Abstract
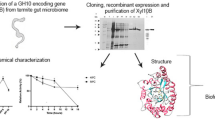
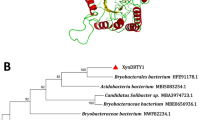
A novel xylanase gene, xyn-lxy, was cloned from a metagenomic fosmid library, which was previously constructed from the rumen contents of Hu sheep and was functionally characterized in Escherichia coli. The open reading frame was composed of 1923 bp and encoded for 640 amino acids, including a catalytic domain of glycosyl hydrolase family 10 and carbohydrate-binding module 9. The gene showed 97 % identity with uncultured bacterium Contig1552 but low similarity with xylanases from known cellulolytic-degrading microorganisms in the rumen. The recombinant XYN-LXY showed a specific activity of 664.7 U mg−1. The optimal temperature and pH of the enzyme were 50 °C and 6.0, respectively. Specifically, XYN-LXY was exclusively activated by Mn2+ among all of the cations and reducing agents tested in this study. An enzymatic hydrolysis assay revealed that XYN-LXY degraded birchwood xylan into xylooligosaccharide with a low degree of polymerization. After incubation for 4 h, the concentration of the dominant product, xylobiose, was 2.297 ± 0.175 mg ml−1 (74.07 % of total product) followed by xylose with a concentration of 0.656 ± 0.010 mg ml−1 (21.14 % of total product). The XYN-LXY exhibited deep degradation effects on the xylan substrate, which were rarely observed with endo-xylanase, making it a promising candidate for industrial application, especially in biofuel production.





Similar content being viewed by others
References
Chiniquy, D., Sharma, V., Schultink, A., Baidoo, E. E., Rautengarten, C., Cheng, K., Carroll, A., Ulvskov, P., Harholt, J., Keasling, J. D., Pauly, M., & Ronald, P. C. (2012). XAX1 from glycosyltransferase family 61 mediates xylosyltransfer to rice xylan. Proceedings of the National Academy of Sciences, 109(42), 17117–17122.
Haki, G. D., & Rakshit, S. K. (2003). Developments in industrially important thermostable enzymes: a review. Bioresource Technology, 89(1), 17–34.
Collins, T., Gerday, C., & Feller, G. (2005). Xylanase, xylanase families and extremophilic xylanases. FEMS Microbiology Reviews, 29(1), 3–23.
Kulkarni, N., Shendye, A., & Rao, M. (1999). Molecular and biotechnological aspects of xylanases. FEMS Microbiology Reviews, 23(4), 411–456.
Watanabe, S., Kodaki, T., & Makino, K. (2005). Complete reversal of coenzyme specificity of xylitol dehydrogenase and increase of thermostability by the introduction of structural zinc. Journal of Biological Chemistry, 280(11), 10340–10349.
Bocchini, D. A., Gomes, E., & Silva, R. D. (2008). Xylanase production by Bacillus circulans Dl using maltose as carbon source. Applied Biochemistry and Biotechnology, 146(1-3), 29–37.
Miyazaki, K., Takenouchi, M., Kondo, H., Noro, N., Suzuku, M., & Tsuda, S. (2006). Thermal stabilization of Bacillus subtilis family-11 xylanase by directed evolution. Journal of Biological Chemistry, 281(15), 10236–10242.
Stephens, D. E., Singh, S., & Rumbold, K. (2009). Error-prone PCR of a fungal xylanase for improvement of its alkaline and thermal stability. FEMS Microbiology Letters, 293(1), 42–47.
Wang, Q., Zhao, L. L., Sun, J. Y., Liu, J. X., & Weng, X. Y. (2012). Enhancing catalytic activity of a hybrid xylanase through single substitution of Leu to Pro near the active site. World Journal of Microbiology and Biotechnology, 28, 929–935.
Silva, J. P. A., Mussatto, S. I., Roberto, I. C., & Teixeira, J. A. (2011). Ethanol production from xylose by Pichia stipitis NRRLY-7124 in a stirred tank bioreactor. Brazilian Journal of Chemical Engineering, 28(3), 151–156.
Cardona, C. A., Quintero, J. A., & Paz, I. C. (2010). Production of bioethanol from sugarcane bagasse: status and perspectives. Bioresource Technology, 101(13), 4754–4766.
Matsushika, A., Watanabe, S., Kodaki, T., Makino, K., & Sawayama, S. (2008). Bioethanol production from xylose by recombinant Saccharomyces cerevisiae expressing xylose reductase, NADP+-dependent xylitol dehydrogenase, and xylulokinase. Journal of Bioscience and Bioengineering, 105(3), 296–299.
Zhang, W., & Geng, A. L. (2012). Improved ethanol production by a xylose fermenting recombinant yeast strain constructed through a modified genome shuffling method. Biotechnology for Biofuels, 5(1), 46–56.
de Almeida, M. N., Guimarães, V. M., Falkoski, D. L., Visser, E. M., Siqueira, G. A., Milagres, A. M., & de Rezende, S. T. (2013). Direct ethanol production from glucose, xylose and sugarcane bagasse by the corn endophytic fungi Fusarium verticillioides and Acremonium zeae. Journal of Biotechnology, 168(1), 71–77.
Shi, J., Zhang, M., Zhang, L., Wang, P., Jiang, L., & Deng, H. (2014). Xylose-fermenting Pichia stipitis by genome shuffling for improved ethanol production. Microbial Biotechnology, 7(2), 90–99.
Wei, N., Quarterman, J., Kim, S. R., Cate, J. H., & Jin, Y. S. (2013). Enhanced biofuel production through coupled acetic acid and xylose consumption by engineered yeast. Nature Communications, 4, 2580.
Sun, J. Y., Liu, M. Q., Xu, Y. L., Xu, Z. R., Pan, L., & Gao, H. (2005). Improvement of the thermostability and catalytic activity of a mesophilic family 11 xylanase by N-terminus replacement. Protein Expression and Purification, 42, 122–130.
Liu, M. Q., & Liu, G. F. (2008). Expression of recombinant Bacillus licheniformis xylanase A in Pichia pastoris and xylooligosaccharides released from xylans by it. Protein Expression and Purification, 57(2), 101–107.
Zhang, M., Jiang, Z., Yang, S., Hua, C., & Li, L. (2010). Cloning and expression of a Paecilomyces thermophila xylanase gene in E. coli and characterization of the recombinant xylanase. Bioresource Technology, 101(2), 688–695.
Zhang, J., Siika-aho, M., Puranen, T., Tang, M., Tenkanen, M., & Viikari, L. (2011). Thermostable recombinant xylanases from Nonomuraea flexuosa and Thermoascus aurantiacus show distinct properties in the hydrolysis of xylans and pretreated wheat straw. Biotechnology for Biofuels, 4(1), 12.
Chen, C. C., Luo, H., Han, X., Lv, P., Ko, T. P., Peng, W., Huang, C. H., Wang, K., Gao, J., Zheng, Y. Y., Yang, Y. Y., Zhang, J. Y., Yao, B., & Guo, R. T. (2014). Structural perspectives of an engineered β-1, 4-xylanase with enhanced thermostability. Journal of Biotechnology, 189, 175–182.
Fan, G., Yang, S., Yan, Q., Guo, Y., Li, Y., & Jiang, Z. (2014). Characterization of a highly thermostable glycoside hydrolase family 10 xylanase from Malbranchea cinnamomea. International Journal of Biological Macromolecules, 70, 482–489.
Gao, H., Yan, P., Zhang, B., & Shan, A. (2014). Expression of Aspergillus niger IA-001 Endo-β-1, 4-xylanase in Pichia pastoris and analysis of the enzymic characterization. Applied Biochemistry and Biotechnology, 173(8), 2028–2041.
Kim do, Y., Shin, D. H., Jung, S., Lee, J. S., Cho, H. Y., Bae, K. S., Sung, C. K., Rhee, Y. H., Son, K. H., & Park, H. Y. (2014). Biocatalytic properties and substrate-binding ability of a modular GH10 β-1,4-xylanase from an insect-symbiotic bacterium, Streptomyces mexicanus HY-14. Journal of Microbiology, 52(10), 863–870.
Morrison, M., Adams, S. E., Nelson, K. E., & Attwood, G. T. (2005). Metagenomic analysis of the microbiomes in ruminants and other herbivores. In P. S. Makkar H & C. S. McSweeney (Eds.), Methods in gut microbial ecology for ruminants (pp. 209–220). Netherlands: Springer.
Wang, J. K., An, P. P., Chen, Z. M., Ye, J. A., & Liu, J. X. (2010). Construction and analysis of fosmid library of rumen microbiota of Hu sheep. Chinese Journal of Animal Nutrition, 22, 341–345.
Wang, J. K., Sun, Z. Y., Zhou, Y., Wang, Q., Ye, J. A., Chen, Z. M., & Liu, J. X. (2012). Screening of a xylanase clone from a fosmid library of rumen microbiota in Hu sheep. Animal Biotechnology, 23(3), 156–173.
Teather, R. M., & Wood, P. J. (1982). Use of Congo red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Applied and Environmental Microbiology, 43(4), 777–780.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227(5259), 680–685.
Bailey, M. J., Biely, P., & Poutanen, K. (1992). Interlaboratory testing of methods for assay of xylanase activity. Journal of Biotechnology, 23(3), 257–270.
Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Analytical Biochemistry, 72(1), 248–254.
Wang, L., Hatem, A., Catalyurek, U. V., Morrison, M., & Yu, Z. (2013). Metagenomic insights into the carbohydrate-active enzymes carried by the microorganisms adhering to solid digesta in the rumen of cows. PLoS ONE, 8(11), e78507.
Han, H., You, S., Zhu, B., Fu, X., Sun, B., Qiu, J., Yu, C., Chen, L., Peng, R., & Yao, Q. (2014). Characterization and high expression of recombinant Ustilago maydis xylanase in Pichia pastoris. Biotechnology Letters, 37(3), 697–703.
Wang, G., Luo, H., Wang, Y., Huang, H., Shi, P., Yang, P., Meng, K., Bai, Y., & Yao, B. (2011). A novel cold-active xylanase gene from the environmental DNA of goat rumen contents: direct cloning, expression and enzyme characterization. Bioresource Technology, 102(3), 3330–3336.
Knob, A., & Carmona, E. C. (2010). Purification and characterization of two extracellular xylanases from Penicillium sclerotiorum: a novel acidophilic xylanase. Applied Biochemistry and Biotechnology, 162(2), 429–443.
Wang, W., Wang, Z., Cheng, B., Zhang, J., Li, C., Liu, X., & Yang, C. (2014). High secretory production of an alkaliphilic actinomycete xylanase and functional roles of some important residues. World Journal of Microbiology and Biotechnology, 30(7), 2053–2062.
Guo, B., Chen, X. L., Sun, C. Y., Zhou, B. C., & Zhang, Y. Z. (2009). Gene cloning, expression and characterization of a new cold-active and salt-tolerant endo-β-1, 4-xylanase from marine Glaciecola mesophila KMM 241. Applied Microbiology and Biotechnology, 84(6), 1107–1115.
Walia, A., Mehta, P., Chauhan, A., Kulshrestha, S., & Shirkot, C. K. (2014). Purification and characterization of cellulase-free low molecular weight endoβ-1, 4 xylanase from an alkalophilic Cellulosimicrobium cellulans CKMX1 isolated from mushroom compost. World Journal of Microbiology and Biotechnology, 30(10), 2597–2608.
Zhang, W., Lou, K., & Li, G. (2010). Expression and characterization of the Dictyoglomus thermophilum Rt46B. 1 xylanase gene (xynB) in Bacillus subtilis. Applied Biochemistry and Biotechnology, 160(5), 1484–1495.
Zhao, L., Meng, K., Bai, Y., Shi, P., Huang, H., Luo, H., Wang, Y., Yang, P., Song, W., & Yao, B. (2013). Two family 11 xylanases from Achaetomium sp. Xz-8 with high catalytic efficiency and application potentials in the brewing industry. Journal of Agricultural and Food Chemistry, 61(28), 6880–6889.
Guo, B., Li, P. Y., Yue, Y. S., Zhao, H. L., Dong, S., Song, X. Y., Sun, C. Y., Zhang, W. X., Chen, X. L., Zhang, X. Y., Zhou, B. C., & Zhang, Y. Z. (2013). Gene cloning, expression and characterization of a novel xylanase from the marine bacterium, Glaciecola mesophila KMM241. Marine Drugs, 11(4), 1173–1187.
Cheng, F., Sheng, J., Dong, R., Men, Y., Gan, L., & Shen, L. (2012). Novel xylanase from a holstein cattle rumen metagenomic library and its application in xylooligosaccharide and ferulic acid production from wheat straw. Journal of Agricultural and Food Chemistry, 60(51), 12516–12524.
Ali, M. K., Rudolph, F. B., & Bennett, G. N. (2005). Characterization of thermostable Xyn10A enzyme from mesophilic Clostridium acetobutylicum ATCC 824. Journal of Industrial Microbiology & Biotechnology, 32(1), 12–18.
Lee, S. F., Forsberg, C. W., & Rattray, M. (1987). Purification and characterization of two endoxylanases from Clostridium acetobutylicum ATCC 824. Applied and Environmental Microbiology, 53(4), 644–650.
Herbers, K., Wilke, I., & Sonnewald, U. (1995). A thermostable xylanase from Clostridium thermocellum expressed at high levels in the apoplast of transgenic tobacco has no detrimental effects and is easily purified. Nature Biotechnology, 13(1), 63–66.
Li, N., Meng, K., Wang, Y., Shi, P., Luo, H., Bai, Y., Yang, P., & Yao, B. (2008). Cloning, expression, and characterization of a new xylanase with broad temperature adaptability from Streptomyces sp. S9. Applied Microbiology and Biotechnology, 80(2), 231–240.
Acknowledgments
The authors acknowledge the financial support from the Innovation Team Program of Zhejiang province (2011R50025) and the Importation and Development of High-Caliber Talents Project of Beijing Municipal Institutions (CIT&CD20130324).
Compliance with Ethical Standards
ᅟ
Conflict of Interest
The authors declare that they have no competing interests.
Authors’ Contribution
Qian Wang drafted the manuscript. Yang Luo and Bo He carried out the studies and contributed to the drafting of the manuscript. Jia-Kun Wang, Jian-Xin Liu, and Lin-Shu Jiang participated in the project design and manuscript preparation. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Qian Wang and Yang Luo contributed equally to this work.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplemental Fig. S1
Screening of positive transformants by PCR. Lane 1-4: different E. coli transformants. Plasmids isolated fromS different colonies served as templates for PCR amplification using Xyn-lxy-F and Xyn-lxy-R primers. (DOC 85 kb)
Supplemental Fig. S2
Lineweaver-Burk plot of XYN-LXY. 0.4 to 15 mg/ml birchwood xylan was used as the substrate for the determination of K m and V max . (DOC 26 kb)
Rights and permissions
About this article
Cite this article
Wang, Q., Luo, Y., He, B. et al. Characterization of a Novel Xylanase Gene from Rumen Content of Hu Sheep. Appl Biochem Biotechnol 177, 1424–1436 (2015). https://doi.org/10.1007/s12010-015-1823-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1823-8