Abstract
Polyketides have been widely used clinically due to their significant biological activities, but the needed structural and functional diversity cannot be achieved by common chemical synthetic methods. The tool of combinatorial biosynthesis provides the possibility to produce “unnatural” natural drugs, which has achieved initial success. This paper provides an overview for the strategies of combinatorial biosynthesis in producing the structural and functional diversity of polyketides, including the redesign of metabolic flow, polyketide synthase (PKS) engineering, and PKS post-translational modification. Although encouraging progress has been made in the last decade, challenges still exist regarding the rational combinatorial biosynthesis of polyketides. In this review, the perspectives of polyketide combinatorial biosynthesis are also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polyketides are a class of natural products with different structures but similar biosynthetic pathways produced by bacteria, fungi or plants through the decarboxylative condensation of the lower carboxylic acids such as acetic acid, malonic acid, and butyric acid CoA derivatives, and the condensation is similar to the synthetic pathway of long-chain fatty acids [1]. Because of the effective biological activities of polyketides, they have been clinically used as antibiotics, such as antimicrobial erythromycin, anti-cancer agent doxorubicin, anti-parasitic agent avermectin, pesticide spinosad, and immunosuppressant rapamycin [2].
Polyketides are synthesized by a multifunctional polyketide synthase (PKS) family, and PKSs can be divided into three types (as shown in Table 1): Type I is called modular PKS, and it is a catalytic subunit module formed by a multi-enzyme complex, such as the 6-deoxyerythronolide B synthase (DEBS) [3]. Figure 1a shows that DEBS is composed of DEBS1, DEBS2, and DEBS3, and each component consists of two modules. Each module contains the following functional domains: acyltransferase (AT), which is responsible for transferring the building blocks from acyl-CoA precursors to acyl carrier protein (ACP); ketosynthase (KS), which is responsible for chain elongation; ACP, which is responsible for providing an intermediate-binding thiol-site; β-2-ketoreductase (KR), which is responsible for the reduction from a β-ketone to a hydroxyl group; dehydratase (DH), which is responsible for the dehydration of β-hydroxy intermediates; and enoyl reductase (ER), which is responsible for the reduction of unsaturated bonds [4]. After the reactions are completed, 6-deoxyerythronolide B is released by thioesterase (TE). Fugal nonreducing PKS (NR-PKS) contains KS, AT, ACP, as well as the product template (PT) domain that controls the aldol cyclization of poly-β-ketone intermediates [5]. Type II is an aromatic compound-forming type, such as the actinorhodin PKS shown in Fig. 1b, which is composed of 4–6 single-functional or bifunctional enzymes, and each type catalyzes the repeated reactions in the biosynthesis. McDaniel et al. [6] determined its minimal functional module that contains KS, the chain length factor (CLF), and ACP. Afterwards, KS was redesignated as KSα, and CLF was redesignated as KSβ (KSβ was previously regarded as the only factor to determine the chain length, but KSα also affects the chain length) [7]. Type III is the chalcone synthase identified from plants, bacteria, and fungi, and is a reusable homodimeric protein that is applied to catalyze the condensation of acyl-CoA without ACP. This type of PKS produces a number of compounds such as chalcones, pyrones, resorcinolic lipids, and stilbenes despite its simple structure. The first reported bacterial Type III PKS RppA comes from Streptomyces griseus and is responsible for flaviolin production as shown in Fig. 1c [8].
Representative reactions catalyzed by three typical bacterial PKSs. a Type I PKSs of DEBS, LD loading domain, TE thioesterase, M modular, KR ketoreductase, DH dehydratase, ER enoylreductase. b Schematic representation of Actinorhodin (ACT) biosynthesis by the act PKS. Eight malonyl-CoA converted to malonyl-ACP by malonyl-CoA:ACP transacylase (MAT) was consumed in the whole process, and the minimal PKS was used iteratively for seven rounds, then the intermediate was modified by various auxiliary PKS subunits and tailoring enzymes into actinorhodin. c Schematic representation of the formation of flaviolin. RppA catalyzes the five molecules of malonyl-CoA into 1,3,6,8-tetrahydroxynaphthalene (THN), and the THN is oxidized to yield flaviolin
Natural polyketides constitute a great proportion of current pharmacopeia; consequently, they have attracted much interest in PKS manipulation for “unnatural” polyketides [9]. The ordinary chemical modification method is difficult to operate due to the complex structure and stereochemistry of polyketides, and recent studies on polyketide biosynthesis by methods of traditional metabolic engineering [10] and systems biology [11,12,13] present the foundation for its combinatorial biosynthesis. The combinatorial biosynthesis is based on the understanding of the biosynthetic pathway to regulate, delete, add, substitute, and recombine the genes from different sources in vitro. Then, these genes can be introduced into an appropriate host to produce a series of compounds through oriented-synthesis. This strategy has been used to change the fixed biosynthetic pathways of many natural products, leading to the formation of many new “unnatural” products with new structures and biological activities.
This review mainly focuses on the combinatorial biosynthesis strategies used recently, including the redesign of the metabolic flow, PKS engineering, and PKS post-translational modifications in “unnatural” polyketide biosynthesis (Fig. 2). This work helps in explaining polyketide combinatorial biosynthesis on a global level. Last, the perspectives of polyketide combinatorial biosynthesis are also discussed.
Redesign of Metabolic Flow
With the development of genetic sequencing, genome mining has become a powerful tool to find new PKS genes [14], especially in Streptomyces, but the genes may be suppressed under standard laboratory growth conditions [15], or the products may have a low yield beyond the detection limit. The metabolic engineering of the biosynthetic pathway in microorganisms provides new starter units and redirects the metabolic flow to target products or new metabolites, which are the starting points of polyketide synthesis [16].
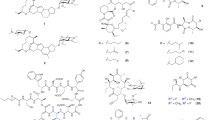
Three policies were mainly taken in metabolic flow redesign: (1) gene knockout to inhibit the metabolic offshoot. Gómez et al. [17] independently disrupted four enzymes (SlgN1, SlgN2, SlgL, which are responsible for condensing the portion of precursor 3-methylaspartate, and SlgO2, which is responsible for the cyclization of the characteristic bicyclic ketal) involved in the synthesis of streptolydigin in Streptomyces lydicus NRRL 2433, while the final mutants could not synthesize any streptolydigin as they could not produce any intermediates or branch products of streptolydigin. However, they synthesized three novel polyketides that were identified as 4-(2-carboxy-propylamino)-3-chloro-benzoic acid, 4-(2-carboxy-propylamino)-3-hydroxy-benzoic acid and 4-(2-carboxy-propylamino)-benzoic acid. Wu et al. [18] identified two putative epigenetic-related genes PfCclA and PfcclA in Pestalotiopsis fici, and the PfCclA or PfcclA deletion mutants were constructed, yielding significantly enhanced polyketide productions of pestaloficiols T–W and ficipyrone C as well as 11 macrodiolide ficiolides A–K, leading to the isolation of 15 novel polyketides (ficiolide K (1) contains a very rare 1,6-anhydro-pyranose moiety, which is shown in Fig. 3). (2) Modification of the precursor pathway. For the Type I and Type II PKSs, each step of the chain extension depends on the choice of module and precursor, significantly affecting the structure and activity of the final polyketide products. Lechner et al. [19] designed the biosynthesis of 36-methyl-FK506 (2 shown in Fig. 3) by polyketide precursor pathway engineering. In Streptomyces sp. KCTC 11604BP, they associated the branched extender isobutyrylmalonyl-CoA with the macrolide ansalactam A from Streptomyces sp. CNH189 by introducing a three-gene operon into FK506 biosynthesis, resulting in the replacement of the natural C-21 allyl side chain of FK506. Katsuyama et al. [20] developed a precursor-directed biosynthesis system with the artificially assembled pathway to produce the curcumin analogs in Escherichia coli, exogenously supplying various carboxylate precursors and analogs of p-coumaric acid, leading to the production of 17 novel curcuminoids. (3) Silence genes activation. Li et al. [21] activated a putative ansamycin gene cluster (nam) of Streptomyces sp. LZ35, and three new naphthalenic octaketide ansamycins (3–5 shown in Fig. 3) were produced with n-pentyl and n-butyl side chains. A similar strategy was also applied to the isolation of lomaiviticin, which is a product of the Type II PKS cluster in Salinispora tropica [22].
PKS Engineering
The PKSs use a series of carbon blocks to construct various polyketides with different stereochemical structures. The malonyl-CoA and methylmalonyl-CoA are the most common extending units for the biosynthesis of the macrolide while some other special modules can be synthesized by some polyketide-producing strains, such as ethylmalonyl-CoA, isobutyrylmalonyl-CoA, and methoxymalonyl-CoA, which are encoded by the partial gene clusters of PKS [23]. This is an efficient way to combine the PKSs derived from different organisms to form the hybrid gene assembly that could lead to the usage of alternative starter units or extending units. Among the three types of PKS, Type I PKSs attract more interest due to the separate modular structure. Type I PKSs are constituted by separate modules, so the recombinant and replacement of specific PKS modules provide the possibility to produce the “unnatural” products and the basis of using combinatorial biosynthesis as a method to recombine these modules (Fig. 3).
Two main strategies are used to manipulate modular PKS based on a catalytic cycle. One method is to change the catalytic cycle such as adding or deleting PKS modules and the other is to modify PKS modules without changing the catalytic cycle such as domain replacement, domain mutation, or modular rearrangement.
Strategies Based on Changing the Catalytic Cycle
The number and the type of modules control the chain length of the macrolide, so the unnatural chain length of the molecule structure could show differences in the insertion or deletion of modules.
The PKS assembly line could be extended by inserting modules, and proved the separate modules would be combined to synthesize novel macrolides. Rowe et al. [24] inserted the module 2 of rapamycin PKS (RAPS) into the mutable site of DEBS1-TE, and two novel tetraketides (6 and 7) were produced. Co-expression of the recombinant DEBS1 (RAPS Module 2)-TE with DEBS2 and DEBS3 in Saccharopolyspora erythraea contributed to the formation of four new octaketide macrolactones (9–12) with the normal product erythronolide B (8) (Fig. 4). In addition, novel macrolides can be obtained by module deletion or inactivation. Collismycin A belongs to the hybrid polyketide nonribosomal peptide that shows cytotoxic activity, and Garcia et al. [25] characterized its gene cluster and used the advantages of insertional inactivation and biocatalysis to obtain 12 collismycin analogs. Interestingly, among these derivatives, some showed potential neuroprotective activity but little cytotoxic activity.
Insertion of an additional module from RAPS Module 2 into the DEBS system [24]. a The bimodular DEBS1-TE, b RAPS Module 2 was inserted into DEBS1 between module 1 and module 2, c Co-expression of DEBS1 with DEBS2 and DEBS3, d Co-expression of the recombinant DEBS1(RAPS Module 2)-TE with DEBS2 and DEBS3
Strategies Based on Modular Modification: Domain Modification
The PKSs of multi-modules such as DEBS are organized in a modular fashion, and each module is responsible for a single extender unit installation. In each module, AT domains are required to select the starter and extender units, while the domains such as ER and KR are significant in stereoisomerism decision. Efforts on modifying such domains would also result in structural diversity.
AT Domain
It has been reported that the KS is remarkably tolerant to a diverse array of extender units, and the remarkable promiscuity of KS significantly expands the potential scope and utility of different starter and extender units [26]. Based on such knowledge, various strategies such as domain swap, domain hybrid, site-directed mutagenesis and trans-AT complementation that were reviewed by Dunn and Khosla [27] can be used to modify the AT domains to select an “unnatural” starter or extender unit. This review summarizes some new cases about AT domain modification in recent years.
AT domains control the extender unit selection and the side-chain diversity of the resulting polyketides. The replacement of native AT domain with heterologous analogs that possess different substrate specificities allows the incorporation of unnatural starter and extender units, providing another way for the diversification of PKS [28]. Rifamycin is the precursor of antibiotics that are effective against tuberculosis, leprosy, and AIDS-related mycobacterial infections. The AT6 domain of rifamycin PKS was substituted by the AT2 domain of rapamycin PKS, and two new rifamycin analogs, 24-desmethylrifamycin B and 24-desmethylrifamycin SV were produced. Then, the analogs were converted to 24-desmethylrifampicin (13 shown in Fig. 5) and showed excellent antibacterial activity against several rifampicin-resistant Mycobacterium tuberculosis strains [29]. By assaying PKS variants using easily accessible acyl-SNAC acceptor substrates, Koryakina et al. [30] created an effective method to discover AT mutations that could shift extender unit specificity toward non-native and non-natural substrates. Multiple mutants of the AT6 domain in DEBS3 were tested, and the Tyr189Arg mutation in combination with Val187Ala in DEBS3 was notably able to utilize extender units not originally screened for and produce novel compounds not detectable via the wild-type enzyme.
Two types of AT domains are known in the PKS assembly line. cis-acting ATs are coupled with the native ACP domains, whereas trans-acting ATs exist as stand-alone enzymes. The latter can transfer the extender unit onto one or more ACPs in a multi-modular PKS assembly line, such as the synthesis of disorazol [31] and bryostatin [32]. Compared to cis-AT, the trans-AT may have a wider substrate library, thus, it can be used to obtain “unnatural” analogs [33]. A promiscuous malonyl-CoA synthase variant was constructed by Koryakina et al. [34], and it can be applied to synthesize a broad range of malonyl-CoA extender units, some of these extender units are not found in natural biosynthetic systems. The utility of the variant in probing the acyl-CoA specificity of several trans-ATs has led to the discovery of poly-specificity toward non-natural extender units. Kinetic analysis and complement experiments of trans-ATs from the disorazole synthase and kirromycin synthase and cis-AT from DEBS were conducted, and the results showed that both carboxyacyl-CoA and ACP specificities are critical to the choice of a trans-AT in combination with a mutant PKS to generate novel polyketides [35].
Modifying Domain (KR, ER)
Modular PKSs bio-synthesize a variety of secondary metabolites by operating on the β-carbon processing degree using various combinations of KR and ER domains.
KR is responsible for determining the chirality of the polyketide product by reducing the β-carboxyl to hydroxyl [36]. Power et al. [37] inactivated the KR domain of amphotericin PKS, replaced the hydroxyl at C-7 of amphotericin B with a ketone group, and produced the 15-deoxy-15-oxo-amphotericin B (14 shown in Fig. 5) and 7-oxo-amphotericin B (15 shown in Fig. 5) analogs. Many researchers have studied the KR domain structure and its stereochemical mechanisms [36, 38, 39]. A mutation (Ser192Ile) in LanV, which is responsible for the 6R stereochemistry of landomycins in Streptomyces cyanogenus S136, led to the accumulation of gaudimycin C with 6S stereochemistry as a minor product [40].
Similar to the KR domain, the ER domain can also influence the configuration of PKS product. It has been reported that a unique tyrosine residue in the ER active site led to S configuration of the methyl branch, otherwise it has R configuration. In vivo, a mutation (Tyr to Val) in an erythromycin PKSs-derived ER caused the methyl branch configuration from S to R of the product, but the resembling mutation in a rapamycin-derived PKSs ER was insufficient to achieve the form switch [41]. Further studies have shown that a lysine residue might serve as a proton donor in enoyl reduction and the action analogously placed lysine in mammalian fatty acid synthases (FASs) [42]. The trans-acting ER in an iterative Type I PKS was also studied, providing a potential target in polyketide combinational biosynthesis [43].
Strategies Based on Modular Modification: Modular Rearrangement
PKS modular rearrangement (or a hybrid) can supply an effective tool to generate “unnatural” products based on the natural PKS genes illustrated. PKSs rational design and assembly by Menzella et al. [44, 45] established an important milestone toward the ultimate goal of making new bioactive polyketides. Liu et al. [46] replaced the starter unit ACP transacylase (SAT) domain of asperfuranone biosynthesis from Aspergillus nidulans with the SAT domain of sterigmatocystin biosynthesis, and the constructed hybrid PKS could utilize the hexanoyl starter unit synthesized by FAS to produce the novel compound (16) with the same length as asperfuranone (Fig. 6). Tautomycetin is a linear polyketide produced in Streptomyces sp. CK4412, while the mutant, in which the tautomycetin TE domain was swapped with the macrocyclic polyketide pikromycin TE domain, could produce an extra cyclized form called tautomycetin [47]. The biosynthesis of benzenediol lactone (BDL) polyketide, an important pharmacophore, is based on a pair of sequentially acting iterative polyketide synthases (iPKSs). Xu et al. [48] also used a plug-and-play combinational approach to express random pairs of iPKS subunits from four BDL model systems in yeast and created a diverse library of BDL congeners, including radilarin with an unnatural DAL14 skeleton and heat-shock response-inducing activity.
Biosynthetic pathway of asperfuranone and the reconstructed hybrid pathway of 16 [46]. Two PKSs AfoE and AfoG are involved in the asperfuranone biosynthesis. To construct the hybrid pathway, the SAT domain was swapped with the sterigmatocystin SAT, and the FAS encoded by stcJ and stcK was responsible for supplying the hexanoyl starter unit. SAT starter unit ACP transacylase, CMeT methyltransferase, R reductase
PKS Post-Translational Modification
The tailoring enzymes of the natural product gene cluster are involved in a series of reactions, such as the glycosylation [49], halogenations [50], hydroxylation [51], and some other modification reactions that make the various molecular structures. As a result, the tailoring enzymes have been used for the combinatorial biosynthesis to produce novel compounds [52].
Glycosylation
The glycosylation of the core ring molecules is one of the structural characteristics of polyketides, and the sugar moiety connected to the macrolide by the glycosyltransferases (GTs) is important to the biological activity. In recent years, many of the GTs have shown substrate flexibility to the glycosyl donor or receptor, and this flexibility is significant for the synthesis of the novel glycosylated macrolide. A Streptomyces venezuelae YJ003 mutant strain bearing the deletion of a desosamine biosynthetic (des) gene cluster was used as the host to accept plasmids with different exogenous deoxysugar gene cassettes and a substrate-flexible GT; as a result, amounts of macrolide antibiotic YC-17 analogs possessing unnatural sugars were produced [53]. The antibacterial assay showed that replacing the native sugar d-desosamine with l-rhamnose led to a better antibacterial activity. Except for macrolide polyketides, Kim et al. [54] constructed the one-pot combinatorial biosynthetic system through the co-cultivation of S. venezuelae mutants to produce aglycones and nucleotide-activated deoxysugars, facilitating the generation of seven novel glycosylated derivatives of aklavinone and ε-rhodomycinone by promoting the facile combination of aglycones with various sugars. YjiC, a substrate-flexible GT, was introduced into Streptomyces chromofuscus and contributed to the generation of two novel derivatives of herboxidiene: 18-O-β-d-glucopyranoside herboxidiene (17 shown in Fig. 7) and 18-O-β-d-glucopyranoside-25-demethyl herboxidiene (18 shown in Fig. 7) [55]. Such examples can also be found in spinosad [56], narbomycin [57], and other polyketides with sugar moieties attached to the aglycone core structures [58].
Oxidation
The oxidation is always important to the PKS post-translation tailoring step that diversifies the structural and biological characteristics of the macrolide, such as the hydroxylation [59] and the epoxidation [60] by the cytochrome P450 monooxygenase. Thus, a series of engineered monooxygenases would produce novel compounds with greater or new activities. Kudo et al. [61] inactivated the cytochrome P450 gfsF gene involved in the biosynthesis of the 16-membered cytotoxic antibiotic macrolide FD-891 in Streptomyces graminofaciens A-8890 that resulted in the production of a novel FD-891 analog 25-O-methyl-FD-892 (19 shown in Fig. 7) without the epoxide and hydroxyl group of FD-891. The novel compound 25-O-methyl-FD-892 is about 800 times less active than FD-891, but 25-O-methyl-FD-892 demonstrates that C8-C9 epoxide is critical for the cytotoxic activity [62]. A cytochrome P450 protein FkbD included in FK506 tailoring is responsible for α-keto amide formation by catalyzing a less common four-electron oxidation at the C-9 position, and the fkbD inactivation strain produced 9-deoxo-FK506, a new intermediate that also displayed antifungal activity [63]. Salcedo et al. [64] characterized the biosynthesis gene cluster for antitumor macrolides PM100117 and PM100118 from the marine actinobacteria Streptomyces caniferus GUA-06-05-006A, including the gonCP gene encoding a putative cytochrome P450 that was responsible for C16 oxygenation. The deletion of gonCP gene conduced to the production of PM100117 and PM100118 analogs that lost the C16 aglycone keto group or the naphthoquinone moiety, and these analogs showed enhanced in vitro cytotoxic activities relative to the parental products.
Perspective
Polyketide synthases are the attractive targets for combinatorial biosynthetic engineering to make “unnatural” products since its discovery. Behind the encouraging advances over the past two decades, there are still challenges that can be summarized as ecological challenges, enzymological challenges, and technological challenges [65]. However, great progress has been recently made in such fields. The development of sequencing technology and its steadily declining price made the availability of DNA sequences from organisms more easily. In addition, with the aid of a new computational method (such as Cluster Finder algorithm and anti-SMASH) [66], new PKS gene clusters can be elucidated more efficiently. Detecting and characterizing the polyketide products were also important in determining the gene cluster, and NMR spectroscopy [67] in such fields was widely used.
Since the PKS gene cluster annotation and its product characteristic are the basic step in PKS combinational biosynthesis, the architecture and states of the catalytic cycle of PKS should be the core part. The explanation of its biosynthesis mechanism can escape the blind manipulation. Technologies such as tandem equilibrium isotope exchange [68], electron cryo-microscopy [69], small-angle X-ray scattering (SAXS) [70, 71], and bottom-up liquid chromatography with Fourier transform ion cyclotron resonance mass spectrometry [72] are of great help. More attention has been paid to the studies of PKS on a systematical level either in vivo or vitro, further accelerating the pace in the rational engineering of polyketide assembly lines [70, 73]. In addition, the total enzymatic reconstruction of PKS biosynthesis pathway shows promising use in PKS biocatalytic analysis [74]. Remarkably, there has been a trend of combining polyketide combinatorial biosynthesis with synthetic biology to exploit and produce “unnatural” compounds of interest, and this strategy appears to have a bright future. Through decades of exploration and development, the rational combinational biosynthesis of polyketides has cut a striking figure. However, rapid advances of polyketide combinatorial biosynthesis still depend on deepening knowledge of modular PKSs and have a long way to go.
References
Jenke-Kodama H, Dittmann E (2009) Evolution of metabolic diversity: insights from microbial polyketide synthases. Phytochemistry 70(15/16):1858–1866
Staunton J, Weissman KJ (2001) Polyketide biosynthesis: a millennium review. Nat Prod Rep 18(4):380–416
Khosla C, Tang Y, Chen AY et al (2007) Structure and mechanism of the 6-deoxyerythronolide B synthase. Annu Rev Biochem 76(1):195–221
Liou GF, Khosla C (2003) Building-block selectivity of polyketide synthases. Curr Opin Chem Biol 7(2):279–284
Barajas JF, Shakya G, Moreno G et al (2017) Polyketide mimetics yield structural and mechanistic insights into product template domain function in nonreducing polyketide synthases. Proc Natl Acad Sci USA 114(21):4142–4148
McDaniel R, Ebert-Khosla S, Fu H et al (1994) Engineered biosynthesis of novel polyketides: influence of a downstream enzyme on the catalytic specificity of a minimal aromatic polyketide synthase. Proc Natl Acad Sci USA 91(24):11542–11546
Meurer G, Gerlitz M, Wendt-Pienkowski E et al (1997) Iterative type II polyketide synthases, cyclases and ketoreductases exhibit context-dependent behavior in the biosynthesis of linear and angular decapolyketides. Chem Biol 4(6):433–443
Funa N, Ohnishi Y, Fujii I et al (1999) A new pathway for polyketide synthesis in microorganisms. Nature 400(6747):897–899
McDaniel R, Thamchaipenet A, Gustafsson C et al (1999) Multiple genetic modifications of the erythromycin polyketide synthase to produce a library of novel “unnatural” natural products. Proc Natl Acad Sci USA 96(5):1846–1851
Xue C, Zhang X, Yu Z et al (2013) Up-regulated spinosad pathway coupling with the increased concentration of acetyl-CoA and malonyl-CoA contributed to the increase of spinosad in the presence of exogenous fatty acid. Biochem Eng J 81(4):47–53
Wang X, Zhang C, Wang M et al (2014) Genome-scale metabolic network reconstruction of Saccharopolyspora spinosa for spinosad production improvement. Microb Cell Fact 13(1):41
Zhang X, Xue C, Zhao F et al (2014) Suitable extracellular oxidoreduction potential inhibit rex regulation and effect central carbon and energy metabolism in Saccharopolyspora spinosa. Microb Cell Fact 13(1):98–108
Zhao F, Xue C, Wang M et al (2013) A comparative metabolomics analysis of Saccharopolyspora spinosa WT, WH124, and LU104 revealed metabolic mechanisms correlated with increases in spinosad yield. Biosci Biotechnol Biochem 77(8):1661–1668
Bachmann BO, Van Lanen SG, Baltz RH (2014) Microbial genome mining for accelerated natural products discovery: is a renaissance in the making? J Ind Microbiol Biotechnol 41(2):175–184
Bertrand S, Bohni N, Schnee S et al (2014) Metabolite induction via microorganism co-culture: a potential way to enhance chemical diversity for drug discovery. Biotechnol Adv 32(6):1180–1204
Xue C, Duan Y, Zhao F et al (2013) Stepwise increase of spinosad production in Saccharopolyspora spinosa by metabolic engineering. Biochem Eng J 72(2):90–95
Gómez C, Olano C, Palomino-Schätzlein M et al (2012) Novel compounds produced by Streptomyces lydicus NRRL 2433 engineered mutants altered in the biosynthesis of streptolydigin. J Antibiot 65(7):341–348
Wu G, Zhou H, Zhang P et al (2016) Polyketide production of pestaloficiols and macrodiolide ficiolides revealed by manipulations of epigenetic regulators in an endophytic fungus. Org Lett 18(8):1832–1835
Lechner A, Wilson MC, Ban YH et al (2013) Designed biosynthesis of 36-methyl-FK506 by polyketide precursor pathway engineering. ACS Synth Biol 2(7):379–383
Katsuyama Y, Hirose Y, Funa N et al (2010) Precursor-directed biosynthesis of curcumin analogs in Escherichia coli. Biosci Biotechnol Biochem 74(3):641–645
Li S, Li Y, Lu C et al (2015) Activating a cryptic ansamycin biosynthetic gene cluster to produce three new naphthalenic octaketide ansamycins with n-pentyl and n-butyl side chains. Org Lett 17(15):3706–3709
Kersten RD, Lane AL, Nett M et al (2013) Bioactivity-guided genome mining reveals the lomaiviticin biosynthetic gene cluster in salinispora tropica. ChemBioChem 14(8):955–962
Chan YA, Podevels AM, Kevany BM et al (2009) Biosynthesis of polyketide synthase extender units. Nat Prod Rep 26(1):90–114
Rowe CJ, Böhm IU, Thomas IP et al (2001) Engineering a polyketide with a longer chain by insertion of an extra module into the erythromycin-producing polyketide synthase. Chem Biol 8(5):475–485
Garcia I, Vior NM, González-Sabín J et al (2013) Engineering the biosynthesis of the polyketide-nonribosomal peptide collismycin a for generation of analogs with neuroprotective activity. Chem Biol 20(8):1022–1032
Koryakina I, McArthur JB, Draelos MM et al (2013) Promiscuity of a modular polyketide synthase towards natural and non-natural extender units. Org Biomol Chem 11(27):4449–4458
Dunn BJ, Khosla C (2013) Engineering the acyltransferase substrate specificity of assembly line polyketide synthases. J R Soc Interface 10(85):20130297
Park SR, Han AR, Ban YH et al (2010) Genetic engineering of macrolide biosynthesis: past advances, current state, and future prospects. Appl Microbiol Biotechnol 85(5):1227–1239
Nigam A, Almabruk KH, Saxena A et al (2014) Modification of rifamycin polyketide backbone leads to improved drug activity against rifampicin-resistant mycobacterium tuberculosis. J Biol Chem 289(30):21142–21152
Koryakina I, Kasey C, Mcarthur JB et al (2017) Inversion of extender unit selectivity in the erythromycin polyketide synthase by acyltransferase domain engineering. ACS Chem Biol 12(1):114–123
Carvalho R, Reid R, Viswanathan N et al (2005) The biosynthetic genes for disorazoles, potent cytotoxic compounds that disrupt microtubule formation. Gene 359(45):91–98
Lopanik NB, Shields JA, Buchholz TJ et al (2008) In vivo and in vitro trans-acylation by BryP, the putative bryostatin pathway acyltransferase derived from an uncultured marine symbiont. Chem Biol 15(11):1175–1186
Walker MC, Thuronyi BW, Charkoudian LK et al (2013) Expanding the fluorine chemistry of living systems using engineered polyketide synthase pathways. Science 341(6150):1089–1094
Koryakina I, McArthur J, Randall S et al (2013) Poly specific trans-acyltransferase machinery revealed via engineered acyl-CoA synthetases. ACS Chem Biol 8(1):200–208
Dunn BJ, Watts KR, Robbins T et al (2014) Comparative analysis of the substrate specificity of trans-versus cis-acyltransferases of assembly line polyketide synthases. Biochemistry 53(23):3796–3806
Caffrey P (2003) Conserved amino acid residues correlating with ketoreductase stereospecificity in modular polyketide synthases. ChemBioChem 4(7):654–657
Power P, Dunne T, Murphy B et al (2008) Engineered synthesis of 7-oxo-and 15-deoxy-15-oxo-amphotericins: insights into structure-activity relationships in polyene antibiotics. Chem Biol 15(1):78–86
Siskos AP, Baerga-Ortiz A, Bali S et al (2005) Molecular basis of Celmer’s rules: stereochemistry of catalysis by isolated ketoreductase domains from modular polyketide synthases. Chem Biol 12(10):1145–1153
Bonnett SA, Whicher JR, Papireddy K et al (2013) Structural and stereochemical analysis of a modular polyketide synthase ketoreductase domain required for the generation of a cis-alkene. Chem Biol 20(6):772–783
Paananen P, Patrikainen P, Kallio P et al (2013) Structural and functional analysis of angucycline C-6 ketoreductase LanV involved in landomycin biosynthesis. Biochemistry 52(31):5304–5314
Kwan DH, Sun Y, Schulz F et al (2008) Prediction and manipulation of the stereochemistry of enoylreduction in modular polyketide synthases. Chem Biol 15(11):1231–1240
Kwan DH, Leadlay PF (2010) Mutagenesis of a modular polyketide synthase enoylreductase domain reveals insights into catalysis and stereospecificity. ACS Chem Biol 5(9):829–838
Ames BD, Nguyen C, Bruegger J et al (2012) Crystal structure and biochemical studies of the trans-acting polyketide enoyl reductase LovC from lovastatin biosynthesis. Proc Natl Acad Sci USA 109(28):11144–11149
Menzella HG, Reid R, Carney JR et al (2005) Combinatorial polyketide biosynthesis by de novo design and rearrangement of modular polyketide synthase genes. Nat Biotechnol 23(9):1171–1176
Menzella HG, Carney JR, Santi DV (2007) Rational design and assembly of synthetic trimodular polyketide synthases. Chem Biol 14(2):143–151
Liu T, Chiang Y, Somoza AD et al (2011) Engineering of an “unnatural” natural product by swapping polyketide synthase domains in Aspergillus nidulans. J Am Chem Soc 133(34):13314–13316
Tripathi A, Choi SS, Sherman DH et al (2016) Thioesterase domain swapping of a linear polyketide tautomycetin with a macrocyclic polyketide pikromycin in Streptomyces sp. CK4412. J Ind Microbiol Biotechnol 43(8):1189–1193
Xu Y, Zhou T, Zhang S et al (2014) Diversity-oriented combinatorial biosynthesis of benzenediol lactone scaffolds by subunit shuffling of fungal polyketide synthases. Proc Natl Acad Sci U. S.A 111(34):12354–12359
Pérez M, Baig I, Braña AF et al (2008) Generation of new derivatives of the antitumor antibiotic mithramycin by altering the glycosylation pattern through combinatorial biosynthesis. ChemBioChem 9(14):2295–2304
Smith DR, Grüschow S, Goss RJ (2013) Scope and potential of halogenases in biosynthetic applications. Curr Opin Chem Biol 17(2):276–283
Perić-Concha N, Borovička B, Long PF et al (2005) Ablation of the otcC gene encoding a post-polyketide hydroxylase from the oxytetracyline biosynthetic pathway in Streptomyces rimosus results in novel polyketides with altered chain length. J Biol Chem 280(45):37455–37460
Olano C, Méndez C, Salas JA (2010) Post-PKS tailoring steps in natural product-producing actinomycetes from the perspective of combinatorial biosynthesis. Nat Prod Rep 27(4):571–616
Shinde PB, Han AR, Cho J et al (2013) Combinatorial biosynthesis and antibacterial evaluation of glycosylated derivatives of 12-membered macrolide antibiotic YC-17. J Biotechnol 168(2):142–148
Kim E, Song MC, Kim MS et al (2017) One-pot combinatorial biosynthesis of glycosylated anthracyclines by co-cultivation of Streptomyces strains producing aglycones and nucleotide deoxysugars. ACS Comb Sci 19(4):262–270
Pokhrel AR, Dhakal D, Jha AK et al (2015) Herboxidiene biosynthesis, production, and structural modifications: prospect for hybrids with related polyketide. Appl Microbiol Biotechnol 99(20):8351–8362
Gaisser S, Carletti I, Schell U et al (2009) Glycosylation engineering of spinosyn analogues containing an L-olivose moiety. Org Biomol Chem 7(8):1705–1708
Han AR, Shinde PB, Park JW et al (2012) Engineered biosynthesis of glycosylated derivatives of narbomycin and evaluation of their antibacterial activities. Appl Microbiol Biotechnol 93(3):1147–1156
Song MC, Kim E, Ban YH et al (2013) Achievements and impacts of glycosylation reactions involved in natural product biosynthesis in prokaryotes. Appl Microbiol Biotechnol 97(13):5691–5704
Han S, Pham TV, Kim JH et al (2015) Functional characterization of CYP107W1 from Streptomyces avermitilis and biosynthesis of macrolide oligomycin A. Arch Biochem Biophys 575(2):1–7
Montemiglio LC, Parisi G, Scaglione A et al (2016) Functional analysis and crystallographic structure of clotrimazole bound OleP, a cytochrome P450 epoxidase from Streptomyces antibioticus involved in oleandomycin biosynthesis. Biochim Biophys Acta Gen Subj 1860(3):465–475
Kudo F, Motegi A, Mizoue K et al (2010) Cloning and characterization of the biosynthetic gene cluster of 16-membered macrolide antibiotic FD-891: involvement of a dual functional cytochrome P450 monooxygenase catalyzing epoxidation and hydroxylation. ChemBioChem 11(11):1574–1582
Itagaki T, Kawamata A, Takeuchi M et al (2016) Synthesis and structure-activity relationship study of FD-891: importance of the side chain and C8–C9 epoxide for cytotoxic activity against cancer cells. J Antibiot 69:287–293
Chen D, Zhang L, Pang B et al (2013) FK506 maturation involves a cytochrome p450 protein-catalyzed four-electron C-9 oxidation in parallel with a C-31 O-methylation. J Bacteriol 195(9):1931–1939
Salcedo RG, Olano C, Gómez C et al (2016) Characterization and engineering of the biosynthesis gene cluster for antitumor macrolides PM100117 and PM100118 from a marine actinobacteria: generation of a novel improved derivative. Microb Cell Fact 15(1):1–19
Wong FT, Khosla C (2012) Combinatorial biosynthesis of polyketides: a perspective. Curr Opin Chem Biol 16(1):117–123
Cimermancic P, Medema MH, Claesen J et al (2014) Insights into secondary metabolism from a global analysis of prokaryotic biosynthetic gene clusters. Cell 158(2):412–421
Wang X, Wang H, Liu T et al (2014) A PKS I gene-based screening approach for the discovery of a new polyketide from Penicillium citrinum Salicorn 46. Appl Microbiol Biotechnol 98(11):4875–4885
Garg A, Xie X, Keatinge-Clay A et al (2014) Elucidation of the cryptic epimerase activity of redox-inactive ketoreductase domains from modular polyketide synthases by tandem equilibrium isotope exchange. J Am Chem Soc 136(29):10190–10193
Dutta S, Whicher JR, Hansen DA et al (2014) Structure of a modular polyketide synthase. Nature 510(7506):512–517
Davison J, Dorival J, Rabeharindranto H et al (2014) Insights into the function of trans-acyl transferase polyketide synthases from the SAXS structure of a complete module. Chem Sci 5(8):3081–3095
Dorival J, Annaval T, Risser F et al (2016) Characterization of intersubunit communication in the virginiamycin trans-acyl tran sferase polyketide synthase. J Am Chem Soc 138(12):4155–4167
Whicher JR, Dutta S, Hansen DA et al (2014) Structural rearrangements of a polyketide synthase module during its catalytic cycle. Nature 510(7506):560–564
Lowry B, Robbins T, Weng CH et al (2013) In vitro reconstitution and analysis of the 6-deoxyerythronolide B synthase. J Am Chem Soc 135(45):16809–16812
Sundaram S, Heine D, Hertweck C (2015) Polyketide synthase chimeras reveal key role of ketosynthase domain in chain branching. Nat Chem Biol 11(12):949–951
Acknowledgements
This work was supported by the Major Research Plan of Tianjin (No. 16YFXTSF00460).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Zhang, C., Ke, D., Duan, Y. et al. The Combinatorial Biosynthesis of “Unnatural” Products with Polyketides. Trans. Tianjin Univ. 24, 501–512 (2018). https://doi.org/10.1007/s12209-018-0151-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12209-018-0151-9