Abstract
Monoclonal antibodies (mAbs) therapeutics are the largest and fastest growing class of biologic drugs, amongst which, the vast majority are immunoglobulin G1 (IgG1). Their antigen binding abilities are used for the treatment of immunologic diseases, cancer therapy, reversal of drug effects, and targeting viruses and bacteria. The high importance of therapeutic mAbs and their derivatives has called for the generation of well-characterized standards for method development and calibration. One such standard, the NISTmAb RM 8621 based on the antibody motavizumab, has been developed by the National Institute of Standards and Technologies (NIST) in the US. Here, we present the resonance assignment of the single chain variable fragment, NISTmAb-scFv, that was engineered by linking the variable domains of the heavy and light chains of the NISTmAb. Also, addition of a peptide, corresponding to the target antigen of motavizumab, to samples of NISTmAb-scFv has induced chemical shift perturbations on residues lining the antigen binding interface thereby indicating proper folding of the NISTmAb-scFv.
Similar content being viewed by others
Explore related subjects
Find the latest articles, discoveries, and news in related topics.Avoid common mistakes on your manuscript.
Biological context
Monoclonal antibody therapeutics are the largest and fastest growing class of protein drugs for human use. Amongst these, single chain fragment variable (scFv) are small versions made by linking the variable regions of the heavy and light chains. ScFvs retain the antigen binding ability with essentially the same affinity as the parent mAb (Huston et al. 1988), and they allow the generation of libraries aimed at optimizing binding specificity and affinity, which is facilitated by the ability to produce scFv in E. coli. In addition, scFvs have been used as drugs, tools for radionuclide delivery on their own or incorporated in larger chimeric biologics for their antigen binding abilities (Ahmad et al. 2012; Monnier et al. 2013; Lu et al. 2020; Ferro Desideri et al. 2021).
The molecular recognition and the binding to the antigen are both functions that are associated with the variable fragment, “Fv”, of a monoclonal antibody (Lozano et al. 2012). As early as 1984, it was demonstrated that fragments from the heavy chain, which were recombinantly expressed in E. coli, had binding affinity for their epitopes (Cabilly et al. 1984), and in 1989 Ward and coworkers showed that a single immunoglobulin domain from the variable fragment (Fv) expressed in E. coli, had the ability to bind to its antigen (Ward et al. 1989). In the mid- and late 1980’s researchers were embarking on the idea of using inter-domain linkers for connecting two immunoglobulin domains from the heavy chain (VH) and the light chain (VL) of the variable region of a mAb, for recombinant production of a “single-chain variable fragment”, “scFv”, which could be used in therapeutic applications that required high specificity for target binding (Bird et al. 1988; Huston et al. 1988). These properties and their size make them interesting models to study the effects of therapeutic product excipients on the protein dynamics by NMR (Ghasriani et al. 2020). Product excipients are additives used to keep the protein active pharmaceutical ingredient stable during product manufacturing, product storage and delivery to patients.
We have focused on the variable fragment of the antigen-binding fragment of the NISTmAb, an IgG1κ monoclonal antibody derived from motavizumab (Schiel et al. 2018), and developed as a standard reference material for the characterization of therapeutic mAbs. Motavizumab was developed to target the fusion protein of the Respiratory Syncytial Virus (RSV-group). Respiratory syncytial virus (RSV-group) is a highly transmissible respiratory virus, which attacks the lower respiratory tract in children and adults and is responsible for the death of an estimate of 20,000 people in the US, and between 66,000 and 239,000 people worldwide every year (Thompson et al. 2003; Falsey et al. 2005; Nair et al. 2010; Lozano et al. 2012; Rha et al. 2020). The NISTmAb has been used in several multi-laboratory studies for assessing various analytical methods including NMR methods (Brinson et al. 2018) aimed for the characterization of therapeutic mAbs. The crystal structure of the antigen-binding fragment (Fab) of NISTmAb was determined (McLellan et al. 2011; Karageorgos et al. 2017). The crystal structure of NISTmAb bound to a 24-amino acid long peptide that corresponds to the epitope from RSV virus has also been determined (McLellan et al. 2010).
Here, we present the backbone and the side chain chemical shifts of the NISTmAb-scFv. In order to test whether the single-chain construct produced in E. coli adopted a biologically active conformation upon refolding, proton-nitrogen correlation maps were recorded in the presence and absence of the target peptide. We aim at using this single-chain variable fragment molecule as a first example of an immunoglobulin protein for in-depth studies of backbone dynamics and other motional properties in the presence of various therapeutic product excipients. Recently, a similar study on the 4-helix bundle filgrastim was conducted in our laboratory (Ghasriani et al. 2020).
Methods and experiments
Design of the scFv construct
The amino acid sequence of scFv was constructed from the sequence of the NIST-mAb reference material 8671 described by Formolo and coworkers (Formolo et al. 2015) (Karageorgos et al. 2017). Residues Q1 to S120 of the heavy chain (underlined) were linked to residues D1 to T108 of the light chain (italicized) using four (GGGGS) elements (Huston et al. 1988). The synthetic gene (Biobasic, Toronto, Canada) optimized for expression in E. coli was inserted in a modified pET15b vector containing ten histidines in the NdeI and BamH1 sites. After cleavage of the polyhistidine tag with thrombin, the resulting polypeptide product had 252 residues (26 kDa) with the following sequence:

where the first four extra residues resulted from the remainder of the thrombin cleavage site (GS) and the NdeI restriction enzyme site (HM) used for cloning. In order to match the numbering of residues in the NISTmAb-scFv with the NISTmAb numbering, Q1 in the NISTmAb heavy chain corresponds to Q4, and D1 in the NISTmAb light chain corresponds to D145.
Expression and purification of scFv
Expression of labelled NISTmAb-scFv was carried out by incubating E. coli BL21(DE3) (Stratagene) harboring the pET15b10-NISTmAb-scFv plasmid in minimal media (M9) using 1 g/l 13C-glucose and 2 g/l 15N-ammonium chloride as sole source of carbon and nitrogen at 37 °C. Protein expression was induced by the addition of isopropyl thio-D-galactopyranoside (IPTG) at an OD600 of 0.8. Cells were harvested 3 h post-induction by centrifugation and frozen at − 80 °C until purification. Expression of labelled NISTmAb-scFv in E. coli resulted in the formation of inclusion bodies. Cell pellets corresponding to a 5 L culture were resuspended in 35 mL of buffer A (10 mM TrisHCl, 100 mM sodium phosphate, 6 M guanidine hydrochloride, 10 mM reduced glutathione, pH 8.0) and disrupted by sonication on ice using a 400 W Branson sonifier (ThermoFisher). After separation of cell debris, lysis was repeated once with 35 mL of buffer A and the supernatants were pooled and added to a slurry of Ni–NTA resin (Qiagen) (80 mL resin, 10 mL buffer A) and gently stirred at room temperature for 30 min before loading into a column. Refolding was accomplished under oxidative condition with a gradient of buffer A to B (Buffer B: 10 mM TrisHCl, 100 mM sodium phosphate, pH 8.0) over 20 column volumes. The column was then washed with three column volumes of Buffer B + 60 mM imidazole pH 8.0 to remove unspecific binding. The protein was eluted off the column with Buffer B + 250 mM imidazole (pH 8.0). The efficiency of the on-column refolding was such that the procedure was repeated several times (7 up to 10 times) to extract properly folded protein by re-equilibrating the column with buffer A followed by the above refolding-washing-elution protocol.
Prior to cleavage of the poly-histidine tag, the buffer was exchanged to 20 mM sodium phosphate (pH 6.0) by ultrafiltration. Cleavage was carried out at a protein concentration of 2 mg/mL using 1U of thrombin (Cytiva) per 100 µg of target protein at room temperature. While almost all starting material was cleaved after 30 min, the reaction was allowed to proceed overnight.
The reaction mixture was then purified on cation exchange chromatography using HiTrap SP FF columns (Cytiva) in 50 mM sodium phosphate buffer pH (6.0) with a 1 M sodium chloride salt gradient. The NISTmAb-scFv eluted at around 200 mM NaCl. Protein concentration was determined by using UV spectroscopy with the theoretical extinction coefficient 18,150 M−1 cm−1 (Swissprot). NMR samples contained 0.25 mM of the uniformly isotope-labeled 13C-15 N– or 15N-NISTmAb-scFv in 20 mM phosphate buffer at pH 6.0, and 5% 2H2O was used for field frequency lock. The sample temperature was kept at 313 K (40 °C).
Peptide binding
The NMR titration of the peptide epitope with the NISTmAb-scFv was carried out with a 24-amino acid long polypeptide chain (NSELLSLINDMPLTNDQKKLMSNN), derived from the X-ray structure (PDBID 3ixt) epitope on the RSV virus fusion protein for motavizumab (McLellan et al. 2010).
The concentration of the stock peptide solution was 5.3 mM. A total of 10 μl of the stock solution was added to 550 μl of 1.65 mg ml−1 (~ 63 μM) protein solution, resulting in 95 μM peptide, and a final molar ratio of peptide-to-scFv of 1:0.65 (peptide being in excess). For recording of 15N-HSQC spectra of both the peptide-free and the peptide-bound scFv, the sample temperatures were kept at 308 K (35 °C).
NMR experiments
Data were collected on Bruker NEO-600 and AVANCE IIIHD-700 MHz NMR spectrometers equipped with cryogenically cooled triple resonance inverse probes fitted with Z-axis gradients. For backbone resonance assignment, the standard double- and triple resonance experiments 2D-15N-HSQC (“hsqcetfpf3gpsi”)(Palmer et al. 1991; Kay et al. 1992; Grzesiek and Bax 1993b; Schleucher et al. 1994), 3D-HNCO (“hncogp3d”)(Grzesiek and Bax 1992; Schleucher et al. 1993; Kay et al. 1994), 3D-HN(CA)CO (“hncacogp3d”)(Clubb et al. 1992), 3D-CBCA(CO)NH (“cbcaconhgp3d”) (Grzesiek and Bax 1993a; Muhandiram and Kay 1994), 3D-HNCACB (“hncacbgp3d”) (Wittekind and Mueller 1993), 3D-HNCA (“hncagp3d”) (Grzesiek and Bax 1992), 3D-HN(CO)CA (“hncocagp3d”) (Grzesiek and Bax 1992), were recorded. For assignment of the side chain 1H and 13C chemical shifts, the double- and triple resonance experiments 2D-13C-HSQC (“hsqcctetgpsisp”) (Palmer et al. 1991; Vuister and Bax 1992), 3D-H(CC)(CO)NH (“hccconhgp3d2”) and 3D-(H)CC(CO)NH (“hccconhgp3d3”) (Montelione et al. 1992; Clowes et al. 1993; Grzesiek et al. 1993; Logan et al. 1993; Lyons and Montelione 1993; Carlomagno et al. 1996), 3D-HA(CO)NH (“haconhgpwg3d”) (Grzesiek and Bax 1993a; Muhandiram and Kay 1994), 3D-HANH (“hanhgpwg3d”) (Kuboniwa et al. 1994; Weisemann et al. 1994), 3D-HBHA(CO)NH (“hbhaconhgp3d”) (Grzesiek and Bax 1993a; Muhandiram and Kay 1994), 3D-HBHANH (“hbhanhgpwg3d”), 3D-(H)N(CA)NNH (“hncannhgpwg3d”) (Weisemann et al. 1993), 3D-CCHTOCSY (“hcchdigp3d2”) (Kay et al. 1993), 3D-HCCH-COSY (“hcchcogp3d”) (Kay et al. 1993), 3D-HCCH-TOCSY (“hcchdigp3d”) (Kay et al. 1993), 3D-13C-NOESY-HSQC (“noesyhsqcetgpsi3d”)(Palmer et al. 1991; Kay et al. 1992; Schleucher et al. 1994), and 3D-15N-NOESY-HSQC (“noesyhsqcf3gpsi3d”)(Palmer et al. 1991; Kay et al. 1992; Schleucher et al. 1994) were recorded. For assignment of aromatic side chain Hδ and Hε, as well as Hδ1Trp, the Yamazaki experiments, 2D-CB(CGCD)HD (“hbcbcgcdhdgp”) and 2D-CB(CGCDCE)HE (“hbcbcgcdcehegp”)(Yamazaki et al. 1993), were recorded. The names of the pulse programs for experiments selected from the standard Bruker library are written in brackets.
Data analysis and the assignment
All NMR data were processed using NMRPipe software (Delaglio et al. 1995). NMRFAM-Sparky(Goddard and Kneller; Lee et al. 2015) was employed for spectral visualization and spectral analysis. Automatic assignment routine PINE-Sparky(Lee et al. 2019) was invoked for probabilistic assignment of the backbone amide 1H, 15N, and backbone carbonyl 13C chemical shifts. Assignment of side chain resonances was done through a semi-automatic approach by initial engagement of PINE-Sparky, followed by implementation of an “inspection–verification” strategy, where PINE result for each assignment was either accepted or rejected based on a holistic approach that included inspection of complementary NOESY data.
Extent of assignments and data deposition
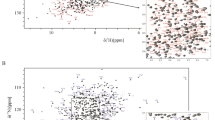
The scFv construct was comprised of 252 residues, which included the 20-amino acid long linker. All but 19 backbone 1H–15N resonances in the 15N-HSQC spectrum were successfully assigned (91% completeness, Fig. 1). The number of missing and unassigned resonances for backbone 13Cα and 13CO were 16 and 42, respectively. These numbers correspond to 94% and 83% assignment completeness for these two heavy backbone atoms, respectively. All the missing resonances reside exclusively in the loops and in the random coil regions (Met148-Ser153, and Gly242-Gly244). Of the total of 123 methyl groups from 84 methyl-containing residues in the 13C-HSQC spectrum (Alaβ, Ileγ, Ileδ, Leuδ, Metε, Thrγ, and Valγ), 118 13C-1H3 pairs were assigned (96% completeness). In total, 207 of 232 residues had their side chains fully, or partially, assigned (89% completeness). In all, the assigned backbone and side chain chemical shifts comprising of 1H (1113), 13C (789), and 15 N (215) were deposited into BMRB database (access code: 51094).
A number of residues, namely Ala36, Ile43, Arg44, Leu51, Asp60, Leu73, Leu86, Cys101, Ala102, Asn108, Tyr110, Phe111, Pro187, Gly207, Leu216, Ser220, and Cys231, displayed unusual chemical shifts that we attributed to the shielding effects afforded by close packing to aromatic rings based on analysis of the X-ray structure (PDBID 5k8a).
Three other types of unusual shifts (deshielding effects) were (a) due to either intra-residual side chain to main chain hydrogen bonding (Eswar and Ramakrishnan 2000) (the case of amide proton of Glu10), (b) due to ring current(Wieloch 1978) from a protonated histidine imidazole ring reaching across from the neighboring strand (the case of nitrogen atom of Asp193), or (c) due to a potential, but unusual, hydrogen bond between methyl protons and backbone carbonyl oxygen of residue i-4,(Yesselman et al. 2015) resulting from helical conformation in the middle of a loop (the case of methyl protons of Leu73). The extreme shift for backbone nitrogen atom of Asp193 can be attributed to a deshielding effect caused by ring current from protonated histidine imidazole ring of His177 (Wieloch 1978). The anomalous chemical shift for backbone nitrogen atom of Gly207 which displayed a down-field shift compared to average glycine chemical shift, can be attributed to side-chain to backbone H-bonding to the adjacent Ser206 (Vijayakumar et al. 1999; Eswar and Ramakrishnan 2000). Two particularly challenging regions for assignment, were the final stretch of beta stand that follows the binding loop (Ser235-Gly244), and the two short loops that are in close proximity of each other from heavy and light chains (Gln45-Lys49, and Lys182-Ala186). These regions account for 15 of the unassigned 13CO, 6 of the unassigned 13Cα, and 3 of the unassigned 15N–1H backbone resonances.
The signals from six methionine methyl groups were well-dispersed in the 13C-HSQC spectrum, such that the 13Cε and 1Hε chemical shifts could be readily identified using a combination of 13C-noesy and 15N-noesy spectra.
Truncation or crystal effect
An interesting finding that emerged while we were examining the 3D crystal structure for plausible explanation of the anomalous and extreme chemical shift of backbone amide nitrogen of Asp224, was the positioning of aromatic ring of Phe226 (Fig. 2a). As evidence, we later found NOE signals, which confirmed that the orientation of the phenylalanine ring in our case is in opposition to the one in the crystal structure of NISTmAb-Fab. For example, we observed NOESY cross peaks from both Pro223 and Ile249 to the aromatic ring of Phe226. Since the loops of the constant fragment in the Fab fragment (residues Glu164.L, Gln165.L, and Asp166.L) would physically restrict the rotation of the aromatic ring of Phe226 (Phe82.L), we speculate that the rotation about the Cα-Cβ bond of Phe226 in NISTmAb-scFv is made possible by the absence of constant domains CH1 and CL. (Fig. 2b). Of course, this does not take into account any “crystal effects” that would have contributed to a dense/strained packing of the loop against the aromatic ring in the full Fab.
a Phe82 (represented in CPK) in the NISTmAb-Fab domain adopts a rotamer that is locked due the proximity of the constant light (CL) domain (green surface).b In the NISTmAb-scFv, the absence of the CL domain allows Phe226 (stick) to adopt a rotamer in NISTmAb-scFv that induces a shielding effect on Asp224 amide resonance (blue sphere)
Peptide binding
In order to verify that refolding of the NISTmAb-scFv polypeptide led to a biologically active conformation similar to the NISTmAb-Fab fragment, we carried out a simple peptide binding experiment. Two dimensional 15N-HSQC spectra were recorded for peptide-free and peptide-bound 15N-NISTmAb-scFv (Fig. 3). The chemical shift perturbations of backbone amide pairs were largest for residues involved in the peptide-binding site, consistent with the residues in close proximity with the peptide in the crystal structure of the complex (PDBID 3IXT) (McLellan et al. 2010). This observation indicates that the NISTmAb-scFv has folded into a biologically active conformation.
a Overlay of 2D-15N-HSQC spectra of 13C-15N-NISTmAb-scFv free (blue) and peptide-bound (red) at 700 MHz recorded at 35 °C. Residues showing chemical shift perturbation upon peptide binding are indicated with a color code (red, green, blue, brown, yellow, grey-white and purple. b Ribbon diagram of the variable domains (VH and CL) of the NISTmAb-Fab X-ray structure (PDBID: 3IXT). Residues experiencing CSP depicted in panel A are mapped on the structure using the same colors
Data availability
Chemical shifts and Bruker raw data ser files were deposited in the BMRB data bank with entry number 51094.
Abbreviations
- mAb:
-
Monoclonal antibody
- scFv:
-
single-chain variable fragment
- VH:
-
heavy chain variable domain
- VL:
-
light chain variable domain
- Fab:
-
antigen-binding fragment
- NISTmAb:
-
National Institute of Standards and Technology monoclonal antibody
- RSV:
-
Respiratory syncitial virus
References
Ahmad ZA, Yeap SK, Ali AM, Ho WY, Alitheen NB, Hamid M (2012) scFv antibody: principles and clinical application. Clin Dev Immunol 2012:980250. https://doi.org/10.1155/2012/980250
Bird RE, Hardman KD, Jacobson JW, Johnson S, Kaufman BM, Lee S-M, Lee T, Pope SH, Riordan GS, Whitlow M (1988) Single-chain antigen-binding proteins. Science 242:423–426
Brinson R, Marino J, Delaglio F, Arbogast L, Evans R, Kearsley A, Gingras G, Ghasriani H, Aubin Y, Pierens G et al (2018) Enabling adoption of 2D-NMR for the higher order structure assessment of monoclonal antibody therapeutics. mAbs. https://doi.org/10.1080/19420862.2018.1544454
Cabilly S, Riggs AD, Pande H, Shively JE, Holmes WE, Rey M, Perry LJ, Wetzel R, Heyneker HL (1984) Generation of antibody activity from immunoglobulin polypeptide chains produced in Escherichia coli. Proc Natl Acad Sci USA 81:3273–3277. https://doi.org/10.1073/pnas.81.11.3273
Carlomagno T, Maurer M, Sattler M, Schwendinger MG, Glaser SJ, Griesinger C (1996) PLUSH TACSY: homonuclear planar TACSY with two-band selective shaped pulses applied to Cα, C′ transfer and Cβ, caromatic correlations. J Biomol NMR 8:161–170. https://doi.org/10.1007/BF00211162
Clowes RT, Boucher W, Hardman CH, Domaille PJ, Laue ED (1993) A 4D HCC(CO)NNH experiment for the correlation of aliphatic side-chain and backbone resonances in 13C/15N-labelled proteins. J Biomol NMR 3:349–354. https://doi.org/10.1007/BF00212520
Clubb RT, Thanabal V, Wagner G (1992) A constant-time three-dimensional triple-resonance pulse scheme to correlate intraresidue 1HN, 15N, and 13C′ chemical shifts in 15N13C-labelled proteins. J Magn Reson 1969(97):213–217. https://doi.org/10.1016/0022-2364(92)90252-3
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Eswar N, Ramakrishnan C (2000) Deterministic features of side-chain main-chain hydrogen bonds in globular protein structures. Protein Eng Des Sel 13:227–238. https://doi.org/10.1093/protein/13.4.227
Falsey AR, Hennessey PA, Formica MA, Cox C, Walsh EE (2005) Respiratory syncytial virus infection in elderly and high-risk adults. N Engl J Med 352:1749–1759. https://doi.org/10.1056/NEJMoa043951
Ferro Desideri L, Traverso CE, Nicolo M (2021) Brolucizumab: a novel anti-VEGF humanized single-chain antibody fragment for treating w-AMD. Expert Opin Biol Ther 21:553–561. https://doi.org/10.1080/14712598.2021.1915278
Formolo T, Ly M, Levy M, Kilpatrick L, Lute S, Phinney K, Marzilli L, Brorson K, Boyne M, Davis D et al (2015) Determination of the NISTmAb primary structure. American Chemical Society, Washington
Ghasriani H, Frahm GE, Johnston MJW, Aubin Y (2020) Effects of excipients on the structure and dynamics of filgrastim monitored by thermal unfolding studies by CD and NMR spectroscopy. ACS Omega 5:31845–31857. https://doi.org/10.1021/acsomega.0c04692
GODDARD, T. D. & KNELLER, D. G. SPARKY 3, University of California, San Francisco. http://www.cgl.ucsf.edu/home/sparky/.
Grzesiek S, Bax A (1992) Improved 3D triple-resonance NMR techniques applied to a 31 kDa protein. J Magn Reson 1969(96):432–440. https://doi.org/10.1016/0022-2364(92)90099-S
Grzesiek S, Bax A (1993a) Amino acid type determination in the sequential assignment procedure of uniformly 13C/15N-enriched proteins. J Biomol NMR 3:185–204. https://doi.org/10.1007/BF00178261
Grzesiek S, Bax A (1993b) The importance of not saturating water in protein NMR. Application to sensitivity enhancement and NOE measurements. J Am Chem Soc 115:12593–12594. https://doi.org/10.1021/ja00079a052
Grzesiek S, Anglister J, Bax A (1993) Correlation of Backbone Amide and Aliphatic Side-Chain Resonances in 13C/15N-Enriched Proteins by Isotropic Mixing of 13C Magnetization. J Magn Reson, Ser B 101:114–119. https://doi.org/10.1006/jmrb.1993.1019
Huston JS, Levinson D, Mudgett-Hunter M, Tai MS, Novotný J, Margolies MN, Ridge RJ, Bruccoleri RE, Haber E, Crea R et al (1988) Protein engineering of antibody binding sites: recovery of specific activity in an anti-digoxin single-chain Fv analogue produced in Escherichia coli. Proc Natl Acad Sci USA 85:5879–5883. https://doi.org/10.1073/pnas.85.16.5879
Karageorgos I, Gallagher ES, Galvin C, Gallagher DT, Hudgens JW (2017) Biophysical characterization and structure of the Fab fragment from the NIST reference antibody, RM 8671. Biologicals 50:27–34. https://doi.org/10.1016/j.biologicals.2017.09.005
Kay L, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/ja00052a088
Kay LE, Xu GY, Singer AU, Muhandiram DR, Formankay JD (1993) A gradient-enhanced HCCH-TOCSY experiment for recording side-chain 1H and 13C correlations in H2O samples of proteins. J Magn Reson 101:333–337. https://doi.org/10.1006/jmrb.1993.1053
Kay LE, Xu GY, Yamazaki T (1994) Enhanced-Sensitivity Triple-Resonance Spectroscopy with Minimal H2O Saturation. J Magn Reson, Ser A 109:129–133. https://doi.org/10.1006/jmra.1994.1145
Kuboniwa H, Grzesiek S, Delaglio F, Bax A (1994) Measurement of HN-Hα J couplings in calcium-free calmodulin using new 2D and 3D water-flip-back methods. J Biomol NMR 4:871–878. https://doi.org/10.1007/BF00398416
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327. https://doi.org/10.1093/bioinformatics/btu830
Lee W, Bahrami A, Dashti HT, Eghbalnia HR, Tonelli M, Westler WM, Markley JL (2019) I-PINE web server: an integrative probabilistic NMR assignment system for proteins. J Biomol NMR 73:213–222. https://doi.org/10.1007/s10858-019-00255-3
Logan TM, Olejniczak ET, Xu RX, Fesik SW (1993) A general method for assigning NMR spectra of denatured proteins using 3D HC(CO)NH-TOCSY triple resonance experiments. J Biomol NMR 3:225–231. https://doi.org/10.1007/BF00178264
Lozano R, Naghavi M, Foreman K, Lim S, Shibuya K, Aboyans V, Abraham J, Adair T, Aggarwal R, Ahn SY et al (2012) Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380:2095–2128. https://doi.org/10.1016/S0140-6736(12)61728-0
Lu RM, Hwang YC, Liu IJ, Lee CC, Tsai HZ, Li HJ, Wu HC (2020) Development of therapeutic antibodies for the treatment of diseases. J Biomed Sci 27:1. https://doi.org/10.1186/s12929-019-0592-z
Lyons BA, Montelione GT (1993) An HCCNH triple-resonance experiment using carbon-13 isotropic mixing for correlating backbone amide and side-chain aliphatic resonances in isotopically enriched proteins. J Magn Reson Ser B 101:206–209. https://doi.org/10.1006/jmrb.1993.1034
McLellan JS, Chen M, Kim A, Yang Y, Graham BS, Kwong PD (2010) Structural basis of respiratory syncytial virus neutralization by motavizumab. Nat Struct Mol Biol 17:248–250. https://doi.org/10.1038/nsmb.1723
McLellan JS, Correia BE, Chen M, Yang Y, Graham BS, Schief WR, Kwong PD (2011) Design and characterization of epitope-scaffold immunogens that present the motavizumab epitope from respiratory syncytial virus. J Mol Biol 409:853–866. https://doi.org/10.1016/j.jmb.2011.04.044
Monnier P, Vigouroux R, Tassew N (2013) In vivo applications of single chain Fv (variable domain) (scFv) fragments. Antibodies 2:193–208. https://doi.org/10.3390/antib2020193
Montelione GT, Lyons BA, Emerson SD, Tashiro M (1992) An efficient triple resonance experiment using carbon-13 isotropic mixing for determining sequence-specific resonance assignments of isotopically-enriched proteins. J Am Chem Soc 114:10974–10975. https://doi.org/10.1021/ja00053a051
Muhandiram DR, Kay LE (1994) Gradient-enhanced triple-resonance three-dimensional NMR experiments with improved sensitivity. J Magn Reson Ser B 103:203–216. https://doi.org/10.1006/jmrb.1994.1032
Nair H, Nokes DJ, Gessner BD, Dherani M, Madhi SA, Singleton RJ, O’Brien KL, Roca A, Wright PF, Bruce N et al (2010) Global burden of acute lower respiratory infections due to respiratory syncytial virus in young children: a systematic review and meta-analysis. The Lancet 375:1545–1555. https://doi.org/10.1016/S0140-6736(10)60206-1
Palmer AG, Cavanagh J, Wright PE, Rance M (1991) Sensitivity improvement in proton-detected two-dimensional heteronuclear correlation NMR spectroscopy. J Magn Reson 1969(93):151–170. https://doi.org/10.1016/0022-2364(91)90036-S
Rha B, Curns AT, Lively JY, Campbell AP, Englund JA, Boom JA, Azimi PH, Weinberg GA, Staat MA, Selvarangan R et al (2020) Respiratory syncytial virus-associated hospitalizations among young children: 2015–2016. Pediatrics. https://doi.org/10.1542/peds.2019-3611
RSV-GROUP (1998) Palivizumab, a humanized respiratory syncytial virus monoclonal antibody, reduces hospitalization from respiratory syncytial virus infection in high-risk infants. Pediatrics 102:531–537. https://doi.org/10.1542/peds.102.3.531
Schiel JE, Turner A, Mouchahoir T, Yandrofski K, Telikepalli S, King J, Derose P, Ripple D, Phinney K (2018) The NISTmAb Reference Material 8671 value assignment, homogeneity, and stability. Anal Bioanal Chem 410:2127–2139. https://doi.org/10.1007/s00216-017-0800-1
Schleucher J, Sattler M, Griesinger C (1993) Coherence selection by gradients without signal attenuation: application to the three-dimensional HNCO experiment. Angew Chem Int Ed Engl 32:1489–1491. https://doi.org/10.1002/anie.199314891
Schleucher J, Schwendinger M, Sattler M, Schmidt P, Schedletzky O, Glaser SJ, Sørensen OW, Griesinger C (1994) A general enhancement scheme in heteronuclear multidimensional NMR employing pulsed field gradients. J Biomol NMR 4:301–306. https://doi.org/10.1007/bf00175254
Thompson WW, Shay DK, Weintraub E, Brammer L, Cox N, Anderson LJ, Fukuda K (2003) Mortality associated with influenza and respiratory syncytial virus in the United States. JAMA 289:179–186. https://doi.org/10.1001/jama.289.2.179
Vijayakumar M, Qian H, Zhou HX (1999) Hydrogen bonds between short polar side chains and peptide backbone: prevalence in proteins and effects on helix-forming propensities. Proteins 34:497–507
Vuister GW, Bax A (1992) Resolution enhancement and spectral editing of uniformly 13C-enriched proteins by homonuclear broadband 13C decoupling. J Magn Reson 1969(98):428–435. https://doi.org/10.1016/0022-2364(92)90144-V
Ward ES, Güssow D, Griffiths AD, Jones PT, Winter G (1989) Binding activities of a repertoire of single immunoglobulin variable domains secreted from Escherichia coli. Nature 341:544–546. https://doi.org/10.1038/341544a0
Weisemann R, Rüterjans H, Bermel W (1993) 3D triple-resonance NMR techniques for the sequential assignment of NH and 15N resonances in 15N- and 13C-labelled proteins. J Biomol NMR 3:113–120. https://doi.org/10.1007/bf00242479
Weisemann R, Rüterjans H, Schwalbe H, Schleucher J, Bermel W, Griesinger C (1994) Determination of H(N), H (α) and H (N), C’ coupling constants in (13)C, (15)N-labeled proteins. J Biomol NMR 4:231–240. https://doi.org/10.1007/bf00175250
Wieloch T (1978) An NMR study of a tyrosine and two histidine residues in the structure of porcine pancreatic colipase. FEBS Lett 85:271–274. https://doi.org/10.1016/0014-5793(78)80471-2
Wittekind M, Mueller L (1993) HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha- and beta-carbon resonances in proteins. J Magn Reson, Ser B 101:201–205. https://doi.org/10.1006/jmrb.1993.1033
Yamazaki T, Forman-Kay JD, Kay LE (1993) Two-dimensional NMR experiments for correlating carbon-13.beta. and proton.delta./.epsilon. chemical shifts of aromatic residues in 13C-labeled proteins via scalar couplings. J Am Chem Soc 115:11054–11055. https://doi.org/10.1021/ja00076a099
Yesselman JD, Horowitz S, BROOKS, ., & TRIEVEL, . CLRC III (2015) Frequent side chain methyl carbon-oxygen hydrogen bonding in proteins revealed by computational and stereochemical analysis of neutron structures. Proteins 83:403–410. https://doi.org/10.1002/prot.24724
Acknowledgements
We thank Geneviève Gingras for the re-sequencing of the NISTmAb-scFv construct, and Drs. Michael Johnston and Roger Tam for critical reading of the manuscript.
Funding
Open Access provided by Health Canada. Not applicable.
Author information
Authors and Affiliations
Contributions
YA: Conceptualization; HG, SA & DJH: Investigation; HG: Formal Analysis; HG&YA: Visualization; HG&YA: Writing-original draft; HG, YA: Writing-review and editing; YA: Supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors have agreed to the publication of the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ghasriani, H., Ahmadi, S., Hodgson, D.J. et al. Backbone and side-chain resonance assignments of the NISTmAb-scFv and antigen-binding study. Biomol NMR Assign 16, 391–398 (2022). https://doi.org/10.1007/s12104-022-10109-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-022-10109-z