Abstract
Near-infrared (NIR) spectroscopy is a proven tool for the determination of food authenticity, mainly because of good classification results and the possibility of industrial use due to its easy and fast application. Since water shows broad absorption bands, the water content of a sample should be as low as possible. Freeze-drying is a commonly used preparatory step for this to reduce the water content in the sample. However, freeze-drying, also known as lyophilization, is very time-consuming impeding the widespread usage of NIR analysis as a rapid method for incoming goods inspections. We used a sample set of 72 almond samples from six economically relevant almond-producing countries to investigate the question of how important lyophilization is to obtain a well-performing classification model. For this approach, the samples were ground and lyophilized for 3 h, 24 h, and 48 h and compared to non-freeze-dried samples. Karl-Fischer titration of non-lyophilized samples showed that water contents ranged from 3.0 to 10.5% and remained constant at 0.36 ± 0.13% after a freeze-drying period of 24 h. The non-freeze-dried samples showed a classification accuracy of 93.9 ± 6.4%, which was in the same range as the samples which were freeze-dried for 3 h (94.2 ± 7.8%), 24 h (92.5 ± 8.7%), and 48 h (95.0 ± 9.0%). Feature selection was performed using the Boruta algorithm, which showed that signals from lipids and proteins are relevant for the origin determination. The presented study showed that samples with low water content, especially nuts, can be analyzed without the time-consuming preparation step of freeze-drying to obtain robust and fast results, which are especially required for incoming goods inspection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NIR spectroscopy has been established as a cost-efficient analytical tool to determine the authenticity of foodstuffs (Biancolillo et al. 2018; Firmani et al. 2019a, b; Karunathilaka et al. 2018; Ruoff et al. 2006). Two key advantages of this technology compared to other commonly used analytical approaches such as nuclear magnetic resonance and mass spectrometry are the rapid sample preparation and short measurement time, as well as the low acquisition and maintenance costs (Duarte et al. 2004; Malik et al. 2010). This makes NIR spectroscopy suitable for incoming goods inspection at food processing companies, which is not the case for most other methods due to the higher personnel and infrastructure requirements. NIR analysis also has the advantages of being a simple and environmentally friendly method, as no solvents are required (Armenta et al. 2008; Blanco and Villarroya 2002; Gałuszka et al. 2013). Forgoing lyophilization would make NIR spectroscopy much more attractive for incoming goods inspection in an industrial setting, as this is the most time-intensive step in sample preparation and without it a result is available within minutes.
NIR spectra of foods with a high water content show broad absorption signals of water in the regions at 6940 cm−1 and 5180 cm−1 (Büning-Pfaue 2003). These signals mask other important information needed for valid interpretation. Because of this, NIR measurements are often preceded by relatively time-consuming freeze-drying, which can take from a few hours to a few days, depending on sample volume and water content (Arndt et al. 2020b, 2021; Büning-Pfaue 2003; Richter et al. 2019; Segelke et al. 2020). However, this negates the advantage of rapid data acquisition. Nevertheless, in some cases, it is possible to generate well-performing classification models of foodstuffs even without freeze-drying (Aykas and Menevseoglu 2021; Ghidini et al. 2019; Ríos-Reina et al. 2018; Zhou et al. 2015). However, the question of whether and how much freeze-drying is necessary to obtain successful classification results in a reasonable amount of time has not yet been adequately answered. There have been studies on the analysis of meat, fish, cheese, and almonds comparing the classification results of non-lyophilized and lyophilized samples, but without investigating the influence of water on the NIR spectra in more depth (Andueza et al. 2013; Arndt et al. 2020a; Bázár et al. 2009; Xiccato et al. 2004). Arndt et al. (2020a) compared different preparation methods of almonds (Prunus dulcis Mill.) and their respective classification results with respect to their geographical origin. Whole, bisected, ground, and lyophilized almond samples were compared, with lyophilized almonds providing the best classification results. The approach was taken up and further investigated in this study by thoroughly including the freeze-drying process and its influence on the classification results.
Food fraud with almonds is observed due to the fundamentally high prices and, in addition, the strong price differences concerning different geographical origins (Food and Agriculture Organization of the United Nations 2021; Manning and Soon 2016). Moreover, almonds from Mallorca have a protected geographical indication, which makes them particularly attractive to conscious consumers (Publications Office of the European Union 2013). Therefore, in order to counteract possible food fraud, methods are needed that allow experimental determination of the geographical origin. Ideally, detection of this fraud should already take place at the point of receipt so that misdeclared goods can be rejected.
To identify the spectral features that are responsible for successful classification, we applied the random forest–based feature selection approach Boruta, which is based on comparing the importance of features with the maximum importance of so-called shadow features obtained by permuting of the data across observations (Kursa and Rudnicki 2010). We selected Boruta from a large number of existing selection techniques because it was shown to be the best performing in a comprehensive comparison study (Degenhardt et al. 2019; Janitza et al. 2018; Kursa and Rudnicki 2010; Seifert et al. 2019).
In order to make NIR spectroscopy more useful for rapid authentication of nuts in the context of incoming goods inspection, we investigated the water content during the freeze-drying process and used supervised multivariate methods to obtain classification accuracies and compare them between non-freeze-dried samples and samples which were freeze-dried for 3, 24, and 48 h. The aim was to investigate whether freeze-drying is required to obtain a well-functioning classification model, or if there is a minimum drying time to achieve this goal.
Materials and Methods
Samples
A total of 72 almond samples, twelve from each of the six most economically relevant growing countries (Australia, Iran, Italy, Morocco, Spain, the USA) were used. Samples were acquired shelled or unshelled. To reflect maximum biological diversity, samples were analyzed from crop years 2016 to 2019, with Australian samples only covering the years 2017 to 2019 due to the seasonal shift between hemispheres. In total, 12 samples from 2016, 19 samples from 2017, 23 samples from 2018, and 18 samples from 2019 were analyzed. More than 20 different varieties were analyzed, including sweet and bitter almonds. Samples were acquired either directly from growers or from personally known importers and were thus considered authentic. An overview about the analyzed samples can be found in Table S1 in the Supplementary Information. In addition, a non-authentic almond sample was purchased in a supermarket in Hamburg (Germany) and used as a quality control sample to track the lyophilization process, since a lot of sample material was necessary for this approach.
Sample Preparation
To study the lyophilization process, the quality control almond sample was first snap-frozen in liquid nitrogen and then ground under dry ice cooling (1/2 wt/wt, almonds/dry ice) using a knife mill (Grindomix GM 300, Retsch, Haan, Germany). Subsequently, the homogenized sample was aliquoted, each containing 4.0 ± 0.1 g homogenized powder. Each aliquot was lyophilized (Beta 1–8 LSCplus, Martin Christ Freeze Dryers GmbH, Osterode, Germany) for different lengths of times to monitor the decrease in water content. The following times were chosen as lyophilization durations: 30 min, 1 h, 2 h, 3 h, 6 h, 9 h, 1 day, 2 days, 3 days, 4 days, 5 days, 6 days, 7 days. Each sample was stirred halfway through each lyophilization duration by shaking the samples vigorously.
The authentic almond samples were prepared analogously to the quality control sample. Additionally, unshelled samples were cracked manually under dry ice cooling to remove the endocarp. The ground powders were then directly analyzed as well as lyophilized for different periods of time. Lyophilization times of 3 h, 24 h, and 48 h were chosen, using 4.0 ± 0.1 g per sample.
Karl-Fischer Titration
For the determination of the water content, approximately 300 mg of the sample was suspended with approximately 50 mL anhydrous methanol (CombiMethanol, Merck KGaA, Darmstadt, Germany) (Gallina et al. 2010). The sample was then titrated automatically to the end point (888 Titrando, Deutsche Metrohm GmbH & Co. KG, Filderstadt, Germany) using Karl-Fischer reagent (CombiTitrant 5, Merck KGaA, Darmstadt, Germany). For the determination of the titer, the consumption of 20 µL of distilled water was measured in triplicate.
NIR Spectroscopy
1.25 ± 0.01 g of each of the non-lyophilized or freeze-dried ground almonds were thawed at 22 °C (± 2 °C) in closed glass vials (52.0 mm × 22.0 mm × 1.2 mm, Nipro Diagnostics Germany GmbH, Ratingen, Germany). The samples were analyzed in reflectance mode using an FT-NIR spectrometer with an integration sphere (TANGO, Bruker Optics, Bremen, Germany). Fifty scans per spectrum were measured with a resolution of 2 cm−1 in a wavenumber range of 11,550–3,950 cm−1. Data were acquired using OPUS software (Bruker Optics, Bremen, Germany). To represent the population of samples as well as possible, five technical replicates were measured by shaking the vials vigorously between measurements.
NIR Spectra Preprocessing
MATLAB R2021b (The MathWorks Inc., Natick, USA) was used for all preprocessing steps. Scattered light effects are triggered by the nature of the samples due to different particle sizes and multiplicative scatter correction is applied to reduce the scattering effects in the NIR spectra (Dhanoa et al. 1994; Geladi et al. 1985). Since adjacent features correlate strongly, the number of features can be reduced, thereby decreasing the computation time (Engel et al. 2013). Therefore, binning was also performed by averaging the intensity of five adjacent wavenumbers, leaving 744 features for classification (Arndt et al. 2021; Shakiba et al. 2021). Finally, the arithmetic mean of the five measured spectra was calculated.
Multivariate Data Analysis
MATLAB R2021b was used for principal component analysis (PCA) as well as to train and validate classification models with various machine learning algorithms. The processed NIR data were used to investigate which machine learning algorithms showed the most promising initial results in the Classification Learner app. Samples were classified according to their country of origin for each of the four data sets. Various methods like decision trees, discriminant analyses, support vector machines, nearest neighbor classifiers, and ensembles of classifiers were applied on the data. The algorithms with the highest classification accuracy were linear discriminant analysis (LDA) and an ensemble of discriminant classifiers using random subspaces, which is a type of bagging (Ho 1998; Rokach 2010; Tharwat et al. 2017). The NIR data of the 72 samples and their corresponding class information were then split into six outer test and training sets with a repeated nested cross-validation (RNCV) stratified by geographical origin to validate the models, which was repeated five times. During model building, the training data was again split into six inner test and training sets (Varma and Simon 2006; Watermann et al. 2021). This approach was independently followed for each of the four data sets. RNCV minimizes the risk of outlier splits and gives an average classification result for each lyophilization method. Another advantage of a RNCV is that all samples are used for validation multiple times. The training and test accuracies are the mean average of all their respective splits from the RNCV. Further measures used were the macro-F1 score as an additional measure of a test’s accuracy and Fleiss’ kappa, which gives information on the conformity of the five classification results of each sample (Davies and Fleiss 1982; Tharwat 2021).
Feature Selection with Boruta
The R package Pomona (version 1.0.1, https://github.com/silkeszy/Pomona) was used for Boruta feature selection with the parameters: ntree = 10,000, min.node.size = 1, mtry = 142, importance = impurity_corrected, maxRuns = 100, and pValue = 0.01 (Degenhardt et al. 2019).
Results and Discussion
Change of the Water Content During Freeze-Drying of a Quality Control Almond Sample
To monitor the freeze-drying process, 4 g of aliquots of the quality control almond sample was lyophilized for different time periods. Figure 1 shows the water contents determined by Karl-Fischer titration and illustrated that the water content of the quality control sample remained at a constant range of 0.36 ± 0.13% after a lyophilization period of 24 h. Additionally, the NIR spectra of the lyophilized samples confirmed these results since the water band at 5300–4950 cm−1 decreased constantly until it stagnated after a lyophilization time of 24 h. Furthermore, the range between 7200 and 6050 cm−1 was also influenced by the drying process but only in the first 2 h of lyophilization (see Fig. S1 in the Supplementary Information).
In previous studies, a freeze-drying time of 48 h was used for 60 g of ground almond sample (Arndt et al. 2020a, 2021). Results from the present study suggested that a lyophilization duration of 24 h for 4 g of sample material might already be sufficient to reduce the water content to a constant level for successful classification.
Change of the Water Content During Freeze-Drying of Authentic Almond Samples
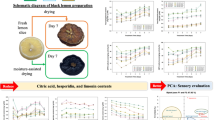
To further investigate the lyophilization duration, we compared the NIR spectra of 72 almond samples freeze-dried for 3 h, 24 h, and 48 h with the non-lyophilized samples. Figure 2a shows the processed and averaged NIR spectra of all samples from the different lyophilization methods. This was also shown for the second derivative of the processed NIR spectra in Fig. 3. The O–H bonds from water absorb at the wavenumbers 9550–9390 cm−1, 7100–6240 cm−1, and 5155–5150 cm−1 (Workman and Weyer 2012). The mean spectra (Fig. 2a) of the non-lyophilized samples (Lyo-0 h), the 3-h freeze-dried samples (Lyo-3 h), the 24-h freeze-dried samples (Lyo-24 h), and the 48-h freeze-dried samples (Lyo-48 h) showed large deviations in the regions between 7200–6050 cm−1 and 5300–4950 cm−1; however, the spectra of samples after 24 h of lyophilization and 48 h showed only minimal differences between each other. Similarly, large differences were observed in the region between 5400 and 5150 cm−1 using the second derivative (Fig. 3), while the differences in the region between 7200 and 6050 cm−1 were much smaller than in the non-derivative spectra. As expected, a longer lyophilization duration led to a decrease in water concentration and thus in absorbance in those spectral regions. Figure 2b shows the boxplots of the signal intensities of the bin at wavenumber 5167 cm−1 from the four different sets of samples, which represented the highest absorption in the water band region from the non-derivative data. The boxplots of the spectra without lyophilization and after 3 h of lyophilization did not overlap with each other and were significantly different (p-value: 3.10 × 10−76). Furthermore, there were also significant differences between 3 h of lyophilization and 24 h (p-value: 6.03 × 10−71) or 48 h (p-value: 7.57 × 10−78) of lyophilization. However, the boxplots of the spectra after freeze-drying durations of 24 and 48 h overlapped (p-value: 1.03 × 10−6), and only small differences occurred in total absorbance values compared to the other lyophilization durations, supporting the results of the Karl-Fischer titration that suggested that a lyophilization duration of 24 h is sufficient to minimize the water content. A total of nine non-lyophilized samples that showed high-, medium-, and low-intensity water bands in the NIR spectra were measured by Karl-Fischer titration to accurately determine the actual water contents. Overall, the water content of the samples ranged from 3.0 to 10.5% which is also reflected by the intensities of the water bands in the respective NIR spectra (see Fig. S2 and Table S2 in the Supplementary Information). In summary, the observed variations in the spectra reflected the different water contents that could be expected at different lyophilization durations. In the following sections, we evaluate the impact that the water signals had in the NIR spectra on the determination of the geographical origin of almonds.
Results of the systematic evaluation of the lyophilization duration on the NIR spectra of almonds: (a) processed (MSC, binning, mean averaging) NIR spectra where the average of all 72 samples from each of the four sample sets was used. (b) Boxplots of the signal intensity of the water signal at 5167 cm−1 from the four sample sets
Principal Component Analysis
PCA of the samples with different lyophilization durations was used as an unsupervised method to analyze whether the main variances in the data set can be exploited for origin separation and to identify outliers. Further, we also investigated the influence of the harvest year with this approach. The scores plots of the PCAs colored and marked by origin are shown in Fig. 4a–d and the loadings of the four different lyophilization methods are shown in Fig. 4e–h. Figure 5 shows the scores plot of the PCA were the samples of the Lyo-48 h data set were colored and marked by harvest year, while the scores plots using the other lyophilization methods can be found in Fig. S3 in the Supplementary Information. The scores plots of freeze-dried samples showed mostly similar results when considering geographical origin. The principal component (PC) 1 explained 56.5–57.7% of the total variance in the freeze-dried samples and 76.8% in the non-freeze-dried samples.
Scores and loadings plots of the processed NIR spectra of almond samples without lyophilization (Lyo-0 h, a, e), as well as after 3 h (Lyo-3 h, b, f), 24 h (Lyo-24 h, c, g), and 48 h (Lyo-48 h, d, h) of lyophilization (Australia: circle, USA: square, Italy: star, Iran: diamond, Spain: asterisk, Morocco: triangle). For the first three, the principal components 1 and 2, while for the latter the principal components 1 and 3 are shown
The Iranian (diamond), Moroccan (triangle), USA (square), and Australian (circle) samples showed a large overlap and clustered to the upper-left in the scores plot of non-lyophilized samples (see Fig. 4a). Spanish (asterisk) and Italian (star) samples clustered to the lower-left, close to each other, but with low overlap. Two Australian samples showed particularly high values for the first PC in the scores plot of non-lyophilized samples. Since the loadings of this PC (see Fig. 4e) were mainly affected by the water bands around 5150 cm−1 and 7000 cm−1, this meant that these samples had a high water content. This was confirmed by the assessment of these two individual NIR spectra (see Fig. S2 in the Supplementary Information) and the water contents were around 10% as determined by the Karl-Fischer titration (see Table S2 in the Supplementary Information). Spanish samples clustered more to the right than any other country, which meant that these samples overall had higher water contents than samples from other countries.
Iranian and Moroccan samples clustered to the lower part of the scores plot of Lyo-3 h (see Fig. 4b), which showed that these samples mostly had lower absorbance at 6000–5000 cm−1 (see Fig. 4f). Italian samples mainly clustered to the right and Iranian samples to the left, with the band around 4400–4200 cm−1 explaining PC1 the most. Spanish, Australian, and USA samples were mostly in the center of the scores plot and showed high intragroup variances. The water signal around 5200 cm−1 had a small impact on the PC1 and a large influence on PC2, which illustrated the smaller effect of the water signal on the total variance of samples.
Scores and loadings plot of Lyo-24 h and Lyo-48 h (see Fig. 4c, d, g, h) were very similar to each other. Spanish and Italian samples clustered more to the lower-right of both scores plots with more influence from the PC1 than the PC2. In both scores plots, Iranian and Moroccan samples clustered in the upper-left and USA samples in the center. Australian samples clustered in the center in Lyo-48 h and more to the right in Lyo-24 h. Bands between 7100 and 6000 cm−1 had negative values and between 6000 and 5200 cm−1 had positive values for PC1 and the reverse was the case for PC2, which was important for the clustering of Italian, Spanish, Iranian, and Moroccan samples.
As the metabolome is also strongly affected by other external factors than the country of origin, such as different climatic conditions in different harvest years, it is important to investigate a balanced sample set with samples from several harvest years to yield a robust model. Still, the influence of the harvest year was investigated by PCA. For this, the Lyo-48 h samples were marked in the scores plots by harvest year in Fig. 5 (see Fig. 4h for the loadings plot). No distinct clusters were apparent in the scores plot corresponding to a harvest year; instead, samples from all harvest years scattered across the plot, demonstrating that the influence of geographic origin is greater than the influence of harvest year since an impact of the harvest year is not apparent in the PC1 or PC3. As Arndt et al. (2021) outlined, several possible influences such as climate, storage, and anthropogenic influences can vary between different harvest years, which makes the impact of the harvest year difficult to evaluate.
Overall, the PCA did not show clear separations between samples originating from different countries, so supervised machine learning methods were used to achieve this goal.
Supervised Approaches
The processed NIR data sets were used to train supervised classification models for each lyophilization duration. First, we used the Classification Learner app from MATLAB R2021b to compare the classification performance of different algorithms. The results of the different algorithms are shown in Table S3 in the Supplementary Information. LDA provided the best results for each data set and was therefore selected for subsequent model building and classification. The results of the different lyophilization methods, i.e., the averaged training and classification accuracy, macro-F1 score, and Fleiss’ kappa, are shown in Table 1. The classification performances of the different lyophilization durations were very similar, as shown by the test accuracies (92.5–95.0%), macro-F1 scores (92.0–94.7%), and Fleiss’ kappa coefficients (88.0–94.5%). Taken together, this suggested that the models predicted the samples well and that the five iterations were largely consistent. More interesting than the specific model performances was that no trend is observed with respect to longer lyophilization durations.
In a previous study, 64 authentic almond samples yielded in an accuracy of 71.9 ± 3.5% for the determination of the geographical origin of homogenized almonds and a slightly higher classification accuracy of 80.2 ± 1.9% for predicting lyophilized almonds (Arndt et al. 2020a). We did not observe differences in classification accuracies of lyophilized and homogenized sample material. Besides the pre-treatment of the data, the main difference was the use of the classifier: we used LDA, while Arndt et al. (2020a) used a support vector machine algorithm. Even though both methods used RNCV to build robust models, the data evaluation could lead to different results. Another possible reason for the different classification results is that we used classes with twelve samples each. Larger class sizes generally lead to better classification performance because they better describe the extremities and centroids of the classes (Foody et al. 2006).
Overall, only three samples in all four sample sets were misclassified at least once. Table S4–S7 in the Supplementary Information show the samples and the countries to which they were misclassified in the five iterations of the RNCV. One of the three frequently misclassified samples was a 2018 Australian sample that was classified as Australian only once, as Spanish twelve times, and as Moroccan seven times. As could be seen in the scores plots of all four data sets (Fig. 4a–d), the reason for this was likely due to the large intragroup variance of the Australian samples. Another sample that was misclassified at least once across all four lyophilization methods was an Iranian sample from 2016 that was classified as Australian and Italian once, as Spanish seven times, as Iranian eight times, and as USA three times. This could be explained by the metadata of the sample set: Almost all samples from Australia, Spain, Italy, and the USA were sweet almonds, while almost all Moroccan and Iranian samples were bitter almonds. Since the only Iranian sweet almond sample was misclassified, it can be concluded that the model was probably only valid for Iranian bitter almonds samples. The third misclassified sample was an Italian sample from 2016, which was classified once as Iranian, four times as Italian, and 15 times as Moroccan. The reason for these misclassifications was again related to the differences in bitter and sweet almonds, as this sample belongs to a bitter variety. Moroccan and Iranian sweet almonds and Italian bitter almonds were underrepresented in their respective sample groups. Thus, classification during RNCV resulted in more misclassifications as their sample groups used for model training were not accurately representative for these subspecies. However, Moroccan and Iranian samples showed few misclassifications between each other. Similarly, countries that mainly contained sweet almonds (Australia, Italy, Spain, USA) also showed few misclassifications. Thus, an influence of the subspecies, although challenging to quantify, did not result in low classification accuracies or a high number of misclassifications between countries mainly containing a specific subspecies.
These results suggested that NIR spectroscopy is able to discriminate between sweet and bitter almonds, which was already demonstrated by Borràs et al. (2014). Cortés et al. (2018) even predicted the amygdalin content of almonds, determined by high-performance liquid chromatography coupled with a diode array detector by applying partial least squares regression on NIR data. This suggests that the amygdalin content had an effect on NIR spectra and the discrimination between sweet and bitter almonds.
Feature Selection with Boruta
A feature selection was conducted to find out which spectral regions and thus which substance classes were relevant for the determination of the geographical origin of almonds. The goal was to obtain more information on the NIR spectra, rather than to develop a targeted approach with fewer features. For this purpose, we used the feature selection algorithm Boruta, which selected 43, 40, 41, and 35 features in the data sets of spectra obtained after lyophilization durations of 0, 3, 24, and 48 h, respectively (see Table S8 in the Supplementary Information). The selected features for each lyophilization method were subsequently used to build classification models. Initially, an LDA was used for this purpose, but classification accuracies of less than 60% were obtained. Therefore, the best classification method was then determined in a similar manner as before utilizing all features with the results shown in Table S9 in the Supplementary Information. We identified an ensemble of discriminant classifiers using random subspaces with test accuracies between 81.1 and 85.0% as the best performing method (Table 2). These classification performances were about 10% lower than those using all 744 features, indicating that Boruta captured the features well that were most important for geographic identification. All selected features of the four preparation sets are listed in Table S8 in the Supplementary Information and are highlighted in Fig. 6 in the processed averaged NIR spectrum of the non-lyophilized samples.
Features were selected from different parts of the spectrum that can be assigned to different classes of molecules, mainly lipids and proteins, but a definite assignment was not possible. Selected features from lipids were apparent in the range between 8700 and 8500 cm−1 and originated from the second overtone of C–H vibrations and the fourth overtone of C = O vibrations. In addition, the features between 6000 and 5800 cm−1 could be assigned to the first overtone of saturated and unsaturated hydrocarbon chains, while features between 7400 and 7000 cm−1 could be assigned to C–H bonds associated with aliphatic hydrocarbons (Burns and Ciurczak 2007; Workman and Weyer 2012).
There were many bands associated with peptides and proteins selected by Boruta. At 4410 cm−1 peptide vibrations of C = O and N–H could be observed in α-helices or β-sheets (Workman and Weyer 2012). Between 7000 and 6000 cm−1, there were several features corresponding to the first overtone of N–H and O–H bonds caused by the peptide bond and side chains of amino acids as well as carbohydrates (Burns and Ciurczak 2007; Workman and Weyer 2012).
Boruta selected many features (26 of 43 for the non-lyophilized samples) in the range of the water signal between 7100 and 6240 cm−1, originating from N–H and O–H bonds of proteins and carbohydrates. The NIR spectra of the non-lyophilized samples showed more intensive water signals, which can be attributed to the higher water content of the samples. This raised the question of whether water content was relevant for the determination of the provenance, which would be problematic since it was affected by post-harvest factors such as drying conditions and transport. Therefore, boxplots were generated depicting the intensity of the water band at 5167 cm−1 for each individual origin of the almond samples (Fig. 7). They showed that the intensity and hence the water content of the samples were comparable, and the samples from different countries showed no significant differences (p-value > 0.05). In addition, the bins around the water signal at 5150 cm−1 were not selected by Boruta. It could be concluded that the water content has no major impact on the classification model. To evaluate the ability of the selected features in this region to discriminate the origin of almonds, an additional classification model was built based on the ensemble algorithm using only the selected features in the range between 7200 and 6000 cm−1 (see Table S8 in the Supplementary Information). The test accuracy of 70.6 ± 13.9% showed that these features carry very important information for classification.
In our analysis, the features at 7194 cm−1 and 5794 cm−1 were selected by Boruta. Borràs et al. (2014) identified these signals as relevant for the differentiation between sweet and bitter almonds. Although the question of distinguishing sweet and bitter almonds was not the issue in this study, it was still relevant because almost all Iranian and Moroccan samples were bitter almonds, and almost all other samples were sweet almonds.
Conclusion
The aim of this work was to investigate how water content affects the classification results of almond geographical origin determination by NIR spectroscopy. Omitting freeze-drying resulted in a test accuracy of 93.9 ± 6.4%, which was in the same range as the samples lyophilized for 3, 24, and 48 h, with no improvements for longer lyophilization durations. Although the water content of the non-freeze-dried samples ranged from 3.0 to 10.5%, it had no effect on classification. Selected features showed that lipids and proteins were responsible for successful discrimination. This study demonstrated that it is possible to determine the geographical origin of non-lyophilized almonds by NIR spectroscopy as a rapid and cost-effective analytical method for incoming goods inspection and food fraud investigation. The results also suggest that freeze-drying may not be necessary for such analytical issues. This is particularly useful for foods with low water content and high price such as nuts and seeds.
Data Availability
The data sets created as part of the current study are available in the Zenodo repository at: https://doi.org/10.5281/zenodo.6241567.
References
Andueza D, Agabriel C, Constant I, Lucas A, Martin B (2013) Using visible or near infrared spectroscopy (NIRS) on cheese to authenticate cow feeding regimes. Food Chem 141:209–214. https://doi.org/10.1016/j.foodchem.2013.02.086
Armenta S, Garrigues S, La Guardia M, de, (2008) Green analytical chemistry. TrAC, Trends Anal Chem 27:497–511. https://doi.org/10.1016/j.trac.2008.05.003
Arndt M, Rurik M, Drees A, Bigdowski K, Kohlbacher O, Fischer M (2020) Comparison of different sample preparation techniques for NIR screening and their influence on the geographical origin determination of almonds (Prunus dulcis MILL.). Food Control 115:107302. https://doi.org/10.1016/j.foodcont.2020.107302
Arndt M, Drees A, Ahlers C, Fischer M (2020b) Determination of the geographical origin of walnuts (Juglans regia L.) using near-infrared spectroscopy and chemometrics. Foods 9. 10.3390/foods9121860
Arndt M, Rurik M, Drees A, Ahlers C, Feldmann S, Kohlbacher O, Fischer M (2021) Food authentication: determination of the geographical origin of almonds (Prunus dulcis Mill.) via near-infrared spectroscopy. Microchemical Journal 160:105702. https://doi.org/10.1016/j.microc.2020.105702
Aykas DP, Menevseoglu A (2021) A rapid method to detect green pea and peanut adulteration in pistachio by using portable FT-MIR and FT-NIR spectroscopy combined with chemometrics. Food Control 121:107670. https://doi.org/10.1016/j.foodcont.2020.107670
Bázár G, Kövér G, Locsmándi L, Andrássy-Baka G, Romvári R (2009) Identification of traditionally reared Mangalica pig’s meat by near infrared spectroscopy using generalised partial least squares in open source R project—a feasibility model study. J near Infrared Spectrosc 17:119–125. https://doi.org/10.1255/jnirs.834
Biancolillo A, de Luca S, Bassi S, Roudier L, Bucci R, Magrì AD, Marini F (2018) Authentication of an Italian PDO hazelnut (“Nocciola Romana”) by NIR spectroscopy. Environ Sci Pollut Res Int 25:28780–28786. https://doi.org/10.1007/s11356-018-1755-2
Blanco M, Villarroya I (2002) NIR spectroscopy: a rapid-response analytical tool. TrAC, Trends Anal Chem 21:240–250. https://doi.org/10.1016/S0165-9936(02)00404-1
Borràs E, Amigo JM, van den Berg F, Boqué R, Busto O (2014) Fast and robust discrimination of almonds (Prunus amygdalus) with respect to their bitterness by using near infrared and partial least squares-discriminant analysis. Food Chem 153:15–19. https://doi.org/10.1016/j.foodchem.2013.12.032
Büning-Pfaue H (2003) Analysis of water in food by near infrared spectroscopy. Food Chem 82:107–115. https://doi.org/10.1016/S0308-8146(02)00583-6
Burns DA, Ciurczak EW (2007) Handbook of near-infrared analysis, 3rd edn. CRC Press, Boca Raton, FL
Cortés V, Talens P, Barat JM, Lerma-García MJ (2018) Potential of NIR spectroscopy to predict amygdalin content established by HPLC in intact almonds and classification based on almond bitterness. Food Control 91:68–75. https://doi.org/10.1016/j.foodcont.2018.03.040
Davies M, Fleiss JL (1982) Measuring agreement for multinomial data. Biometrics 38:1047–1051. https://doi.org/10.2307/2529886
Degenhardt F, Seifert S, Szymczak S (2019) Evaluation of variable selection methods for random forests and omics data sets. Brief Bioinform 20:492–503. https://doi.org/10.1093/bib/bbx124
Dhanoa MS, Lister SJ, Sanderson R, Barnes RJ (1994) The link between multiplicative scatter correction (MSC) and standard normal variate (SNV) transformations of NIR spectra. J near Infrared Spectrosc 2:43–47. https://doi.org/10.1255/jnirs.30
Duarte IF, Barros A, Almeida C, Spraul M, Gil AM (2004) Multivariate analysis of NMR and FTIR data as a potential tool for the quality control of beer. J Agric Food Chem 52:1031–1038. https://doi.org/10.1021/jf030659z
Engel J, Gerretzen J, Szymańska E, Jansen JJ, Downey G, Blanchet L, Buydens LMC (2013) Breaking with trends in pre-processing? TrAC, Trends Anal Chem 50:96–106. https://doi.org/10.1016/j.trac.2013.04.015
Firmani P, Bucci R, Marini F, Biancolillo A (2019a) Authentication of “Avola almonds” by near infrared (NIR) spectroscopy and chemometrics. J Food Compos Anal 82:103235. https://doi.org/10.1016/j.jfca.2019.103235
Firmani P, de Luca S, Bucci R, Marini F, Biancolillo A (2019b) Near infrared (NIR) spectroscopy-based classification for the authentication of Darjeeling black tea. Food Control 100:292–299. https://doi.org/10.1016/j.foodcont.2019.02.006
Food and Agriculture Organization of the United Nations (2021) Production of crops. http://www.fao.org/faostat/en/#data. Accessed 25 February 2021
Foody GM, Mathur A, Sanchez-Hernandez C, Boyd DS (2006) Training set size requirements for the classification of a specific class. Remote Sens Environ 104:1–14. https://doi.org/10.1016/j.rse.2006.03.004
Gallina A, Stocco N, Mutinelli F (2010) Karl Fischer titration to determine moisture in honey: a new simplified approach. Food Control 21:942–944. https://doi.org/10.1016/j.foodcont.2009.11.008
Gałuszka A, Migaszewski Z, Namieśnik J (2013) The 12 principles of green analytical chemistry and the significance mnemonic of green analytical practices. TrAC, Trends Anal Chem 50:78–84. https://doi.org/10.1016/j.trac.2013.04.010
Geladi P, MacDougall D, Martens H (1985) Linearization and scatter-correction for near-infrared reflectance spectra of meat. Appl Spectrosc 39:491–500. https://doi.org/10.1366/0003702854248656
Ghidini S, Varrà MO, Dall’Asta C, Badiani A, Ianieri A, Zanardi E, (2019) Rapid authentication of European sea bass (Dicentrarchus labrax L.) according to production method, farming system, and geographical origin by near infrared spectroscopy coupled with chemometrics. Food Chem 280:321–327. https://doi.org/10.1016/j.foodchem.2018.12.075
Ho TK (1998) The random subspace method for constructing decision forests. IEEE Trans Pattern Anal Machine Intell 20:832–844. https://doi.org/10.1109/34.709601
Janitza S, Celik E, Boulesteix A-L (2018) A computationally fast variable importance test for random forests for high-dimensional data. Adv Data Anal Classif 12:885–915. https://doi.org/10.1007/s11634-016-0276-4
Karunathilaka SR, Yakes BJ, He K, Chung JK, Mossoba M (2018) Non-targeted NIR spectroscopy and SIMCA classification for commercial milk powder authentication: a study using eleven potential adulterants. Heliyon 4:e00806. https://doi.org/10.1016/j.heliyon.2018.e00806
Kursa MB, Rudnicki WR (2010) Feature selection with the Boruta package. J. Stat. Soft. 36. https://doi.org/10.18637/jss.v036.i11
Malik AK, Blasco C, Picó Y (2010) Liquid chromatography-mass spectrometry in food safety. J Chromatogr A 1217:4018–4040. https://doi.org/10.1016/j.chroma.2010.03.015
Manning L, Soon JM (2016) Food safety, food fraud, and food defense: a fast evolving literature. J Food Sci 81:R823–R834. https://doi.org/10.1111/1750-3841.13256
Publications Office of the European Union (2013) COUNCIL REGULATION (EC) No 510/2006 on the protection of geographical indications and designations of origin for agricultural products and foodstuffs ‘ALMENDRA DE MALLORCA’/‘ALMENDRA MALLORQUINA’/‘AMETLLA DE MALLORCA’/ ‘AMETLLA MALLORQUINA.’ Official Journal of the European Union C 317(56):8–12
Richter B, Rurik M, Gurk S, Kohlbacher O, Fischer M (2019) Food monitoring: screening of the geographical origin of white asparagus using FT-NIR and machine learning. Food Control 104:318–325. https://doi.org/10.1016/j.foodcont.2019.04.032
Ríos-Reina R, García-González DL, Callejón RM, Amigo JM (2018) NIR spectroscopy and chemometrics for the typification of Spanish wine vinegars with a protected designation of origin. Food Control 89:108–116. https://doi.org/10.1016/j.foodcont.2018.01.031
Rokach L (2010) Ensemble-based classifiers. Artif Intell Rev 33:1–39. https://doi.org/10.1007/s10462-009-9124-7
Ruoff K, Luginbühl W, Bogdanov S, Bosset JO, Estermann B, Ziolko T, Amado R (2006) Authentication of the botanical origin of honey by near-infrared spectroscopy. J Agric Food Chem 54:6867–6872. https://doi.org/10.1021/jf060770f
Segelke T, Schelm S, Ahlers C, Fischer M (2020) Food authentication: truffle (Tuber spp.) Species differentiation by FT-NIR and chemometrics. Foods 9. 10.3390/foods9070922
Seifert S, Gundlach S, Szymczak S (2019) Surrogate minimal depth as an importance measure for variables in random forests. Bioinformatics 35:3663–3671. https://doi.org/10.1093/bioinformatics/btz149
Shakiba N, Gerdes A, Holz N, Wenck S, Bachmann R, Schneider T, Seifert S, Fischer M, Hackl T (2021) Determination of the geographical origin of hazelnuts (Corylus avellana L.) by near-infrared spectroscopy (NIR) and a low-level fusion with nuclear magnetic resonance (NMR). Microchemical Journal. https://doi.org/10.1016/j.microc.2021.107066
Tharwat A (2021) Classification assessment methods. ACI 17:168–192. https://doi.org/10.1016/j.aci.2018.08.003
Tharwat A, Gaber T, Ibrahim A, Hassanien AE (2017) Linear discriminant analysis: a detailed tutorial. AIC 30:169–190. https://doi.org/10.3233/AIC-170729
Varma S, Simon R (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics 7:91. https://doi.org/10.1186/1471-2105-7-91
Watermann S, Schmitt C, Schneider T, Hackl T (2021) Comparison of regular, pure shift, and fast 2D NMR experiments for determination of the geographical origin of walnuts. Metabolites 11. https://doi.org/10.3390/metabo11010039
Workman J, Weyer L (2012) Practical guide and spectral atlas for interpretive near-infrared spectroscopy, 2nd edn. CRC Press, Boca Raton, FL
Xiccato G, Trocino A, Tulli F, Tibaldi E (2004) Prediction of chemical composition and origin identification of European sea bass (Dicentrarchus labrax L.) by near infrared reflectance spectroscopy (NIRS). Food Chem 86:275–281. https://doi.org/10.1016/j.foodchem.2003.09.026
Zhou X, Yang Z, Haughey SA, Galvin-King P, Han L, Elliott CT (2015) Classification the geographical origin of corn distillers dried grains with solubles by near infrared reflectance spectroscopy combined with chemometrics: a feasibility study. Food Chem 189:13–18. https://doi.org/10.1016/j.foodchem.2014.09.104
Acknowledgements
We thank Tobias Schneider for aiding in data preparation. We thank René Bachmann, Marie Sophie Müller, Johannes Brockelt, Felix Schmauder, and Stephanie Watermann for providing expertise. We would also like to thank the almond suppliers for the samples.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was developed within the project “Food Profiling – Development of analytical tools for experimental verification of the origin and identity of food”. This project (funding code: 2816500914) is funded by the Federal Ministry of Food and Agriculture (BMEL) based on a resolution of the German Bundestag. Project funding was provided by the Federal Agency for Agriculture and Food (BLE) within the framework of the Innovation Promotion Program.
Author information
Authors and Affiliations
Contributions
Henri Lösel: conceptualization, investigation, visualization, methodology, writing—original draft.
Navid Shakiba: conceptualization, investigation, visualization, methodology, writing—original draft.
Soeren Wenck: investigation, writing—review and editing.
Phat Le Tan: investigation, writing—review and editing.
Maike Arndt: investigation, writing—review and editing.
Stephan Seifert: supervision, writing—review and editing.
Thomas Hackl: supervision, resources, writing—review and editing.
Markus Fischer: conceptualization, supervision, resources, writing—review and editing.
Corresponding author
Ethics declarations
Ethical Approval
This article does not contain any studies with human or animal subjects.
Informed Consent
Not applicable.
Conflict of Interest
Henri Lösel declares that he has no conflict of interest. Navid Shakiba declares that he has no conflict of interest. Soeren Wenck declares that he has no conflict of interest. Phat Le Tan declares that he has no conflict of interest. Maike Arndt declares that she has no conflict of interest. Stephan Seifert declares that he has no conflict of interest. Thomas Hackl declares that he has no conflict of interest. Markus Fischer declares that he has no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lösel, H., Shakiba, N., Wenck, S. et al. Impact of Freeze-Drying on the Determination of the Geographical Origin of Almonds (Prunus dulcis Mill.) by Near-Infrared (NIR) Spectroscopy. Food Anal. Methods 15, 2847–2857 (2022). https://doi.org/10.1007/s12161-022-02329-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-022-02329-9