Abstract
Plasmid vectors can be optimized by including specific signals that promote antigen targeting to the major antigen presentation and processing pathways, increasing the immunogenicity and potency of DNA vaccines. A pVAX1-based backbone was used to encode the Green Fluorescence Protein (GFP) reporter gene fused either to ISG (Invariant Surface Glycoprotein) or to TSA (trans-sialidase) Trypanosoma brucei genes. The plasmids were further engineered to carry antigen-targeting sequences, which promote protein transport to the extracellular space (secretion signal), lysosomes (LAMP-1) and to the endoplasmic reticulum (adenovirus e1a). Transfection efficiency was not affected by differences in the size between each construct as no differences in the plasmid copy number per cell were found. This finding also suggests that the addition of both ISG gene and targeting sequences did not add sensitive regions prone to nuclease attack to the plasmid. Cells transfected with pVAX1GFP had a significant higher number of transcripts. This could be a result of lower mRNA stability and/or a lower transcription rate associated with the bigger transcripts. On the other hand, no differences were found between transcript levels of each ISG-GFP plasmids. Therefore, the addition of these targeting sequences does not affect the maturation/stability of the transcripts. Microscopy analysis showed differences in protein localization and fluorescent levels of cells transfected with pVAX1GFP and ISG constructs. Moreover, cells transfected with the lamp and secretory sequences presented a distinct distribution pattern when compared with ISG protein. Protein expression was quantified by flow cytometry. Higher cell fluorescence was observed in cells expressing the cytoplasmic fusion protein (ISG-GFP or TSA-GFP) compared with cells where the protein was transported to the lysosomal pathway. Protein transport to the endoplasmic reticulum does not lead to a decrease in the mean fluorescence values. The secretion signal was only effective when used in conjunction with TSA gene. Therefore, the characteristics of each protein (e.g., presence of transmembrane domains) might influence the efficacy of its cellular transport. This analysis constitutes a useful tool for the optimization of the design of DNA vaccines.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA vaccine technology has been successfully applied to the treatment of cancer [1], autoimmune diseases [2], and allergies [3], as well as a prophylactic approach against several infectious diseases from bacterial and viral origin [4]. DNA vaccines are effective at eliciting a T-cell response as well as the production of antibodies against the protein encoded by the plasmid [5]. Protective immunity has also been observed following DNA immunization [6]. Additionally, DNA vaccination presents commercial advantages compared to conventional approaches since plasmids can be manufactured by generic processes, including standardized quality control and storage conditions that are independent of the encoded sequence [7].
Regardless of their potential, an effort has been made to improve their performance through the use of several optimization strategies such as the (1) improvement of the immunological properties of the vector itself through the addition of CpG sequences, capable of acting as an adjuvant [8]; (2) maximization of antigen expression through the use of improved vectors using strong constitutive promoters, addition of regulatory elements that work as transcriptional enhancers, or through codon optimization of the antigen gene sequence [9–11]; (3) co-administration of DNA with cytokines and chemokines to amplify the immune response and modulate the Th1 and Th2 profile [12–14]; (4) combined administration of DNA with lipid complexes, microparticles or other delivery formulations to increase transfection and expression efficiencies [15–17].
Furthermore, improvement of antigen delivery to the immune system is also considered to be a fundamental aspect for DNA vaccine development, since several studies report that the localization of the expressed protein can significantly influence the efficacy of DNA vaccines [18]. Therefore, several molecular approaches have been attempted including the use of signal sequences which target the antigen to intracellular compartments involved in the processing and presentation pathways MHC I [19] and MHC II [20, 21].
The invariant surface glycoprotein (ISG) [22] and the N-terminal trans-sialidase (nTSA) [23] of Trypanosoma brucei were used as model proteins in this study. These antigens have been selected for the construction of DNA vaccine prototypes against African Trypanosomiasis [23]. The two proteins were fused to GFP and to three different targeting sequences: (i) a secretion signal (sc) composed of the first 21 amino acids of the human tissue plasminogen activator signal sequence [24] that promotes protein secretion to the extracellular space, (ii) the lysosomal-associated membrane protein-1 (LAMP) sequence [25] that directs protein to the lysosomes, and (iii) the adenovirus e1a sequence [26] that allows protein transport to the endoplasmic reticulum (ER).
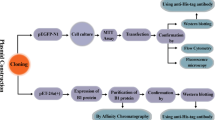
It has been reported that introduction of DNA sequences into a vector backbone can significantly affect plasmid stability and resistance to nuclease attack, reducing the number of active plasmid copies that are able to reach the nucleus [27, 28]. The addition of DNA sequences can also influence mRNA stability and compromise transgene expression. Therefore, plasmid copy number and mRNA levels were determined in order to assess the effect of addition of targeting sequences on plasmid stability, transfection, and transcription efficiencies. Protein expression was analyzed by microscopy and quantified by flow cytometry. Here we present a systematic analysis of the performance of vectors encoding three targeting sequences used in vaccine development. This study is an important contribution for a rational design of DNA vaccines.
Materials and Methods
DNA Vaccine Prototype Design and Purification
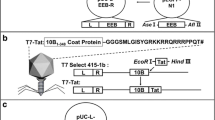
A detailed description of the plasmids used in this study is outlined in Table 1. The plasmids were initially constructed using the pVAX1/lacZ (Invitrogen) as the backbone vector, to be used as candidate vaccines against African Trypanosomiasis. For the purpose of this study, the lacZ reporter gene was replaced by the Green Fluorescence Protein (GFP), generating pVAX1GFP [27] (3697 bp). Firstly, the ISG [22] gene from T. brucei was amplified by PCR (1571 nucleotides) and inserted into pVAX1/lacZ. This construct (pISGlacZ) produces a cytoplasmic protein that was used as a control of the experiments. The second construction (pscISGlacZ) was generated through the addition of the coding sequence for the first 21 amino acids of the human tissue plasminogen activator signal sequence [24] upstream of the ISG gene. To generate the third construct (pISGlacZlamp) the LAMP [25] (Lysosomal-associated membrane protein-1) nucleotide sequence was inserted downstream into the pISGlacZ plasmid. The fourth plasmid (pe1aISGlacZ) was constructed to include the adenovirus e1a ER signal sequence [26] upstream and in frame with ISG. Since there are reports in the literature that suggest that an efficient transport to the lysosomes is dependent on protein loading to the secretory pathway by an ER targeting signal [29], the e1a sequence was also added to the pISGlamplacZ plasmid to obtain a fifth construct (pe1aISGlacZlamp). Finally, in all plasmids the reporter gene lacZ was replaced by the GFP gene (pISG, pISGlamp, pscISG, pe1aISG, and pe1aISGlamp).
The above plasmids were used to construct a second set of vaccines prototypes (pTSA, pTSAlamp, pscTSA, pe1aTSA, pe1aTSAlamp) where ISG was replaced with the N-terminal sequence of the TSA gene (1416 nucleotides) [23], also amplified from T. brucei genomic DNA by PCR.
Automated DNA sequencing was performed (Stab Vida Inc., Portugal) on these constructs to check correct sequence and alignment between GFP, antigen genes, and targeting sequences.
Plasmids were amplified in Escherichia coli DH5alpha and purified using the High Pure Isolation Kit (Roche). One microgram of pDNA was added to the lipofectamine solution. Each plasmid was diluted to the same concentration in order to use the same volume of purified DNA in each transfection assay.
Cell Culture and Transfection Assays
Transfection assays were performed using Chinese Hamster Ovary (CHO) cells. This cell line was selected as a model to analyze if differences in transfection were introduced by the addition of targeting sequences in the parameters studied in this work. CHO cells are a standard mammalian cell line used for transfection and expression of a broad range of recombinant proteins [30]. Moreover, they are considered to be an appropriate model for studies designed to assess plasmid content in eukaryotic cells [27, 31]. However, results present here may not necessarily translate to other cell lines since protein expression, plasmid half life, and transfection levels vary using different models [27].
The starting culture was seeded in 75 cm2 flasks using F-12 (Ham) nutrient mixture (Gibco, UK), containing 10% (v/v) fetal bovine serum (FBS) (Gibco, UK) and supplemented with 1× non-essential amino acids (Gibco, UK), 1× gentamicin (Gibco, UK), 1× sodium pyruvate (Gibco, UK), and 1× antibiotic–antimycotic (Gibco, UK). Cultures were incubated in 5% CO2 humidified environment at 37ºC. Cells were trypsinized after reaching confluence and then seeded into 24-well culture plates (4 × 104 cells per well). The cells were then incubated for 48 h and transfected using the combination of 1.5 μl of lipofectamine (Lipofectamine 2000TM, Invitrogen, UK) with 1 μg of each plasmid per well. Lipofectamine is a cationic lipid that forms a complex with plasmid DNA. The negative control of this experiment was set using cells transfected with plasmid DNA to which no lipofectamine was added. Finally, the medium was replaced 12 h after the transfection to remove plasmids not internalized by cells.
Protein Expression and Intracellular Localization
Protein expression was first assessed by fluorescence microscopy (Olympus 6X40 with an Olympus U-MWB mirror cube unit; excitation filter BP 450–480, barrier filter BA 515). Cells were transfected as described above and observed at 24, 30, and 48 h post-transfection. It was concluded that the best time point to visualize protein expression was at around 30 h. Therefore, all other parameters investigated in this study were assessed at this time point.
Protein expression was quantified by flow cytometry, a more precise method when compared to fluorescence microscopy. At 30 h post-transfection, cells were washed three times with phosphate buffered saline (PBS), trypsinized, and resuspended in 2 ml of PBS supplemented with 10% of FBS. The cells were analyzed using the cytometer FACscan Scalibur (Becton–Dickinson, USA). In each run the forward scatter (FSC), side scatter (SSC), and green fluorescence (FL1) parameters were registered. When FSC is plotted against SSC, it is possible to isolate the debris and dead cells from the population of cells to be analyzed. Subsequently, using the FL1 parameter, the selected population of cells was divided between the non-transfected cells, displaying background FL1 (determined using the auto-fluorescence of the negative cells), and the transfected cells, emitting green fluorescence. A direct correlation between the transfection level (defined as the percentage of cells expressing green fluorescence above the threshold level defined by the natural autofluorescence of the control non-transfected cells) and the mean fluorescence was found. As the fluorescence levels are more relevant to the analysis performed here, only the mean fluorescence data are presented in the “Results” section. The CellQuest Pro software (Becton–Dickinson, USA) was used for the analysis of these parameters.
Plasmid Copy Number
The plasmid copy number in cells transfected with each one of the constructs was analyzed by quantitative real time PCR [31]. Thirty hours post-transfection, the cells were washed three times with 1 ml of PBS in order to remove non-specifically bound pDNA. Cells were then trypsinized and centrifuged for 8 min at 160g. Subsequently, the cell pellet was washed and resuspended in 2 ml of the PBS. The number of cells was determined by the average of two performed microscope counts using a Neubauer chamber. Finally, cells were centrifuged to remove the supernatant and stored at −80ºC until use.
Immediately upon thawing, cell pellets were resuspended in PCR-grade water to obtain a final concentration of 6,250 cells/μl [27]. The set temperature of the first cycle of the amplification PCR programme is enough to induce cell lysis. Two microliters of this solution was added to the PCR reaction mixture (1× SYBR Green mixture, 0.4 μM of each primer, and 3 mM of MgCl2 in a total volume of 20 μl). A reaction containing 2 μl of control cells (cells transfected with pVAX1GFP plasmid to which no lipofectamine was added) was included in each PCR run to be used as a negative control of the assay. A second negative control of the PCR reaction was set up using PCR-grade water instead of the control cells.
Plasmid quantification was performed by promoting the amplification of a 108 bp sequence from the eGFP gene using the 5′-TCGAGCTGGACGGCGACGTAA A-3′ forward primer, and the 5′-TGCCGGTGGTGCAGATGAAC-3′ reverse primer. PCRs were carried out in a Roche LightCycler® detection system using the FastStart DNA Master SYBR Green I kit (Roche, Germany). Reactions were incubated at 95ºC for 10 min to activate FastStart DNA polymerase, followed by 30 cycles of 10 s at 95ºC, 5 s at 55ºC, and 7 s at 72ºC. After the final extension cycle, reaction mixtures were kept at 70ºC for 30 s, and heat-denatured over a temperature gradient at 0.1ºC/s from 70 to 95ºC.
The plasmid copy number was determined based on a calibration curve, constructed using a range of concentrations from 2 pg up to 2,000 pg of pVAX1GFP. Each plasmid dilution was prepared using PCR-grade water that was then added to 2 μl of control cells.
GFP mRNA Content
The GFP mRNA levels in cells transfected with each one of the plasmids were quantified using a quantitative reverse transcription RT-PCR method [27]. Cells were harvested 30 h after transfection and stored at −80ºC until use. Total RNA was purified using the Pure RNA Isolation Kit (Roche, Germany) and quantified by absorption at 260 nm. Subsequently, 300 ng of total mRNA purified was used as template for the reverse transcription reaction performed using the First Strand cDNA Synthesis Kit for RT-PCR (AMV) (Roche, Germany). Samples were treated with DNase to ensure removal of any DNA contamination. Two microliters of the cDNA reaction was then used in the quantitative real time PCR reaction, performed as described above for the quantification of the plasmid copy number per cell.
Statistics
Differences in the plasmid copy number, mRNA and mean fluorescence levels between plasmids were analyzed performing an ANOVA (General Linear Model, Tukey’s comparison test) with the level of confidence being P < 0.05. Analyses were performed with the Minitab statistical package, version 15 (Minitab Inc.).
Results
In this work, ISG or nTSA genes from T. brucei were fused with GFP reporter and with sequences targeting proteins to extracellular space (sc), to lysosome (LAMP or the combination of LAMP and e1a) or to the ER (e1a). Even though these sequences have been included in several vectors used as DNA vaccines, the effect of the selected targeting sequences in cell plasmid uptake and stability, mRNA transcription and protein expression are examined and discussed on a comparative basis.
Effect of the Signal Sequences on Plasmid Uptake/Stability and GFP mRNA Transcription
No significant differences were found between the number of plasmid copies in cells transfected with each of the constructs (Fig. 1). Therefore, plasmid uptake was not affected by the differences in the size of the different plasmids analyzed. Furthermore, the addition of ISG gene and targeting sequences appears not to have introduced sensitive regions to nuclease attack into the vector backbone.
Plasmid copy number CHO cells transfected with pVAX1GFP- and ISG-encoding plasmids at 30 h post-transfection. Data shown resulted from the analysis of three independent transfection assays, each one performed in duplicate. One way ANOVA (Tukey’s comparison test) was performed in order to compare pDNA copies in cells transfected with each construct. No significant differences were observed (P > 0.05)
mRNA determination showed that cells transfected with the plasmid pVAX1GFP, which codes for the smaller protein (GFP), had significantly more transcripts than cells expressing the ISG-GFP fusion proteins (Fig. 2). No significant differences in the levels of GFP mRNA were detected in cells transfected with pISG and with each one of the plasmids encoding a targeted protein (Fig. 2).
mRNA quantification of CHO cells transfected with pVAX1GFP- and ISG-encoding plasmids at 30 h post-transfection. Data shown resulted from the analysis of three independent transfection assays, each one performed in duplicate. One way ANOVA (Tukey’s comparison test) was performed in order to compare the number of copies of GFP mRNA in cells transfected with each construct. pVAX1GFP has a significant higher number of mRNA GFP copies compared with the rest of the groups (P < 0.05). However, no significant differences between ISG constructs were observed (P > 0.05)
Effect of the Signal Sequences on Protein Expression
The expression of ISG fused to GFP was analyzed by fluorescence microscopy (Fig. 3). Fluorescence of cells transfected with pVAX1GFP is significantly higher than cells transfected with any of the other constructs. Moreover, in this case, GFP is found throughout the cytoplasm and nucleus as described previously elsewhere [32]. Nuclear fluorescence was not observed in cells that received the ISG constructs.
Differences in fluorescence intensities were also observed between the ISG encoding plasmids, especially between pe1aISGlamp and pISGlamp and cells transfected with pISG plasmid. The later cells were brighter and presented an even distribution of GFP in the cytoplasm. The low fluorescence levels of cells expressing the lamp sequence made it difficult to point out the exact localization of e1aISGlamp and ISGlamp proteins. However, in some of these cells it was possible to observe fluorescence not only in the cytoplasm, but also localized in vesicles. In the case of cells transfected with the secretory sequence, the fluorescence is more intense close to the cellular membrane. Lastly, no obvious differences were found between fluorescence of e1aISG and ISG proteins.
Quantification of protein expression was performed using flow cytometry. This method is presented here as an alternative method to immunofluorescence assays to assess protein expression and targeting. CHO cells transfected with both ISG and nTSA coding plasmids were analyzed. Fluorescent green cells were observed in cells transfected with each new construct, confirming that protein expression was successful. In addition, cells expressing the different fusion proteins were markedly lower (up to 100 times) when compared with the fluorescence of cells expressing the reporter gene GFP. Differences between the fluorescence levels of the different ISG proteins were also observed. The mean fluorescence of cells transfected with pISG was significantly higher (P < 0.05) when compared with cells transfected with pISGlamp and pe1aISGlamp (Fig. 4a). The fluorescence pattern of cells transfected with pTSAlamp and pe1aTSAlamp is also reduced when compared to the cytoplasmic protein encoded by pTSA (Fig. 4b). No statistically significant differences (P > 0.05) were observed in the mean fluorescence obtained when transfecting cells with plasmids pe1aISG (Fig. 4a) or pe1aTSA (Fig. 4b). Finally, a decrease in the mean fluorescence of cells transfected with plasmids that encode a secreted protein was only observed in cells transfected with pscTSA (Fig. 4b) and not with pscISG (Fig. 4a).
Mean fluorescence of cells transfected with a plasmid ISG and b nTSA constructs. Cells were collected at 30 h post-transfection. The averages and standard error of the mean shown are from three (ISG) and two (nTSA) independent assays, each one performed in duplicate. ANOVA (General Linear Model, Tukey’s comparison test) was performed in order to compare differences on the fluorescence levels between transfected plasmids. Fluorescence levels that are significantly different (P < 0.05) from the ones detected in cells transfected with pISG or pTSA plasmids are signed (*). The mean fluorescence values in cells transfected with pVAX1GFP were 1336 ± 238 and 1573 ± 30, respectively, used as control experiments to assess the fluorescence of cells transfected with ISG and TSA constructs. AU arbitrary units
Discussion
When combined with plasmid DNA, cationic lipids (lipoplexes), as the one used in this study, are internalized by cells using an endocytic pathway including a clathrin-dependent and -independent pathway and a caveolae-mediated uptake. Upon internalization, lipoplexes are trapped in endocytic vesicles that converge to late endosomes and finally to lysosomes where degradation occurs [33]. This topic was extensively reviewed elsewhere [34, 35]. The timing and the exact mechanisms responsible for plasmid DNA release from endo-lysosomal compartments into the cytoplasm are not clear, but this is considered to be one of the major limitation steps to transgene expression. Once in the cytoplasm, plasmid DNA is degraded by Ca2+ sensitive cytosolic nucleases that reduce even more the number of plasmid copies capable of reaching the nucleus [36]. In fact, 106 naked plasmid molecules are needed to transform a single cell, from which 102 to 104 are detected in the nucleus following plasmid delivery using cationic lipids [37]. The fact that only a minority of the total number of plasmid copies internalized by cells is detected in the nucleus is also a result of the processes that regulate plasmid uptake to this compartment. Plasmid DNA can be trapped inside the nucleus following the reorganization of the nuclear envelope in active dividing cells. However, since plasmid DNA is also found in non-mitotic cells other processes have to regulate nuclear import. In fact, nuclear entry can be mediated by DNA sequences containing binding sites for transcription factors [38, 39]. In conclusion, a number of physical and biochemical barriers determine the efficiency of gene expression. As demonstrative of the complexity of these processes, a recent study has shown that intracellular elimination and DNA release from the different non-viral vectors correlate poorly with transgene expression [40].
One of the objectives of this study was to assess if the addition of targeting sequences had introduced sensitive spots to nuclease attack to the vector backbone. The time point selected for the performed analysis is within the period when plasmid DNA is more vulnerable. Our group and others have demonstrated that the half-life of pDNA in cells following transfection using cationic lipids is approximately 20 h [31, 35, 41]. However, this time-frame can change depending on the carrier system used [40].
The addition of extra sequences to plasmids can introduce secondary structures more nuclease labile that may lead to a decrease in intracellular plasmid stability. The decreased resistance to nucleases action ultimately results in a lower transgene copy number, and therefore compromises gene expression [28]. Plasmid susceptibility to nuclease attack due to the addition of the selected targeting sequences, as well as the ISG gene, was unknown. Previous work has demonstrated that the GFP gene did not introduce sensitive regions, such as single-stranded secondary structures, into the pVAX1 vector (Invitrogen) [27, 28]. Since no differences in plasmid copy number were observed between cells transfected with each construct is reasonable to conclude that the addition of ISG gene sequence and the targeting sequences did not affect plasmid susceptibility to nuclease degradation. On the other hand, the increase in plasmid size up to 1.6 kb did not compromise plasmid DNA uptake.
As transfected cells presented similar plasmid content an approximate number of GFP transcripts were expected to be detected. However, a higher number of GFP transcripts were determined in cells transfected with pVAX1GFP, possibly due to the fact that the transcript is much smaller, ultimately contributing to a higher transcription rate and stability. Importantly, no significant differences were found between the ISG constructs. This means that these plasmids have similar transcription efficiencies and comparable mRNA stability, even though they contain different signal sequences.
The amount of expressed protein was determined indirectly by the amount of molecules with biological activity as measured by the fluorescence of GFP. Since LAMP sequence or its combination with e1a signal promotes protein transport to the lysosome where protein degradation occurs, the decrease in the fluorescence levels was expected. The data obtained suggest that protein sorting to the lysosomes was achieved. Microscopy analysis also indicates that this transport was effective, since fluorescence appeared to be localized inside vesicles, possibly part of the lysosomal pathway. On the other hand, transport to the endoplasmic reticulum does not result in protein degradation. Therefore, no significant decrease of fluorescence was likely to be observed. Again, this is supported by flow cytometry and microscopy data.
At the time of the analysis, only cells expressing the plasmid encoding TSA in conjunction with the secretory signal showed a decrease in the fluorescent levels. The intrinsic characteristics of each protein might help to explain these differences. The ISG protein has a predicted transmembrane domain located between the amino acids 469 and 491, as predicted by the TMHMM tool (available on http://www.cbs.dtu.dk/services/TMHMM/) [42]. The presence of this domain might retain the protein in the cell membrane, hampering or delaying its secretion to the extracellular space. In fact, a more intense fluorescence was observed near the cell membrane, which corroborates this hypothesis. On the other hand, no such type of domains is found in the TSA protein.
Since cells transfected with the different plasmids show similar levels of plasmid content and GFP transcripts, it is reasonable to assume that the differences observed in the fluorescence levels relatively to the control is the result of an effective transport of the model proteins to the expected compartments (extracellular space, lysosomes, and endoplasmic reticulum).
It has been demonstrated that an efficient antigen sorting to the MHC I and MHC II pathways improves DNA vaccines immunogenicity. The endogenous protein encoded by the DNA vector should not have access to the MHC II pathway that primarily processes exogenous antigens [43]. This limitation can be overcome if a targeting signal is used that promotes protein sorting to the lysosomes [44], compartments involved in the generation of epitopes to be presented to the immune system when complexed with MHC II molecules. On the other hand, when the antigen encoded by a DNA vaccine is expressed in the cytoplasm it can be processed in the context of MHC I pathway. However, protein sorting to the proteasomes [45] and to the endoplasmic reticulum has also been engineered to increase the efficiency of DNA vaccines [19] by increasing antigen loading to compartments involved in the MHC I pathway.
The sequences selected for this study have been used in conjunction with various antigens of both bacterial [46] and viral [21, 47, 48] origins, as well as part of therapeutic DNA vaccines against cancer [49]. This is illustrative of their general applicability in DNA vaccine design. However, it should be noted that differences might be observed in protein expression levels and transfection efficiencies if different cell lines and animal models had been used. Additionally, mRNA expression levels have been considered to be construct specific. Nevertheless, this work presents a systematic in vitro study on the influence of signal sequences with potential interest for DNA vaccines development at the level of plasmid uptake and susceptibility to degradation, mRNA transcription and protein expression.
References
Bodles-Brakhop, A. M., & Draghia-Akli, R. (2008). DNA vaccination and gene therapy: Optimization and delivery for cancer therapy. Expert Review of Vaccines, 7(7), 1085–1101.
Okura, Y., Miyakoshi, A., Kohyama, K., Park, I. K., Staufenbiel, M., & Matsumoto, Y. (2006). Nonviral Abeta DNA vaccine therapy against Alzheimer’s disease: Long-term effects and safety. Proceedings of the National Academy of Sciences of the United States of America, 103(25), 9619–9624.
Li, G., Liu, Z., Zhong, N., Liao, B., & Xiong, Y. (2006). Therapeutic effects of DNA vaccine on allergen-induced allergic airway inflammation in mouse model. Cellular and Molecular Immunology, 3(5), 379–384.
Laddy, D. J., & Weiner, D. B. (2006). From plasmids to protection: a review of DNA vaccines against infectious diseases. International Reviews of Immunology, 25(3–4), 99–123.
Chikhlikar, P., de Arruda, L. B., Maciel, M., et al. (2006). DNA encoding an HIV-1 Gag/human lysosome-associated membrane protein-1 chimera elicits a broad cellular and humoral immune response in Rhesus macaques. PLoS ONE, 1, e135.
Hoft, D. F., Eickhoff, C. S., Giddings, O. K., Vasconcelos, J. R. C., & Rodrigues, M. M. (2007). Trans-sialidase recombinant protein mixed with CpG motif-containing oligodeoxynucleotide induces protective mucosal and systemic trypanosoma cruzi immunity involving CD8+ CTL and B cell-mediated cross-priming. Journal of Immunology, 179(10), 6889–6900.
Carvalho, J. A., Prazeres, D. M., & Monteiro, G. A. (2009). Bringing DNA vaccines closer to commercial use. IDrugs, 12(10), 642–647.
Klinman, D. M., Yamshchikov, G., & Ishigatsubo, Y. (1997). Contribution of CpG motifs to the immunogenicity of DNA vaccines. Journal of Immunology, 158(8), 3635–3639.
Cheung, Y. K., Cheng, S. C.-S., Sin, F. W.-Y., & Xie, Y. (2004). Plasmid encoding papillomavirus Type 16 (HPV16) DNA constructed with codon optimization improved the immunogenicity against HPV infection. Vaccine, 23(5), 629–638.
Moreau, P., Hen, R., Wasylyk, B., Everett, R., Gaub, M. P., & Chambon, P. (1981). The SV40 72 base repair repeat has a striking effect on gene expression both in SV40 and other chimeric recombinants. Nucleic Acids Research, 9(22), 6047–6068.
Barouch, D. H., Zy, Yang, Wp, Kong, et al. (2005). A human T-cell leukemia virus type 1 regulatory element enhances the immunogenicity of human immunodeficiency virus type 1 DNA vaccines in mice and nonhuman primates. Journal of Virology, 79(14), 8828–8834.
Kim, J. J., Simbiri, K. A., Sin, J. I., et al. (1999). Cytokine molecular adjuvants modulate immune responses induced by DNA vaccine constructs for HIV-1 and SIV. Journal of Interferon and Cytokine Research, 19(1), 77–84.
Xu, R., Megati, S., Roopchand, V., et al. (2008). Comparative ability of various plasmid-based cytokines and chemokines to adjuvant the activity of HIV plasmid DNA vaccines. Vaccine, 26(37), 4819–4829.
Barouch, D. H., Santra, S., Schmitz, J. E., et al. (2000). Control of viremia and prevention of clinical AIDS in Rhesus monkeys by cytokine-augmented DNA vaccination. Science, 290(5491), 486–492.
Wang, D., Christopher, M. E., Nagata, L. P., et al. (2004). Intranasal immunization with liposome-encapsulated plasmid DNA encoding influenza virus hemagglutinin elicits mucosal, cellular and humoral immune responses. Journal of Clinical Virology, 31(Supplement 1), 99–106.
Otten, G. R., Schaefer, M., Doe, B., et al. (2005). Enhanced potency of plasmid DNA microparticle human immunodeficiency virus vaccines in rhesus macaques by using a priming-boosting regimen with recombinant proteins. Journal of Virology, 79(13), 8189–8200.
Minigo, G., Scholzen, A., Tang, C. K., et al. (2007). Poly-l-lysine-coated nanoparticles: A potent delivery system to enhance DNA vaccine efficacy. Vaccine, 25(7), 1316–1327.
Boyle, J. S., Koniaras, C., & Lew, A. M. (1997). Influence of cellular location of expressed antigen on the efficacy of DNA vaccination: Cytotoxic T lymphocyte and antibody responses are suboptimal when antigen is cytoplasmic after intramuscular DNA immunization. International Immunology, 9(12), 1897–1906.
Xu, W., Chu, Y., Zhang, R., Xu, H., Wang, Y., & Xiong, S. (2005). Endoplasmic reticulum targeting sequence enhances HBV-specific cytotoxic T lymphocytes induced by a CTL epitope-based DNA vaccine. Virology, 334(2), 255–263.
Arruda, L. B., Sim, D., Chikhlikar, P. R., et al. (2006). Dendritic cell-lysosomal-associated membrane protein (LAMP) and LAMP-1-HIV-1 gag chimeras have distinct cellular trafficking pathways and prime T and B cell responses to a diverse repertoire of epitopes. Journal of Immunology, 177(4), 2265–2275.
Henriques, A. M., Fevereiro, M., Prazeres, D. M. F., & Monteiro, G. A. (2007). Development of a candidate DNA vaccine against Maedi-Visna virus. Veterinary Immunology and Immunopathology, 119(3–4), 222–232.
Overath, P., Chaudhri, M., Steverding, D., & Ziegelbauer, K. (1994). Invariant surface proteins in bloodstream forms of Trypanosoma brucei. Parasitology Today, 10(2), 53–58.
Silva, M. S., Prazeres, D. M. F., Lança, A., Atouguia, J., & Monteiro, G. A. (2009). Trans-sialidase from Trypanosoma brucei as a potential target for DNA vaccine development against African trypanosomiasis. Parasitology Research, 105(5), 53–58.
Qiu, J. T., Liu, B., Tian, C., Pavlakis, G. N., & Yu, X. F. (2000). Enhancement of primary and secondary cellular immune responses against human immunodeficiency virus type 1 gag by using DNA expression vectors that target gag antigen to the secretory pathway. Journal of Virology, 74(13), 5997–6005.
Ruff, A. L., Guarnieri, F. G., Staveley-O’Carroll, K., Siliciano, R. F., & August, J. T. (1997). The enhanced immune response to the HIV gp160/LAMP chimeric gene product targeted to the lysosome membrane protein trafficking pathway. Journal of Biological Chemistry, 272(13), 8671–8678.
Persson, H., Jornvall, H., & Zabielski, J. (1980). Multiple mRNA species for the precursor to an adenovirus-encoded glycoprotein: Identification and structure of the signal sequence. Proceedings of the National Academy of Sciences of the United States of America, 77(11), 6349–6353.
Azzoni, A. R., Ribeiro, S. C., Monteiro, G. A., & Prazeres, D. M. (2007). The impact of polyadenylation signals on plasmid nuclease-resistance and transgene expression. Journal of Gene Medicine, 9(5), 392–402.
Ribeiro, S. C., Monteiro, G. A., & Prazeres, D. M. (2004). The role of polyadenylation signal secondary structures on the resistance of plasmid vectors to nucleases. Journal of Gene Medicine, 6(5), 565–573.
Thomson, S. A., Burrows, S. R., Misko, I. S., Moss, D. J., Coupar, B. Eá. H., & Khanna, R. (1998). Targeting a polyepitope protein incorporating multiple class II-restricted viral epitopes to the secretory/endocytic pathway facilitates immune recognition by CD4+ cytotoxic T lymphocytes: A novel approach to vaccine design. Journal of Virology, 72(3), 2246–2252.
Werner, R. G., Noe, W., Kopp, K., & Schluter, M. (1998). Appropriate mammalian expression systems for biopharmaceuticals. Arzneimittelforschung, 48(8), 870–880.
Carapuca, E., Azzoni, A. R., Prazeres, D. M., Monteiro, G. A., & Mergulhao, F. J. (2007). Time-course determination of plasmid content in eukaryotic and prokaryotic cells using real-time PCR. Molecular Biotechnology, 37(2), 120–126.
Seibel, N. M., Eljouni, J., Nalaskowski, M. M., & Hampe, W. (2007). Nuclear localization of enhanced green fluorescent protein homomultimers. Analytical Biochemistry, 368(1), 95–99.
Vázquez, E., Ferrer-Miralles, N., & Villaverde, A. (2008). Peptide-assisted traffic engineering for nonviral gene therapy. Drug Discovery Today, 13(23–24), 1067–1074.
Hoekstra, D., Rejman, J., Wasungu, L., Shi, F., & Zuhorn, I. (2007). Gene delivery by cationic lipids: In and out of an endosome. Biochemical Society Transactions, 35(Pt 1), 68–71.
Lechardeur, D., Verkman, A. S., & Lukacs, G. L. (2005). Intracellular routing of plasmid DNA during non-viral gene transfer. Advanced Drug Delivery Reviews, 57(5), 755–767.
Pollard, H., Toumaniantz, G., Amos, J. L., et al. (2001). Ca2+-sensitive cytosolic nucleases prevent efficient delivery to the nucleus of injected plasmids. Journal of Gene Medicine, 3(2), 153–164.
Tachibana, R., Harashima, H., Ide, N., et al. (2002). Quantitative analysis of correlation between number of nuclear plasmids and gene expression activity after transfection with cationic liposomes. Pharmaceutical Research, 19(4), 377–381.
Vacik, J., Dean, B. S., Zimmer, W. E., & Dean, D. A. (1999). Cell-specific nuclear import of plasmid DNA. Gene Therapy, 6(6), 1006–1014.
Dean, D. A. (1997). Import of plasmid DNA into the nucleus is sequence specific. Experimental Cell Research, 230(2), 293–302.
Ruponen, M., Arkko, S., Urtti, A., Reinisalo, M., & Ranta, V. P. (2009). Intracellular DNA release and elimination correlate poorly with transgene expression after non-viral transfection. Journal of Controlled Release, 136(3), 226–231.
Roth, C. M., & Sundaram, S. (2004). Engineering synthetic vectors for improved DNA delivery: Insights from intracellular pathways. Annual Review of Biomedical Engineering, 6(1), 397–426.
Krogh, A., Larsson, Br., von Heijne, G., & Sonnhammer, E. L. L. (2001). Predicting transmembrane protein topology with a hidden markov model: Application to complete genomes. Journal of Molecular Biology, 305(3), 567–580.
Rocha, N., & Neefjes, J. (2008). MHC class II molecules on the move for successful antigen presentation. EMBO Journal, 27(1), 1–5.
Marques, E. T. A., Jr., Chikhlikar, P., De Arruda, L. B., et al. (2003). HIV-1 p55Gag encoded in the lysosome-associated membrane protein-1 as a DNA plasmid vaccine chimera is highly expressed, traffics to the major histocompatibility class II compartment, and elicits enhanced immune responses. Journal of Biological Chemistry, 278(39), 37926–37936.
Andersson, H. A., & Barry, M. A. (2004). Maximizing antigen targeting to the proteasome for gene-based vaccines. Molecular Therapy, 10(3), 432–446.
Kamath, A. T., Feng, C. G., Macdonald, M., Briscoe, H., & Britton, W. J. (1999). Differential protective efficacy of DNA vaccines expressing secreted proteins of Mycobacterium tuberculosis. Infection and Immunity, 67(4), 1702–1707.
Kim, S. J., Suh, D., Park, S. E., et al. (2003). Enhanced immunogenicity of DNA fusion vaccine encoding secreted hepatitis B surface antigen and chemokine RANTES. Virology, 314(1), 84–91.
Suk Kim, M., & Sin, J. I. (2005). Both antigen optimization and lysosomal targeting are required for enhanced anti-tumour protective immunity in a human papillomavirus E7-expressing animal tumour model. Immunology, 116(2), 255–266.
Sherritt, M., Cooper, L., Moss, D. J., Kienzle, N., Altman, J., & Khanna, R. (2001). Immunization with tumor-associated epitopes fused to an endoplasmic reticulum translocation signal sequence affords protection against tumors with down-regulated expression of MHC and peptide transporters. International Immunology, 13(3), 265–271.
Acknowledgments
The authors thank the Portuguese Fundação para a Ciência e a Tecnologia for fund support (POCI/BIO/55799/2004), and doctoral and post-doctoral Grants, respectively, to JA Carvalho (SFRH/BD/21423/2005) and AR Azzoni (BPD/14879/2003). The authors would like to thank to MS Silva for providing pVAX1-TSA construct. We would also like to acknowledge A.M. Henriques for providing the targeting sequences used in this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article can be found at http://dx.doi.org/10.1007/s12033-010-9327-9
Rights and permissions
About this article
Cite this article
Carvalho, J.A., Azzoni, A.R., Prazeres, D.M.F. et al. Comparative Analysis of Antigen-Targeting Sequences Used in DNA Vaccines. Mol Biotechnol 44, 204–212 (2010). https://doi.org/10.1007/s12033-009-9229-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-009-9229-x