Abstract
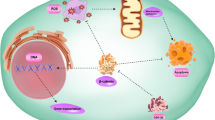
Notch signaling pathway is evolutionarily conserved in mammals, which plays an important role in cell development and differentiation. In recent years, increasing evidence has shown that aberrant activation of Notch is associated with tumor process. Aberrant activation of Notch signaling pathway has been found in many different solid tumors can induce cell proliferation, metastasis and epithelial-mesenchymal transition. Notch receptor and its ligand are both single transmembrane protein, and Notch is activated when it binds to the Notch ligand of neighbor cells. The signal transduction of Notch signaling pathway is only between cells that are in contact with each other, which is independent of second messengers. Thus, Notch needs to cross talk with other signaling pathways, including PI3K/AKT, NF-κB, integrin and miRNAs, to precisely regulate cell fate. In this review, we summarize the roles of Notch signaling pathway in tumor metastasis and its regulatory mechanisms and discuss the current treatment strategies targeting Notch signal pathway.


Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin. 2017;67(1):7–30. doi:10.3322/caac.21387.
Charpentier M, Martin S. Interplay of stem cell characteristics, EMT, and microtentacles in circulating breast tumor cells. Cancers. 2013;5(4):1545–65. doi:10.3390/cancers5041545.
Andersson M, Olsen JH. Malignant mesotheliomas in Denmark 1943–1980. Cancer statistics 9. Ugeskr Laeger. 1984;146(14):1085–7.
Zardawi SJ, Zardawi I, McNeil CM, Millar EK, McLeod D, Morey AL, et al. High Notch1 protein expression is an early event in breast cancer development and is associated with the HER-2 molecular subtype. Histopathology. 2010;56(3):286–96. doi:10.1111/j.1365-2559.2009.03475.x.
Danza G, Di Serio C, Ambrosio MR, Sturli N, Lonetto G, Rosati F, et al. Notch3 is activated by chronic hypoxia and contributes to the progression of human prostate cancer. Int J Cancer. 2013;133(11):2577–86. doi:10.1002/ijc.28293.
Ai Q, Ma X, Huang Q, Liu S, Shi T, Zhang C, et al. High-level expression of Notch1 increased the risk of metastasis in T1 stage clear cell renal cell carcinoma. PLoS ONE. 2012;7(4):e35022. doi:10.1371/journal.pone.0035022.
Yang Y, Ahn Y-H, Gibbons DL, Zang Y, Lin W, Thilaganathan N, et al. The Notch ligand Jagged2 promotes lung adenocarcinoma metastasis through a miR-200—dependent pathway in mice. J Clin Investig. 2011;121(4):1373–85. doi:10.1172/jci42579.
Liu H, Wang J, Liu Z, Wang L, Liu S, Zhang Q. Jagged1 modulated tumor-associated macrophage differentiation predicts poor prognosis in patients with invasive micropapillary carcinoma of the breast. Medicine. 2017;96(16):e6663. doi:10.1097/MD.0000000000006663.
Reedijk M, Odorcic S, Chang L, Zhang H, Miller N, McCready DR, et al. High-level coexpression of JAG1 and NOTCH1 is observed in human breast cancer and is associated with poor overall survival. Cancer Res. 2005;65(18):8530–7. doi:10.1158/0008-5472.CAN-05-1069.
Chance O. Value of statistics in the study of cancer of the uterine cervix. Comptes rendus de la Societe francaise de gynecologie. 1951;21(7):305–11.
Dotta JS, Delporte TV. Statistics on the treatment of prostatic cancer. Revista argentina de urologia. 1951;20(9–11):255–7.
Mumm J. Notch signaling: from the outside in. Dev Biol. 2000;228(2):151–65. doi:10.1006/dbio.2000.9960.
Lai EC. Notch signaling: control of cell communication and cell fate. Development. 2004;131(5):965–73. doi:10.1242/dev.01074.
Cesarani F, Garbagnoli E. Local recurrence and lymphatic and osseous metastases following surgery of breast cancer; radiotherapy department statistics for 1944-50. Athena; rassegna mensile di biologia, clinica e terapia. 1951;17(7–8):189–92.
Kopan R, Ilagan MX. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell. 2009;137(2):216–33. doi:10.1016/j.cell.2009.03.045.
Weinmaster G. Notch signal transduction a real rip and more. Curr Opin Genet Dev. 2000;10:363–9.
Nahum AM. Biting the bullet: minimum standards for reporting cancer treatment statistics. Head Neck Surg. 1979;1(3):201.
Upton AC. Survey: reporting practices for cancer treatment statistics. Head Neck Surg. 1979;1(6):500.
Ranganathan P, Weaver KL, Capobianco AJ. Notch signalling in solid tumours: a little bit of everything but not all the time. Nat Rev Cancer. 2011;11(5):338–51. doi:10.1038/nrc3035.
Ellisen LW, Bird J, West PC, et al. TAN+1, the human hormolog of the Drosophila notch gene, is broken by chromosomal translocations in T lymphoblastie neoplasms. Cell. 1991;66(4):649–61.
Weng AP, Ferrando AA, Lee W, Morris JP IV, Silverman LB, Sanchez-Irizarry C, Blacklow SC, Thomas Look A, Aster JC. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science. 2004;306(5694):269–71. doi:10.1126/science.1102160.
Espinosa L, Cathelin S, D’Altri T, Trimarchi T, Statnikov A, Guiu J, et al. The Notch/Hes1 pathway sustains NF-kappaB activation through CYLD repression in T cell leukemia. Cancer Cell. 2010;18(3):268–81. doi:10.1016/j.ccr.2010.08.006.
Wu F, Stutzman A, Mo YY. Notch signaling and its role in breast cancer. Front Biosci. 2007;12:4370–83.
Ma YC, Shi C, Zhang YN, Wang LG, Liu H, Jia HT, et al. The tyrosine kinase c-Src directly mediates growth factor-induced Notch-1 and Furin interaction and Notch-1 activation in pancreatic cancer cells. PLoS ONE. 2012;7(3):e33414. doi:10.1371/journal.pone.0033414.
Song LL, Peng Y, Yun J, Rizzo P, Chaturvedi V, Weijzen S, et al. Notch-1 associates with IKKalpha and regulates IKK activity in cervical cancer cells. Oncogene. 2008;27(44):5833–44. doi:10.1038/onc.2008.190.
Zhou W, Fu XQ, Zhang LL, Zhang J, Huang X, Lu XH, et al. The AKT1/NF-kappaB/Notch1/PTEN axis has an important role in chemoresistance of gastric cancer cells. Cell Death Dis. 2013;4:e847. doi:10.1038/cddis.2013.375.
Zhang Y, Li B, Ji ZZ, Zheng PS. Notch1 regulates the growth of human colon cancers. Cancer. 2010;116(22):5207–18. doi:10.1002/cncr.25449.
Lefort K, Mandinova A, Ostano P, Kolev V, Calpini V, Kolfschoten I, et al. Notch1 is a p53 target gene involved in human keratinocyte tumor suppression through negative regulation of ROCK1/2 and MRCKalpha kinases. Genes Dev. 2007;21(5):562–77. doi:10.1101/gad.1484707.
Viatour P, Ehmer U, Saddic LA, Dorrell C, Andersen JB, Lin C, et al. Notch signaling inhibits hepatocellular carcinoma following inactivation of the RB pathway. J Exp Med. 2011;208(10):1963–76. doi:10.1084/jem.20110198.
Gupta A, Wang Y, Browne C, Kim S, Case T, Paul M, et al. Neuroendocrine differentiation in the 12T-10 transgenic prostate mouse model mimics endocrine differentiation of pancreatic beta cells. Prostate. 2008;68(1):50–60. doi:10.1002/pros.20650.
Konishi J, Yi F, Chen X, Vo H, Carbone DP, Dang TP. Notch3 cooperates with the EGFR pathway to modulate apoptosis through the induction of bim. Oncogene. 2010;29(4):589–96. doi:10.1038/onc.2009.366.
Parr C, Watkins G, Jiang WG. The possible correlation of Notch-1 and Notch-2 with clinical outcome and tumour clinicopathological parameters in human breast cancer. Int J Mol Med. 2004;14(5):779–86.
PRESENTATION of results in the treatment of cancer. V. World Health Organization Expert Committee on Health Statistics. Br J Radiol. 1951;24(282):311–314. doi:10.1259/0007-1285-24-282-311.
Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331(6024):1559–64. doi:10.1126/science.1203543.
Li S, Zhang J, Yang H, Wu C, Dang X, Liu Y. Copper depletion inhibits CoCl2-induced aggressive phenotype of MCF-7 cells via downregulation of HIF-1 and inhibition of Snail/Twist-mediated epithelial-mesenchymal transition. Sci Rep. 2015;5:12410. doi:10.1038/srep12410.
Sahlgren C, Gustafsson MV, Jin S, Poellinger L, Lendahl U. Notch signaling mediates hypoxia-induced tumor cell migration and invasion. Proc Natl Acad Sci. 2008;105(17):6392–7. doi:10.1073/pnas.0802047105.
Kao SH, Wu KJ, Lee WH. Hypoxia, epithelial-mesenchymal transition, and TET-mediated epigenetic changes. J Clin Med. 2016;. doi:10.3390/jcm5020024.
Ahmad A, Wang Z, Kong D, Ali R, Ali S, Banerjee S, et al. Platelet-derived growth factor-D contributes to aggressiveness of breast cancer cells by up-regulating Notch and NF-kappaB signaling pathways. Breast Cancer Res Treat. 2011;126(1):15–25. doi:10.1007/s10549-010-0883-2.
Xie M, Zhang L, He CS, Xu F, Liu JL, Hu ZH, et al. Activation of Notch-1 enhances epithelial-mesenchymal transition in gefitinib-acquired resistant lung cancer cells. J Cell Biochem. 2012;113(5):1501–13. doi:10.1002/jcb.24019.
Erler JT, Bennewith KL, Nicolau M, Dornhofer N, Kong C, Le QT, et al. Lysyl oxidase is essential for hypoxia-induced metastasis. Nature. 2006;440(7088):1222–6. doi:10.1038/nature04695.
Peinado H, Portillo F, Cano A. Switching on-off Snail: LOXL2 versus GSK3beta. Cell Cycle. 2005;4(12):1749–52. doi:10.4161/cc.4.12.2224.
Yuan XW, Wang DM, Hu Y, Tang YN, Shi WW, Guo XJ, et al. Hepatocyte nuclear factor 6 suppresses the migration and invasive growth of lung cancer cells through p53 and the inhibition of epithelial-mesenchymal transition. J Biol Chem. 2013;288(43):31206–16. doi:10.1074/jbc.M113.480285.
Ramms L, Fabris G, Windoffer R, Schwarz N, Springer R, Zhou C, et al. Keratins as the main component for the mechanical integrity of keratinocytes. Proc Natl Acad Sci USA. 2013;110(46):18513–8. doi:10.1073/pnas.1313491110.
Mierke CT. The fundamental role of mechanical properties in the progression of cancer disease and inflammation. Rep Prog Phys. 2014;77(7):076602. doi:10.1088/0034-4885/77/7/076602.
Diaz B, Yuen A, Iizuka S, Higashiyama S, Courtneidge SA. Notch increases the shedding of HB-EGF by ADAM12 to potentiate invadopodia formation in hypoxia. J Cell Biol. 2013;201(2):279–92. doi:10.1083/jcb.201209151.
Higashiyama S, Abraham JA, Miller J, Fiddes JC, Klagsbrun M. A heparin-binding growth factor secreted by macrophage-like cells that is related to EGF. Science. 1991;251(4996):936–9.
Higashiyama S, Iwamoto R, Goishi K, Raab G, Taniguchi N, Klagsbrun M, et al. The membrane protein CD9/DRAP 27 potentiates the juxtacrine growth factor activity of the membrane-anchored heparin-binding EGF-like growth factor. J Cell Biol. 1995;128(5):929–38.
Miyamoto S, Hirata M, Yamazaki A, Kageyama T, Hasuwa H, Mizushima H, et al. Heparin-binding EGF-like growth factor is a promising target for ovarian cancer therapy. Cancer Res. 2004;64(16):5720–7. doi:10.1158/0008-5472.CAN-04-0811.
Jezierska A, Motyl T. Matrix metalloproteinase-2 involvement in breast cancer progression: a mini-review. Med Sci Monit. 2009;15(2):RA32–40.
Li L, Zhao F, Lu J, Li T, Yang H, Wu C, et al. Notch-1 signaling promotes the malignant features of human breast cancer through NF-kappaB activation. PLoS ONE. 2014;9(4):e95912. doi:10.1371/journal.pone.0095912.
Wang Z, Zhang Y, Li Y, Banerjee S, Liao J, Sarkar FH. Down-regulation of Notch-1 contributes to cell growth inhibition and apoptosis in pancreatic cancer cells. Mol Cancer Ther. 2006;5(3):483–93. doi:10.1158/1535-7163.MCT-05-0299.
Inoue J, Gohda J, Akiyama T, Semba K. NF-kappaB activation in development and progression of cancer. Cancer Sci. 2007;98(3):268–74. doi:10.1111/j.1349-7006.2007.00389.x.
Dolcet X, Llobet D, Pallares J, Matias-Guiu X. NF-kB in development and progression of human cancer. Virchows Arch. 2005;446(5):475–82. doi:10.1007/s00428-005-1264-9.
Yamamoto M, Taguchi Y, Ito-Kureha T, Semba K, Yamaguchi N, Inoue J. NF-kappaB non-cell-autonomously regulates cancer stem cell populations in the basal-like breast cancer subtype. Nat Commun. 2013;4:2299. doi:10.1038/ncomms3299.
Zhang X, Chen T, Zhang J, Mao Q, Li S, Xiong W, et al. Notch1 promotes glioma cell migration and invasion by stimulating beta-catenin and NF-kappaB signaling via AKT activation. Cancer Sci. 2012;103(2):181–90. doi:10.1111/j.1349-7006.2011.02154.x.
Li L, Zhang J, Xiong N, Li S, Chen Y, Yang H, et al. Notch-1 signaling activates NF-kappaB in human breast carcinoma MDA-MB-231 cells via PP2A-dependent AKT pathway. Med Oncol. 2016;33(4):33. doi:10.1007/s12032-016-0747-7.
Hales EC, Orr SM, Larson Gedman A, Taub JW, Matherly LH. Notch1 receptor regulates AKT protein activation loop (Thr308) dephosphorylation through modulation of the PP2A phosphatase in phosphatase and tensin homolog (PTEN)-null T-cell acute lymphoblastic leukemia cells. J Biol Chem. 2013;288(31):22836–48. doi:10.1074/jbc.M113.451625.
Zhang W, Grivennikov SI. Top Notch cancer stem cells by paracrine NF-kappaB signaling in breast cancer. Breast Cancer Res: BCR. 2013;15(5):316. doi:10.1186/bcr3565.
Zhao F, Li L, Guan L, Yang H, Wu C, Liu Y. Roles for GP IIb/IIIa and alphavbeta3 integrins in MDA-MB-231 cell invasion and shear flow-induced cancer cell mechanotransduction. Cancer Lett. 2014;344(1):62–73. doi:10.1016/j.canlet.2013.10.019.
Giancotti FG, Ruoslahti E. Integrin signaling. Science. 1999;285(5430):1028–32.
Ruoslahti E. RGD and other recognition sequences for integrins. Annu Rev Cell Dev Biol. 1996;12:697–715. doi:10.1146/annurev.cellbio.12.1.697.
Deford P, Brown K, Richards RL, King A, Newburn K, Westover K, et al. MAGP2 controls Notch via interactions with RGD binding integrins: Identification of a novel ECM-integrin-Notch signaling axis. Exp Cell Res. 2016;341(1):84–91. doi:10.1016/j.yexcr.2016.01.011.
Hodkinson PS, Elliott PA, Lad Y, McHugh BJ, MacKinnon AC, Haslett C, et al. Mammalian NOTCH-1 activates beta1 integrins via the small GTPase R-Ras. J Biol Chem. 2007;282(39):28991–9001. doi:10.1074/jbc.M703601200.
Hodkinson PS, Elliott PA, Lad Y, McHugh BJ, MacKinnon AC, Haslett C, et al. Mammalian NOTCH-1 activates beta1 integrins via the small GTPase R-Ras. J Biol Chem. 2007;282(39):28991–9001. doi:10.1074/jbc.M703601200.
Kim KH, Chen CC, Alpini G, Lau LF. CCN1 induces hepatic ductular reaction through integrin alphavbeta(5)-mediated activation of NF-kappaB. J Clin Invest. 2015;125(5):1886–900. doi:10.1172/JCI79327.
Haque I, De A, Majumder M, Mehta S, McGregor D, Banerjee SK, et al. The matricellular protein CCN1/Cyr61 is a critical regulator of Sonic Hedgehog in pancreatic carcinogenesis. J Biol Chem. 2012;287(46):38569–79. doi:10.1074/jbc.M112.389064.
Legate KR, Montanez E, Kudlacek O, Fassler R. ILK, PINCH and parvin: the tIPP of integrin signalling. Nat Rev Mol Cell Biol. 2006;7(1):20–31. doi:10.1038/nrm1789.
Hsu EC, Kulp SK, Huang HL, Tu HJ, Chao MW, Tseng YC, et al. Integrin-linked kinase as a novel molecular switch of the IL-6-NF-kappaB signaling loop in breast cancer. Carcinogenesis. 2016;37(4):430–42. doi:10.1093/carcin/bgw020.
Sethi N, Dai X, Winter CG, Kang Y. Tumor-derived JAGGED1 promotes osteolytic bone metastasis of breast cancer by engaging Notch signaling in bone cells. Cancer Cell. 2011;19(2):192–205. doi:10.1016/j.ccr.2010.12.022.
Hsu EC, Kulp SK, Huang HL, Tu HJ, Salunke SB, Sullivan NJ, et al. Function of integrin-linked kinase in modulating the stemness of IL-6-abundant breast cancer cells by regulating gamma-secretase-mediated Notch1 activation in caveolae. Neoplasia. 2015;17(6):497–508. doi:10.1016/j.neo.2015.06.001.
Sansone P, Storci G, Tavolari S, Guarnieri T, Giovannini C, Taffurelli M, et al. IL-6 triggers malignant features in mammospheres from human ductal breast carcinoma and normal mammary gland. J Clin Invest. 2007;117(12):3988–4002. doi:10.1172/JCI32533.
Tahir SA, Yang G, Goltsov A, Song KD, Ren C, Wang J, et al. Caveolin-1-LRP6 signaling module stimulates aerobic glycolysis in prostate cancer. Cancer Res. 2013;73(6):1900–11. doi:10.1158/0008-5472.CAN-12-3040.
Lee C, Lee C, Lee S, Siu A, Ramos DM. The cytoplasmic extension of the integrin beta6 subunit regulates epithelial-to-mesenchymal transition. Anticancer Res. 2014;34(2):659–64.
Ding Y, Shen Y. Notch increased vitronection adhesion protects myeloma cells from drug induced apoptosis. Biochem Biophys Res Commun. 2015;467(4):717–22. doi:10.1016/j.bbrc.2015.10.076.
Masia A, Almazan-Moga A, Velasco P, Reventos J, Toran N, Sanchez de Toledo J, et al. Notch-mediated induction of N-cadherin and alpha9-integrin confers higher invasive phenotype on rhabdomyosarcoma cells. Br J Cancer. 2012;107(8):1374–83. doi:10.1038/bjc.2012.411.
Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11(9):597–610. doi:10.1038/nrg2843.
Sun DW, Zhang HD, Mao L, Mao CF, Chen W, Cui M, et al. Luteolin inhibits breast cancer development and progression in vitro and in vivo by suppressing Notch signaling and regulating MiRNAs. Cell Physiol Biochem. 2015;37(5):1693–711. doi:10.1159/000438535.
Wang Z, Li Y, Kong D, Ahmad A, Banerjee S, Sarkar FH. Cross-talk between miRNA and Notch signaling pathways in tumor development and progression. Cancer Lett. 2010;292(2):141–8. doi:10.1016/j.canlet.2009.11.012.
Mansour MR, Sanda T, Lawton LN, Li X, Kreslavsky T, Novina CD, et al. The TAL1 complex targets the FBXW7 tumor suppressor by activating miR-223 in human T cell acute lymphoblastic leukemia. J Exp Med. 2013;210(8):1545–57. doi:10.1084/jem.20122516.
Dorhoi A, Iannaccone M, Farinacci M, Fae KC, Schreiber J, Moura-Alves P, et al. MicroRNA-223 controls susceptibility to tuberculosis by regulating lung neutrophil recruitment. J Clin Invest. 2013;123(11):4836–48. doi:10.1172/JCI67604.
Barbarulo A, Grazioli P, Campese AF, Bellavia D, Di Mario G, Pelullo M, et al. Notch3 and canonical NF-kappaB signaling pathways cooperatively regulate Foxp3 transcription. J Immunol. 2011;186(11):6199–206. doi:10.4049/jimmunol.1002136.
Kumar V, Palermo R, Talora C, Campese AF, Checquolo S, Bellavia D, et al. Notch and NF-kB signaling pathways regulate miR-223/FBXW7 axis in T-cell acute lymphoblastic leukemia. Leukemia. 2014;28(12):2324–35. doi:10.1038/leu.2014.133.
Vandewalle C, Van Roy F, Berx G. The role of the ZEB family of transcription factors in development and disease. Cell Mol Life Sci: CMLS. 2009;66(5):773–87. doi:10.1007/s00018-008-8465-8.
Brabletz S, Bajdak K, Meidhof S, Burk U, Niedermann G, Firat E, et al. The ZEB1/miR-200 feedback loop controls Notch signalling in cancer cells. EMBO J. 2011;30(4):770–82. doi:10.1038/emboj.2010.349.
Yang Y, Ahn YH, Gibbons DL, Zang Y, Lin W, Thilaganathan N, et al. The Notch ligand Jagged2 promotes lung adenocarcinoma metastasis through a miR-200-dependent pathway in mice. J Clin Invest. 2011;121(4):1373–85. doi:10.1172/JCI42579.
Ohashi S, Natsuizaka M, Naganuma S, Kagawa S, Kimura S, Itoh H, et al. A NOTCH3-mediated squamous cell differentiation program limits expansion of EMT-competent cells that express the ZEB transcription factors. Cancer Res. 2011;71(21):6836–47. doi:10.1158/0008-5472.CAN-11-0846.
Shimono Y, Mukohyama J, Nakamura S, Minami H. MicroRNA regulation of human breast cancer stem cells. J Clin Med. 2015;. doi:10.3390/jcm5010002.
Xu YF, Hannafon BN, Ding WQ. microRNA regulation of human pancreatic cancer stem cells. Stem Cell Investig. 2017;4:5. doi:10.21037/sci.2017.01.01.
Chen J, Zhang H, Chen Y, Qiao G, Jiang W, Ni P, et al. miR-598 inhibits metastasis in colorectal cancer by suppressing JAG1/Notch2 pathway stimulating EMT. Exp Cell Res. 2017;352(1):104–12. doi:10.1016/j.yexcr.2017.01.022.
Wang XW, Xi XQ, Wu J, Wan YY, Hui HX, Cao XF. MicroRNA-206 attenuates tumor proliferation and migration involving the downregulation of NOTCH3 in colorectal cancer. Oncol Rep. 2015;33(3):1402–10. doi:10.3892/or.2015.3731.
Rizzo P, Miao H, D’Souza G, Osipo C, Song LL, Yun J, et al. Cross-talk between Notch and the estrogen receptor in breast cancer suggests novel therapeutic approaches. Cancer Res. 2008;68(13):5226–35. doi:10.1158/0008-5472.CAN-07-5744.
Clementz AG, Rogowski A, Pandya K, Miele L, Osipo C. NOTCH-1 and NOTCH-4 are novel gene targets of PEA3 in breast cancer: novel therapeutic implications. Breast Cancer Res: BCR. 2011;13(3):R63. doi:10.1186/bcr2900.
Stoeck A, Lejnine S, Truong A, Pan L, Wang H, Zang C, et al. Discovery of biomarkers predictive of GSI response in triple-negative breast cancer and adenoid cystic carcinoma. Cancer Discov. 2014;4(10):1154–67. doi:10.1158/2159-8290.CD-13-0830.
Samon JB, Castillo-Martin M, Hadler M, Ambesi-Impiobato A, Paietta E, Racevskis J, et al. Preclinical analysis of the gamma-secretase inhibitor PF-03084014 in combination with glucocorticoids in T-cell acute lymphoblastic leukemia. Mol Cancer Ther. 2012;11(7):1565–75. doi:10.1158/1535-7163.MCT-11-0938.
Bhagat TD, Zou Y, Huang S, Park J, Palmer MB, Hu C, et al. Notch pathway is activated via genetic and epigenetic alterations and is a therapeutic target in clear cell renal cancer. J Biol Chem. 2017;292(3):837–46. doi:10.1074/jbc.M116.745208.
Notch inhibitor shows modest efficacy. Cancer Discov. 2017;7(2):OF3. doi:10.1158/2159-8290.CD-NB2016-159.
Gavai AV, Quesnelle C, Norris D, Han WC, Gill P, Shan W, et al. Discovery of clinical candidate BMS-906024: a potent pan-Notch inhibitor for the treatment of leukemia and solid tumors. ACS Med Chem Lett. 2015;6(5):523–7. doi:10.1021/acsmedchemlett.5b00001.
Knoechel B, Bhatt A, Pan L, Pedamallu CS, Severson E, Gutierrez A, et al. Complete hematologic response of early T-cell progenitor acute lymphoblastic leukemia to the gamma-secretase inhibitor BMS-906024: genetic and epigenetic findings in an outlier case. Cold Spring Harb Mol Case Stud. 2015;1(1):a000539. doi:10.1101/mcs.a000539.
Acknowledgements
We would like to thank the National Natural Science Foundation of China (11772088, 31470906, 31700811, 11502049, 81671821, 31470959, 81471785), the Basic Research Program of Sichuan Science and Technology (2017JY0019, 2017JY0217), the China Postdoctoral Science Foundation (2016M592657), the Fundamental Research Funds for the Central Universities (ZYGX2016Z001, ZYGX2015J143) and the Postdoctoral Science Foundation of University of Electronic Science and Technology of China (Y0200623601737) for financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Li, L., Tang, P., Li, S. et al. Notch signaling pathway networks in cancer metastasis: a new target for cancer therapy. Med Oncol 34, 180 (2017). https://doi.org/10.1007/s12032-017-1039-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-017-1039-6