Abstract
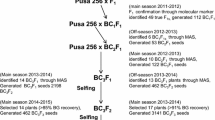
Cotton leaf curl disease (CLCuD), caused by a geminivirus complex, is the most serious disease of upland cotton in northwest India and Pakistan. It results in substantial losses in cotton yield and fibre quality. Due to continuous appearance of new viral strains, all the established CLCuD resistant stocks, extant and obsolete cultivars of upland cotton have become susceptible. Therefore, it became crucial to explore the novel sources of CLCuD resistance, as development of CLCuD resistant varieties is the most practical approach to manage this menace. Here, for the first time, we report introgression and mapping of CLCuD resistance from a ‘synthetic cotton polyploid’ to upland cotton. A backcross population (synthetic polyploid / Gossypium hirsutum Acc. PIL 43/G. hirsutum Acc. PIL 43) was developed for studying inheritance and mapping of CLCuD resistance. Dominance of CLCuD resistance was observed over its susceptibility. Two dominant genes were found to confer resistance to CLCuD. Molecular analysis through genotyping-by-sequencing revealed that chromosomes A01 and D07 harboured one CLCuD resistance gene each.



Similar content being viewed by others
References
Ahmad G., Malik S. A., Mamood Z., Iqbal M. Z. and Ahmad S. 2002 Effect of cotton leaf curl virus disease on morphology, yield and fibre characteristics of susceptible lines/cultivars of cotton (Gossypium hirsutum L.). Asian. J. Plant. Sci. 1, 705–707.
Ahmad S., Mahmood K., Hanif M., Nazeer W., Malik W., Qayyum A. et al. 2011 Introgression of cotton leaf curl virus resistance genes from Asiatic cotton (Gosssypium arboreum) into upland cotton (G. hirsutum). Genet. Mol. Res. 10, 2404–2414.
Ahuja S. L., Monga D. and Dhayal L. S. 2007 Genetics of resistance to cotton leaf curl disease in Gossypium hirsutum L. under field conditions. J. Hered. 98, 79–83.
Ajmera B. D. 1994 Occurrence of leaf curl virus on American cotton (G. hirsutum) in north Rajasthan. Poster presentation, National Seminar on Cotton Production Challenges in 21st Century, April 18–20, Hisar, India.
Ali M. 1997 Breeding of cotton varieties for resistance to cotton leaf curl virus. Pak. J. Phytopathol. 9, 1–7.
Anjum Z. I., Azhar M. T., Hayat K., Ashraf F., Shahzad U. and Azam M. 2014 Development of high yielding and CLCuV resistant upland cotton variety “CIM-608.” Pak. J. Phytopathol. 26, 23–32.
Aslam M., Jiang C., Wright R. and Paterson A. H. 2000 Identification of molecular markers linked to leaf curl virus disease resistance in cotton. J. Sci. I. R. Iran. 11, 277–280.
Beasley J. O. 1942 Meiotic chromosome behavior in species, species hybrids, haploids, and induced polyploids of Gossypium. Genetics 27, 25–54.
Bell A. and Robinson A. F. 2004 Development and characteristics of triple species hybrids used to transfer reniform nematode resistance from Gossypium longicalyx to Gossypium hirsutum, pp. 422–426. Beltwide Cotton Conferences, San Antonio.
Briddon R. W. and Markham P. G. 2000 Cotton leaf curl virus disease. Virus Res. 71, 151–159.
Briddon R. W., Mansoor S., Bedford I. D., Pinner M. S., Saunders K., Stanley J. et al. 2001 Identification of DNA components required for induction of cotton leaf curl disease. Virology 285, 234–243.
Brubaker C. L. and Brown A. H. D. 2003 The use of multiple alien chromosome addition aneuploids facilitates genetic linkage mapping of the Gossypium G genome. Genome 46, 774–791.
Cai J. H., Xie K., Lin L., Qin B. X., Chen B. S., Meng J. R. and Liu Y. L. 2010 Cotton leaf curl Multan virus newly reported to be associated with cotton leaf curl disease in China. Plant Pathol. 59, 794–795.
Chakrabarty P. K., Kumar P., Kalbande B. B., Chavhan R. L., Koundal V., Monga D. et al. 2020 Recombinant variants of cotton leaf curl Multan virus is associated with the breakdown of leaf curl resistance in cotton in northwestern India. Virus Dis. 31, 45–55.
Chen Y., Wang Y., Zhao T., Yang J., Feng S., Nazeer W. et al. 2015 A new synthetic amphiploid (AADDAA) between Gossypium hirsutum and G. arboreum lays the foundation for transferring resistances to Verticillium and drought. PLoS One 10, e0128981.
Ellango R., Singh S. T., Rana V. S., Priya N. G., Raina H., Chaubey R. et al. 2015 Distribution of Bemisia tabaci genetic groups in India. Environ. Entomol. 44, 1258–1264.
Farooq J., Farooq A., Rizwan M., Petrescu-Mag V. I., Ali M. A., Mahmood K. et al. 2015 Cotton fibers: attributes of specialized cells and factors affecting them. Adv. Environ. Sci. Int. J. Bioflux Soc. 7, 369–382.
Haider S., Khan I. A. and Mansoor S. 2003 Genetics of cotton leaf curl virus disease in upland cotton. Sarhad J. Agric. 19, 207–210.
Hussain T. and Ali M. 1975 A review of cotton diseases of Pakistan. Pak. Cottons 19, 71–86.
Hussain M., Azhar F. M., Khan A. A. and Ali Z. 2012 Expression of genes controlling the inheritance of resistance to cotton leaf curl virus disease (CLCuD) in Gossypium hirsutum L.: a quantitative analysis. Pak. J. Bot. 44, 247–254.
Iqbal M., Chang M. A., Mahmood A., Khumber M. B., Nasir A. and Hassan M. A. 2003 Inheritance of response to cotton leaf curl virus (CLCuV) infection in cotton. Asian J. Plant Sci. 2, 261–264.
Javed M., Hussain S. B. and Baber M. 2019 Role of QTL mapping to circumscribe various diseases in different crops with special emphasis on cotton. J. Genet. Mol. Biol. 3, 23–33.
Khan N. U. 2013 Diallel analysis of cotton leaf curl virus (CLCuV) disease, earliness, yield and fiber traits under CLCuV infestation in upland cotton. Aust. J. Crop Sci. 7, 1955–1966.
Kranthi K. R. 2019 ICAC cotton data book 2020. International Cotton Advisory Committee, Washington.
Mahmood Z. 2004 Inheritance of cotton leaf curl virus resistance in cotton (Gossypium hirsutum L.). J. Res. Sci. 15, 297–299.
Mansoor S., Bedford I., Pinner M. S., Stanley J. and Markham P. G. 1993 A whitefly-transmitted geminivirus associated with cotton leaf curl disease in Pakistan. Pak. J. Bot. 25, 105–107.
Mansoor S., Khan S. H., Bashir A., Saeed M., Zafar Y., Malik K. A. et al. 1999 Identification of a novel circular single-stranded DNA associated with cotton leaf curl disease in Pakistan. Virology 259, 190–199.
Margarido G. R. A., Souza A. P. and Garcia A. A. F. 2007 OneMap: software for genetic mapping in outcrossing species. Hereditas 144, 78–79.
Monga D. 2014 Status of cotton leaf curls virus disease in India. Presented in 6th meeting of Asian Cotton Research and Development Network held in Dhaka, Bangladesh, from 18–20 June 2014.
Monga D., Chakrabarty P. K. and Kranthi K. R. 2011 Cotton leaf curl virus disease in India—Recent status and management strategies. ICAC Recorder XXIX, 6–7.
Monga D., Narula A. M. and Raj S. 2001 Management of cotton leaf curl virus—A dreaded disease in north India. In National Seminar on Sustainable Cotton Production to Meet the Future Requirement of Industry, pp.112–115, Directorate of Cotton Development, Mumbai.
Monga D., Raj S. and Mayee C. D. 2004 Strategies for cotton leaf curl virus disease management. In National symposium on changing world order-cotton research development and policy in context, pp. 205–213. ANGRAU, Hyderabad.
Monga D. and Sain S. K. 2021 Incidence and severity of cotton leaf curl virus disease on different BG II hybrids and its effect on the yield and quality of cotton crop. J. Environ. Biol. 42, 90–98.
Naveen N. C., Chaubey R., Kumar D., Rebijith K. B., Rajagopal R., Subrahmanyam B. et al. 2017 Insecticide resistance status in the whitefly, Bemisia tabaci genetic groups Asia-I, Asia-II-1 and Asia-II-7 on the Indian subcontinent. Sci. Rep. 7, 40634.
Pathak D., Bala S., Rathore P., Sekhon P. S. and Singh K. 2016 Identification of new sources of resistance to cotton leaf curl disease and its introgression in American cotton. Proceedings 6th World Cotton Research Conf. pp 50–51. Goiania, Brazil.
Pathak D., Rathore P. and Gumber R. K. 2009 Cytoplasmic effects in relation to cotton leaf curl disease resistance in Gossypium hirsutum L Icfai Univ. J. Genet. Evol. 2, 31–35.
Pathak D., Suneja Y. and Gill A. K. 2019 Global status of cotton genomics and utilization in improving trait value. ICAC Recorder XXXVII, 5–18.
Peterson B. K., Weber J. N., Kay E. H., Fisher H. S. and Hoekstra H. E. 2012 Double digest RADseq: an inexpensive method for de novo SNP discovery and genotyping in model and non-model species. PLoS One 7, e37135.
Rahman M., Hussain D., Malik T. A. and Zafar Y. 2005 Genetics of resistance to cotton leaf curl disease in Gossypium hirsutum. Plant Pathol. 54, 764–772.
Rahman M., Khan A. Q., Rahmat Z., Iqbal M. A. and Zafar Y. 2017 Genetics and genomics of cottonleaf curl disease, its viral causal agents and whitefly vector: a way forward to sustain cotton fiber security. Front. Plant Sci. 8, 1157.
Rahman M. and Zafar Y. 2007 Registration of ‘NIBGE-2’ Cotton. J. Plant Regist. 1, 113–114.
Rajagopalan P. A., Naik A., Katturi P., Kurulekar M., Kankanallu R. S. and Anandalakshmi R. 2012 Dominance of resistance-breaking cotton leaf curl Burewala virus (CLCuBuV) in northwestern India. Arch. Virol. 157, 855–868.
RStudio Team 2020 RStudio: Integrated development environment for R. RStudio, PBC, Boston, MA. http://www.rstudio.com/.
Sacks E. J. and Robinson A. F. 2009 Introgression of resistance to reniform nematode (Rotylenchulus reniformis) into upland cotton (Gossypium hirsutum) from Gossypium arboreum and a G. hirsutum/Gossypium aridum bridging line. Field Crops Res. 112, 1–6.
Saeed M., Behjatnia S. A., Mansoor S., Zafar Y., Hasnain S. and Rezaian M. A. 2005 A single complementary-sense transcript of a geminiviral DNA beta satellite is determinant of pathogenicity. Mol. Plant-Microbe Interact. 18, 7–14.
Saghai-Maroof M. A., Soliman K. M., Jorgensen R. A. and Allard R. W. 1984 Ribosomal DNA spacer-length polymorphism in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc. Natl. Acad. Sci. USA 81, 8014–8018.
Singh D., Gill J. S., Gumber R. K., Singh R. and Singh S. 2013 Yield and fibre quality associated with cotton leaf curl disease of Bt cotton in Punjab. J. Environ. Biol. 34, 113–116.
Suthar T., Gupta N., Pathak D., Sharma S. and Rathore P. 2021 Morpho-anatomical characterization of interspecific derivatives of Gossypium hirsutum L × G armourianum Kearney cross for whitefly tolerance. Phytoparasitica, https://doi.org/10.1007/s12600-021-00963-3.
Vij S., Pathak D., Rathore P. and Nikhanj P. 2020 Genetic analysis of some morphological traits in synthetic × naturally polyploid cotton derivatives. J. Genet. 99, 73.
Voorrips R. E. 2002 MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 93, 77–78.
Vyskocilova S., Tay W. T., Brunschot S. V., Susan S. and Colvin J. 2018 An integrative approach to discovering cryptic species within the Bemisia tabaci whitefly species complex. Sci. Rep. 8, 10886.
Zaidi S. S., Naqvi R. Z., Asif M., Strickler S., Shakir S. and Shafiq M. 2020 Molecular insight into cotton leaf curl geminivirus disease resistance in cultivated cotton (Gossypium hirsutum). Plant Biotechnol. J. 18, 691–706.
Zhang X., Zhai C., He L., Guo Q., Zhang X., Xu P. et al. 2014 Morphological, cytological and molecular analyses of a synthetic hexaploid derived from an interspecific hybrid between Gossypium hirsutum and Gossypium anomalum. Crop J. 2, 272–277.
Acknowledgements
The research work was carried out under the Programme Support on ‘Enhancing Durability of Resistance to Biotic Stresses in Selected Cereal and Fibre Crops through Biotechnological Approaches (BT/01/CE1B/121/01)’ funded by the Department of Biotechnology, Government of India. Ministry of Science and Technology (Grant No. 102/IFD/SAN/1307/2014-15).
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Manoj Prasad
Rights and permissions
About this article
Cite this article
Vij, S., Pathak, D., Rathore, P. et al. Molecular mapping of CLCuD resistance introgressed from synthetic cotton polyploid in upland cotton. J Genet 101, 25 (2022). https://doi.org/10.1007/s12041-022-01365-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12041-022-01365-y