Abstract
KKT4 is a kinetoplastid-specific microtubule-binding kinetochore protein that lacks significant similarity to any known kinetochore or microtubule-binding proteins. Here we present the 1H, 13C and 15N resonance assignments for several fragments from the microtubule-binding domain of KKT4 (KKT4115–343) from Trypanosoma brucei. These assignments provide the starting point for detailed investigations of the structure, dynamics and interactions of the microtubule-binding region of KKT4.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
During chromosome segregation, cells must accurately transmit their genetic material into two daughter cells in an organised manner. Errors in this process lead to genetic abnormalities that can lead to cancer or cell death. Chromosome segregation is therefore a critical requirement for the survival and development of all organisms (McIntosh 2016). In eukaryotes, a key structure involved in this process is the kinetochore, a dynamic protein complex that assembles onto centromeric DNA of each chromosome and captures spindle microtubules (Brinkley and Stubblefield 1966; Cheeseman 2014). Many kinetochore components, such as CENP-A and Ndc80, are widely conserved among eukaryotes (Biggins 2013; Cheeseman and Desai 2008; Meraldi et al. 2006; Santaguida and Musacchio 2009; van Hooff et al. 2017) and it was thought that all eukaryotes utilise the same set of kinetochore proteins.
However, none of the canonical kinetochore proteins has been found in kinetoplastids, an evolutionarily divergent group of unicellular flagellated eukaryotes including parasitic trypanosomatids (e.g. Trypanosoma brucei, Trypanosoma cruzi, and Leishmania species). Instead, a number of unique kinetochore proteins, KKT1–25 and KKIP1–12, were identified in T. brucei (Akiyoshi 2016; Akiyoshi and Gull 2014; Brusini et al. 2019; D’Archivio and Wickstead 2017; Nerusheva and Akiyoshi 2016; Nerusheva et al. 2019). No significant similarity to conventional kinetochore proteins is found in these proteins and very little is known about their structure, function and interactions.
We recently showed that the KKT4 protein, and in particular a fragment containing residues 115–343, has microtubule-binding activities in vitro (Llauro et al. 2018). Further investigation demonstrated that KKT4 can track depolymerising microtubule tips and form load-bearing attachments with microtubules. Therefore, KKT4 has all the functions required for a microtubule-binding kinetochore protein. Interestingly, sequence analysis of the protein failed to reveal significant similarity to any known kinetochore or microtubule-binding proteins. Structural characterisation of KKT4 is therefore essential for understanding the molecular mechanism of its function. Here we present 1H, 13C and 15N resonance assignments for the microtubule-binding region of the kinetoplastid kinetochore protein KKT4 (KKT4115–343) from T. brucei. These NMR assignments provide the starting point for detailed investigations of the structure and dynamics of KKT4.
Methods and experiments
Protein expression and purification
KKT4 fragments characterised in this study were amplified from T. brucei genomic DNA that was cloned into the pNIC28-Bsa4 expression vector using a ligation-independent cloning method (Gileadi et al. 2008). Transformed E. coli BL21(DE3) cells were plated on agar plates containing 50 µg/ml kanamycin and incubated at 37 °C overnight. After overnight incubation, a few colonies were inoculated into 5 ml of 2xTY medium containing 50 µg/ml kanamycin and grown at 37 °C for 6 h. Next, 50 ml of M9 minimal medium containing 50 µg/ml kanamycin supplemented with 1 g/l 15NH4Cl and 4 g/l [13C]-D-glucose (CIL) as the sole nitrogen and carbon sources was inoculated with 500 μl of bacterial culture. Cell growth was continued overnight at 37 °C. Next, 5 ml of overnight culture was inoculated into 1 l of M9 minimal medium supplemented with 1 g/l 15NH4Cl, 4 g/l [13C]-D-glucose and 50 µg/ml kanamycin. Cells were grown at 37 °C to an OD600 of ∼ 0.8. Protein expression was induced by 0.4 mM IPTG and incubated overnight at 16 °C with shaking (200 rpm). Expression of uniformly labelled 2H/13C/15N-KKT4145–232 was facilitated by cell growth in D2O-based M9 minimal medium, with 1 g/l 15NH4Cl, 4 g/l [13C]-D-glucose and 50 µg/ml kanamycin, according to the protocol described in (Tugarinov et al. 2006).
Cells were pelleted at 3400g at 4 °C and resuspended in lysis buffer (50 mM sodium phosphate pH 7.5, 500 mM NaCl, and 10% glycerol) supplemented with protease inhibitors (20 μg/ml leupeptin, 20 μg/ml pepstatin, 20 μg/ml E-64, 2 mM benzamidine, and 0.4 mM PMSF) and 0.5 mM TCEP. All subsequent extraction steps were performed at 4 °C. Cell lysis was facilitated by mechanical cell disruption (French press, 1 passage at 20,000 psi). Lysed cells were spun at 48,000g for 30 min and the supernatant was loaded on a gravity column with TALON beads (Takara Clontech) pre-equilibrated in lysis buffer. After loading, the beads were washed extensively with lysis buffer and proteins were eluted with elution buffer (50 mM sodium phosphate pH 7.5, 500 mM NaCl, 10% glycerol, 250 mM imidazole, 0.5 mM TCEP). To remove the N-terminal 6xHis tag, the proteins were incubated with TEV protease and dialysed overnight against buffer containing 25 mM sodium phosphate pH 7.5, 250 mM NaCl, 5% glycerol, 5 mM imidazole and 0.5 mM TCEP. After overnight incubation, samples were diluted with 25 mM HEPES pH 7.5 with 0.5 mM TCEP (buffer A) to the final NaCl concentration of 50 mM and loaded on a cation exchange RESOURCE S column (GE Healthcare), pre-equilibrated with 5% of buffer B (25 mM HEPES pH 7.5, 1 M NaCl and 0.5 mM TCEP). Proteins were eluted from the column with a linear gradient from 5 to 100% of buffer B. Fractions containing KKT4 fragments were pooled, concentrated and loaded on a gel filtration column SD200 or SD75 16/60 (GE Healthcare) to purify further and buffer exchanged into 25 mM HEPES pH 7.5, 150 mM NaCl and 0.5 mM TCEP. The samples were concentrated using a 10-kD MW Amicon concentrator (Millipore), and aliquots were flash-frozen in liquid nitrogen and stored at − 80 °C.
NMR spectroscopy
15N or 13C/15N-labelled samples of KKT4115–174, KKT4145–232 and KKT4115–343 were used for resonance assignment using standard triple-resonance protocols (Redfield 2015). For KKT4145–232 a 2H/13C/15N-labelled sample was also required. All samples contained 300–500 µM protein in a 25 mM HEPES buffer at pH 7.2 with 150 mM NaCl, 0.5 mM TCEP and 5% D2O. NMR experiments were carried out at 20 °C for KKT4115–174 and KKT4115–343 and at 30 °C for KKT4145–232, using 750 and 950 MHz spectrometers equipped with Oxford Instruments Company magnets, Bruker Avance III HD consoles and 5 mm TCI CryoProbes.
The KKT4115–174, KKT4145–232 and KKT4115–343 samples were initially characterised using 2D 1H–15N HSQC (Kay et al. 1992; Palmer et al. 1991; Schleucher et al. 1994) SOFAST-HMQC (Schanda and Brutscher 2005) and BEST-TROSY (Lescop et al. 2007; Schulte-Herbruggen and Sorensen 2000) experiments. The latter two experiments allow rapid data collection or more extensive signal averaging than the HSQC. In crowded regions of the spectra, particularly for KKT4115–343, the resolution in the SOFAST-HMQC was significantly worse than that in the BEST-TROSY. Many residues in all the KKT4 constructs, particularly in KKT4145–232, gave rise to relatively weak peaks due to unfavourable relaxation properties (Supplementary Fig. 1); for these residues the highest quality data were obtained using the BEST-TROSY experiment. In order to be consistent between the three constructs, the assignments for all three are illustrated below using BEST-TROSY spectra.
Resonance assignments for KKT4115–174, KKT4145–232 and KKT4115–343 were obtained using 2D 1H–13C HSQC and 1H–15N BEST-TROSY experiments and 3D experiments including 15N-edited NOESY-HSQC, 15N-edited TOCSY-HSQC, HNCA, CBCANH, CBCA(CO)NH, HNCACB, HN(CO)CACB, HNCO, HN(CA)CO, HBHA(CBCACO)NH, HCA(CO)N, HCAN, (H)N(CA)NNH, (H)N(COCA)NNH, (H)CC(CO)NH, and HCCH-TOCSY. For KKT4115–174 and KKT4145–232 BEST-TROSY versions of the HNCA, HNCO, HN(CA)CO, HNCACB and HN(CO)CACB experiments were collected (Lescop et al. 2007; Schulte-Herbruggen and Sorensen 2000); for KKT4145–232 these incorporated 2H-decoupling. All 3D experiments were collected with 25% non-uniform sampling in the two indirect dimensions using standard Bruker sampling schedules. 2D NMR data were processed using NMRPipe (Delaglio et al. 1995) and 3D NUS data were processed with the hmsIST software (Hyberts et al. 2012) and NMRPipe. Spectra were analysed and assignments recorded using CcpNmr Analysis version 2.4 (Vranken et al. 2005). 1H and 13C chemical shifts were referenced using DSS and 15N chemical shifts were referenced indirectly. Details of the specific experiments and sample conditions used for each construct can be found in the BMRB deposition files.
Extent of assignments and data deposition
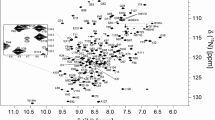
The 1H–15N BEST-TROSY spectrum of KKT4115–343 shows a large variation in peak intensities (Fig. 1a). Approximately 160 peaks are observed; this is less than expected for the 218 non-proline residues. For ~ 30 of the weakest peaks no 13C correlations were observed in triple resonance experiments making assignment of these impossible. The range of peak intensities for KKT4115–343 suggests a mixture of structured and disordered regions and/or differences in backbone dynamics on a µs–ms timescale leading to significant broadening and disappearance of many of the expected peaks. The strongest peaks visible in the 1H–15N BEST-TROSY spectrum of KKT4115–343 were assigned at pH 7.2 and 20 °C (Fig. 1b). 1HN and 15N backbone resonances for 90 of the 218 non-proline residues of KKT4115–343 were assigned. These all correspond to residues between N231 and E343, the C-terminal residue. In this region of the sequence, 87.4% of 1HN and 88.5% of 15N were assigned; further statistics for the extent of 1H and 13C assignments are presented in Table 1. Most of the missing 1HN/15N assignments correspond to Asn/Ser/Thr residues which either have fast intrinsic solvent exchange rates or are located in regions of low sequence complexity (for example, TTTSS). In addition to backbone resonance assignments, some side chain 1H and 13C assignments were obtained from HBHA(CBCACO)NH, (H)CC(CO)NH and 15N-TOCSY-HSQC experiments (Table 1).
750 MHz 1H–15N BEST-TROSY spectra of KKT4115–343 in 25 mM HEPES, 150 mM NaCl and 0.5 mM TCEP (95% H2O/5% D2O), at pH 7.2, 20 °C. a This spectrum is contoured at a low level to highlight the range of peak intensities observed for KKT4115–343. b The peak assignments for backbone amides of the C-terminal region (residues 231–343) of KKT4115–343 are annotated. The inset is an expansion of the area outlined with dashed lines. Peaks in the region of 111–114 ppm and upfield of ~ 7.6 ppm are artefacts in the BEST-TROSY arising from incomplete cancellation of signals from the side chain amides of Asn and Gln which have not been assigned. The three circled peaks in a correspond to Y278 (15N = 120 ppm), T279 (15N = 117 ppm) and L280 (15N = 124 ppm) in a minor species involving P281 in a cis peptide bond. The three peaks in squares in a correspond to Y278 (15N = 123 ppm), T279 (15N = 118 ppm) and L280 (15N = 127 ppm) in a minor species involving P277 in a cis peptide bond
In order to obtain assignments for residues in the N-terminal part of KKT4, NMR data were collected for a shorter fragment, KKT4115–232, containing the residues that could not be assigned in KKT4115–343. However, the spectrum of KKT4115–232 did not show the expected number of peaks (not shown). To overcome this, two shorter overlapping constructs, KKT4115–174 and KKT4145–232, were used. KKT4115–174 has been shown previously to bind to microtubules, albeit less efficiently than KKT4115–343 (Llauro et al. 2018). KKT4145–232 has been identified as a stable fragment in trypsin digests of KKT4115–343 (Supplementary Fig. 2). The 1H–15N BEST-TROSY spectra of KKT4115–174 and KKT4145–232 show the expected number of peaks, more uniform peak intensities and better dispersion of 1HN chemical shifts than observed for KKT4115–343.
For KKT4115–174 extensive backbone and side-chain assignments were completed at pH 7.2 and 20 °C (Fig. 2). This includes 1H/13C assignments for 82.6% of methyl groups and all three tyrosine side chains; detailed assignment statistics are presented in Table 1. Assignment of the KKT4145–232 spectrum proved more challenging; although the expected number of peaks were observed in the BEST-TROSY spectrum, triple resonance data were of poor quality. Deuteration of KKT4145–232 was required in addition to 13C/15N labelling and triple resonance spectra were collected at 30 °C (Fig. 3). Because of deuteration, only 1HN, 15N, 13Cα, 13Cβ and 13Cʹ assignments were possible for KKT4145–232; assignment statistics are presented in Table 1. For both KKT4115–174 and KKT4145–232, assignments for all Asn and Gln side chain NH2 groups were obtained using HSQC and triple-resonance spectra (Supplementary Fig. 3). The quality of the spectra obtained for KKT4115–174 and KKT4145–232 are significantly poorer than would be expected for monomeric, globular proteins of 60 and 88 residues. This suggests that these constructs may exist as oligomers and/or have overall shapes that give rise to anisotropic tumbling resulting in challenging relaxation properties; this will be explored in future studies of their structure and dynamics.
750 MHz 1H–15N BEST-TROSY spectrum of KKT4115–174 in 25 mM HEPES, 150 mM NaCl and 0.5 mM TCEP (95% H2O/5% D2O), at pH 7.2, 20 °C. The peak assignments for backbone amides are annotated. Peaks in the region of 111–114 ppm and upfield of ~ 7.6 ppm are artefacts in the BEST-TROSY arising from incomplete cancellation of signals from the side chain amides of Gln; these NH2 groups have been assigned in the HSQC spectrum shown in Supplementary Fig. 3A
750 MHz 1H–15N BEST-TROSY spectrum of KKT4145–232 in 25 mM HEPES, 150 mM NaCl and 0.5 mM TCEP (95% H2O/5% D2O), at pH 7.2, 30 °C. The peak assignments for backbone amides are annotated. Peaks in the region of 111–114 ppm and upfield of ~ 7.6 ppm are artefacts in the BEST-TROSY arising from incomplete cancellation of signals from the side chain amides of Asn and Gln; these NH2 groups have been assigned in the HSQC spectrum shown in Supplementary Fig. 3B
Assignments were obtained for all ten proline residues in the C-terminal region of KKT4115–343. This included 15N assignments obtained from HCA(CO)N and HCAN experiments which correlate the proline 15N with the preceding 1Hα/13Cα, and the proline 15N with both the preceding 1Hα/13Cα and the proline 1Hα/13Cα and 1Hδ/13Cδ, respectively. A previous study of the ID4 linker region of CREB-binding protein showed a clustering of peaks observed in the proline-fingerprint region of 2D CON spectra according to the type of residue preceding the proline (highlighted for GP, LP, PP, SP, TP and VP motifs) (Murrali et al. 2018). For KKT4115–343, the 13Cʹ(i-1)/15N(i) chemical shifts for G273/P274, L280/P281, T320/P321 and T332/P333 fall within the regions highlighted for these motifs in ID4 (Supplementary Table 1). Murrali et al. observe two clusters each for the AP and QP motifs; in both cases the proline 15N chemical shift is ~ 1.5–2 ppm further downfield if the proline is followed by a second proline (APP/QPP instead of APX/QPX) (Murrali et al. 2018). For KKT4115–343, the chemical shifts for A299/P300, Q253/P254 and Q276/P277, which are not followed by proline, fall within the regions observed for the APX and QPX motifs in ID4 (Supplementary Table 1).
Analysis of the proline 13Cβ and 13Cγ chemical shifts shows that the major conformation for all ten prolines involves a trans peptide bond (Schubert et al. 2002). However, a number of weak peaks are observed in the BEST-TROSY spectrum and in many cases these can be assigned to residues in the vicinity of a cis proline. For example, the three peaks highlighted with circles in Fig. 1a have been assigned to the sequence Y278-T279-L280 where P277 and P281 are in the trans and cis conformations, respectively; while the three peaks highlighted with squares are assigned to Y278-T279-L280 where P277 and P281 are in the cis and trans conformations, respectively. Analysis of the peak intensities indicates populations of ~ 10% and ~ 13% for the P277cis and P281cis conformers, respectively. A recent study of proline conformation in the intrinsically disordered protein Osteopontin suggested that cis proline populations of over 10% were only observed for proline residues directly preceded or followed by an aromatic residue (Phe or Trp) (Mateos et al. 2020). Interestingly, in KKT4115–343, P277 and P281 are also followed by aromatic residues, Y278 and H282.
The 1H, 13C and 15N chemical shift assignments presented here for KKT4115–174, KKT4145–232 and KKT4115–343 have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under the accession numbers 50215, 50228 and 50229, respectively. These assignments provide the starting point for detailed investigations of the structure, dynamics and interactions of the microtubule-binding domain of KKT4 (KKT4115–343) from T. brucei.
References
Akiyoshi B (2016) The unconventional kinetoplastid kinetochore: from discovery toward functional understanding. Biochem Soc Trans 44:1201–1217. https://doi.org/10.1042/Bst20160112
Akiyoshi B, Gull K (2014) Discovery of unconventional kinetochores in kinetoplastids. Cell 156:1247–1258. https://doi.org/10.1016/j.cell.2014.01.049
Biggins S (2013) The composition, functions, and regulation of the budding yeast kinetochore. Genetics 194:817–846. https://doi.org/10.1534/genetics.112.145276
Brinkley BR, Stubblefield E (1966) Fine structure of kinetochore of a mammalian cell in vitro. Chromosoma 19:28–43. https://doi.org/10.1007/Bf00332792
Brusini L, D’Archivio S, McDonald J, Wickstead B (2019) Ndc80/Nuf2-like protein KKIP1 connects a stable kinetoplastid outer kinetochore complex to the inner kinetochore and responds to metaphase tension. bioRxiv 764829
Cheeseman IM (2014) The kinetochore. Cold Spring Harb Perspect Biol 6: a015826. https://doi.org/10.1101/cshperspect.a015826
Cheeseman IM, Desai A (2008) Molecular architecture of the kinetochore–microtubule interface. Nat Rev Mol Cell Biol 9:33–46. https://doi.org/10.1038/nrm2310
D’Archivio S, Wickstead B (2017) Trypanosome outer kinetochore proteins suggest conservation of chromosome segregation machinery across eukaryotes. J Cell Biol 216:379–391. https://doi.org/10.1083/jcb.201608043
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Gileadi O, Burgess-Brown NA, Colebrook SM, Berridge G, Savitsky P, Smee CE, Loppnau P, Johansson C, Salah E, Pantic NH (2008) High throughput production of recombinant human proteins for crystallography. Methods Mol Biol 426:221–246. https://doi.org/10.1007/978-1-60327-058-8_14
Hyberts SG, Milbradt AG, Wagner AB, Arthanari H, Wagner G (2012) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson Gap scheduling. J Biomol NMR 52:315–327. https://doi.org/10.1007/s10858-012-9611-z
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/ja00052a088
Lescop E, Schanda P, Brutscher B (2007) A set of BEST triple-resonance experiments for time-optimized protein resonance assignment. J Magn Reson 187:163–169. https://doi.org/10.1016/j.jmr.2007.04.002
Llauro A, Hayashi H, Bailey ME, Wilson A, Ludzia P, Asbury CL, Akiyoshi B (2018) The kinetoplastid kinetochore protein KKT4 is an unconventional microtubule tip-coupling protein. J Cell Biol 217:3886–3900. https://doi.org/10.1083/jcb.201711181
Mateos B, Conrad-Billroth C, Schiavina M, Beier A, Kontaxis G, Konrat R, Felli IC, Pierattelli R (2020) The ambivalent role of proline residues in an intrinsically disordered protein: from disorder promoters to compaction facilitators. J Mol Biol 432:3093–3111. https://doi.org/10.1016/j.jmb.2019.11.015
McIntosh JR (2016) Mitosis. Cold Spring Harb Perspect Biol 8: a023218. https://doi.org/10.1101/cshperspect.a023218
Meraldi P, McAinsh AD, Rheinbay E, Sorger PK (2006) Phylogenetic and structural analysis of centromeric DNA and kinetochore proteins. Genome Biol 7:R23. https://doi.org/10.1186/gb-2006-7-3-r23
Murrali MG, Piai A, Bermel W, Felli IC, Pierattelli R (2018) Proline fingerprint in intrinsically disordered proteins. ChemBioChem 19:1625–1629. https://doi.org/10.1002/cbic.201800172
Nerusheva OO, Akiyoshi B (2016) Divergent polo box domains underpin the unique kinetoplastid kinetochore. Open Biol 6:150206. https://doi.org/10.1098/rsob.150206
Nerusheva OO, Ludzia P, Akiyoshi B (2019) Identification of four unconventional kinetoplastid kinetochore proteins KKT22–25 in Trypanosoma brucei. Open Biol 9:190236. https://doi.org/10.1098/rsob.190236
Palmer AG, Cavanagh J, Wright PE, Rance M (1991) Sensitivity improvement in proton-detected 2-dimensional heteronuclear correlation NMR-spectroscopy. J Magn Reson 93:151–170. https://doi.org/10.1016/0022-2364(91)90036-S
Redfield C (2015) Assignment of protein NMR spectra using heteronuclear NMR—a tutorial. Biol Magn Reson 32:1–42. https://doi.org/10.1007/978-1-4899-7621-5_1
Santaguida S, Musacchio A (2009) The life and miracles of kinetochores. EMBO J 28:2511–2531. https://doi.org/10.1038/emboj.2009.173
Schanda P, Brutscher B (2005) Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J Am Chem Soc 127:8014–8015. https://doi.org/10.1021/ja051306e
Schleucher J, Schwendinger M, Sattler M, Schmidt P, Schedletzky O, Glaser SJ, Sorensen OW, Griesinger C (1994) A general enhancement scheme in heteronuclear multidimensional NMR employing pulsed-field gradients. J Biomol NMR 4:301–306
Schubert M, Labudde D, Oschkinat H, Schmieder P (2002) A software tool for the prediction of Xaa-Pro peptide bond conformations in proteins based on C-13 chemical shift statistics. J Biomol NMR 24:149–154. https://doi.org/10.1023/A:1020997118364
Schulte-Herbruggen T, Sorensen OW (2000) Clean TROSY: compensation for relaxation-induced artifacts. J Magn Reson 144:123–128. https://doi.org/10.1006/jmre.2000.2020
Tugarinov V, Kanelis V, Kay LE (2006) Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat Protoc 1:749–754. https://doi.org/10.1038/nprot.2006.101
van Hooff JJE, Tromer E, van Wijk LM, Snel B, Kops GJPL (2017) Evolutionary dynamics of the kinetochore network in eukaryotes as revealed by comparative genomics. EMBO Rep 18:1559–1571. https://doi.org/10.15252/embr.201744102
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696. https://doi.org/10.1002/prot.20449
Acknowledgements
We thank Svenja Hester and the Advanced Proteomics Facility for mass spectrometry analysis. P.L. received funding from Boehringer Ingelheim Fonds. B.A. acknowledges support from a Wellcome Trust Senior Research Fellowship (Grant No. 210622/Z/18/Z) and the European Molecular Biology Organisation Young Investigator Program. The 950 MHz spectrometer was upgraded with funding from the University of Oxford Wellcome Institutional Strategic Support Fund, the John Fell Fund, and the Edward Penley Abraham Cephalosporin Fund, and from the Engineering and Physical Sciences Research Council (Grant Ref: EP/R029849/1).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ludzia, P., Akiyoshi, B. & Redfield, C. 1H, 13C and 15N resonance assignments for the microtubule-binding domain of the kinetoplastid kinetochore protein KKT4 from Trypanosoma brucei. Biomol NMR Assign 14, 309–315 (2020). https://doi.org/10.1007/s12104-020-09968-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-020-09968-1