Abstract
Declining cerebral blood flow leads to chronic cerebral hypoperfusion which can induce neurodegenerative disorders, such as vascular dementia. The reduced energy supply of the brain impairs mitochondrial functions that could trigger further damaging cellular processes. We carried out stepwise bilateral common carotid occlusions on rats and investigated long-term mitochondrial, mitochondria-associated membrane (MAM), and cerebrospinal fluid (CSF) proteome changes. Samples were studied by gel-based and mass spectrometry-based proteomic analyses. We found 19, 35, and 12 significantly altered proteins in the mitochondria, MAM, and CSF, respectively. Most of the changed proteins were involved in protein turnover and import in all three sample types. We confirmed decreased levels of proteins involved in protein folding and amino acid catabolism, such as P4hb and Hibadh in the mitochondria by western blot. We detected reduced levels of several components of protein synthesis and degradation in the CSF as well as in the subcellular fractions, implying that hypoperfusion-induced altered protein turnover of brain tissue can be detected in the CSF by proteomic analysis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chronic cerebral hypoperfusion (CCH) is a pathological state that is characterized by declining cerebral blood flow (CBF). One of the potential risk factors for vascular dementia (VD) and sporadic Alzheimer’s disease (AD) is the decreased blood flow of the brain [1, 2]. Bilateral common carotid artery occlusion (BCCAO) induces reduced cerebral blood flow, which results in oxidative stress, inflammatory response, and disturbed lipid metabolism leading to cognitive impairment [3]. In the stepwise BCCAO model, rats have 1 week of regeneration time between the bilateral common carotid artery occlusions, leading to a gradual decrease in cerebral blood flow. After 8 weeks of occlusion, the CBF is almost completely recovered in the frontal cortex [4]; however, cognitive deficits and oxidative stress are apparent [5]. In this study, we performed stepwise BCCAO and investigated the long-term effects of the model.
Brain plasticity is the ability of the brain to modify its structure and function in response to the alterations of its environment. Many subcellular organelles can adapt to changing environments of neurons such as synapses and mitochondria. In our previous studies, we have investigated mitochondrial [6] and mitochondrial-associated membrane (MAM) proteome [7] alterations in Alzheimer’s disease model animals, revealing the relevance of these organelles in neurodegeneration.
Since the brain is one of the most energy-demanding organs and its ATP source relies mostly on oxidative phosphorylation, mitochondrial proteome changes can give insight into its molecular adaptation to CCH. The mitochondria are vital, dynamic, and plastic organelles that are essential for maintaining membrane ion gradients, neurotransmission, and synaptic plasticity, requiring a large amount of energy. Moreover, mitochondria participate in other neuronal processes such as calcium buffering and intracellular signaling. Dysfunctions of mitochondria were implicated in VD and its animal models [5, 8, 9] and several other neurodegenerative disorders [10,11,12,13]. In the BCCAO model, increased mitochondrial DNA deletion and structural damage were observed [14]. Despite the known role of mitochondria in CCH, our study is the first proteomic analysis of this organelle in the BCCAO model.
Furthermore, mitochondrial dynamics are disturbed in several neurodegenerative disorders [11, 15]. MAM also participates in the protein supply of mitochondria and regulates mitochondrial dynamics [16]. Besides mitochondrial dynamics, MAM also regulates essential cellular processes, such as fatty acid metabolism and calcium homeostasis [17]. Also, it has a main role in the processing of amyloid precursor protein which is dysregulated in AD [18]. In addition, MAM dysfunction is involved in other neurodegenerative diseases [17]; however, its role in CCH is poorly studied.
Cerebrospinal fluid (CSF) has a vital role in the clearance of brain interstitial fluid [19]. Due to its direct contact with the brain, its content can reflect biochemical changes in the brain. CSF has detectable protein content; therefore, proteomic alterations in the CSF might provide potential biomarkers that can indicate early, pre-symptomatic pathological alterations, and CSF sampling is feasible in human translational studies.
Additionally, decreased brain metabolism is one of the earliest clinical symptoms of AD which induces mitochondrial dysfunction in the brain [20]. Detecting molecular shifts of mitochondria or MAM in the CSF can provide early and detectable pathological signs of neurodegeneration. In the current study, we aimed to monitor changes induced by BCCAO in the MAM and mitochondria of the frontal cortex which is one of the brain areas responsible for higher-order cognitive functions and affected in VD [21, 22]. We also analyzed the proteome of the CSF to reveal whether organellar changes could be detected in body fluids. We applied mass spectrometry-based (MS-based) and gel-based proteomic methods to study the proteomic alterations - induced by stepwise BCCAO - of CSF and the organelles of the frontal cortex, respectively.
Materials and Methods
Animals
Adult male Wistar rats (aged 3 months, weighing ~ 220 g) were obtained from Toxi-Coop Ltd. (Budapest, Hungary). Rats were housed under standard laboratory conditions (lights on at 9:00 AM, lights off at 9:00 PM) in temperature- and humidity-controlled rooms with ad libitum access to food and water. All animal care and experimental procedures were following the Council Directive 86/609/EEC, the Hungarian Act of Animal Care and Experimentation (1998, XXVIII). All the procedures conformed to the National Institutes of Health guidelines were in accordance with the guidelines of the local Animal Care and Use Committee and were approved by the local Ethical Committee of Gedeon Richter Plc. (PE/EA/2885–6/2016). All efforts were carried out to minimize the animals’ pain and suffering and to reduce the number of animals used. A total of 16 rats were used in the experiments; they were assigned randomly to operated and sham-operated groups. We guaranteed the blindness of the experimenters whenever it was possible.
Stepwise Bilateral Occlusion of Common Carotid Arteries
Stepwise bilateral common carotid artery occlusion was performed on rats as previously described [7, 23, 24]. Briefly, rats were anesthetized with isoflurane (1.5–2% in air), and a ventral midline incision was placed on the neck. First, the left common carotid artery was exposed, gently separated from the vagus nerve, and occluded by three ligatures (2–0). The animals were allowed to regenerate for a week in their home cages. Then, the same surgical procedure was performed on the right common carotid artery too. Rats in the control group underwent a sham operation as they received the same surgical procedures in both steps, but only a thread was placed around the vessels without ligation of the arteries.
Magnetic Resonance Angiography
The efficacy of the occlusions and the changes in cerebral blood flow were monitored with MRA, on the second and fifth weeks after the second occlusion, based on our previous studies [23, 24]. Anesthetized animals were scanned in a 9.4 T MRI system (Varian, Medical Systems Inc., Palo Alto, CA, USA) with a free bore of 210 mm, containing a 120 mm inner size gradient coil. Three-dimensional time-of-flight angiography (3D gradient echo) was performed at TR/TE = 30/2.8 ms, resolution = 0.42 × 0.42 × 0.46 mm. The MRA studies were performed to evaluate cervical and intracranial arteries and parenchymal injury. For proteomic experiments, the animals had to meet the criteria as follows: (i) lack of gross anatomical abnormalities (i.e., pathologically large or asymmetrical anatomical structures) and (ii) lack of extensive lesions or any sign of extended tissue impairment. The MRA revealed ischemic lesions in none of the animals. Maximum intensity projection was applied on the volumetric MRA recordings to create 2D images. The images were analyzed by ImageJ2; to discard background noise, the low threshold of 110 was applied on the intensity values. Then, the images were inverted, and mean gray values were obtained. A nonparametric, Mann–Whitney test was applied to the mean gray values of the MRA images with a significance level of 0.05.
Collection of CSF and Preparation of Mitochondria and MAM Samples
We collected CSF and prepared mitochondria and the subcellular fraction containing MAM from the frontal cortices of sham-operated and operated rats based on the protocol of Wieckowski et al. [25]. Animals were anesthetized with isoflurane (1.5–2% in air), and CSF was collected from cisterna magna, then rats were sacrificed and their brains were quickly removed from the skull. Brain samples were washed in ice-cold isolation buffer-1 (IB-1) (225 mM mannitol, 75 mM saccharose, 0.5% BSA, 0.5 mM EGTA, and 30 mM Tris–HCl pH 7.4), then three times in ice-cold isolation buffer-3 (IB-3) (225 mM mannitol, 75 mM saccharose, and 30 mM Tris–HCl pH 7.4). Brain samples were cut into smaller pieces and washed in IB-1. Brain samples were homogenized in IB-1 supplemented with protease and phosphatase inhibitor cocktail with a Dounce tissue grinder (Sigma-Aldrich) manually (15 strokes per sample). Homogenized samples were centrifuged at 740 × g for 7 min at 4 °C. Supernatants were centrifuged at 9000 × g for 10 min at 4 °C, and pellets were suspended in isolation buffer-2 (IB-2) (225 mM mannitol, 75 mM saccharose, 0.5% BSA, and 30 mM Tris–HCl pH 7.4). Samples were centrifuged at 10,000 × g for 10 min at 4 °C, and pellets were suspended in IB-3. Samples were centrifuged again at 10,000 × g for 10 min at 4 °C, and pellets were suspended in mitochondria resuspending buffer (MRB) (250 mM mannitol, 5 mM HEPES (pH 7.4), and 0.5 mM EGTA); this is the crude mitochondrial fraction that contains MAM. One-fourth of each sample was centrifuged at 10,000 × g for 10 min at 4 °C, and pellets were resuspended in lysis buffer (7 M urea, 2 M thiourea, 4% CHAPS, 20 mM Tris, 5 mM magnesium-acetate). The other three-fourths of the fractions were used for pure mitochondria preparation. Samples were layered onto Percoll medium and centrifuged at 95,000 × g for 35 min at 4 °C with SW-40 Ti rotor. The pure mitochondria fraction of the sample was collected from the interface and centrifuged at 6300 × g for 10 min at 4 °C. Pellets were resuspended and centrifuged again at 6300 × g for 10 min at 4 °C with 70.1 Ti rotor, and pellets were resuspended in lysis buffer. All the samples were stored at − 80 °C until use.
Proteomic Investigation of Subcellular Fractions
We investigated the proteome alterations of MAM-containing and mitochondria samples from the frontal cortex of 6 sham-operated and 6 BCCAO rats, using two-dimensional differential gel electrophoresis (2-D DIGE). The samples were adjusted to pH 8 and pH 8.5, respectively. Then, their protein concentration was measured by the 2D-Quant kit (GE Healthcare, Chicago, IL, USA). The fluorescent labeling of the mitochondrial proteins was conducted with a CyDye DIGE Fluor Minimal dye labeling kit (Cytiva). From BCCAO and control rats, 50 µg of protein of each sample were randomly labeled with either Cy5 or Cy3 dyes, while the internal sample (containing equal protein amounts (25 μg)) was labeled with Cy2. MAM-containing samples were labeled with CyDye DIGE Fluor labeling kit for Scarce sample (Cytiva). Five µg of protein from each sample and the internal standard were labeled with Cy3 and Cy5, respectively. The differently labeled samples were mixed and rehydrated passively onto Immobiline DryStrip gel strips (24 cm, pH 3–10 NL, GE Healthcare) overnight. Isoelectric focusing (IEF) was performed in an EttanIPGphor 3 IEF unit for 24 h to attain a total of 100 kVh (GE Healthcare). Following IEF, the mitochondrial proteins were reduced and carbamidomethylated using an equilibration buffer containing 1% mercaptoethanol and 2.5% iodoacetamide, respectively. In the case of MAM, proteins were only reduced as previously mentioned. SDS-PAGE separation was performed on 24 × 20 cm, 10% polyacrylamide gels in an EttanDALTsix Electrophoresis System (GE Healthcare). Then, the gels were scanned with a TyphoonTRIO + scanner (GE Healthcare) using appropriate lasers and filters with the photomultiplier tube biased at 600 V. Differential protein analysis was performed using the DeCyder v7.0 software package (GE Healthcare), employing its differential analysis and biological variance analysis modules. The fluorescence intensities of the Cy3 and Cy5 dyes on a particular gel were normalized to the intensity of the Cy2 dye, or Cy3 intensities were normalized to the Cy5 dye. Quantitation of the fluorescence intensities of the protein spots and statistical analyses were carried out using the software. Independent two-tailed Student’s t-test was performed on spots that are present on at least 80% of the gels and have fold changes of more than ± 1.2. The Benjamini–Hochberg procedure was applied with a false discovery rate of 0.25. In the results, we show the original P-values of only those spots that remained significant after the procedure. Statistically significantly altered protein spots (P < 0.05) were picked for further protein identification. For the identification of proteins in spots of interest, preparative 2-D gel electrophoresis was performed separately using a total of 800 μg of synaptic proteins per gel. Resolved protein spots were visualized by the Colloidal Coomassie Blue G-250 stain (Merck Millipore, Billerica, MA, USA). One preparative gel for each brain region was run, and the selected spots were manually excised from the gels with pipette tips for protein identification. Excised spots were placed in a 1% acetic acid solution.
Mass Spectrometry-Based Protein Identification from 2-D Gel Spots
Proteins in the selected 2-D gel spots were in-gel digested as described in the protocol available online (http://msf.ucsf.edu/proto cols.html). Briefly, gel spots were cut into smaller cubes, washed with 25 mM ammonium-bicarbonate/50% acetonitrile, reduced using 10 mM TCEP and alkylated with 55 mM MMTS. After dehydration, the gel pieces were rehydrated with 100 ng, trypsin )sequencing grade, side chain protected porcine trypsin, Promega) in 20 μl of 25 mM ammonium-bicarbonate. Samples were digested for 4 h at 37 °C. Tryptic peptides were extracted and dried in a vacuum centrifuge.
Samples were reconstructed in 20 µl of 0.1% formic acid before mass spectrometric analysis. Five µl of the digest was injected for LC–MS/MS analyses onto an LTQ-Orbitrap Elite (Thermo Fisher Scientific) mass spectrometer online coupled with a nanoAcquity UPLC (waters) system. In order to shorten the injection time, the sample was injected with a high flow rate to a trap column (Symmetry C18, nanoACQUITY UPLC 2D, V/M 0.180 mm × 20 mm, 5 µm, 100 Å, waters,) and a nano column (BEH130, C18 Acquityuplc column, 0.100 mm × 100 mm, 1.7 µm,130 Å, waters) was used for the analysis. Gradient elution was applied from 3 to 40% of eluent B (0.1% formic acid in acetonitrile) in 37 min. Mass spectrometry data were collected in a data-dependent manner; a high-resolution survey scan was followed by a maximum of 20 dependent CID spectra analyzed in the ion-trap. Only multiple charged precursor ions were selected for fragmentation, and after that, they were excluded for 30 s from the repeated selection. A PAVA script (UCSF, MSF, San Francisco, CA) was used for peak picking and our in-cloudProteinProspector (version: 5.22.0) server (https://cloud.mta.hu/) was used for database search, with the following parameters: UniProtKB.2019.6.12.random.concat database was filtered for the rat sequences concatenated with the most frequent contaminant proteins (36,319 sequences; with 209 additional contaminant protein sequences were searched). Only tryptic peptides were considered, with a maximum of one missed cleavage site. Several variable modifications were set as acetyl (protein N-term), acetyl + oxidation (protein N-term M), Gln- > pyro-Glu (N-term Q), Met-loss (protein N-term M), Met-loss + acetyl (protein N-term M), oxidation (M), and carbamidomethyl- and methyltio-cysteine. Mass accuracy was set to 10 ppm for the parent and 0.6 Da for fragment ions. Proteins and peptides were accepted with a maximum of 1% FDR. Proteins were rejected with less than 10 unique peptides, 30 peptide counts, or 40% of sequence coverage; accepted proteins are shown in Table 1.
CSF Proteome Analysis Using LC–MS/MS
The protein content of the collected rat cerebrospinal fluid samples was treated with trypsin using the S-Trap micro spin columns according to the vendor’s protocol (https://protifi.com/pages/protocols), and the resulting peptide mixture was analyzed using a Waters MClass nUPLC-Thermo Orbitrap Fusion Lumos Tribrid LC–MS system in a data-dependent fashion. Proteins were identified using the Protein Prospector BatchTag Web software applying score-based acceptance criteria. In more detail, the tryptic digest was injected onto a trapping column (MClass Symmetry Waters Acquity UPLC Trap Col 2G V/M C18 column, 0.180 mm ID * 20 mm L, 5 m particle size, 100 Å pore size; loading time: 3 min with 1% B at 5 l/min) and after desalting was separated using a nonlinear gradient of 10–50% B in 80 min (solvent A: 0.1% formic acid/water, solvent B: 0.1% formic acid/acetonitrile, flow rate: 400 nl/min) on a waters separating nanocolumn (nanoAcquity UPLC BEH130 C18 column, 0.075 mm ID*250 mm L, 1.7 m particle size, 130 Å pore size; column temperature: 40 °C). Peptides eluting from the column were analyzed in 2-s cycles selecting the most abundant multiply charged ions (z = 2–6, m/z range: 380–1580) for HCD fragmentation (normalized collision energy: 35%) following each MS1 scan. Both MS and MS/MS spectra were collected in the Orbitrap analyzer with a resolution of 120,000 or 15,000, respectively. Raw data were converted into peaklists using the Proteome Discoverer (v2.4 SP1) software. Proteins were identified using the BatchTag Web software of Protein Prospector (v6.3.1.) with the following parameters: database: Uniprot Rattus norvegicus sequences (2020.10.07. version, 36,457 sequences) concatenated with a randomized version for each entry and also supplemented with 172 additional sequences representing the most common contaminant proteins (such as trypsin and human keratins); enzyme: trypsin allowing maximum one missed cleavage site; modifications: static: methylthio on Cys, variable: cleavage of Met and/or acetylation of protein N-termini, oxidation of Met, pyroglutamic acid formation from peptide N-terminal Gln, deamidation of Gln or Asn, allowing up to two variable modifications/peptide; mass accuracy: 5 and 20 ppm for precursor and fragment ions, respectively defined as monoisotopic values. Acceptance parameters: minimum score: 22 and 15, maximum E-value: 0.01 and 0.05 on the protein and peptide level, respectively; minimum protein best discriminant score: 0. Peptide level false discovery rate was below 1% for all samples as estimated by the incidence of randomized sequence identifications.
Quantification of the proteins across the sample groups was performed using spectral counting. Three sham-operated control and nine BCCAO-operated rats were compared. Peptide counts of proteins were normalized to the total peptide count found in each sample. Proteins were removed from analysis with more than 1 or 6 missing values among the control or operated rats, respectively. Protein abundance mean ratios of BCCAO and control samples were calculated. An independent two-tailed Student’s t-test was performed on proteins of more than 1.5 or less than 0.5 mean ratios. The Benjamini–Hochberg procedure was applied with a false discovery rate of 0.25. In Online resource 1, we show the original P-values of only those proteins that remained significant after the procedure.
Hierarchical Clustering and Principal Component Analysis
Hierarchical clustering and principal component analysis (PCA) were based on significantly altered proteins. PCA was performed in STASTICA 8.0 (StatSoft Inc., Tulsa, OK, USA); the calculations of gel-based proteomics and MS-based proteomics data were based on the correlations of normalized fluorescence intensities and normalized protein abundances, respectively. For the clustering of gel-based proteomics and MS-based proteomics data, z-scores of normalized fluorescence intensities and normalized protein abundances were calculated, respectively. Then, hierarchical clustering was performed, and heatmaps were created with R v4.0.5 programming language in RStudio v2022.07.1 with the heatmap.2 function of gplots package which uses agglomerative hierarchical clustering.
Enrichment Analyses of Altered Proteins
Enrichment analysis was performed on altered proteins of MAM, mitochondria, and CSF, using the DAVID Bioinformatics Resources v2022q1 [26, 27] with the background of the rat genome. The functional annotation tool of DAVID was used to evaluate the enrichment of the KEGG pathway, GO biological process, cellular component, and molecular function terms with the EASE score and count threshold of 0.01 and 2, respectively. Benjamini–Hochberg correction was applied for controlling the false discovery rate.
Validations of Protein Changes by Western Blot
We carried out western blot experiments on mitochondrial (n = 8) samples to confirm our proteomic results. Samples (30 µg) were diluted with sample loading buffer (1 M Tris–HCl pH = 6.8, 8 w/v% SDS, 24 v/v% glycerol, 200 mM DTT, 0.2 v/v% bromophenol blue) and incubated at 96 °C for 5 min. Protein samples were separated on 10% acrylamide gel by SDS-PAGE. Then, the gel was transferred to a PVDF membrane. The membrane was washed with distilled water and stained with Ponceau S stain (0.5 (w/v)% in 1 (v/v)% acetic acid), after washing with distilled water; it was photographed when it air dried. The membrane was destained in 200 µM NaOH and 20% acetonitrile and washed again in distilled water, then blocked with 3% BSA solution (in TBS-T for 1 h at RT). The membrane was stained with primary antibodies: anti-Hibadh (at 1:1000 dilution) (Sigma-Aldrich, Cat# HPA021002), anti-P4hb (at 1:1000 dilution) (Sigma-Aldrich, Cat# HPA018884), anti-Hspa5 (at 1:1000 dilution) (Thermo Fisher Scientific, Cat# PA1-014A), and anti-Trap1 (at 1:2000 dilution) (Thermo Fisher Scientific, Cat# MA1-010) at 4 °C overnight and washed in TBS-T. Secondary antibody staining with A647-conjugated anti-rabbit (Jackson ImmunoResearch, Cat# 711–605-152) and A594-conjugated anti-mouse (Jackson ImmunoResearch, Cat# 715–585-151) was applied in 1:800 dilution for 2 h at RT and washed in TBS-T then TBS. The antibody-labeled membrane was scanned with Typhoon Trio + scanner (Amersham) with appropriate filter settings and 50-micron resolution. Images were analyzed with ImageJ (version 1.53c); densitometric values of protein bands were determined and normalized to the highest intensity band on the blot and then to the total protein content. A two-sample test for variance in OriginPro 9 was performed to test that the variances of the groups do not differ significantly (P < 0.05). Independent two-tailed Student’s t-test (P < 0.05) was applied to confirm altered levels of proteins in OriginPro 9.
Results
Magnetic Resonance Angiography (MRA) Confirmed the Efficacy of the Occlusions
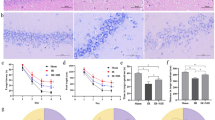
MRA confirmed that the experiment did not affect sham-operated animals, while the blood flow was bilaterally blocked in the common carotid artery in each CCH animal. As a result of the occlusions, the basilar artery thickened in the operated rats (Fig. 1a), implying it partially overtook the functions of the occluded arteries. Based on image analysis, the mean pixel density of blood vessels significantly (P-value = 0.0026) decreased in CCH animals 5 weeks after the surgery (Fig. 1b).
MRA recordings of animals 5 weeks after the operations. Representative images of sham-operated (left) and BCCAO (right) rats are shown (a). The common carotid arteries (CC) are occluded and the basilar artery (B) is thickened in the BCCAO animals. Image analysis of MRA recordings (b), statistical evaluation was performed with non-parametric, Mann–Whitney test (*: P-value < 0.01)
Mitochondrial and MAM Proteome Alterations Induced by CCH
The current gel-based proteomics study was performed on the MAM and mitochondria containing subcellular fractions of frontal cortices from stepwise BCCAO rats. We have separated 725 and 708 protein spots on the gels of MAM and mitochondria samples, respectively. PCA and hierarchical clustering showed that CCH and sham-operated animals could be distinguished based on significantly altered proteins in the MAM (Fig. 2a and b). However, one sham-operated animal prominently differed from the rest in the mitochondrial samples (Fig. 2c and d). Seventeen and 6 spots showed significant change in the MAM and mitochondria sample of the CCH rats compared to the sham-operated group, respectively (Fig. 2e and f; Online Resources 1 and 2). In the MAM samples, altered proteins showed fold changes between − 1.69 to 2.38. Three spots showed elevation while 14 decreased compared to the control group. Strikingly, most of the altered proteins in the mitochondria decreased with fold changes (FC) varying between − 1.31 and − 1.2, and one spot increased with the fold change of 1.27. From altered spots, HPLC–MS/MS identified 19 proteins from the mitochondria sample and 35 from MAM (Table 1). It is known that several proteins can be present in one spot and one protein can be found in several spots, thus the number of significantly altered spots is not equal to the number of identified proteins. Taken together, we observed a modest alteration in the levels of mitochondrial proteins and a more complex proteome modification in the MAM-containing sample.
PCA (a, b) and hierarchical clustering (c, d) of MAM and mitochondria, respectively, are shown. Volcano plots of MAM (e) and mitochondria (f) proteomic results. Significantly decreased and elevated spots are shown in blue and red, respectively. Pie charts of significantly altered proteins in MAM (g) and mitochondria (h) represent functional annotation with a legend (i)
Since the MAM comprises the interacting surfaces of the endoplasmic reticulum (ER) and mitochondrial membrane, the subcellular fraction of MAM will inevitably contain some mitochondrial proteins. Here, the spot with the highest fold change (FC = 2.38) consisted of NADH dehydrogenase (Ubiquinone) Fe-S protein 1 (Ndufs1) (the subunit of complex I of the electron transport chain), and heat shock proteins, namely stress-70 protein, mitochondrial (Hspa9 or mortalin), and heat shock cognate 71 kDa protein (Hspa8). While, in the most decreasing spot (FC = − 1.69), we detected protein disulfide isomerase A3 (Pdia3) chaperone among others with lower peptide count. The other two increased spots, with fold changes of 1.67, are comprised of methylcrotonoyl-CoA carboxylase subunit alpha (Mccc1) and vesicle-fusing ATPase (Nsf) which are involved in branch-chained amino acid catabolism and vesicle transport, respectively. All the other protein spots decreased; T-complex protein 1 subunit epsilon (Cct5), proteasome subunit alpha type-6 (Psma6), and neuronal-specific septin-3 (Septin3) had fold changes of − 1.62, − 1.55, and − 1.41, respectively. In the mitochondrial fraction, the spot decreasing and increasing the most (FC = − 1.31 and 1.27) comprised protein disulfide isomerase (P4hb or Pdia1) chaperon and cytoskeletal protein tropomodulin-2 (Tmod2), respectively. Other altered spots changed between − 1.20 and − 1.29; these contained endoplasmic reticulum chaperone BiP (Hspa5), ATP synthase subunit beta (Atp5f1b), heat shock protein 75 kDa (Trap1), and cytoskeletal protein dihydropyrimidinase-related protein 2 (Dpysl2).
The annotation of significantly altered proteins can reveal the disturbed cellular processes in the MAM and mitochondria induced by the stepwise BCCAO (Fig. 2g–i). Protein turnover and import showed the highest ratio of altered proteins in the MAM (n = 11; 32%) and mitochondria samples (n = 6; 32%). Energy and carbohydrate metabolism-associated proteins were highly represented in the MAM (n = 10; 29%) and less robustly in the mitochondria (n = 3; 16%) of the frontal cortex. Additionally, we identified cytoskeletal proteins in both mitochondria (n = 5; 26%) and MAM (n = 6; 18%) subcellular fractions. Proteins of fatty acid metabolism (n = 1; 3%) and ion transport (n = 1; 3%) were present only in MAM. Proteins involved in membrane dynamics and mitochondrial processes were also present among the altered proteins of MAM (n = 2; 6%) and mitochondria (n = 2; 10%). In conclusion, our data show a wide disturbance of the protein turnover process and mitochondrial energy metabolism in response to stepwise BCCAO.
Functional Proteomic Alterations in the CSF Reflect Cellular Organelle Changes in the Frontal Cortex
PCA and hierarchical clustering could distinguish CCH and sham-operated animals based on significantly altered proteins in the CSF (Fig. 3a and b). We found 12 proteins with significantly altered abundance with mean ratios between 0.32 and 2.06, using MS-based proteomics (Fig. 3c, Table 2, and Online Resource 1). Two of them, CD59 glycoprotein (Cd59) and WAP four-disulfide core domain protein 1 (Wfdc1), increased by 1.65 and 2.06 fold, respectively. Annexin A2 (Anxa2) showed the lowest mean ratio of 0.32. Functional annotations (Fig. 3b) showed that 17% of altered proteins were involved in redox state regulation, namely glutaredoxin-1 (Glrx) and glutathione S-transferase P (Gstp1) decreased by 0.53 and 0.52. Protein turnover and import had the highest ratio of altered proteins in the CSF (n = 5; 42%) similar to the MAM and mitochondria. Four of them were less than halved compared to control namely, proteasome subunit beta (Psmb4), ubiquitin carboxyl-terminal hydrolase isozyme L3 (Uchl3), eukaryotic translation initiation factor 3 subunit J (Eif3j), and importin subunit beta-1 (Kpnb1)); and Wfdc1, an endopeptidase inhibitor, increased.
PCA (a) and hierarchical clustering (b) of CSF could separate sham-operated and CCH animals. Volcano plots of CSF proteomic results (c) and pie chart (d) of functional annotation with a legend (e) are shown. Significantly decreased and elevated spots are shown in blue and red in the volcano plot, respectively
Enrichment Analysis of Altered Proteins Upon Stepwise BCCAO Is in Accordance with Functional Annotation
The enrichment of gene ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways can give further insight into the disturbed cellular process and related genes affected by stepwise BCCAO (Online Resource 3). Enrichment maps of the top five terms of GO biological process, molecular function, and KEGG pathways are shown in Fig. 4. Regarding biological process (Fig. 4a), ATP metabolic process was enriched with five related genes Hspa8, Atp5f1b, v-type proton ATPase subunit B (Atp6v1b2), Ndufs1, adenylate kinase 5 (Ak5) (32.2 fold, adj. P-value = 4.2E-03). Also, protein folding was significantly overrepresented (20.8 fold, adj. P-value = 5.44E-05). In accordance, protein binding involved in protein folding molecular function term was highly enriched (32.2 fold, adj. P-value = 9.02E-04) with five related genes (Hspa9, Hspa8, Cct2, Hspa5, Cct5) (Fig. 4b). Among KEGG pathways (Fig. 4c), the most enriched terms were glycolysis/gluconeogenesis (14.3 fold, adj. P-value = 4.83E-03) and biosynthesis of amino acids (12.5 fold, adj. P-value = 7.25E-03). Also, pathways of neurodegeneration in multiple diseases were enriched (4.5 fold, adj. P-value = 2.24E-03) with 11 related genes (such as Trap1, Psma6, Psmb4, Atp5f1b, Hspa5, Ndufs1a, and tubulins) (Online Resource 3).
Enrichment map of top five terms of GO biological process (a), GO molecular function (b), and KEGG pathway (c) is connected to their related proteins labeled with gene names. –Log(adjusted P-value) of terms and fold changes of related proteins are color coded. Fold enrichments of terms are represented with node size. Altered proteins of MAM, mitochondria, and CSF are depicted with circle, hexagon, and rhombus, respectively
Altered Proteins of Organellar Quality Control Validated by Western Blot
We analyzed the levels of four proteins involved in mitochondrial and ER quality control and protein folding by western blot to confirm our proteomic results (Fig. 5a–e). We performed a two-sample test for a variance to ensure equal variances of the two experimental groups, and we did not find significant changes in variances (for Hibadh, P = 0.74; for P4hb, P = 0.16; for Hspa5, P = 0.41; and for total protein, P = 0.57). P4hb (P-value = 0.026) and Hibadh (P-value = 0.031) levels decreased significantly in the mitochondria (Fig. 5a–b). While the decline of the Hspa5 level in the mitochondria was not significant (P-value = 0.11) (Fig. 5c). We did not find significant changes in total protein content in the blot (P-value = 0.93) (Fig. 5d).
Discussion
Limitations
High-throughput methods are powerful tools to track molecular changes upon different treatments in animals, tissues, or cell lines. Two-D DIGE proteomics enables the separation of two or three protein samples labeled by different fluorescent dyes simultaneously on a single gel. This technique makes spot comparison and protein quantification more reliable and reproducible [28].
At the same time, 2-D DIGE has some limitations that must be taken into account. Hydrophobic membrane proteins solubilize poorly in polar detergent-free solvents applied in isoelectric focusing (IEF). This effect aggravates their migration into the gel; thus, their detection is limited. Additionally, we could not separate and detect the higher molecular weight proteins and the strongly acidic or alkaline ones since applied IEF and PAGE allows protein separation in the range of ~ 3 to ~ 10 isoelectric points (pI) and 10–100 kDa, respectively. In addition, highly abundant proteins can mask lower-abundant protein alterations if more than one protein runs into the same spot. This can be improved by subcellular fractionation, such as mitochondria and MAM isolation in this study. However, contaminations from other cellular organelles are inevitable and not negligible [29]. At the same time, the applied preparations are highly reproducible and validated [25].
Chaperons of the ER Protein Quality Control
MAM is a specialized area of the ER, communicating directly with mitochondria. ER is essential in protein synthesis, folding, and trafficking, and several ER chaperones localize to the MAM [30]. Upon acute energy restriction, misfolded and unfolded proteins accumulate in the ER lumen [31]. In response to ER stress, the unfolded protein response (UPR) is triggered to restore proteostasis by reducing protein synthesis, promoting proper protein folding, and the degradation of misfolded proteins. ER-associated degradation (ERAD) induces misfolded protein clearance by proteasomal degradation. Thus, ERAD and UPR serve to restore normal proteostasis upon stress.
A widely used marker of ER stress is the Hspa5 chaperone; we detected its decrease in the mitochondria. Although the Hspa5 level showed a declining trend analyzed by western blot, its decrease was not significant. Overexpressing Hspa5 in primary cultured astrocytes protected mitochondria against ischemic stress [32]. Additional ER chaperones implicated in UPR, Pdia3, and P4hb, also changed in our study, and the latter was validated by western blot. Pdia3 reduced ER stress upon ischemia [33], and its mRNA level was upregulated upon glucose depletion [34]. P4hb is upregulated in short-term hypoxic astrocytes and protects neurons from apoptotic cell death in vitro [35]. Chaperone-containing T-complex (TRiC) protein subunits are chaperones that compose the CCT-β complex upon interacting with β-tubulin. Disruption of the CCT-β complex reduced ER stress response through altering mitochondrial membrane potential and inducing heat shock protein 75 kDa (Trap1) dependent protein degradation and ERAD [36]. In our study, we detected decreased levels of T-complex protein subunits 2, 5 (Cct2, Cct5), several β-tubulin subunits (Tubb2a, Tubb4a), and Trap1. It was found that a component of ERAD localizes proteasome subunits to the ER [37], and we detected decreased proteasome subunit alpha type-6 (Psma6) levels in the MAM and proteasome subunit beta (Psmb4) in CSF.
In a CCH model, EPTA staining of hippocampal slices revealed abnormal protein aggregation of newly synthesized proteins in the cytosol, 3 months after the operation [38]. In addition, elevated levels of ubiquitinated proteins were shown in the chronic BCCAO model [39], implying that disturbed proteostasis is a long-term effect of CCH. In conclusion, most of the proteins of the quality control mechanism decreased in our study, implying that ER stress response is hindered in response to chronic hypoperfusion, and its protective role in maintaining normal proteostasis might be diminished.
We detected dihydropyrimidinase-related proteins 2 and 3 (Dpysl2 and 3) which were also identified in the MAM fraction previously [29, 40]. However, their role in the regulation of MAM is poorly studied.
Altered Proteins of Mitochondrial Oxidative Phosphorylation and Quality Control
The occlusion of the common carotid arteries reduces cerebral blood flow for several weeks in BCCAO rats. Although the blood flow recovers to the control level [3], subsequently, mitochondrial damage develops [41]. Additionally, BCCAO augmented the level of reactive oxygen species and reduced the activity of Mn-superoxide dismutase (Sod2), even 8 weeks after the surgery [5]. Two components of the oxidative phosphorylation (OXPHOS) system were altered in our study; one of them is the ATP synthase subunit beta (Atp5f1b), and the other is Ndufs1. Altered levels of subunits of the electron transport chain can hinder the correct assembly of the OXPHOS system which can lead to oxidative [42] and possibly proteotoxic stress [43].
Mitochondrial quality control is crucial in maintaining normal cellular metabolism. Several proteins and molecular pathways serve to maintain or restore mitochondrial functions such as chaperones, proteases, and import proteins involved in mitochondrial UPR (mtUPR). In the mitochondria, misfolded or damaged proteins are digested, and the peptides are transported to the cytosol where they induce mtUPR [44]. In our proteomic study, we found 60 kDa heat shock protein (Hspd1 also known as Hsp60) and Hspa9 that are involved in mtUPR. Hspa9 is a matrix chaperone and the only component of the mitochondrial import complex that has an ATPase function [45]. Since most of the mitochondrial proteins are encoded in the nucleus, the import and refolding of proteins are essential to maintain mitochondrial proteostasis. Loss of Hspa9 induced proteotoxic stress and autophagic clearance of damaged mitochondria in vitro [46].
Trap1 is a mitochondrial chaperone, involved in oxidative stress response. It reduces mitochondrial respiration, promotes glycolysis, and reduces reactive oxygen species [47]. We have detected decreased level of Trap1 in mitochondria in response to long-term hypoperfusion by proteomics analyses.
Cathepsin D (Ctsd) is a lysosomal endopeptidase and its release from lysosomes triggers bax activation [48], mitochondrial cytochrome C release, and transmembrane potential loss during apoptosis upon oxidative stress [49]. We detected Ctsd elevation in the mitochondrial fraction.
Surprisingly, several cytoskeletal proteins were detected in the mitochondrial samples. However, α- and β-tubulin were shown to associate with mitochondrial membranes [50]. Furthermore, tubulins decreased mitochondrial respiration by reducing the permeability of voltage-dependent anion channel for ADP [51, 52]. Reducing mitochondrial respiration protects against oxidative stress; in our study, levels of several tubulin subunits dropped in mitochondria and MAM samples. This might indicate increasing oxidative damage and increased level of ROS found in the BCCAO model [5].
Metabolism of Branch-Chained Amino Acids (BCAAs) in Dementia
BCAAs (Leu, Ile, Val) are essential amino acids necessary for protein synthesis and also have roles in the regulation of metabolic functions. BCAAs serve as nitrogen donors for amino acids (e.g., glutamine) and neurotransmitter synthesis (e.g., glutamate, gamma-aminobutyric acid).
Decreased levels of BCAAs were detected in the serum levels of patients with Alzheimer’s disease and dementia [53]. Furthermore, decreased level of leucine (Leu) was also detected in the saliva of Alzheimer’s disease and vascular dementia patients [54]. We have detected an increased level of methylcrotonoyl-CoA carboxylase subunit alpha (Mccc1) which is responsible for the catabolism of Leu. However, we also measured a decreased level of 3-hydroxyisobutyrate dehydrogenase (Hibadh), an enzyme of valine catabolism, and its decrease was validated by western blot. Thus, our results suggest unbalanced BCAA catabolism in CCH.
Disturbed Protein Turnover and Redox State Regulation are Reflected in the CSF
While we could not detect significant changes of the same proteins in the CSF and the subcellular fractions, several proteins of protein turnover decreased in the CSF and subcellular fraction proteomes upon stepwise BCCAO, such as Eif3j, Uchl3, Psmb4 in the CSF, Psma6 in the MAM, and glycine–tRNA ligase (Gars) in the mitochondria.
Glrx and Gstp1 have main roles in the detoxification of reactive oxygen species which levels’ are elevated upon BCCAO [5]. Although we did not detect significant alterations of Glrx and Gstp1 in the subcellular fraction of the frontal cortex, both proteins decreased in the CSF in our study, and they were shown to localize to mitochondria [55, 56]. Furthermore, we previously showed altered Gstp1 level in the synaptosomal fraction of the occipital cortex in CCH rats compared to sham-operated controls [23].
There were several proteins that we could only detect in sham-operated animals (see Online Resource 1). For instance, proteasome subunit beta type-7, exportin-1, and electron transfer flavoprotein subunit beta have roles in protein degradation, protein transport from the nucleus, and mitochondrial electron transport, respectively. Lack of their detection can indicate that their levels are below the detection limit or missing from the samples of BCCAO animals. Thus, it further suggests that protein turnover impairment and mitochondrial damage of the brain can be detected in the CSF.
Conclusions
Here, we studied the long-term effects of oxidative stress in the brain by inducing CCH and investigating the proteomic alterations in two subcellular fractions of the frontal cortex and the CSF. Our results suggest declining cellular response against oxidative and proteotoxic stress since several key proteins of these processes showed decreasing levels, such as Glrx, Gstp1, T-complex protein subunits, Hspd1, and P4hb. Stress responses have a protective effect on cells; therefore, we could speculate that the long-term decline of stress responses can be one of the causes of cognitive impairment during chronic cerebral hypoperfusion. We can also suggest impaired protein turnover by detecting reduced levels of a proteasome subunit and a eukaryotic translation initiation factor in the CSF and amino acid-tRNA ligases in the MAM and mitochondria. Reduced protein turnover might be a compensatory mechanism to reduce energy consumption and oxidative damage in response to impaired blood flow. In conclusion, we have detected reduced levels of several components of protein turnover in the mitochondria, MAM, and CSF of the stepwise BCCAO model, implying that altered cellular processes of brain tissue can be detected in the cerebrospinal fluid by proteomic analysis.
Data Availability
The datasets supporting the conclusions of this article are available upon request.
Abbreviations
- CCH :
-
Chronic cerebral hypoperfusion
- CBF :
-
Cerebral blood flow
- VD :
-
Vascular dementia
- AD :
-
Alzheimer’s disease
- BCCAO :
-
Bilateral common carotid artery occlusion
- MAM :
-
Mitochondrial-associated membrane
- 2-D DIGE :
-
2-Dimensional differential gel electrophoresis
- HPLC–MS/MS :
-
High performance liquid chromatography-tandem mass spectroscopy
- MRA :
-
Magnetic resonance angiography
- IB-1 :
-
Isolation buffer-1
- IB-2 :
-
Isolation buffer-2
- IB-3 :
-
Isolation buffer-3
- MRB :
-
Mitochondria resuspending buffer
- IEF :
-
Isoelectric focusing
- FC :
-
Fold change
References
Feng T, Yamashita T, Zhai Y, Shang J, Nakano Y, Morihara R et al (2018) Chronic cerebral hypoperfusion accelerates Alzheimer’s disease pathology with the change of mitochondrial fission and fusion proteins expression in a novel mouse model. Brain Res 1696:63–70. https://doi.org/10.1016/j.brainres.2018.06.003
Montaldi D, Brooks DN, McColl JH, Wyper D, Patterson J, Barron E et al (1990) Measurements of regional cerebral blood flow and cognitive performance in Alzheimer’s disease. J Neurol Neurosurg Psychiatry 53:33–38. https://doi.org/10.1136/jnnp.53.1.33
Farkas E, Luiten PGM, Bari F (2007) Permanent bilateral common carotid artery occlusion in the rat: a model for chronic cerebral hypoperfusion-related neurodegenerative diseases. Brain Res Rev 54:162–180. https://doi.org/10.1016/j.brainresrev.2007.01.003
Otori T, Katsumata T, Muramatsu H, Kashiwagi F, Katayama Y, Terashi A (2003) Long-term measurement of cerebral blood flow and metabolism in a rat chronic hypoperfusion model. Clin Exp Pharmacol Physiol 30:266–272. https://doi.org/10.1046/j.1440-1681.2003.03825.x
Yadav A, Sunkaria A, Sinhal N, Sandhir R (2018) Resveratrol loaded solid lipid nanoparticles attenuate mitochondrial oxidative stress in vascular dementia by activating Nrf2/HO-1 pathway. Neurochem Int 112:239–254. https://doi.org/10.1016/j.neuint.2017.08.001
Völgyi K, Háden K, Kis V, Gulyássy P, Badics K, Györffy BA et al (2017) Mitochondrial proteome changes correlating with β-amyloid accumulation. Mol Neurobiol 54:2060–2078. https://doi.org/10.1007/s12035-015-9682-4
Völgyi K, Badics K, Sialana FJ, Gulyássy P, Udvari EB, Kis V et al (2018) Early presymptomatic changes in the proteome of mitochondria-associated membrane in the APP/PS1 mouse model of Alzheimer’s disease. Mol Neurobiol 55:7839–7857. https://doi.org/10.1007/s12035-018-0955-6
Du J, Ma M, Zhao Q, Fang L, Chang J, Wang Y et al (2013) Mitochondrial bioenergetic deficits in the hippocampi of rats with chronic ischemia-induced vascular dementia. Neuroscience 231:345–352. https://doi.org/10.1016/j.neuroscience.2012.11.062
Li H, Liu Y, Lin LT, Wang XR, Du SQ, Yan CQ et al (2016) Acupuncture reversed hippocampal mitochondrial dysfunction in vascular dementia rats. Neurochem Int 92:35–42. https://doi.org/10.1016/j.neuint.2015.12.001
Islam MT (2017) Oxidative stress and mitochondrial dysfunction-linked neurodegenerative disorders. Neurol Res 39:73–82. https://doi.org/10.1080/01616412.2016.1251711
Burté F, Carelli V, Chinnery PF, Yu-Wai-Man P (2015) Disturbed mitochondrial dynamics and neurodegenerative disorders. Nat Rev Neurol 11:11–24. https://doi.org/10.1038/nrneurol.2014.228
Bhat AH, Dar KB, Anees S, Zargar MA, Masood A, Sofi MA et al (2015) Oxidative stress, mitochondrial dysfunction and neurodegenerative diseases; a mechanistic insight. Biomed Pharmacother 74:101–110. https://doi.org/10.1016/j.biopha.2015.07.025
Mancuso C, Scapagini G, Currò D, Giuffrida Stella AM, De Marco C, Butterfield DA et al (2007) Mitochondrial dysfunction, free radical generation and cellular stress response in neurodegenerative disorders. Front Biosci 12:1107–1123. https://doi.org/10.2741/2130
He J, Huang Y, Du G, Wang Z, Xiang Y, Wang Q (2019) Lasting spatial learning and memory deficits following chronic cerebral hypoperfusion are associated with hippocampal mitochondrial aging in rats. Neuroscience 415:215–229. https://doi.org/10.1016/j.neuroscience.2019.04.044
Calkins MJ, Manczak M, Mao P, Shirendeb U, Reddy PH (2011) Impaired mitochondrial biogenesis, defective axonal transport of mitochondria, abnormal mitochondrial dynamics and synaptic degeneration in a mouse model of Alzheimer’s disease. Hum Mol Genet 20:4515–4529. https://doi.org/10.1093/hmg/ddr381
Area-Gomez E, Scon EA (2017) On the pathogenesis of Alzheimer’s disease: the MAM hypothesis. FASEB J 31:864–867. https://doi.org/10.1096/fj.201601309
Krols M, van Isterdael G, Asselbergh B, Kremer A, Lippens S, Timmerman V et al (2016) Mitochondria-associated membranes as hubs for neurodegeneration. Acta Neuropathol 131:505–552. https://doi.org/10.1007/s00401-015-1528-7
Area-Gomez E, Castillo MDCL, Tambini MD, Guardis-Laguarta C, de Groof AJC, Madra M et al (2012) Upregulated function of mitochondria-associated ER membranes in Alzheimer disease. EMBO J 31:4106–4123. https://doi.org/10.1038/emboj.2012.202
Iliff JJ, Wang M, Liao Y, Plogg BA, Peng W, Gundersen GA, Benveniste H, Vates GE et al (2012) A paravascular pathway facilitates CSF flow through the brain parenchyma and the clearance of interstitial solutes, including amyloid β. Sci Transl Med 4:147ra111. https://doi.org/10.1126/scitranslmed.3003748
Cardoso S, Carvalho C, Correia SC, Seica RM, Moreira PI (2016) Alzheimer’s disease: from mitochondrial perturbations to mitochondrial medicine. Brain Pathol 26:632–647. https://doi.org/10.1111/bpa.12402
Tatemichi TK, Desmond DW, Prohovnik I (1995) Strategic infarcts in vascular dementia. A clinical and brain imaging experience. Arzneimittelforschung 45:371–385
Thomas T, Miners S, Love S (2015) Post-mortem assessment of hypoperfusion of cerebral cortex in Alzheimer’s disease and vascular dementia. Brain 138:1059–1069. https://doi.org/10.1093/brain/awv025
Tukacs V, Mittli D, Györffy BA, Hunyady-Gulyás É, Hlatky D, Tóth V et al (2020) Chronic stepwise cerebral hypoperfusion differentially induces synaptic proteome changes in the frontal cortex, occipital cortex, and hippocampus in rats. Sci Rep 10(1):15999. https://doi.org/10.1038/s41598-020-72868-w
Nyitrai G, Spisák T, Spisák Z, Gajári D, Diószegi P, Kincses TZ et al (2018) Stepwise occlusion of the carotid arteries of the rat: MRI assessment of the effect of donepezil and hypoperfusion-induced brain atrophy and white matter microstructural changes. PLoS One 13(5):e0198265. https://doi.org/10.1371/journal.pone.0198265
Wieckowski MR, Giorgi C, Lebiedzinska M, Duszynski J, Pinton P (2009) Isolation of mitochondria-associated membranes and mitochondria from animal tissues and cells. Nat Protoc 4:1582–1590. https://doi.org/10.1038/nprot.2009.151
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4(1):44–57. https://doi.org/10.1038/nprot.2008.211
Sherman BT, Hao M, Qiu J, Jiao X, Baseler MW, Lane HC, Imamichi T, Chang W (2022) DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res 23:gkac194. https://doi.org/10.1093/nar/gkac194
Marouga R, David S, Hawkins E (2005) The development of the DIGE system: 2D fluorescence difference gel analysis technology. Anal Bioanal Chem 382:669–678. https://doi.org/10.1007/s00216-005-3126-3
Poston CN, Krishnan SC, Bazemore-Walker CR (2013) In-depth proteomic analysis of mammalian mitochondria-associated membranes (MAM). J Proteomics 79:219–230. https://doi.org/10.1016/j.jprot.2012.12.018
Simmen T, Lynes EM, Gesson K, Thomas G (2010) Oxidative protein folding in the endoplasmic reticulum: tight links to the mitochondria-associated membrane (MAM). Biochim Biophys Acta 1798:1465–1473. https://doi.org/10.1016/j.bbamem.2010.04.009
Wang M, Kaufman RJ (2016) Protein misfolding in the endoplasmic reticulum as a conduit to human disease. Nature 529:326–335. https://doi.org/10.1038/nature17041
Ouyang B, Xu LJ, Emery JF, Lee AS, Giffard RG (2011) Overexpressing GRP78 influences Ca2+ handling and function of mitochondria in astrocytes after ischemia-like stress. Mitochondrion 11:279–286. https://doi.org/10.1016/j.mito.2010.10.007
Yoo DY, Cho SB, Jung HY, Kim W, Lee KY, Kim JW et al (2019) Protein disulfide-isomerase A3 significantly reduces ischemia-induced damage by reducing oxidative and endoplasmic reticulum stress. Neurochem Int 122:19–30. https://doi.org/10.1016/j.neuint.2018.11.002
Zhou Y, Lee J, Reno CM, Sun C, Park SW, Chung J et al (2011) Regulation of glucose homeostasis through a XBP-1-FoxO1 interaction. Nat Med 17:356–365. https://doi.org/10.1038/nm.2293
Tanaka S, Uehara T, Nomura Y (2000) Up-regulation of protein-disulfide isomerase in response to hypoxia/brain ischemia and its protective effect against apoptotic cell death. J Biol Chem 275:10388–10393. https://doi.org/10.1074/jbc.275.14.10388
Lin YF, Lee YF, Liang PH (2012) Targeting β-tubulin: CCT-β complexes incurs Hsp90- and VCP-related protein degradation and induces ER stress-associated apoptosis by triggering capacitative Ca2+ entry, mitochondrial perturbation and caspase overactivation. Cell Death Dis 3:e434. https://doi.org/10.1038/cddis.2012.173
Tcherpakov M, Broday L, Delaunay A, Kadoya T, Khurana A, Erdjument-Bromage H et al (2008) JAMP optimizes ERAD to protect cells from unfolded proteins. Mol Biol Cell 19:5019–5028. https://doi.org/10.1091/mbc.e08-08-0839
Hai J, Qi L, Wu Y-F, Huang X-S, Zhang G-Y, Wang F (2013) Effects of N-stearoyl-L-tyrosine on the hippocampal ubiquitin-proteasome system in rats with chronic cerebral hypoperfusion. Neurol Res 35:734–743. https://doi.org/10.1179/1743132812Y.0000000154
Hai J, Lin Q, Su SH, Zhang L, Wan JF, Lu Y (2011) Chronic cerebral hypoperfusion in rats causes proteasome dysfunction and aggregation of ubiquitinated proteins. Brain Res 1374:73–81. https://doi.org/10.1016/j.brainres.2010.12.030
Földi I, Tóth AM, Szabó Z, Mózes E, Berkecz R, Datki ZL et al (2013) Proteome-wide study of endoplasmic reticulum stress induced by thapsigarginin N2a neuroblastoma cells. Neurochem Int 62:58–69. https://doi.org/10.1016/j.neuint.2012.11.003
He X-L, Wang Y-H, Gao M, Li X-X, Zhang T-T, Du G-H (2008) Baicalein protects rat brain mitochondria against chronic cerebral hypoperfusion-induced oxidative damage. Brain res 1249:212–221. https://doi.org/10.1016/j.brainres.2008.10.005
Miwa S, Jow H, Baty K, Johnson A, Czapiewski R, Saretzki G et al (2014) Low abundance of the matrix arm of complex I in mitochondria predicts longevity in mice. Nat commun 5:3837. https://doi.org/10.1038/ncomms4837
Callegari S, Dennerlein S (2018) Sensing the stress: a role for the UPRmt and UPRam in the quality control of mitochondria. Front Cell Dev Biol 6:1–10. https://doi.org/10.3389/fcell.2018.00031
Haynes CM, Fiorese CJ, Lin Y-F (2013) Evaluating and responding to mitochondrial dysfunction: the mitochondrial unfolded-protein response and beyond. Trends in Cell Biol 23:311–318. https://doi.org/10.1016/j.tcb.2013.02.002
Chacinska A, Koehler CM, Milenkovic D, Lithgow T, Pfanner N (2009) Importing mitochondrial proteins: machineries and mechanisms. Cell 138:628–644. https://doi.org/10.1016/j.cell.2009.08.005
Burbulla LF, Fitzgerald JC, Stegen K, Westermeier J, Thost A-K, Kato H et al (2014) Mitochondrial proteolytic stress induced by loss of mortalin function is rescued by Parkin and PINK1. Cell Death and Dis 5:1180. https://doi.org/10.1038/cddis.2014.103
Yoshida S, Tsutsumi S, Muhlebach G, Sourbier C, Lee M-J, Lee S et al (2013) Molecular chaperone TRAP1 regulates a metabolic switch between mitochondrial respiration and aerobic glycolysis. Proc Natl Acad Sci U S A 110:E1604–E1612. https://doi.org/10.1073/pnas.1220659110
Bidère N, Lorenzo HK, Carmona S, Laforge M, Harper F, Dumont C, Senik A (2003) Cathepsin D triggers Bax activation, resulting in selective apoptosis-inducing factor (AIF) relocation in T lymphocytes entering the early commitment phase to apoptosis. J Biol Chem 278(33):31401–31411. https://doi.org/10.1074/jbc.M301911200
Roberg K, Johansson U, Ollinger K (1999) Lysosomal release of cathepsin D precedes relocation of cytochrome c and loss of mitochondrial transmembrane potential during apoptosis induced by oxidative stress. Free Radic Biol Med 15(11–12):1228–1237. https://doi.org/10.1016/s0891-5849(99)00146-x
Carré M, André N, Carles G, Borghi H, Brichese L, Briand C et al (2002) Tubulin is an inherent component of mitochondrial membranes that interacts with the voltage-dependent anion channel. J Biol Chem 277:33664–33669. https://doi.org/10.1074/jbc.M203834200
Rostovtseva TK, Sheldon KL, Hassanzadeh E, Monge C, Saks V, Bezrukov SM et al (2008) Tubulin binding blocks mitochondrial voltage-dependent anion channel and regulates respiration. Proc Natl Acad Sci U S A 105:18746–18751. https://doi.org/10.1073/pnas.0806303105
Monge C, Beraud N, Kuznetsov AV, Rostovtseva T, Sackett D, Schlattner U et al (2008) Regulation of respiration in brain mitochondria and synaptosomes: restrictions of ADP diffusion in situ, roles of tubulin, and mitochondrial creatine kinase. Mol Cell Biochem 318:147–165. https://doi.org/10.1007/s11010-008-9865-7
Tynkkynen J, Chouraki V, van der Lee SJ, Hernesniemi J, Yang Q, Li S et al (2018) Association of branched-chain amino acids and other circulating metabolites with risk of incident dementia and Alzheimer’s disease: a prospective study in eight cohorts. Alzheimers Dement 14:723–733. https://doi.org/10.1016/j.jalz.2018.01.003
Figueira J, Jonsson P, Nordin Adolfsson A, Adolfsson R, Nyberg L, Öhman A (2016) NMR analysis of the human saliva metabolome distinguishes dementia patients from matched controls. Mol Biosyst 12:2562–2571. https://doi.org/10.1039/c6mb00233a
Goto S, Kawakatsu M, Izumi S, Urata Y, Kageyama K, Ihara Y, Koji T, Kondo T (2009) Glutathione S-transferase pi localizes in mitochondria and protects against oxidative stress. Free Radic Biol Med 46(10):1392–1403. https://doi.org/10.1016/j.freeradbiomed.2009.02.025
Pai HV, Starke DW, Lesnefsky EJ, Hoppel CL, Mieyal JJ (2007) What is the functional significance of the unique location of glutaredoxin 1 (GRx1) in the intermembrane space of mitochondria? Antioxid Redox Signal 9(11):2027–2033. https://doi.org/10.1089/ars.2007.1642
Acknowledgements
We are grateful to Agnes Arva for her technical assistance, and we thank for the usage of MTA Cloud (https://cloud.mta.hu/) that significantly helped us achieve the results published in this paper.
Funding
Open access funding provided by Eötvös Loránd University. Supported by the ÚNKP-21–3 New National Excellence Program of the Ministry for Innovation and Technology from the source of the National Research, Development and Innovation Fund (to VT). This study was supported by the National Research, Development, and Innovation Office of Hungary Grants 2017–1.2.1-NKP-2017–00002, FIEK_16-1–2016-0005, and National Brain Research Program NAP 3.0 of the Hungarian Academy of Sciences (NAP2022-I-3/2022) (to VT, DM, JG, JK, and KAK). This work was supported by the 2019–1.2.1-Egyetemi-ÖKO-2019–0004 ELTE Proof of Concept (POC) grant (VT, DM, KAK). Mass spectrometry analyses were supported by the following grants: the Economic Development and Innovation Operative Programs GINOP-2.3.2–15-2016–00001 and GINOP-2.3.2–15-2016–00020 (ÉHG, ZD). HCEMM has received funding from the EU’s Horizon 2020 research and innovation program under grant agreement No. 739593 (ÉHG, ZD).
Author information
Authors and Affiliations
Contributions
VT performed most of the experiments and wrote the manuscript. DH and GN performed magnetic resonance imaging. ÉHG, KMF, and ZD carried out mass spectrometry protein identification experiments. DM took part in the analysis of proteomic data. GJ, JK, AC, and KAK conceived, designed, supervised the studies, and wrote the manuscript.
Corresponding author
Ethics declarations
Ethics Approval
All animal care and experimental procedures were following the Council Directive 86/609/EEC, the Hungarian Act of Animal Care and Experimentation (1998, XXVIII). All the procedures conformed to the National Institutes of Health guidelines were in accordance with the guidelines of the local Animal Care and Use Committee and were approved by the local Ethical Committee of Gedeon Richter Plc. (PE/EA/2885–6/2016). All efforts were carried out to minimize the animals’ pain and suffering and to reduce the number of animals used.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Online resource 1.
The list of significantly altered proteins in the CSF and identified proteins of spots in the mitochondria and MAM (XLSX 80 kb)

Online resource 2.
Representative gel images of the proteomics analysis with significantly altered protein spots in the (a) MAM and (b) mitochondria (red and blue spots indicate increasing and decreasing results, respectively) (PNG 3036 kb)
Online resource 3.
Result of enrichment analysis using DAVID Bioinformatics Resources v2022q1 (EASE score threshold: 0.01, Count threshold: 4, Background: rat genome) (XLSX 20 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tukacs, V., Mittli, D., Hunyadi-Gulyás, É. et al. Chronic Cerebral Hypoperfusion-Induced Disturbed Proteostasis of Mitochondria and MAM Is Reflected in the CSF of Rats by Proteomic Analysis. Mol Neurobiol 60, 3158–3174 (2023). https://doi.org/10.1007/s12035-023-03215-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-023-03215-z