Abstract
Antibiotic resistance is a growing problem and a global threat for modern healthcare. New approaches complementing the traditional antibiotic drugs are urgently needed to secure the ability to treat bacterial infections also in the future. Among the promising alternatives are bacteriolytic enzymes, such as the cell wall degrading peptidoglycan hydrolases. Staphylococcus aureus LytM, a Zn2+-dependent glycyl-glycine endopeptidase of the M23 family, is one of the peptidoglycan hydrolases. It has a specificity towards staphylococcal peptidoglycan, making it an interesting target for antimicrobial studies. LytM hydrolyses the cell wall of S. aureus, a common pathogen with multi-resistant strains that are difficult to treat, such as the methicillin-resistant S. aureus, MRSA. Here we report the 1H, 15N and 13C chemical shift assignments of S. aureus LytM N-terminal domain and linker region, residues 26–184. These resonance assignments can provide the basis for further studies such as elucidation of structure and interactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
The increasing prevalence of antimicrobial resistance in bacteria is among the most pressing threats to public health (Murray et al. 2022). The higher mortality and morbidity associated with antibiotic resistant bacterial infections burden the operation of healthcare as well as impose significant economic and societal costs (World Health Organization 2014). The background of the problem is multifaceted, stemming from the overuse and misuse of current antimicrobial drugs combined with the scarcity of new prominent antibiotics (Gould and Bal 2013; Ventola 2015). Thus, development of alternative solutions is urgently needed to avoid a future in which antibiotic drugs are no longer an effective treatment for bacterial infections.
One of the most problematic bacterial species is Staphylococcus aureus, a major human pathogen causing a wide variety of infections of different severity. S. aureus is acknowledged as the leading cause in skin and soft tissue infections, particularly infections occurring in surgical sites and associated with medical implants (Lowy 1998; Tong et al. 2015; Diekema et al. 2019). S. aureus has several features which make it a challenging pathogen to treat, and among them is the capability to rapidly acquire resistance to antimicrobial drugs. Conformingly, the methicillin-resistant S. aureus (MRSA) is reported to have gained resistance against many of the first-line antibiotics, including the traditional β-lactam antibiotics (Monaco et al. 2017). Therefore, it is particularly alarming that an emerging resistance to last-resort antibiotics such as vancomycin has been observed in MRSA strains, making MRSA one of the most serious infectious disease threats globally (Guo et al. 2020).
Among the promising approaches to combat antibiotic resistant strains of bacteria is the utilization of bacteriolytic enzymes, such as the peptidoglycan hydrolases (PGHs). PGHs kill bacteria via degradation of peptidoglycan, an essential component of the bacterial cell wall (Szweda et al. 2012). PGHs have multiple beneficial attributes, including their efficacy against biofilms and the low likelihood of inducing the development of resistance in bacteria due to their specificity and the conserved nature of their target, the peptidoglycan (Pastagia et al. 2013).
LytM is a Zn2+-dependent glycyl-glycine endopeptidase belonging to the M23 family of metallopeptidases. The target of its catalytic activity is the pentaglycine bridge specific for S. aureus peptidoglycan, making it an interesting candidate for treating S. aureus infections. The structure of this enzyme has been studied by the means of X-ray crystallography, yielding a two-domain structure (Odintsov et al. 2004). The C-terminal domain with a conserved catalytic site harbours the catalytic activity of the enzyme, whereas the function of the N-terminal domain is currently unknown. The unique N-terminal domain sets LytM apart from the other enzymes of the same family, such as S. simulans lysostaphin (Sabala et al. 2014; Tossavainen et al. 2018) and S. aureus LytU (Raulinaitis et al. 2017). LytM has been reported to be inactive in its full form, but the catalytic domain alone has been shown to cleave peptidoglycan (Odintsov et al. 2004). The activation mechanism of LytM, however, remains unclear. In the latent form the catalytic site is blocked by an occluding residue Asn117 residing in the linker, and consequently an “asparagine switch”, analogous to the cysteine switch of matrix metalloproteases, has been proposed for the regulation of the activity (Odintsov et al. 2004). This highlights the importance of understanding the role of the N-terminal domain and the linker region in the activation. To this end, we have characterized the LytM N-terminal domain and linker region by assigning the chemical shifts of residues 26–184, including the previously uncharacterized, potentially disordered regions in the crystal structure (Odintsov et al. 2004).
Methods and experiments
Protein expression and purification
Two constructs of S. aureus LytM, including the N-terminal domain (residues 26–107) and N-terminal domain with the linker region (residues 26–184) were cloned in pET-15b vectors. To improve protein solubility the LytM constructs were tagged with GB1 in the N-terminus connected with linker region containing the TEV protease site. Proteins were expressed using Escherichia coli strain BL21(DE3). 50 ml precultures were grown in standard M9 supplemented with 100 µg/ml ampicillin at 30 °C with 230 rpm shaking for 20 h. Cultures were expanded to 1 l and the cells were grown at 37 °C with 250 rpm shaking in standard M9 minimal medium supplemented with 100 µg/ml ampicillin and 15NH4Cl (1 g/l) and 13 C-D-glucose (2 g/l) as the nitrogen and carbon sources for uniform 15N and 13C labelling, respectively, until the OD at 600 nm was 0.55–0.6. Cells were cooled down to 16 °C, and the protein expression was induced by adding 0.5 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) at 16 °C for 20 h, at 230 rpm.
Cells were harvested by centrifugation, resuspended in 50 mM sodium phosphate pH 8, 300 mM NaCl buffer and lysed by sonication (three times 1 min, 1 s pulse, 30% amplitude). Clarified lysates were purified with 1 ml His GraviTrap (Cytiva) column according to manufacturer’s instructions. GB1 was cleaved using TEV protease, overnight incubation in dialysis (buffer 25 mM sodium phosphate pH 8, 150 mM NaCl, 10% glycerol, 1 mM β-mercaptoethanol) at 4 °C. Cleaved proteins were purified in 20 mM sodium phosphate pH 6.5, 50 mM NaCl, 1 mM DTT buffer by size exclusion chromatography using ÄKTA pure chromatography system (GE Healthcare) with HiLoad Superdex S75 (16/60) column (GE Healthcare). Proteins were concentrated using Amicon Ultra-15 centrifugal filter units (Millipore).
NMR spectroscopy
The NMR data acquisition was done using uniformly 15N,13C labelled LytM(26–107) and LytM(26–184) protein fragments in a buffer composed of 20 mM sodium phosphate, 50 mM NaCl, 1 mM DTT, pH 6.5 in 95/5% H2O/ D2O. The concentrations of LytM(26–107) and LytM(26–184) used for the data collection were 1.0 mM and 0.2 mM, respectively. NMR spectra were acquired at 308 K using a Bruker Avance III HD 800 MHz NMR spectrometer equipped with a 1H, 13C, 15N TCI cryoprobe.
Full chemical shift assignment was performed for LytM(26–107). Backbone resonances were assigned using 1H,15N HSQC (Kay et al. 1992) and 1H,13C-CT-HSQC (Vuister and Bax 1992) experiments for aliphatic and aromatic regions, and a suite of triple resonance NMR experiments including HNCO (Muhandiram and Kay 1994) and i(HCA)CO(CA)NH aka iHNCO (Mäntylahti et al. 2009), and HNCACB and HN(CO)CACB (Yamazaki et al. 1994). Aliphatic side chain resonances were assigned using H(CC)(CO)NH, (H)CC(CO)NH, HBHA(CO)NH and HCCH-COSY, whereas aromatic side chains were assigned using (HB)CB(CGCD)HD, (HB)CB(CGCDCE)HE and NOESY-13C-HSQC for aliphatic and aromatic carbons (Sattler et al. 1999).
The Hα, Cα, N and C′ chemical shifts of the LytM linker region (residues 108–184) were assigned using the longer construct LytM(26–184) and by employing Hα-detected 4D HACANCOi and HACACON (Tossavainen et al. 2020; Karjalainen et al. 2020), 3D HA(CA)NCOi (Karjalainen et al. 2020) and HA(CA)CON (Mäntylahti et al. 2010), together with HNCO and iHNCO experiments.
The NMR data were processed with TopSpin 3.5 pl 7 (Bruker Corporation) and analysed using CcpNmr Analysis v.2.4.2 software (Vranken et al. 2005).
Extent of assignment and data deposition
The amino acid sequence of LytM N-terminal domain and linker region, shown in Fig. 1a, was analyzed using bioinformatical tools to evaluate intrinsic disorder. A mean charge against mean scaled hydropathy plot (Fig. 1b), based on a PONDR (Xue et al. 2010) prediction, categorized both LytM N-terminal domain and N-terminal domain with linker region among intrinsically disordered proteins. For further evaluation, an IUPred3 (Erdos et al. 2021) analysis describing the probability of disorder within the sequence was performed. The results (Fig. 1c) show that most of the sequence has an IUPred3 score higher than 0.5 and is thus expected to be disordered. When only the N-terminal domain (yellow in Fig. 1c) is considered, the N- and C-terminal regions appear disordered, whereas the middle part scores lower values. Residues 41–106 have scores below 0.5, indicating ordered structure. The entire linker region (light purple in Fig. 1c) appears disordered, with the lowest probability for disorder being found locally around residue Val138.
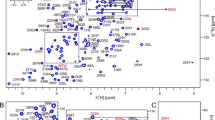
The conclusion of intrinsic disorder is supported also by the experimental data. An overview of the 1H,15N HSQC spectrum of LytM(26–184) (Fig. 1d) shows that while some of the cross-peaks are well-dispersed in the 1HN dimension, the majority are located between 7.5 and 8.5 ppm. All the well-dispersed correlations arise from the middle part of the N-terminal domain, particularly residues 51–94. The peaks assigned to the linker region (108–184), on the other hand, are all found within 7.8–8.3 ppm in the 1HN dimension, and the signals overlap considerably. This discrepancy in the distribution of peaks between the two regions of the protein was also reflected in the extent of chemical shift assignment.
a The amino acid sequence of LytM N-terminal domain and linker region. The N-terminal region (26–107) is marked in black and the linker region (108–184) is marked in blue. The N-terminal Gly is a cloning artefact, and not a part of the native sequence. b The charge-hydropathy plot of LytM N-terminal domain and linker region, showing LytM(26–107) (yellow triangle) and LytM(26–184) (green diamond) compared to a set of ordered (blue squares) and disordered (red circles) proteins. c Results of structural analyses of LytM N-terminal domain and linker region, plotted against the residue numbers. The upper part of the panel shows TALOS-N predictions. The predicted RCI S2 value is plotted as a line chart (axis on the left) and the probabilities of helical (in red, as positive values) and strand (blue, negative) secondary structures are plotted in the predicted secondary structure bar chart (axis on the right). The lower part is a combined graph of IUPred3 prediction of intrinsic disorder (line chart, axis on the left) and secondary structure prediction (bar chart, axis on the right). In the IUPred3 prediction, the probability of disorder is presented as a score from 0 to 1, with higher value indicating higher propensity for disorder. LytM N-terminal domain (26–107) is marked with a yellow line, the linker region (108–184) with a light purple line. The secondary structure prediction is based on the difference in Cα (dark purple bars) and C’ (green) chemical shift values and random coil shifts generated by POTENCI. Positive values denote helical structures and negative values strand structures. Secondary structure elements observed in the crystal structure of full-length LytM (PDB ID 1QWY) are highlighted on both sections, beta strands in light blue and alpha helices in pink. d Assigned 1H,15N HSQC spectrum of LytM N-terminal construct with full linker (26–184), acquired at 308 K and 800 1H MHz frequency, showing one-letter codes and residue numbers. Amides with peaks at contour levels lower than shown here are marked with crosses. Side chain assignments for Asn δ2, Gln ε2, His ε2 and Trp ε1 are marked in red. Asn δ2 and Gln ε2 side chain resonances are connected with horizontal lines. Insets on the right show the enlarged view of the areas indicated with dashed rectangles, encompassing the crowded regions. Inset on the left shows an additional peak outside the shown area
The chemical shifts of the N-terminal domain (residues 26–107) were assigned more comprehensively. The backbone chemical shifts were assigned nearly completely, with an assignment percentage of 98.8%. Out of the non-proline residues, 97.5% of 1H-15N pairs were assigned, omitting the N-terminal glycine not belonging to the native sequence. 98.8% of C′ and all Cα, Hα, Cβ and Hβ resonances were assigned. The aliphatic side chain assignment was 73.6% complete, and aromatic side chain assignment completeness was 93.2%. The low aliphatic side chain assignment percentage was largely due to the unassigned Asn δ2 (55.6% assigned) and Gln ε2 (12.6%) resonances. These resonances could not be reliably identified due to overlap of signals in the 1H,15N HSQC spectrum.
The backbone chemical shifts of the disordered linker region (residues 108–184) were assigned using additional 3D HA(CA)NCOi and HA(CA)CON, and 4D HACANCOi and HACACON experiments to overcome the issue of signal overlap and missing amides in the 1H,15N HSQC spectrum. The process is described in Fig. 2. The assignment percentage for the backbone of the complete LytM N-terminal domain with the linker region was 94.9%. The missing resonances were mainly amides, including residues Ala26, His35, Pro63, Asn151 and His157, emphasizing the decision to opt for HA-detected experiments. Additionally, assignment percentage was affected by C’ missing from 46Ser, and Cα from His147 and Ser156. Residues Asn151 and His157 remained completely unassigned.
Strip plots showing the sequential walk through residues 123-127 of LytM using a 4D HACANCOi (blue) and HACACON (red) experiments and b 3D HNCACB (Cα magenta, Cβ cyan) and HN(CO)CACB (Cα grey, Cβ green) experiments. The spectra of 4D experiments are less crowded and no peaks are missing, unlike in the 3D experiments
These data have been deposited in BioMagResBank (www.bmrb.wisc.edu) under accession number 51662.
For further characterization, the chemical shifts were used to investigate the secondary structure content of LytM N-terminal domain and linker region. Two approaches, TALOS-N (Shen and Bax 2013) and secondary structure prediction based on the deviations of the chemical shifts of Cα and C′ from random coil chemical shifts predicted by the POTENCI (Nielsen and Mulder 2018) tool are shown in the upper and lower halves of Fig. 1c, respectively. Similar results were obtained from both analyses, showing that the N-terminal domain has indications for secondary structure mainly on residues 45–95. This conclusion is reciprocated also in the results of IUPred3 analysis and predicted random coil index order parameter (RCI S2) values derived from TALOS-N. These analyses outline a similar increase of order and decrease of flexibility in the mid-N-terminal domain.
Comparison with the secondary structure elements of the crystal structure of full-length LytM (PDB ID 1QWY, Odintsov et al. 2004) agrees well with these findings (Fig. 1c). The alpha helix at residues 68–74 is detected in both secondary structure predictions but TALOS-N appears more sensitive for the beta strand elements in the N-terminal domain. On the other hand, residues 26–44 which are missing from the crystal structure show less propensity for secondary structure in the predictions, conforming to the expectation. Similar observation is made regarding the linker (108–184), out of which residues 147–182 are not defined in the crystal structure. When the characterized residues 108–146 are considered, they appear mostly disordered with the exceptions of transient helical structures that are observed in residues 126–133 and 141–145, and propensity to extended structure in residues 124–125. Both secondary structure predictions agree on the disordered nature of the linker region. Helical elements can be detected in both analyses; based on the results at least residues 120–131 and 142–146 would be expected to show transient helical structure. The presence of the transient helical elements in the truncated protein indicates that their structure is not completely dependent on the interaction with the catalytic domain. The beta strand structure of residues 124–125, on the other hand, does not show clear pattern in the prediction, suggesting that it is formed through the interaction.
Here we have presented the chemical shift assignment of LytM residues 26–184. A nearly complete assignment was obtained for N-terminal residues 26–107, which are likely to form a structured domain. Based on chemical shift dispersion and bioinformatics, residues downstream 107 are mainly disordered. Hα, HN, Cα, Cβ, C, and NH resonances were assigned for these residues. These assignments allow further studies such as structural characterization and investigation of interactions.
Data Availability
The chemical shifts have been deposited in BioMagResBank (www.bmrb.wisc.edu) under Accession Number 51662.
References
Diekema DJ, Pfaller MA, Shortridge D et al (2019) Twenty-year trends in antimicrobial susceptibilities among Staphylococcus aureus from the SENTRY antimicrobial surveillance program. Open forum infectious diseases. Oxford University Press, Oxford, pp S47–S53. https://doi.org/10.1093/ofid/ofy270
Erdos G, Pajkos M, Dosztányi Z (2021) IUPred3: prediction of protein disorder enhanced with unambiguous experimental annotation and visualization of evolutionary conservation. Nucleic Acids Res 49:W297–W303. https://doi.org/10.1093/nar/gkab408
Gould IM, Bal AM (2013) New antibiotic agents in the pipeline and how hey can help overcome microbial resistance. Virulence 4:185–191. https://doi.org/10.4161/viru.22507
Guo Y, Song G, Sun M et al (2020) Prevalence and therapies of antibiotic-resistance in Staphylococcus aureus. Front Cell Infect Microbiol 10:107. https://doi.org/10.3389/fcimb.2020.00107
Karjalainen M, Tossavainen H, Hellman M, Permi P (2020) HACANCOi: a new Hα-detected experiment for backbone resonance assignment of intrinsically disordered proteins. J Biomol NMR 74:741–752. https://doi.org/10.1007/s10858-020-00347-5
Kay L, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/ja00052a088
Lowy FD (1998) Staphylococcus aureus infections. N Engl J Med 339:520–532. https://doi.org/10.1056/NEJM199808203390806
Mäntylahti S, Tossavainen H, Hellman M, Permi P (2009) An intraresidual i(HCA)CO(CA)NH experiment for the assignment of main-chain resonances in 15N, 13C labeled proteins. J Biomol NMR 45:301–310. https://doi.org/10.1007/s10858-009-9373-4
Mäntylahti S, Aitio O, Hellman M, Permi P (2010) HA-detected experiments for the backbone assignment of intrinsically disordered proteins. J Biomol NMR 47:171–181. https://doi.org/10.1007/s10858-010-9421-0
Monaco M, Pimentel de Araujo F, Cruciani M et al (2017) Worldwide epidemiology and antibiotic resistance of Staphylococcus aureus. Current topics in microbiology and immunology. Springer, Berlin, pp 21–56.https://doi.org/10.1007/82_2016_3
Muhandiram DR, Kay LE (1994) Gradient-enhanced triple-resonance three-dimensional NMR experiments with improved sensitivity. J Magn Reson B 103:203–216. https://doi.org/10.1006/jmrb.1994.1032
Murray CJ, Ikuta KS, Sharara F et al (2022) Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet 399:629–655. https://doi.org/10.1016/S0140-6736(21)02724-0
Nielsen JT, Mulder FAA (2018) Potenci: prediction of temperature, neighbor and ph-corrected chemical shifts for intrinsically disordered proteins. J Biomol NMR 70:141–165. https://doi.org/10.1007/s10858-018-0166-5
Odintsov SG, Sabala I, Marcyjaniak M, Bochtler M (2004) Latent LytM at 1.3 Å resolution. J Mol Biol 335:775–785. https://doi.org/10.1016/j.jmb.2003.11.009
Pastagia M, Schuch R, Fischetti VA, Huang DB (2013) Lysins: the arrival of pathogen-directed anti-infectives. J Med Microbiol 62:1506–1516. https://doi.org/10.1099/jmm.0.061028-0
Raulinaitis V, Tossavainen H, Aitio O et al (2017) Identification and structural characterization of LytU, a unique peptidoglycan endopeptidase from the lysostaphin family. Sci Rep 7:6020. https://doi.org/10.1038/s41598-017-06135-w
Sabala I, Jagielska E, Bardelang PT et al (2014) Crystal structure of the antimicrobial peptidase lysostaphin from Staphylococcus simulans. FEBS J 281:4112–4122. https://doi.org/10.1111/febs.12929
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Magn Reson Spectrosc 34:93–158. https://doi.org/10.1016/S0079-6565(98)00025-9
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241. https://doi.org/10.1007/s10858-013-9741-y
Szweda P, Schielmann M, Kotlowski R et al (2012) Peptidoglycan hydrolases-potential weapons against Staphylococcus aureus. Appl Microbiol Biotechnol 96:1157–1174. https://doi.org/10.1007/s00253-012-4484-3
Tong SYC, Davis JS, Eichenberger E et al (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28:603–661. https://doi.org/10.1128/CMR.00134-14
Tossavainen H, Raulinaitis V, Kauppinen L et al (2018) Structural and functional insights into lysostaphin-substrate interaction. Front Mol Biosci 5:60. https://doi.org/10.3389/fmolb.2018.00060
Tossavainen H, Salovaara S, Hellman M et al (2020) Dispersion from Cα or NH: 4D experiments for backbone resonance assignment of intrinsically disordered proteins. J Biomol NMR 74:147–159. https://doi.org/10.1007/s10858-020-00299-w
Ventola CL (2015) The antibiotic resistance crisis: part 1: causes and threats. Pharm Ther 40:277–283
Vranken WF, Boucher W, Stevens TJ et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Vuister GW, Bax A (1992) Resolution enhancement and spectral editing of uniformly 13 C-enriched proteins by homonuclear broadband 13 C decoupling. J Magn Reson 98:428–435. https://doi.org/10.1016/0022-2364(92)90144-V
World Health Organization (2014) Antimicrobial resistance: global report on surveillance. World Health Organization, Geneva
Xue B, Dunbrack RL, Williams RW et al (2010) PONDR-FIT: a meta-predictor of intrinsically disordered amino acids. Biochim Biophys Acta Proteins Proteom 1804:996–1010. https://doi.org/10.1016/j.bbapap.2010.01.011
Yamazaki T, Lee W, Arrowsmith CH et al (1994) A suite of triple resonance NMR experiments for the Backbone assignment of 15N, 13C, 2H labeled proteins with high sensitivity. J Am Chem Soc 116:11655–11666. https://doi.org/10.1021/ja00105a005
Acknowledgements
We thank Laura Pitkänen and Dr. Maarit Hellman for excellent technical assistance. This work was supported by the Academy of Finland and Jane ja Aatos Erkon Säätiö.
Funding
Open Access funding provided by University of Jyväskylä (JYU). Academy of Finland, (Grant number 323435). Jane ja Aatos Erkon Säätiö.
Author information
Authors and Affiliations
Contributions
IP expressed and purified proteins, prepared all figures and wrote the initial draft of the manuscript. IP, HT, and PP performed experiments and data analyses. HT and PP conceived of and designed the experiments. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing conflict of interest.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pitkänen, I., Tossavainen, H. & Permi, P. 1H, 13C, and 15N NMR chemical shift assignment of LytM N-terminal domain (residues 26–184). Biomol NMR Assign 17, 257–263 (2023). https://doi.org/10.1007/s12104-023-10151-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10151-5