Abstract
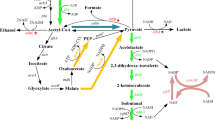
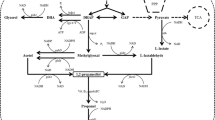
Biofuel offers a promising solution to the adverse environmental problems and depletion in reserves of fossil fuels. Higher alcohols including 3-methyl-1-butanol were paid much more attention as fuel substitute in recent years, due to its similar properties to gasoline. In the present work, 3-methyl-1-butanol production in engineered Corynebacterium glutamicum was studied. α-Ketoisovalerate decarboxylase gene (kivd) from Lactococcus lactis combined with alcohol dehydrogenase gene (adh2, adhA, and adh3) from three organisms were overexpressed in C. glutamicum. Enzymatic assay and alcohol production results showed that adh3 from Zymomonas mobilis was the optimum candidate for 3-methyl-1-butanol production in C. glutamicum. The recombinant with kivd and adh3 could produce 0.182 g/L of 3-methyl-1-butanol and 0.144 g/L of isobutanol after 12 h of incubation. Further inactivation of the E1 subunit of pyruvate dehydrogenase complex gene (aceE) and lactic dehydrogenase gene (ldh) in the above C. glutamicum strain would improve the 3-Methyl-1-butanol titer to 0.497 g/L after 12 h of incubation.





Similar content being viewed by others
References
Atsumi, S., Cann, A. F., Connor, M. R., Shen, C. R., Smith, K. M., Brynildsen, M. P., et al. (2008). Metabolic engineering of Escherichia coli for 1-butanol production. Metabolic Engineering, 10, 305–311.
Atsumi, S., Hanai, T., & Liao, J. C. (2008). Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature, 451, 86–89.
Atsumi, S., & Liao, J. C. (2008). Directed evolution of Methanococcus jannaschii citramalate synthase for biosynthesis of 1-propanol and 1-butanol by Escherichia coli. Applied and Environment Microbiology, 74, 7802–7808.
Atsumi, S., Wu, T. Y., Eckl, E. M., Hawkins, S. D., Buelter, T., & Liao, J. C. (2010). Engineering the isobutanol biosynthetic pathway in Escherichia coli by comparison of three aldehyde reductase/alcohol dehydrogenase genes. Applied Microbiology and Biotechnology, 85, 651–657.
Blombach, B., Arndt, M., & Auchter, B. (2009). Eikmanns l-Valine production during growth of pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum in the presence of ethanol or by inactivation of the transcriptional regulator SugR. Applied and Environment Microbiology, 75, 1197–1200.
Blombach, B., Riester, T., Wieschalka, S., Ziert, C., Youn, J. W., Wendisch, V. F., & Eikmanns, B. J. (2010). Corynebacterium glutamicum tailored for efficient isobutanol production. Applied and Environment Microbiology, 77, 3300–3310.
Blombach B, Schreiner ME, Holátko J, Bartek T, Oldiges M, Eikmanns BJ (2007) l-Valine production with pyruvate dehydrogenase complex-deficient Corynebacterium glutamicum. Appl Environ Microbiol 73:2079–2084
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochemn, 72, 248–254.
Candy, J. M., & Duggleby, R. G. (1998). Structure and properties of pyruvate decarboxylase and site-directed mutagenesis of the Zymomonas mobilis enzyme. Biochimica et Biophysica Acta, 1385, 323–338.
Chen, X. Y., Xu, J. L., Yang, L., Yuan, Z. H., Xiao, S. Y., Zhang, Y., et al. (2015). Production of C4 and C5 branched-chain alcohols by engineered Escherichia. Coli J Ind Microbiol Biotechnol, 42, 1473–1479.
Connor, M. R., Cann, A. F., & Liao, J. C. (2010). 3-Methyl-1-butanol production in Escherichia coli: random mutagenesis and two-phase fermentation. Applied Microbiology and Biotechnology, 86, 1155–1164.
Connor, M. R., & Liao, J. C. (2008). Engineering of an Escherichia coli strain for the production of 3-methyl-1-butanol. Applied and Environment Microbiology, 74(18), 5769–5775.
De la Plaza, M., Fernández de Palencia, P., Peláez, C., & Requena, T. (2004). Biochemical and molecular characterization of a-ketoisovalerate decarboxylase, an enzyme involved in the formation of aldehydes from amino acids by Lactococcus lactis. FEMS Microbiology Letters, 238, 367–374.
Dürre P (2007) Biobutanol: an attractive biofuel. Biotechnol J 2(12):1525–1534
Iding, H., Siegert, P., & Mesch, K. (1998). Application of a-keto acid decarboxylases in biotransformations. Biochimica et Biophysica Acta, 1385, 307–322.
Ingram, L. O., Aldrich, H. C., Borges, A. C., Causey, T. B., Martinez, A., Morales, F., et al. (1999). Enteric bacterial catalysts for fuel ethanol production. Biotechnology Progress, 15, 855–866.
Konig, S. (1998). Subunit structure, function and organisation of pyruvate decarboxylases from various organisms. Biochimica et Biophysica Acta, 1385, 271–286.
Kirchner, O., & Tauch, A. (2003). Tools for genetic engineering in the amino acid-producing bacterium Corynebacterium glutamicum. Journal of Biotechnology, 104, 287–299.
Mutalik, V. K., Guimaraes, J. C., Cambray, G., Lam, C., Christoffersen, M. J., Mai, Q. A., et al. (2013). Precise and reliable gene expression via standard transcription and translation initiation elements. Nature Methods, 10(4), 354–360.
Lee, W. H., Seo, S. O., Bae, Y. H., Nan, H., Jin, Y. S., & Seo, J. H. (2012). Isobutanol production in engineered Saccharomyces cerevisiae by overexpression of 2-ketoisovalerate decarboxylase and valine biosynthetic enzymes. Bioprocess and Biosystems Engineering, 35, 1467–1475.
Leuchtenberger, W., Huthmacher, K., & Drauz, K. (2005). Biotechnological production of amino acids and derivates: current status and prospects. Applied Microbiology and Biotechnology, 69, 1–8.
Mary, V. M., Maurille, J. F., & David, A. T. (2002). Depletion of free 30S ribosomal subunits in Escherichia coli by expression of RNA containing Shine-Dalgarno-like sequences. Journal of Bacteriology, 184, 494–502.
Neil, S. (2011). The ideal biofuel. Nature, 474, 9–11.
Nicole, E. N., Shuchi, H. D., Anna, E. C., & Atsumi, S. (2014). Metabolic engineering for higher alcohol production. Metabolic Engineering, 25, 174–182.
Schäfer, A., Tauch, A., Jäger, W., Kalinowski, J., Thierbach, G., & Pühler, A. (1994). Small mobilizable multi-purpose cloning vectors derived from the E. coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene, 145, 69–73.
Schreiner ME, Fiur D, Holátko J, Pátek M, Eikmanns BJ (2005) E1 enzyme of the pyruvate dehydrogenase complex in Corynebacterium glutamicum: molecular analysis of the gene and phylogenetic aspects. J Bacteriol 187:6005–6018
Seong, H. P., Su, J. K., & Ji, S. H. (2014). Metabolic engineering of Saccharomyces cerevisiae for the production of isobutanol and 3-methyl-1-butanol. Applied Microbiology and Biotechnology, 98, 9139–9147.
Shine, J., & Dalgarno, L. (1974). The 3′-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proceedings of the National Academy of Sciences of the United States of America, 4, 1342–1346.
Smith, K. M., Cho, K. M., & Liao, J. C. (2010). Engineering Corynebacterium glutamicum for isobutanol production. Applied Microbiology and Biotechnology, 87, 1045–1055.
Takors, R., Bathe, B., Rieping, M., Hans, S., Kelle, R., & Huthmacher, K. (2007). Systems biology for industrial strains and fermentation processes—example: amino acids. Journal of Biotechnology, 129(2), 181–190.
Vander, R. M. E., Lange, C., & Molenaar, D. (1999). A heat shock following electroporation induces highly efficient of corynebacterium glutamicum with xenogeneic plasmid DNA. Applied Microbiology and Biotechnology, 52, 541–545.
Acknowledgments
This research was financially supported by the National Natural Science Foundation of China (No. 2117623 and No.21211140237), National High Technology Research and Development Program of China (863 Program) (2013AA065803), Guangdong science and technology research program (2013B010403021).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author(s) Shiyuan Xiao, Jingliang Xu, Xiaoyan Chen, Xiekun Li, Yu Zhang, Zhenhong Yuan declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Xiao, S., Xu, J., Chen, X. et al. 3-Methyl-1-butanol Biosynthesis in an Engineered Corynebacterium glutamicum . Mol Biotechnol 58, 311–318 (2016). https://doi.org/10.1007/s12033-016-9929-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-016-9929-y