achieve more efficient retrograde traffic and neuronal targeting [58]. Due to their excellent performance, these lentivirus pseudotypes have been selected for pathway-selective manipulation and connectome mapping in NHPs [41, 59, 60].
Canine Adenoviruses 2 (CAV-2)
Canine adenoviruses-2 (CAV-2) are double-stranded DNA adenoviruses with a payload capacity of approximately 30 kb. In comparison to naturally-occurring AAV serotypes, CAV is taken up preferentially by terminals and then retrogradely transported from dendrites to soma [61]. Engineered CAV-2 has largely reduced neurotoxicity thanks to the removal of replication-related viral genes, which enables long-term "exogenous cargo" loading in a pathway-selective manner [37]. CAV-2 has been used to investigate neural circuits in primates. Bohlen et al. have shown that CAV-2 can mediate retrograde transgene expression of fluorescent proteins in motoneurons following craniofacial intramuscular injections in rhesus monkeys [62]. A subsequent study showed that CAV-2 is promising for circuit-specific [e.g., primary visual cortex (V1) to secondary visual cortex (V2)] delivery of Cre recombinase in primates [63]. CAV-2 provides another option to target and investigate specific cell types, but more research is required in order to optimize the performance of its transduction system.
Rabies Virus (RbV)
Wild-type RbV, a negative-sense, single-stranded RNA rhabdovirus, can be trafficked in a retrograde manner across polysynaptic connections. The advent of the glycoprotein (G)-deleted rabies virus (RVdG) pseudotype, in which the propagation-related sequence “G” is replaced with a required sequence, such as GFP, led to the onset of a new era in monosynaptic input connectome research in rodents [64, 65]. RVdG has also been used in monkeys to depict the morphologies and connectomes of neurons in V1 [66]. Currently, the most widely-used RVdG is the avian ASLV type A envelope protein (EnvA)-pseudotype RVdG system, which can target specific cell types via TVA-EnvA recognition and monosynaptic retrograde transduction. In this system, the helper virus incorporates TVA and G, and EnvA-RVdG is injected into the desired region sequentially. RVdG particles only enter cells that co-express TVA and G (“starter cells”), then they replicate and propagate in a transsynaptic fashion by trans-complementation with G. Using the RVdG-mediated monosynaptic tracing system, researchers have mapped the direct monosynaptic inputs of V1 neurons and demonstrated the existence of brain regions and functional-specific feedforward to feedback (FF→FB) loops in the primate cortex [63]. Yet, RVdG presents several limitations in terms of labeling efficiency of presynaptic neurons and of potential cell toxicity, especially in higher mammals. The RVdG system needs to be engineered to achieve high efficiency and safety in order to study monosynaptic input connectomes in NHPs.
Genetic Tools Used for Neuronal Activity Manipulation
In order to understand the assembly and inner workings of the mammalian brain, we need to decode complex dynamic brain networks by manipulating neural circuits artificially under normal and pathological conditions. The recent continuous development of genetic modulators has triggered a growth spurt of neural circuit research in small animals. Despite falling behind against rodents, some progress has also been made in primate optogenetics and chemogenetics. Here, we summarize the genetic manipulation tools that have been used in monkeys (Table 1).
Primate Optogenetics
Optogenetics is a revolutionary neuroscience approach, which realizes temporally precise control of neural activity by genetically overexpressing opsins in neurons [67]. Since the discovery of the first light-gated protein, the halorhodopsin family (NpHR) in 1977 and channelrhodopsin-2 (ChR2) in 2002 [68], a large arsenal of naturally-occurring and synthetic opsins that enable optical control of neuronal activity with millisecond precision has been engineered.
Three main light-activated opsin types are used in monkeys: ChR2, a classic blue light-gated (470 nm) opsin with fast kinetics derived from Chlamydomonas reinhardtii; C1V1, a red-shifted opsin with enhanced photocurrent and fast kinetics; and step-function opsins (SFO), a variant of hChR2 (C128S) that switches on and off in response respectively to 470 nm and 542 nm light. The most widely used optical depolarizing opsins in monkeys are ChR2 and its enhanced variant ChR2 (H134R). Given that red light penetrates deeper into tissues than other visible wavelengths, C1V1 has also been chosen several times for optical stimulation in monkeys [69,70,71]. SFO and red-shifted CnChR1 (ChrimsonR) have also been selected for a few studies [13, 72]. More sensitive opsins with fast kinetics have been applied in mice with good results (e.g., Chrger2.0), and are expected to be useful in future monkey studies [73, 74].
Three main light-inhibitory types of opsins are used in monkeys: ArchT (archaerhodopsin-3), an outward proton pump that responds to yellow light (mostly 450-550 nm) [75]; eNpHR3, a yellow-light-sensitive chloride pump engineered from NpHR; and Jaws, a red-shifted opsin from Halobacterium salinarum, that enables robust and deeper neuronal silencing in red light (635 nm) [73, 76]. Jaws currently appears to be the most effective neuronal silencer for NHPs.
Han et al. first used ChR2 in 2009 to activate excitatory neurons in the frontal cortex of macaques, proving the feasibility of applying optogenetics to primates [55], and they transduced ArchT to neuronal membranes and axons in the visual cortex again in 2011, which resulted in the rapid and complete silencing of recorded neurons [56]. Similar opsin-mediated neuron depolarization/hyperpolarization has been further confirmed in the motor cortex, the somatosensory cortex, the parietal cortex, the visual cortex, the frontal eye field (FEF), the superior colliculus (SC), and in thalamic subnuclei [13, 57, 75, 77]. At this stage, the optogenetic perturbations are not yet able to induce any significant behavior. The work of Ohayon et al. has ruled out the hypothesis that optical stimulation in monkeys is subtle and subthreshold [78].
Researchers have faced enormous challenges due to many obstacles, including limited transfection efficiency, inefficient light delivery and readout equipment, severe immune responses, and deficient injection protocols. After continuous improvement, NHP optogenetics has been transformed, fueled by developments in viral transudation efficiency (e.g., AAV2.1), surgical procedures (e.g., the convection-enhanced delivery (CED) method and window implantation), light delivery devices (e.g., the coaxial optrode), and readout approaches (e.g., fMRI and Ca2+ imaging) [43, 70, 79,80,81,82,83]. In the last decade, primates optogenetics have started an era of mesoscopic neuronal circuit neuroanatomy and functional studies with cell-type specificity (e.g., dopamine neurons and Purkinje cells) and pathway selectivity (e.g., FEF to SC and LGN to V1). Such studies have focused on face gender discrimination, direction discrimination, decision confidence, value-based learning, working memory, reward prediction, visual perception, saccade movement, and forelimb movements. Most of those studies have been completed in the primary visual system and the frontal system [15, 16, 27, 40, 57, 68, 69, 71, 72, 75,76,77, 83,84,85,86,87,88,89,90,91,92,93,94,95,96,97] (Fig. 3 A).
Application summary of genetic tools used for neuron manipulation in primates. A Opsins used for primate optogenetics. B Genetic tools used for primate chemogenetics and neurotoxins. The size of the dots is proportional to the injection frequency in the brain region for optogenetics and chemogenetics; the PFC, motor cortex, V1, and thalamus are popular regions. ChR2, channelrhodopsin-2; C1V1, red-shifted chimera; SFO, step-function opsins; ChrimsonR, red-shifted CnChR1; ArchT, archaerhodopsin-3; eNpHR3.0, halorhodopsin-kir2.1; Jaws, red-shifted cruxhalordoposin; eTeNT, tetanus neurotoxin; hM4Di, human Gi-coupled M4 muscarinic receptor; hM3Dq, human Gi-coupled M3 muscarinic receptor; PFC, prefrontal cortex; LIP, lateral intraparietal; FEF, frontal eye field; VIP, ventral intraparietal area; Cd, caudate nucleus; SC, superior colliculus; M1, primary motor cortex; GPe, globus pallidus, external segment; LGN, lateral geniculate nucleus; MT, middle temporal area; IT, inferior temporal cortex; V1, primary visual cortex; CB, cerebellum; MB, midbrain; NAc, nucleus accumbens, core; VTA, ventral tegmental area; SNr, substantia nigra pars reticulate; MDL, lateral mediodorsal thalamus; LC, locus coeruleus; POA, preoptic area.
Today, primate optogenetics is undergoing a phase of rapid development, while still facing many challenges. The stability and efficiency of genetic modulators delivery largely depend on viral vectors. Engineering more useful viral vectors to ensure intense and stable expression of opsins is key to the modulation of neuronal activity. In addition, cell-type-specific manipulation would be a significant step toward decoding complex brain dynamics. The screening of specific cis-regulatory elements can lead us to achieve this goal [25, 26]. Developed opsins are also urgently needed to upgrade primate optogenetics; for instance, ChRs with enhanced light sensitivity 2.0 (Chrger2.0), a newly engineered opsin, enable neuron activation in mice without fiber-optic implantation, extending application directions for optogenetics experiments [74]. We believe that its use in monkeys would greatly reduce heat absorption in the brain and engage more neurons at lower power to induce behaviors.
Primate Chemogenetics
Chemogenetics is another neuronal manipulation approach, in which engineered exogenous receptors expressed in target cells interact with exogenous ligands via the system or local drug administration [98]. The DREADDs used in chemogenetics are almost exclusively the human Gi-coupled M4 muscarinic receptor (hM4Di, inhibitory DREADDs) and human Gi-coupled M3 muscarinic receptor (hM3Dq, excitatory DREADDs), which are activated by clozapine-N-oxide (CNO), compound 21, and perlapine [98]. Once activated, hM4Di and hM3Dq attenuate or enhance neuronal excitation by binding to Gq/s or Gi proteins [99]. Compared to optogenetics, the development of primate chemogenetics is relatively lagging. The biggest disadvantage of chemogenetics is the inferior temporal resolution, which ranges from minutes to hours and does not allow investigation of the rapid dynamics of brain operations. In addition, chemogenetics has a relatively low spatial resolution. Despite the poor temporal resolution, chemogenetics is more powerful in driving robust behaviors (even motor behavior) than optogenetics. As such, DREADD technology is sometimes more desirable for reversible and large-scale control of cell populations, while being less invasive than optogenetics [99, 100].
Chemogenetics studies on NHPs are becoming increasingly popular. In these studies, hM4Di has been used to selectively block the motor cortex-propriospinal neurons (PNs) pathway and affect hand dexterity [100]; to disrupt the connection between the lateral prefrontal cortex (lPFC) and the striatum involved in inhibitory control [101]; to silence the dorsolateral prefrontal cortex (dlPFC)-lateral mediodorsal thalamus (MDL) or dlPFC-dorsal caudate nucleus (dCD) pathways involved in decision-making and working memory [102]; to inactivate the rostromedial caudate (rmCD) and PFC through the positron emission tomography (PET)-guided systemic administration of CNO [103, 104]. It has also been used to silence the PFC and induce spatial working memory deficits [105, 106]; and to reversibly inactivate the locus coeruleus (LC) and affect cognitive functions [107]. hM4Di, equipped with Cre-dependent viral tools, provides a powerful pharmacogenetic tool to selectively inactivate neuronal circuits. Oguchi et al. have used AAV5, which carries the "Cre-On" FLEX double floxed sequence, transferred hM4Di to the PFC, and then injected a retrograde vector that incorporated Cre-recombinase into the caudate nucleus (Cd). In this study, researchers have successfully suppressed specific neural circuits on a time scale ranging from minutes to weeks [101] (Fig. 3B). In a more recent study, the high-affinity and selective agonist deschloroclozapine (DCZ) has been reported for DREADD activation. This agonist displays high selectivity and fast kinetics in mice and monkeys [105]. There are also a few reports about chemogenetic activation using hM3Dq. Mimura et al. have used AAV1-encoded hM3Dq to activate TH+ neurons of the SNc in marmosets and observed rotation behaviors in a contralateral direction relative to the activated side [108]. Another study used AAV8-encoded HM3Dq to activate a group of neurons in the preoptic area (POA) and successfully drive hypothermia and cold defense in macaques [109] (Fig. 3 B). Overall, chemogenetic technologies are very promising, but there is still considerable room for improvement in terms of safety, off-target effects, efficiency, and specificity. For example, DREADD actuators have different potential effects on animals. One study showed that the monkey's working memory is impaired after olanzapine and DCZ injections [110]. Another example is that the subcellular localization of DREADDs depends on the DREADD/tag combination (e.g., the hM4Di‐mCherry is mostly expressed in the intracellular space, and the hM4Di‐haemagglutinin (HA) tag is mostly confined to the plasma membrane) [111]. A systematic parameter test before chemogenetic experiments is recommended.
Neurotoxins
Neurotoxins are substances that can modulate the electrical activity and neurotransmitter release of neurons and destroy the nervous system by means of binding to specific membrane acceptors. Common examples of neurotoxin tools include nitric oxide (NO), botulinum neurotoxins (BoNT), and tetanus toxin (TeNT) [112]. Since most neurotoxins have a high affinity for specific molecular targets, they have long been used to study the anatomy, physiology, function, and disorders of the brain. And yet, the application of neurotoxins has some limitations, including slow inactivation time, off-target activity, the difficulty of extraction, and unclear effective concentration [113]. To date, two studies have reported a tetracycline-dependent enhanced tetanus toxin (eTeNT) system to realize selective and reversible lesions in the spinal cord and ventral tegmental area (VTA) respectively in NHPs. In this system, the virus that incorporates the Tet-on sequence is injected into the target brain region, while another retrograde virus, carrying the eTeNT sequence driven by the tetracycline (TRE) promoter, is injected into the downstream regions of interest. During the "doxycycline (Dox) on" period, Dox binds to the tetracycline transactivator (tTA) and activates the TRE promoter. The expression of eTeNT is then turned up in double-infected neurons that send signals to specific downstream regions. Then the connections between specific neural circuits are reversibly blocked in a Dox-dependent manner; these events do not occur in the absence of Dox [41, 59] (Fig. 3 B). Due to the natural deficits of neurotoxins, and the immaturity of the associated manipulation approach, their application for neural circuit studies in NHPs is relatively limited.
Genetic Tools Used for Neuronal Activity Monitoring
The ability to monitor neuronal populations or single neurons with high spatiotemporal fidelity is crucial for a comprehensive understanding of the mammalian brain. Traditional approaches for neuron activity recording in primates are electrophysiology, fMRI, voltage-sensitive dye imaging (VSDI) and intrinsic optical imaging. Electrophysiology, a classic method in neuroscience, has high temporal fidelity but is limited by its low resolution regarding cell number, identity, and spatial localization. The oldest intrinsic optical imaging offers a minimally or non-invasive method to look inside the monkey brain, but its cell-type resolution, temporal fidelity, and signal-to-noise ratio are still lacking [114,115,, 115]. VSDI can track neuronal activity with fast kinetics on the millisecond scale. However, it cannot be used for long-term recording due to high phototoxicity and low cell-type resolution due to the non-selective infection pattern of VSDI dyes [116,116,117,118,118].
The advent of genetically encoded optical indicators has enabled researchers to simultaneously record neurotransmitter concentrations, transmembrane voltage, the intracellular Ca2+ dynamics of specific neuron populations, and even fine structure. Novel genetically encoded optical tools have been used in small animals for years, but their application in primates lags behind. Recent advances in optical imaging in primates have shed new light on cortical information processing. Here, we summarize the genetically encoded tools used in primate neuronal circuit imaging to provide a reference for future research. Most of these tools are still at the proof-of-concept stage (Table 2).
Genetically Encoded Calcium Indicators (GECIs)
Genetically Encoded Calcium Indicators (GECIs) have proven to be powerful imaging tools in NHPs, as they are capable of monitoring neurons via Ca2+ changes in neuron depolarization, during which the cytoplasmic Ca2+ concentration increases 10 or even 100 times. Two major types of GECIs have been used in monkeys: calmodium-based indicators, and troponinC-based indicators. memTNXL, a troponinC-based indicator, was used to monitor neurons as early as 2010 [119]. Yet memTNXL did not result in robust and fast fluorescence changes, and there have been no follow-up reports. Instead, improved versions of calmodium-based indicators have been applied in primates. The most widely used calmodium-based indicators in primates are GCaMP5G, GCaMP6s, and GCaMP6f [120,121,122]. Since the tTA-TRE inducible GCaMP6f expression system was first used for long-term imaging in anesthetized marmosets, several researchers have focused on this indicator [122]. GCaMP5G, GCaMP6s, and GCaMP6f, equipped with improved microscopes and window implantation, have been widely used to dissect neural circuits involved in social interactions, motor planning and execution, visual information processing, and sequential working memory [120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136]. Recent research has realized two-photon recording of interneurons in the marmoset cortex via the human DLX5/6 enhancer-mediated GCaMP expression strategy. This approach, based on specific gene expression mediated by cis-regulatory elements, may be useful in revealing the functional characteristics of distinct neuron types [28].
In addition to the classical GCaMP version, a series of new Ca2+ indicators have been engineered to meet different experimental needs in rodents. For instance, jGCaMP7 improves single-spike detection; soma-targeted GCaMP by the endoplasmic reticulum and ribosome tethering reduces the level of crosstalk from the neuropil and improves imaging quality [137,138,139]. Newly engineered GECIs will provide novel and interesting insights for future primate studies.
Genetically Encoded Transmitter Indicators (GETIs)
Genetically Encoded Transmitter Indicators (GETIs) can help us monitor the release of specific neurotransmitters and neuromodulators in the nervous system with high sensitivity and faster kinetics than Ca2+ signals [140]. A large arsenal of GETIs has been developed over the years. Among these, iGluSnFR is the most responsive sensor to a single action potential and dendritic boutons, with a high signal-to-noise ratio and temporal resolution when excited at 488 nm. Ju et al. have performed two-photon dendritic imaging with iGluSnFR in V1 of macaques and mapped a fine dendritic excitatory input atlas of neurons in the superficial layers of V1 [127]. The application of GETIs in primates is still in the early stage. Given the importance of neurotransmitters and neuromodulators in information processing, the development and application of GETIs for NHPs studies are crucial for research.
Other Genetically Encoded Indicators
Genetically Encoded pH Indicators (GEPIs), such as super ecliptic pHluorin (SEP), can detect the release of synaptic vesicles mediated by pH changes between the vesicle lumen (pH 5.5) and the extracellular environment (pH 7.0) [140]. Genetically encoded voltage indicators (GEVIs) can also be used to investigate neuronal dynamics based on membrane potential changes in postsynaptic targets. Recent advances have led to the development of new voltage indicators with improved performance in voltage sensitivity, activation and inactivation speed, brightness, and cell toxicity. For example, mNeonGreen, a recently developed GEVI-fused proton-pumping rhodopsin, enables bright and fast neuron recording in mice and flies [141]. In addition, GEVIs can be coupled with other reporters, like opsin and neurotransmitter indicators, to enable all-optical interrogation of neuronal circuits [142]. These genetically encoded indicators have not yet been applied to the dissection of neural circuits in primates, and given their inherent properties, their application remains a challenge for studies in NHPs.
Nowadays, neural circuit functional research, either perturbing or observing, faces a fundamental and unavoidable dilemma. On one hand, even with the enhanced delivery efficiency of engineered AAVs, regulatory or recording units induced by local viral injection are still relatively small compared to the sizes of the brains of NHPs. On the other hand, multiple sophisticated neural circuits are mobilized systemically to guide behaviors in higher mammals. This makes it difficult to induce behaviors by relying solely on simple neuron manipulation as has been done in mice. Considering the particularity of monkey models, decoding and re-encoding information processing based on large numbers of cells would be perhaps more convincing. We recommend the further development of large optical control and imaging fields with high resolution and cell-type specificity in NHPs.
In order to address these limitations, we can improve the following aspects: (1) increased transfection range through surgical improvements and new technologies; (2) the development of BBB-penetrant AAVs to target brain cells in adult primates; and (3) the generation of transgenic optogenetic and calcium imaging monkey models. It is worth noting that transgenic monkey models can not only express the desired proteins in the whole brain, but their ability to indicate neuronal activity is also maintained over time by the constant synthesis of new proteins, meaning this method can support long-term studies for particular behaviors and diseases. To date, the first GCaMP marmoset study has been reported [143]. The use of optogenetics and GCaMP transgenic NHPs for functional neuroscience studies would be a milestone in this field.
Closed-loop investigation is another possible approach to quickly and intuitively build encoding and decoding models in primates. This concept has already been proven feasible in macaques [126]. Closed-loop interrogation of cortical neurons by optical activation and imaging still has a lot of room for technological improvement; for instance, improved opsins should be compatible with Ca2+ indicators, and they both should be expressed in the same cells.
Conclusions
Genetic approaches offer powerful tools for studying the dynamics and connectivity of neural circuits in different cortical areas, and linking them to particular behaviors. The genome-editing technologies developed so far (e.g., the CRISPR/Cas system) have made it feasible to precisely generate gene-modified monkey models for neuroscience research [144]. In parallel, advances in single-cell transcriptomic and epi-genomic sequencing technologies have facilitated the understanding of gene expression regulatory networks and the cellular makeup of the primate brain, leading to the development of cell-type-specific tools in monkeys. Engineered viral capsids enable uniform and stable exogenous gene delivery. Neuronal activity modulators and indicators, driven by innovations in surgical procedures, light delivery devices, and readout equipment, are beginning to overcome past limitations and provide novel approaches to mesoscopic neural circuit research in primates.
Recently, the scientific community worldwide has undertaken the project of mapping the comprehensive brain cell atlas and mesoscopic connectivity atlas of primates [145, 146]. We believe that this project will help to develop improved genetic targeting tools and explore the neurobiological basis within neural circuits, including cellular components, wiring diagrams, and inner workings. In turn, efforts in the genetic functional analysis of neural circuits will facilitate a better understanding of the molecular-cellular and cell properties of the nervous system in NHPs, which are similar to those in humans. Progress in NHPs brain studies will enable people to better understand how the human brain works under normal physiological and pathological conditions.
References
Yin S, Lu K, Tan T, Tang J, Wei J, Liu X. Transcriptomic and open chromatin atlas of high-resolution anatomical regions in the rhesus macaque brain. Nat Commun 2020, 11: 474.
Chansel-Debordeaux L, Bezard E. Local transgene expression and whole-body transgenesis to model brain diseases in nonhuman primate. Animal Model Exp Med 2019, 2: 9–17.
Krienen FM, Goldman M, Zhang Q, Del Rosario RCH, Florio M, Machold R, et al. Innovations present in the primate interneuron repertoire. Nature 2020, 586: 262–269.
Fuentealba P, Begum R, Capogna M, Jinno S, Márton LF, Csicsvari J, et al. Ivy cells: A population of nitric-oxide-producing, slow-spiking GABAergic neurons and their involvement in hippocampal network activity. Neuron 2008, 57: 917–929.
Daigle TL, Madisen L, Hage TA, Valley MT, Knoblich U, Larsen RS, et al. A suite of transgenic driver and reporter mouse lines with enhanced brain-cell-type targeting and functionality. Cell 2018, 174: 465-480.e22.
Madisen L, Garner AR, Shimaoka D, Chuong AS, Klapoetke NC, Li L, et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 2015, 85: 942–958.
Izpisua Belmonte JC, Callaway EM, Caddick SJ, Churchland P, Feng G, Homanics GE, et al. Brains, genes, and Primates. Neuron 2015, 86: 617–631.
Li T, Ai Z, Ji W. Primate stem cells: Bridge the translation from basic research to clinic application. Sci China Life Sci 2019, 62: 12–21.
Lu Z, He S, Jiang J, Zhuang L, Wang Y, Yang G, et al. Base-edited cynomolgus monkeys mimic core symptoms of STXBP1 encephalopathy. Mol Ther 2022, 30: 2869–2873.
Yang W, Liu Y, Tu Z, Xiao C, Yan S, Ma X, et al. CRISPR/Cas9-mediated PINK1 deletion leads to neurodegeneration in rhesus monkeys. Cell Res 2019, 29: 334–336.
Watakabe A, Ohtsuka M, Kinoshita M, Takaji M, Isa K, Mizukami H, et al. Comparative analyses of adeno-associated viral vector serotypes 1, 2, 5, 8 and 9 in marmoset, mouse and macaque cerebral cortex. Neurosci Res 2015, 93: 144–157.
Tremblay S, Acker L, Afraz A, Albaugh DL, Amita H, Andrei AR, et al. An open resource for non-human primate optogenetics. Neuron 2020, 108: 1075-1090.e6.
Diester I, Kaufman MT, Mogri M, Pashaie R, Goo W, Yizhar O, et al. An optogenetic toolbox designed for Primates. Nat Neurosci 2011, 14: 387–397.
El-Shamayleh Y, Ni AM, Horwitz GD. Strategies for targeting primate neural circuits with viral vectors. J Neurophysiol 2016, 116: 122–134.
Stauffer WR, Lak A, Yang A, Borel M, Paulsen O, Boyden ES, et al. Dopamine neuron-specific optogenetic stimulation in Rhesus macaques. Cell 2016, 166: 1564-1571.e6.
El-Shamayleh Y, Kojima Y, Soetedjo R, Horwitz GD. Selective optogenetic control of Purkinje cells in monkey cerebellum. Neuron 2017, 95: 51-62.e4.
Shima Y, Sugino K, Hempel CM, Shima M, Taneja P, Bullis JB, et al. A Mammalian enhancer trap resource for discovering and manipulating neuronal cell types. Elife 2016, 5: e13503.
Rickels R, Shilatifard A. Enhancer logic and mechanics in development and disease. Trends Cell Biol 2018, 28: 608–630.
Andersson R, Sandelin A. Determinants of enhancer and promoter activities of regulatory elements. Nat Rev Genet 2020, 21: 71–87.
Blankvoort S, Witter MP, Noonan J, Cotney J, Kentros C. Marked diversity of unique cortical enhancers enables neuron-specific tools by enhancer-driven gene expression. Curr Biol 2018, 28: 2103-2114.e5.
Gray LT, Yao Z, Nguyen TN, Kim TK, Zeng H, Tasic B. Layer-specific chromatin accessibility landscapes reveal regulatory networks in adult mouse visual cortex. Elife 2017, 6: e21883.
Joo JY, Schaukowitch K, Farbiak L, Kilaru G, Kim TK. Stimulus-specific combinatorial functionality of neuronal c-fos enhancers. Nat Neurosci 2016, 19: 75–83.
Kim TK, Hemberg M, Gray JM, Costa AM, Bear DM, Wu J, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010, 465: 182–187.
Duba-Kiss R, Niibori Y, Hampson DR. GABAergic gene regulatory elements used in adeno-associated viral vectors. Front Neurol 2021, 12: 745159.
Graybuck LT, Daigle TL, Sedeño-Cortés AE, Walker M, Kalmbach B, Lenz GH, et al. Enhancer viruses for combinatorial cell-subclass-specific labeling. Neuron 2021, 109: 1449-1464.e13.
Mich JK, Graybuck LT, Hess EE, Mahoney JT, Kojima Y, Ding Y, et al. Functional enhancer elements drive subclass-selective expression from mouse to primate neocortex. Cell Rep 2021, 34: 108754.
De A, El-Shamayleh Y, Horwitz GD. Fast and reversible neural inactivation in macaque cortex by optogenetic stimulation of GABAergic neurons. Elife 2020, 9: e52658.
Mehta P, Kreeger L, Wylie DC, Pattadkal JJ, Lusignan T, Davis MJ, et al. Functional access to neuron subclasses in rodent and primate forebrain. Cell Rep 2019, 26: 2818-2832.e8.
Kaseniit KE, Katz N, Kolber NS, Call CC, Wengier DL, Cody WB, et al. Author Correction: Modular, programmable RNA sensing using ADAR editing in living cells. Nat Biotechnol 2023, 41: 577.
Qian Y, Li J, Zhao S, Matthews EA, Adoff M, Zhong W, et al. Programmable RNA sensing for cell monitoring and manipulation. Nature 2022, 610: 713–721.
Jiang K, Koob J, Chen XD, Krajeski RN, Zhang Y, Volf V, et al. Programmable eukaryotic protein synthesis with RNA sensors by harnessing ADAR. Nat Biotechnol 2022, https://doi.org/10.1038/s41587-022-01534-5.
Reautschnig P, Wahn N, Wettengel J, Schulz AE, Latifi N, Vogel P, et al. CLUSTER guide RNAs enable precise and efficient RNA editing with endogenous ADAR enzymes in vivo. Nat Biotechnol 2022, 40: 759–768.
Burgess DJ. RADARs and READRs for programmable RNA sensing. Nat Rev Genet 2022, 23: 711.
Keaveney MK, Tseng HA, Ta TL, Gritton HJ, Man HY, Han X. A microRNA-based gene-targeting tool for virally labeling interneurons in the rodent cortex. Cell Rep 2018, 24: 294–303.
Hordeaux J, Buza EL, Jeffrey B, Song C, Jahan T, Yuan Y, et al. MicroRNA-mediated inhibition of transgene expression reduces dorsal root ganglion toxicity by AAV vectors in Primates. Sci Transl Med 2020, 12: eaba9188.
Challis RC, Ravindra Kumar S, Chen X, Goertsen D, Coughlin GM, Hori AM, et al. Adeno-associated virus toolkit to target diverse brain cells. Annu Rev Neurosci 2022, 45: 447–469.
Nectow AR, Nestler EJ. Viral tools for neuroscience. Nat Rev Neurosci 2020, 21: 669–681.
Markakis EA, Vives KP, Bober J, Leichtle S, Leranth C, Beecham J, et al. Comparative transduction efficiency of AAV vector serotypes 1–6 in the substantia nigra and striatum of the primate brain. Mol Ther 2010, 18: 588–593.
Masamizu Y, Okada T, Kawasaki K, Ishibashi H, Yuasa S, Takeda S, et al. Local and retrograde gene transfer into primate neuronal pathways via adeno-associated virus serotype 8 and 9. Neuroscience 2011, 193: 249–258.
Watanabe H, Sano H, Chiken S, Kobayashi K, Fukata Y, Fukata M, et al. Forelimb movements evoked by optogenetic stimulation of the macaque motor cortex. Nat Commun 2020, 11: 3253.
Tohyama T, Kinoshita M, Kobayashi K, Isa K, Watanabe D, Kobayashi K, et al. Contribution of propriospinal neurons to recovery of hand dexterity after corticospinal tract lesions in monkeys. Proc Natl Acad Sci U S A 2017, 114: 604–609.
Weiss AR, Liguore WA, Domire JS, Button D, McBride JL. Intra-striatal AAV2.retro administration leads to extensive retrograde transport in the rhesus macaque brain: Implications for disease modeling and therapeutic development. Sci Rep 2020, 10: 6970.
Kimura K, Nagai Y, Hatanaka G, Fang Y, Tanabe S, Zheng A, et al. A mosaic adeno-associated virus vector as a versatile tool that exhibits high levels of transgene expression and neuron specificity in primate brain. bioRxiv 2021. doi: https://doi.org/10.1101/2021.07.18.452859.
Chuapoco MR, Flytzanis NC, Goeden N, Octeau JC, Roxas KM, Chan KY, et al. Intravenous gene transfer throughout the brain of infant Old World primates using AAV. bioRxiv 2022: 2022.2001.2008.475342.
Matsuzaki Y, Konno A, Mochizuki R, Shinohara Y, Nitta K, Okada Y, et al. Intravenous administration of the adeno-associated virus-PHP.B capsid fails to upregulate transduction efficiency in the marmoset brain. Neurosci Lett 2018, 665: 182–188.
Naidoo J, Stanek LM, Ohno K, Trewman S, Samaranch L, Hadaczek P, et al. Extensive transduction and enhanced spread of a modified AAV2 capsid in the non-human primate CNS. Mol Ther 2018, 26: 2418–2430.
Li C, Samulski RJ. Engineering adeno-associated virus vectors for gene therapy. Nat Rev Genet 2020, 21: 255–272.
Hordeaux J, Wang Q, Katz N, Buza EL, Bell P, Wilson JM. The neurotropic properties of AAV-PHP.B are limited to C57BL/6J mice. Mol Ther 2018, 26: 664–668.
Hordeaux J, Yuan Y, Clark PM, Wang Q, Wang Q, Sims JJ, et al. The GPI-linked protein LY6A drives AAV-PHP.B transport across the blood-brain barrier. Mol Ther 2019, 27: 912–921.
Huang Q, Chan KY, Tobey IG, Chan YA, Poterba T, Boutros CL, et al. Delivering genes across the blood-brain barrier: LY6A, a novel cellular receptor for AAV-PHP.B capsids. PLoS One 2019, 14: e0225206.
Chen X, Ravindra Kumar S, Adams CD, Yang D, Wang T, Wolfe DA, et al. Engineered AAVs for non-invasive gene delivery to rodent and non-human primate nervous systems. Neuron 2022, 110: 2242-2257.e6.
Yao Y, Wang J, Liu Y, Qu Y, Wang K, Zhang Y, et al. Variants of the adeno-associated virus serotype 9 with enhanced penetration of the blood-brain barrier in rodents and Primates. Nat Biomed Eng 2022, 6: 1257–1271.
Büning H, Srivastava A. Capsid modifications for targeting and improving the efficacy of AAV vectors. Mol Ther Methods Clin Dev 2019, 12: 248–265.
Hislop JN, Islam TA, Eleftheriadou I, Carpentier DCJ, Trabalza A, Parkinson M, et al. Rabies virus envelope glycoprotein targets lentiviral vectors to the axonal retrograde pathway in motor neurons. J Biol Chem 2014, 289: 16148–16163.
Han X, Qian X, Bernstein JG, Zhou HH, Franzesi GT, Stern P, et al. Millisecond-timescale optical control of neural dynamics in the nonhuman primate brain. Neuron 2009, 62: 191–198.
Han X, Chow BY, Zhou H, Klapoetke NC, Chuong A, Rajimehr R, et al. A high-light sensitivity optical neural silencer: Development and application to optogenetic control of non-human primate cortex. Front Syst Neurosci 2011, 5: 18.
Galvan A, Hu X, Smith Y, Wichmann T. In vivo optogenetic control of striatal and thalamic neurons in non-human Primates. PLoS One 2012, 7: e50808.
Kobayashi K, Kato S, Kobayashi K. Genetic manipulation of specific neural circuits by use of a viral vector system. J Neural Transm 2018, 125: 67–75.
Vancraeyenest P, Arsenault JT, Li X, Zhu Q, Kobayashi K, Isa K, et al. Selective mesoaccumbal pathway inactivation affects motivation but not reinforcement-based learning in macaques. Neuron 2020, 108: 568-581.e6.
Oguchi M, Okajima M, Tanaka S, Koizumi M, Kikusui T, Ichihara N, et al. Double virus vector infection to the prefrontal network of the macaque brain. PLoS One 2015, 10: e0132825.
Liu Q, Wu Y, Wang H, Jia F, Xu F. Viral tools for neural circuit tracing. Neurosci Bull 2022, 38: 1508–1518.
Bohlen MO, El-Nahal HG, Sommer MA. Transduction of craniofacial motoneurons following intramuscular injections of canine adenovirus type-2 (CAV-2) in Rhesus macaques. Front Neuroanat 2019, 13: 84.
Siu C, Balsor J, Merlin S, Federer F, Angelucci A. A direct interareal feedback-to-feedforward circuit in primate visual cortex. Nat Commun 2021, 12: 4911.
Callaway EM, Luo L. Monosynaptic Circuit Tracing with Glycoprotein-Deleted Rabies Viruses. J Neurosci 2015, 35: 8979–8985.
Kim EJ, Jacobs MW, Ito-Cole T, Callaway EM. Improved monosynaptic neural circuit tracing using engineered rabies virus glycoproteins. Cell Rep 2016, 15: 692–699.
Nhan HL, Callaway EM. Morphology of superior colliculus- and middle temporal area-projecting neurons in primate primary visual cortex. J Comp Neurol 2012, 520: 52–80.
Deisseroth K. Optogenetics. Nat Methods 2011, 8: 26–29.
Adamantidis AR, Zhang F, de Lecea L, Deisseroth K. Optogenetics: Opsins and optical interfaces in neuroscience. Cold Spring Harb Protoc 2014, 2014: 815–822.
Dai J, Brooks DI, Sheinberg DL. Optogenetic and electrical microstimulation systematically bias visuospatial choice in Primates. Curr Biol 2014, 24: 63–69.
Ozden I, Wang J, Lu Y, May T, Lee J, Goo W, et al. A coaxial optrode as multifunction write-read probe for optogenetic studies in non-human Primates. J Neurosci Methods 2013, 219: 142–154.
Lu Y, Truccolo W, Wagner FB, Vargas-Irwin CE, Ozden I, Zimmermann JB, et al. Optogenetically induced spatiotemporal gamma oscillations and neuronal spiking activity in primate motor cortex. J Neurophysiol 2015, 113: 3574–3587.
McGregor JE, Godat T, Dhakal KR, Parkins K, Strazzeri JM, Bateman BA, et al. Optogenetic restoration of retinal ganglion cell activity in the living primate. Nat Commun 2020, 11: 1703.
Chuong AS, Miri ML, Busskamp V, Matthews GAC, Acker LC, Sørensen AT, et al. Noninvasive optical inhibition with a red-shifted microbial rhodopsin. Nat Neurosci 2014, 17: 1123–1129.
Bedbrook CN, Yang KK, MacKey ED, Gradinaru V, Arnold FH. Machine learning-guided channelrhodopsin engineering enables minimally invasive optogenetics. Nat Methods 2019, 16: 1176–1184.
Cavanaugh J, Monosov IE, McAlonan K, Berman R, Smith MK, Cao V, et al. Optogenetic inactivation modifies monkey visuomotor behavior. Neuron 2012, 76: 901–907.
Acker L, Pino EN, Boyden ES, Desimone R. FEF inactivation with improved optogenetic methods. Proc Natl Acad Sci U S A 2016, 113: E7297–E7306.
Gerits A, Farivar R, Rosen BR, Wald LL, Boyden ES, Vanduffel W. Optogenetically induced behavioral and functional network changes in Primates. Curr Biol 2012, 22: 1722–1726.
Ohayon S, Grimaldi P, Schweers N, Tsao DY. Saccade modulation by optical and electrical stimulation in the macaque frontal eye field. J Neurosci 2013, 33: 16684–16697.
Ruiz O, Lustig BR, Nassi JJ, Cetin A, Reynolds JH, Albright TD, et al. Optogenetics through windows on the brain in the nonhuman primate. J Neurophysiol 2013, 110: 1455–1467.
Tamura K, Ohashi Y, Tsubota T, Takeuchi D, Hirabayashi T, Yaguchi M, et al. A glass-coated tungsten microelectrode enclosing optical fibers for optogenetic exploration in primate deep brain structures. J Neurosci Methods 2012, 211: 49–57.
Ziv Y, Ghosh KK. Miniature microscopes for large-scale imaging of neuronal activity in freely behaving rodents. Curr Opin Neurobiol 2015, 32: 141–147.
Chen LL, Goffart L, Sparks DL. A simple method for constructing microinjectrodes for reversible inactivation in behaving monkeys. J Neurosci Methods 2001, 107: 81–85.
Oguchi M, Sakagami M. Dissecting the prefrontal network with pathway-selective manipulation in the macaque brain-a review. Front Neurosci 2022, 16: 917407.
Afraz A, Boyden ES, DiCarlo JJ. Optogenetic and pharmacological suppression of spatial clusters of face neurons reveal their causal role in face gender discrimination. Proc Natl Acad Sci U S A 2015, 112: 6730–6735.
Amita H, Kim HF, Inoue KI, Takada M, Hikosaka O. Optogenetic manipulation of a value-coding pathway from the primate caudate tail facilitates saccadic gaze shift. Nat Commun 1876, 2020: 11.
Chernov MM, Friedman RM, Chen G, Stoner GR, Roe AW. Functionally specific optogenetic modulation in primate visual cortex. Proc Natl Acad Sci U S A 2018, 115: 10505–10510.
El-Shamayleh Y, Horwitz GD. Primate optogenetics: Progress and prognosis. Proc Natl Acad Sci U S A 2019, 116: 26195–26203.
Fetsch CR, Odean NN, Jeurissen D, El-Shamayleh Y, Horwitz GD, Shadlen MN. Focal optogenetic suppression in macaque area MT biases direction discrimination and decision confidence, but only transiently. Elife 2018, 7: e36523.
Fortuna MG, Hüer J, Guo H, Gruber J, Gruber-Dujardin E, Staiger JF, et al. Histological assessment of optogenetic tools to study fronto-visual and fronto-parietal cortical networks in the rhesus macaque. Sci Rep 2020, 10: 11051.
Galvan A, Caiola MJ, Albaugh DL. Advances in optogenetic and chemogenetic methods to study brain circuits in non-human Primates. J Neural Transm 2018, 125: 547–563.
Galvan A, Hu X, Smith Y, Wichmann T. Effects of optogenetic activation of corticothalamic terminals in the motor thalamus of awake monkeys. J Neurosci 2016, 36: 3519–3530.
Inoue KI, Takada M, Matsumoto M. Neuronal and behavioural modulations by pathway-selective optogenetic stimulation of the primate oculomotor system. Nat Commun 2015, 6: 8378.
Jazayeri M, Lindbloom-Brown Z, Horwitz GD. Saccadic eye movements evoked by optogenetic activation of primate V1. Nat Neurosci 2012, 15: 1368–1370.
Klein C, Evrard HC, Shapcott KA, Haverkamp S, Logothetis NK, Schmid MC. Cell-targeted optogenetics and electrical microstimulation reveal the primate koniocellular projection to supra-granular visual cortex. Neuron 2016, 90: 143–151.
Nurminen L, Merlin S, Bijanzadeh M, Federer F, Angelucci A. Top-down feedback controls spatial summation and response amplitude in primate visual cortex. Nat Commun 2018, 9: 2281.
Tang S, Zhang Y, Li Z, Li M, Liu F, Jiang H, et al. Large-scale two-photon imaging revealed super-sparse population codes in the V1 superficial layer of awake monkeys. Elife 2018, 7.
Deng C, Yuan H, Dai J. Behavioral manipulation by optogenetics in the nonhuman primate. Neuroscientist 2018, 24: 526–539.
Roth BL. DREADDs for neuroscientists. Neuron 2016, 89: 683–694.
Raper J, Galvan A. Applications of chemogenetics in non-human Primates. Curr Opin Pharmacol 2022, 64: 102204.
Kinoshita M, Matsui R, Kato S, Hasegawa T, Kasahara H, Isa K, et al. Genetic dissection of the circuit for hand dexterity in primates. Nature 2012, 487: 235–238.
Oguchi M, Tanaka S, Pan X, Kikusui T, Moriya-Ito K, Kato S, et al. Chemogenetic inactivation reveals the inhibitory control function of the prefronto-striatal pathway in the macaque brain. Commun Biol 2021, 4: 1088.
Oyama K, Hori Y, Nagai, Miyakawa N, Mimura K, Hirabayashi T, et al. Chemogenetic dissection of the primate prefronto-subcortical pathways for working memory and decision-making. Sci Adv 2021, 7: eabg4246.
Nagai Y, Kikuchi E, Lerchner W, Inoue KI, Ji B, Eldridge MA, et al. PET imaging-guided chemogenetic silencing reveals a critical role of primate rostromedial caudate in reward evaluation. Nat Commun 2016, 7: 13605.
Oyama K, Hori Y, Nagai Y, Miyakawa N, Mimura K, Hirabayashi T, et al. Chronic behavioral manipulation via orally delivered chemogenetic actuator in macaques. J Neurosci 2022, 42: 2552–2561.
Nagai Y, Miyakawa N, Takuwa H, Hori Y, Oyama K, Ji B, et al. Deschloroclozapine, a potent and selective chemogenetic actuator enables rapid neuronal and behavioral modulations in mice and monkeys. Nat Neurosci 2020, 23: 1157–1167.
Upright NA, Brookshire SW, Schnebelen W, Damatac CG, Hof PR, Browning PGF, et al. Behavioral effect of chemogenetic inhibition is directly related to receptor transduction levels in Rhesus monkeys. J Neurosci 2018, 38: 7969–7975.
Perez P, Chavret-Reculon E, Ravassard P, Bouret S. Using inhibitory DREADDs to silence LC neurons in monkeys. Brain Sci 2022, 12: 206.
Mimura K, Nagai, Inoue KI, Matsumoto J, Hori Y, Sato C, et al. Chemogenetic activation of nigrostriatal dopamine neurons in freely moving common marmosets. iScience 2021, 24: 103066.
Zhang Z, Shan L, Wang Y, Li W, Jiang M, Liang F, et al. Primate preoptic neurons drive hypothermia and cold defense. Innovation (Camb) 2023, 4: 100358.
Upright NA, Baxter MG. Effect of chemogenetic actuator drugs on prefrontal cortex-dependent working memory in nonhuman Primates. Neuropsychopharmacology 2020, 45: 1793–1798.
Galvan A, Raper J, Hu X, Paré JF, Bonaventura J, Richie CT, et al. Ultrastructural localization of DREADDs in monkeys. Eur J Neurosci 2019, 50: 2801–2813.
Humeau Y, Doussau F, Grant NJ, Poulain B. How botulinum and tetanus neurotoxins block neurotransmitter release. Biochimie 2000, 82: 427–446.
Israel MR, Morgan M, Tay B, Deuis JR. Toxins as tools: Fingerprinting neuronal pharmacology. Neurosci Lett 2018, 679: 4–14.
Macknik SL, Martinez-Conde S, Haglund MM. The role of spatiotemporal edges in visibility and visual masking. Proc Natl Acad Sci U S A 2000, 97: 7556–7560.
Watanabe M, Tanaka H, Uka T, Fujita I. Disparity-selective neurons in area V4 of macaque monkeys. J Neurophysiol 2002, 87: 1960–1973.
Grinvald A, Hildesheim R. VSDI: A new era in functional imaging of cortical dynamics. Nat Rev Neurosci 2004, 5: 874–885.
Chemla S, Chavane F. Voltage-sensitive dye imaging: Technique review and models. J Physiol Paris 2010, 104: 40–50.
Slovin H, Arieli A, Hildesheim R, Grinvald A. Long-term voltage-sensitive dye imaging reveals cortical dynamics in behaving monkeys. J Neurophysiol 2002, 88: 3421–3438.
Heider B, Nathanson JL, Isacoff EY, Callaway EM, Siegel RM. Two-photon imaging of calcium in virally transfected striate cortical neurons of behaving monkey. PLoS One 2010, 5: e13829.
Kondo T, Saito R, Otaka M, Yoshino-Saito K, Yamanaka A, Yamamori T, et al. Calcium transient dynamics of neural ensembles in the primary motor cortex of naturally behaving monkeys. Cell Rep 2018, 24: 2191-2195.e4.
Li M, Liu F, Jiang H, Lee TS, Tang S. Long-term two-photon imaging in awake macaque monkey. Neuron 2017, 93: 1049-1057.e3.
Sadakane O, Masamizu Y, Watakabe A, Terada SI, Ohtsuka M, Takaji M, et al. Long-term two-photon calcium imaging of neuronal populations with subcellular resolution in adult non-human Primates. Cell Rep 2015, 13: 1989–1999.
Guan SC, Ju NS, Tao L, Tang SM, Yu C. Functional organization of spatial frequency tuning in macaque V1 revealed with two-photon calcium imaging. Prog Neurobiol 2021, 205: 102120.
Guan SC, Zhang SH, Zhang YC, Tang SM, Yu C. Plaid detectors in macaque V1 revealed by two-photon calcium imaging. Curr Biol 2020, 30: 934-940.e3.
Jiang R, Andolina IM, Li M, Tang S. Clustered functional domains for curves and corners in cortical area V4. Elife 2021, 10: e63798.
Ju N, Jiang R, Macknik SL, Martinez-Conde S, Tang S. Long-term all-optical interrogation of cortical neurons in awake-behaving nonhuman Primates. PLoS Biol 2018, 16: e2005839.
Ju N, Li Y, Liu F, Jiang H, Macknik SL, Martinez-Conde S, et al. Spatiotemporal functional organization of excitatory synaptic inputs onto macaque V1 neurons. Nat Commun 2020, 11: 697.
Ju NS, Guan SC, Tao L, Tang SM, Yu C. Orientation tuning and end-stopping in macaque V1 studied with two-photon calcium imaging. Cereb Cortex 2021, 31: 2085–2097.
Kalaska JF. Emerging ideas and tools to study the emergent properties of the cortical neural circuits for voluntary motor control in non-human primates. F1000Research 2019, 8: F1000FacultyRev–749.
Macknik SL, Alexander RG, Caballero O, Chanovas J, Nielsen KJ, Nishimura N, et al. Advanced circuit and cellular imaging methods in nonhuman Primates. J Neurosci 2019, 39: 8267–8274.
Matsuzaki M, Ebina T. Optical deep-cortex exploration in behaving rhesus macaques. Nat Commun 2021, 12: 4656.
O’Shea DJ, Trautmann E, Chandrasekaran C, Stavisky S, Kao JC, Sahani M, et al. The need for calcium imaging in nonhuman Primates: New motor neuroscience and brain-machine interfaces. Exp Neurol 2017, 287: 437–451.
Oguchi M, Jiang J, Yoshioka TW, Tanaka YR, Inoue K, Takada M, et al. Microendoscopic calcium imaging of the primary visual cortex of behaving macaques. Sci Rep 2021, 11: 17021.
Santisakultarm TP, Kersbergen CJ, Bandy DK, Ide DC, Choi SH, Silva AC. Two-photon imaging of cerebral hemodynamics and neural activity in awake and anesthetized marmosets. J Neurosci Methods 2016, 271: 55–64.
Tang S, Zhang Y, Li Z, Li M, Liu F, Jiang H, et al. Large-scale two-photon imaging revealed super-sparse population codes in the V1 superficial layer of awake monkeys. Elife 2018, 7: e33370.
Trautmann EM, O’Shea DJ, Sun X, Marshel JH, Crow A, Hsueh B, et al. Dendritic calcium signals in rhesus macaque motor cortex drive an optical brain-computer interface. Nat Commun 2021, 12: 3689.
Dana H, Sun Y, Mohar B, Hulse BK, Kerlin AM, Hasseman JP, et al. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat Methods 2019, 16: 649–657.
Shemesh OA, Linghu C, Piatkevich KD, Goodwin D, Celiker OT, Gritton HJ, et al. Precision calcium imaging of dense neural populations via a cell-body-targeted calcium indicator. Neuron 2020, 107: 470-486.e11.
Chen Y, Jang H, Spratt PWE, Kosar S, Taylor DE, Essner RA, et al. Soma-targeted imaging of neural circuits by ribosome tethering. Neuron 2020, 107: 454-469.e6.
Lin MZ, Schnitzer MJ. Genetically encoded indicators of neuronal activity. Nat Neurosci 2016, 19: 1142–1153.
Gong Y, Huang C, Li JZ, Grewe BF, Zhang Y, Eismann S, et al. High-speed recording of neural spikes in awake mice and flies with a fluorescent voltage sensor. Science 2015, 350: 1361–1366.
Xu Y, Zou P, Cohen AE. Voltage imaging with genetically encoded indicators. Curr Opin Chem Biol 2017, 39: 1–10.
Park JE, Zhang XF, Choi SH, Okahara J, Sasaki E, Silva AC. Generation of transgenic marmosets expressing genetically encoded calcium indicators. Sci Rep 2016, 6: 34931.
Feng G, Jensen FE, Greely HT, Okano H, Treue S, Roberts AC, et al. Opportunities and limitations of genetically modified nonhuman primate models for neuroscience research. Proc Natl Acad Sci U S A 2020, 117: 24022–24031.
Ngai J. BRAIN 2.0: Transforming neuroscience. Cell 2022, 185: 4–8.
Wang L. Mu-Ming poo: China brain project and the future of Chinese neuroscience. Natl Sci Rev 2017, 4: 258–263.
Acknowledgements
This review was supported by grants from the Shanghai Municipal Science and Technology Major Project, the Strategic Priority Research Program of the Chinese Academy of Sciences, and the Lingang National Laboratory Key Project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, L., Liu, Z. Genetic Approaches for Neural Circuits Dissection in Non-human Primates. Neurosci. Bull. 39, 1561–1576 (2023). https://doi.org/10.1007/s12264-023-01067-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-023-01067-0