Abstract
Photoactive yellow protein (PYP) is involved in the negative phototactic response towards blue light of the bacterium Halorhodospira halophila. Here, we report nearly complete backbone and side chain 1H, 13C and 15N resonance assignments at pH 5.8 and 20 °C of PYP in its electronic ground state.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological context
Photoactive yellow protein (PYP) is a 125 amino acid (14 kDa) water-soluble, blue-light sensor protein, first found in the halophilic bacterium Halorhodospira halophila (Meyer 1985). PYP is a photoreceptor, believed to be responsible for the negative phototactic response of its host organism (Sprenger et al. 1993). This kind of response is required for organisms to evade potentially harmful short-wavelength light. Based on this observation, PYP has become a suitable model to understand the signal-transduction mechanism in Per-Arnt-Sim (PAS) domain signaling (Crosthwaite et al. 1997; Nambu et al. 1991). Several PYP-like proteins have meanwhile been found in other organisms, where they are also thought to act as light sensors. In addition, PYP-like proteins found in purple bacteria are involved in cell buoyancy or sensing bacteriophytochromes (Jiang et al. 1999; Kyndt et al. 2004).
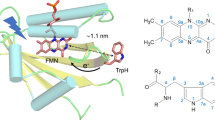
Understanding of light transduction in PYP requires structural information in atomic detail. A 1.4 Å crystallographic structure was determined in 1995 by Borgstahl et al. and in 1998 Düx and coworkers revealed the solution structure and backbone dynamics of PYP by NMR spectroscopy. The reaction center of PYP is protected from solvent by R52, which is believed to function as a gateway in the photocycle (Borgstahl et al. 1995; Genick et al. 1997). The chromophore, para-coumaric acid (pCA), is covalenty bound to C69 with a thioester bond and pCA participates in two short hydrogen bonds with E46 and Y42 to stabilize the negative charge of pCA in the electronic ground state, pG. Upon blue-light capture, the chromophore undergoes trans–cis isomerisation and the intermediate pR is formed, which subsequently relaxes to the proposed signaling state, pB. In the latter state, the reaction center is exposed and the two short hydrogen bonds are broken (Borgstahl et al. 1995; Sigala et al. 2009; Yamaguchi et al. 2009).
In this paper, we present the nearly complete assignment of the backbone and side chain resonances of the pG state of PYP.
Methods and experiments
Uniformly 13C,15N-labeled wild type PYP was overexpressed and purified as described previously (Düx et al. 1998). The NMR sample contained 1.2 mM doubly labeled [13C, 15N] PYP, 5 mM potassium phosphate buffer pH 5.8, 10 % D2O and 0.15 mM 2,2-dimethyl-2-silapentane-5-sulfonic acid (DSS). All 1D 1H, 2D and 3D NMR experiments were carried out at 20 °C using a Varian Unity INOVA 600 MHz spectrometer equipped with a field-gradient probe. Referencing was performed according to the method described by Wishart et al. (1995) using DSS. To obtain the sequential backbone resonance assignments, 2D 15N-HSQC equipped with sensitivity enhancement and water flip back pulses (Kay et al. 1992; Zhang et al. 1994), 3D CBCA(CO)NH/HNCACB (Wittekind and Mueller 1993) and 3D HNCO/HN(CA)CO (Ikura et al. 1990; Yamazaki et al. 1994) experiments were used.
To accomplish most of the side chain assignments, the strategy outlined by Oktaviani, et al. (2011) was performed. 3D (H)C(CO)NH-TOCSY and H(CCO)NH-TOCSY were used to assign carbon and proton aliphatic side chain resonances, respectively (Grzesiek et al. 1993; Logan et al. 1993; Montelione et al. 1992). For aromatic side chains, CB(CGCD)HD, CB(CGCDCE)HE (Yamazaki et al. 1993a), 1H–13C HSQC CParo (Zuiderweg et al. 1996), 2D 13C-1H CT HSQC, and 2D 13C–1H CT HSQCaro were recorded. Aromatic Cγ–Hβ correlations were detected using CG(CB)HB experiments (Prompers et al. 1998). Carbonyl 13C side chain resonances of Asx and Glx residues were assigned using H2(C)CO (Kay et al. 1990; Powers et al. 1991; Yamazaki et al. 1993b) and (HBGCBG)CO(CBGCABCON)H (Tollinger et al. 2002) experiments. Nζ–Hε (Lys) and Nε–Hδ (Arg) correlations were detected using H2(C)N (André et al. 2007).
All spectra were processed using NMRPipe (Delaglio et al. 1995) and they were analyzed with Sparky (Goddard and Kneller 2008).
Completeness of assignments and data deposition
The obtained backbone assignment is over 96 % complete (see Fig. 1). Missing assignments are the amide backbone 15N frequencies of the four Pro residues, and the amide nitrogen and proton frequencies of M1 and G7. At this pH no backbone assignment was possible for E12 and the 13C’ frequencies of F6, D10 and I11 were not found.
The entire 1H, 13C and 15N side chain resonance assignment is over 85 % complete. Unassigned side chain resonances mainly pertain to labile protons and their attached heteroatoms, such as Nη/Hη of the two Arg residues, protons connected to oxygen (i.e. Tyr, Ser and Thr OH), protons bound to nitrogen and their corresponding nitrogens in the imidazole moiety of His residues, Hζ of Lys, and also resonances due to several non-labile groups, such as methyl Cε/Hε of Met residues, and a number of signals due to Phe and Trp ring systems.
The Cζ frequency of the tyrosine residues were found in the 1H–13C HSQC CParo experiment (run overnight) and could be assigned by using a combination of CB(CGCD)HD and CB(CGCDCE)HE spectra. However, Y76 and Y118 have too similar Hε frequencies to be able to determine which Hε-Cζ peak corresponds to which residue. Therefore this information was obtained from pH-titration experiments, using the fact that tyrosine Cγ and Cζ chemical shifts change simultaneously by large amounts during this titration. As the Y76 signals start moving at a lower pH than those of Y118, it was possible to assign the Cζ nuclei in this fashion. We also can detect signals due to two short hydrogen bonds, which belong to Y42 and E46, using a 1D proton water flip-back sequence that suppresses the water signal without saturating it (see Fig. 2). These NMR signals have been assigned by Sigala et al. (2009).
Previously, 1H and 15N chemical shift assignments have been made at 37 °C by Düx et al. (1998), but these have not been submitted to the BMRB. Our results agree well with these, with a few exceptions. 1H, 13C and 15N assignments for the N-terminal deletion variant Δ25-PYP at 20 °C are available under BMRB accession number 6321 (Bernard et al. 2005), and these show significant differences as a result of the removal of part of the native protein structure.
The 1H, 13C and 15N assignment can be found in the BioMagResBank under accession number 18122.
References
André I, Linse S, Mulder FAA (2007) Residue-specific pK a determination of lysine and arginine side chains by indirect N-15 and C-13 NMR spectroscopy: application to apo calmodulin. J Am Chem Soc 129:15805–15813
Bernard C, Houben K, Derix N, Marks D, van der Horst M, Hellingwerf K, Boelens R, Kaptein R, van Nuland N (2005) The solution structure of a transient photoreceptor intermediate: delta25 photoactive yellow protein. Structure 13:953–962
Borgstahl GEO, Williams DR, Getzoff ED (1995) 1.4 angstrom structure of photoactive yellow protein, a cytosolic photoreceptor—unusual fold, active-site, and chromophore. Biochemistry 34:6278–6287
Crosthwaite SK, Dunlap JC, Loros JJ (1997) Neurospora wc-1 and wc-2: transcription, photoresponses, and the origins of circadian rhythmicity. Science 276:763–769
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe - a multidimensional spectral processing system based on Unix pipes. J Biomol NMR 6:277–293
Düx P, Rubinstenn G, Vuister GW, Boelens R, Mulder FAA, Hard K, Hoff WD, Kroon AR, Crielaard W, Hellingwerf KJ, Kaptein R (1998) Solution structure and backbone dynamics of the photoactive yellow protein. Biochemistry 37:12689–12699
Genick UK, Borgstahl GEO, Ng K, Ren Z, Pradervand C, Burke PM, Srajer V, Teng TY, Schildkamp W, Mcree DE, Moffat K, Getzoff ED (1997) Structure of a protein photocycle intermediate by millisecond time-resolved crystallography. Science 275:1471–1475
Goddard TD, Kneller DG (2008) SPARKY 3. University of California, San Francisco
Grzesiek S, Anglister J, Bax A (1993) Correlation of backbone amide and aliphatic side-chain resonances in 13C/15N-enriched proteins by isotropic mixing of 13C magnetization. J Magn Reson B 101:114–119
Ikura M, Kay LE, Bax A (1990) A novel-approach for sequential assignment of H-1, C-13, and N-15 spectra of larger proteins—heteronuclear triple-resonance 3-dimensional NMR-spectroscopy—application to calmodulin. Biochemistry 29:4659–4667
Jiang ZY, Swem LR, Rushing BG, Devanathan S, Tollin G, Bauer CE (1999) Bacterial photoreceptor with similarity to photoactive yellow protein and plant phytochromes. Science 285:406–409
Kay LE, Ikura M, Tschudin R, Bax A (1990) Three-dimensional triple-resonance NMR spectroscopy of isotopically enriched proteins. J Magn Reson 89:496–514
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665
Kyndt JA, Meyer TE, Cusanovich MA (2004) Photoactive yellow protein, bacteriophytochrome, and sensory rhodopsin in purple phototrophic bacteria. Photochem Photobiol Sci 3:519–530
Logan TM, Olejniczak ET, Xu RX, Fesik SW (1993) A general-method for assigning NMR-spectra of denatured proteins using 3D HC(CO)NH-TOCSY triple resonance experiments. J Biomol NMR 3:225–231
Meyer TE (1985) Isolation and characterization of soluble cytochromes, ferredoxins and other chromophoric proteins from the halophilic phototrophic bacterium Ectothiorhodospira-Halophila. Biochim Biophys Acta 806:175–183
Montelione GT, Lyons BA, Emerson SD, Tashiro M (1992) An efficient triple resonance experiment using carbon-13 isotropic mixing for determining sequence-specific resonance assignments of isotopically-enriched proteins. J Am Chem Soc 114:10974–10975
Nambu JR, Lewis JO, Wharton KA, Crews ST (1991) The Drosophila single-minded gene encodes a helix-loop-helix protein that acts as a master regulator of CNS midline development. Cell 67:1157–1167
Oktaviani NA, Otten R, Dijkstra K, Scheek RM, Thulin E, Akke M, Mulder FAA (2011) 100 % complete assignment of non-labile 1H, 13C, and 15N signals for calcium-loaded Calbindin D9k P43G. Biomol NMR Assign 5:79–84
Powers R, Gronenborn AM, Marius Clore G, Bax A (1991) Three-dimensional triple-resonance NMR of 13C/15N-enriched proteins using constant-time evolution. J Magn Reson 94:209–213
Prompers JJ, Groenewegen A, Hilbers CW, Pepermans HAM (1998) Two-dimensional NMR experiments for the assignment of aromatic side chains in C-13-labeled proteins. J Magn Reson 130:68–75
Sigala PA, Tsuchida MA, Herschlag D (2009) Hydrogen bond dynamics in the active site of photoactive yellow protein. Proc Natl Acad Sci USA 106:9232–9237
Sprenger WW, Hoff WD, Armitage JP, Hellingwerf KJ (1993) The eubacterium Ectothiorhodospira-Halophila is negatively phototactic, with a wavelength dependence that fits the absorption-spectrum of the photoactive yellow protein. J Bacteriol 175:3096–3104
Tollinger M, Forman-Kay JD, Kay LE (2002) Measurement of side-chain carboxyl pK a values of glutamate and aspartate residues in an unfolded protein by multinuclear NMR spectroscopy. J Am Chem Soc 124:5714–5717
Wishart DS, Bigam CG, Yao J, Abildgaard F, Dyson HJ, Oldfield E, Markley JL, Sykes BD (1995) H-1, C-13 and N-15 chemical-shift referencing in biomolecular NMR. J Biomol NMR 6:135–140
Wittekind M, Mueller L (1993) HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha-carbon and beta-carbon resonances in proteins. J Magn Reson B 101:201–205
Yamaguchi S, Kamikubo H, Kurihara K, Kuroki R, Niimura N, Shimizu N, Yamazaki Y, Kataoka M (2009) Low-barrier hydrogen bond in photoactive yellow protein. Proc Natl Acad Sci USA 106:440–444
Yamazaki T, Formankay JD, Kay LE (1993a) 2-dimensional NMR experiments for correlating C-13-beta and H-1-delta/epsilon chemical-shifts of aromatic residues in C-13-labeled proteins via scalar couplings. J Am Chem Soc 115:11054–11055
Yamazaki T, Yoshida M, Nagayama K (1993b) Complete assignments of magnetic resonances of ribonuclease-H from Escherichia-Coli by double-resonance and triple-resonance 2D and 3D NMR spectroscopies. Biochemistry 32:5656–5669
Yamazaki T, Lee W, Arrowsmith CH, Muhandiram DR, Kay LE (1994) A suite of triple-resonance NMR experiments for the backbone assignment of N-15, C-13, H-2 labeled proteins with high-sensitivity. J Am Chem Soc 116:11655–11666
Zhang O, Kay LE, Olivier JP, Forman-Kay JD (1994) Backbone 1H and 15N resonance assignments of the N-terminal SH3 domain of drk in folded and unfolded states using enhanced-sensitivity pulsed field gradient NMR techniques. J Biomol NMR 4:845–858
Zuiderweg ERP, Zeng L, Brutscher B, Morshauser RC (1996) Band-selective hetero- and homonuclear cross-polarization using trains of shaped pulses. J Biomol NMR 8:147–160
Acknowledgments
This work was supported by a VIDI career development award to F.A.A.M from The Netherlands Organization for Scientific Research (NWO).
Conflict of interest
The authors declare that they have no conflict of interest.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Pool, T.J., Oktaviani, N.A., Kamikubo, H. et al. 1H, 13C, and 15N resonance assignment of photoactive yellow protein. Biomol NMR Assign 7, 97–100 (2013). https://doi.org/10.1007/s12104-012-9387-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-012-9387-9