Abstract
Exploration of avian gametologous genes, i.e., homologous genes located on both the Z and W chromosomes, provides a crucial information about the underlying mechanism pertaining to the evolution of these chromosomes. The domestic chicken (Gallus gallus (Linnaeus 1758); GGA) traditionally serves as the primary reference subject of these comparative cytogenomic studies. Using bioinformatic, molecular (overgo BAC library scanning), and cytogenetic (BAC-based FISH) techniques, we have investigated in detail a pair of UBE2R2/UBE2R2L gametologs. By screening a gridded genomic jungle fowl BAC library, CHORI-261, with a short labeled UBE2R2L gene fragment called overgo probe, we detected seven specific clones. For three of them, CH261-019I23, CH261-105E16, and CH261-114G22, we identified their precise cytogenetic location on the Gallus gallus W chromosome (GGAW). They also co-localized with the UBAP2L2 gene on the, as was shown previously, along with the CH261-053P09 BAC clone also containing the GGAW-specific UBE2R2L DNA sequence. The fine mapping of the UBE2R2/UBE2R2L homologs in the chicken genome also shed the light on comparative cytogenetic aspects in birds. Our findings provided further evidence that bird genomes moderately changed only during evolution and are suitable for successful use of interspecies hybridization using both overgo-based BAC library screen and BAC-based FISH.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
It has been suggested that avian Z- and W-chromosomes originated from one autosome pair, but a different one from which the mammalian X- and Y-chromosomes evolved (Schmid et al. 2000). Some lines of evidence, however, lead to alternative suggestions concerning the evolution of bird and mammal chromosomes (Stiglec et al. 2007a). The W-chromosome in neognathous avian species has some features similar to the mammalian Y-chromosome, i.e., it is gene-poor and largely heterochromatic. Highly repetitive sequences belonging to XhoI-, EcoRI- and SspI-families represent about 80% of the DNA in the chicken (Gallus gallus (Linnaeus, 1758); GGA) W chromosome (GGAW). Only about 10 Mb is represented by non-repetitive DNA (Itoh and Mizuno 2002).
In all neognathous birds studied to date, the Z–W pair of sex chromosomes (gonosomes) shows strictly localized recombination in a very short pseudoautosomal region (PAR; reviewed in Pigozzi and Solari 2005). Differentiation of the Z and W chromosomes in modern birds is thought to be a result of progressive and stepwise cessation of meiotic recombination (Lawson Handley et al. 2004; Schmid et al. 2005).
A significant part of GGAW was misassembled in initial sequencing efforts, that is, it was based exclusively on repetitive sequences, which were subsequently found to be present on other chromosomes (Stiglec et al. 2007b). FISH mapping of the bacterial artificial chromosome (BAC) clones, which were putatively thought to be GGAW-specific, to its counterpart Z chromosome (GGAZ) allowed researchers to clarify the genic composition of GGAW (Stiglec et al. 2007b). Thus, several W-linked genes in this region have respective homologs on GGAZ (e.g., CHD1, HINT, SPIN, UBAP2, and ATP5A1), forming pairs of gametologous genes and reflecting their common origin from ancestral homologous chromosome pairs (Schmid et al. 2005).
One of these gametologous genes is UBE2R2 encoding the ubiquitin conjugating enzyme E2 R2 that is involved in modification of proteins with ubiquitin or ubiquitin-like proteins via an E1–E2–E3 cascade, which is crucial in many signaling networks (Jin et al. 2007). Being originally unmapped in the chicken, it was presumably localized on GGAZ judging from the comparative map analysis: the human ortholog maps to the HSA9 region that corresponds to GGAZ. In the recent chicken genome assembly GRCg6a (GRCg6a 2018), it is located on GGAZ, while its W-linked counterpart, UBE2R2L, is mapped to GGAW within the chicken PAR (Fig. 1).
Here, we investigated the chicken UBE2R2 genes located on GGAZ and GGAW. For this purpose, we implemented an effective genome analysis and mapping pipeline using the appropriate bioinformatic tools, molecular techniques (e.g., overgo hybridization), and cytogenetic methods such as a BAC-based fluorescence in situ hybridization (FISH). The relevant implications for, and insights into, evolutionary aspects of bird gonosome organization were inferred providing an additional information for further discussion and research in this area of avian biology.
Materials and methods
Avian species
In this work, we conducted bioinformatic, molecular and cytogenetic analyses focusing on the chicken (Gallus gallus; order Galliformes) genome best studied among all birds. For further multifaceted comparative investigation, data obtained from our previous published and unpublished research on the following five bird species (with their respective Latin name and order in the parentheses) were also used: turkey (Meleagris gallopavo Linnaeus, 1758; Galliformes), Japanese quail (Coturnix japonica Temminck & Schlegel, 1848; Galliformes), Sunda zebra finch (Taeniopygia guttata (Vieillot, 1817); Passeriformes), white-throated sparrow (Zonotrichia albicollis (Gmelin, 1789); Passeriformes), California condor (Gymnogyps californianus (Shaw, 1797); Cathartiformes; alternatively, Falconiformes, Ciconiiformes or Accipitriformes), and black stork (Ciconia nigra (Linnaeus, 1758); Ciconiiformes).
Genome analysis and mapping pipeline
To explore the pair of UBE2R2 genes, we developed and used an in-house genome analysis and mapping pipeline that encompassed the following components and applications: bioinformatics analyses → overgo hybridization → BAC-based FISH. The bioinformatic toolbox involved use of databases, sequence alignment tools, in silico overgo probe design, etc. Mining for UBE2R2 and related sequences was performed using NCBI- (e.g., Altschul et al. 1997; Wheeler et al. 2000; Cuff et al. 2000; Maglott et al. 2011; Sayers et al. 2021), UCSC- (Navarro Gonzalez et al. 2021) and Ensembl-based (Howe et al. 2021) genome browsers. Retrieved sequences were aligned with ClustalW (Thompson et al. 1994) and COBALT (Papadopoulos and Agarwala 2007). Contig with BACs were detected in the Chicken FPC (ChickFPC) database (Schmid et al. 2005; Kerstens and Groenen 2004–2008). The employed molecular (overgo hybridization) and cytogenetic (FISH) procedures are outlined below (see also Online resources 1 and 2 for more detail).
BAC library scanning
The chicken genomic BAC library CHORI-261 was prepared from a red jungle fowl DNA sample (Nefedov et al. 2003; BACPAC Genomics 2020) and screened using the overgo probe hybridization approach as described in detail elsewhere (Romanov et al. 2003; Romanov and Dodgson 2006). Briefly, ~ 40-bp overgo probes comprised of two synthetic oligonucleotides with a complementary 8-bp region were designed in silico using consequently RepeatMarker (Smit et al. 1996–2010) and Overgo (Cai et al. 1998) web tools (Online resource 2). These probes were radioactively labeled with [32P]-deoxynucleotide triphosphates and hybridized at 60 °C to BAC clone array filters. The screening resulted in positively hybridized BAC clones with their IDs deposited electronically in the Michigan State University-hosted Database of BACs Assigned to Chicken Genes and Markers (Resources 2000–2013). Other avian BAC libraries were also screened using cross-species hybridization approach (Romanov and Dodgson 2006; Romanov et al. 2006, 2009, 2011).
FISH procedure
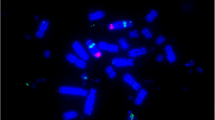
The FISH experiments were performed following the protocol as specified elsewhere (e.g., Sazanov et al. 2006; Blagoveschensky et al. 2011; see also Online resource 3 for further detail). In short, the 96-h Brown Leghorn chick embryo cells were used to generate preparations of mitotic chromosomes using hypotonic treatment, fixation, and colchicine incubation. The double-color FISH using the respective BAC clone DNA was carried out basically in accordance with the instructions reported elsewhere (Florijn et al. 1995). A fluorescent microscopic workstation (Ista, St. Petersburg, Russia) equipped with a CCD camera and the software program VideoTest-FISH was used to record hybridization signals.
Results
Overgo design and hybridization using chicken BAC library
For a starting point of the in silico overgo probe design, we used the mRNA sequence NM_017811.4 of the human orthologous UBE2R2 gene (located on chromosome 9p13.3) as a BLAST® search query and identified the best match amongst available chicken (Expressed Sequence Tags) EST sequences that was the chicken cDNA clone BU122359 (see for more detail Online resource 1: Fig. S1-1). In the Ensembl database conforming to an older chicken assembly, BU122359 matched one W_random and one Z_random contigs that might be indicative of two UBE2R2 homologs that exist on both GGAZ and GGAW (see for more detail Online resource 1: Table S1).
The designed 38-mer overgo (GCAATGAGGAGTCGTGACCTGCTCTCTATGCTGTTGTA) only matched one W_random contig suggesting that it uniquely conformed to the W-linked UBE2R2L gene (see Online resource 1). The overgo probe was labeled and applied to screening the gridded genomic BAC library CHORI-261. As a result, we found the following seven specific clones: CH261-019I23, CH261-053P09, CH261-105E16, CH261-114G22, CH261-124O24, CH261-082I17, and CH261-095J14 (Table 1, Fig. 2).
Results of the CHORI-261 BAC library screen using the W-linked UBE2R2L-specific overgo hybridized to three BAC array filters. Arrows indicate the positive hybridization signals that corresponded to the respective BAC clones: a CH261-019I23; b CH261-053P09; and c CH261-114G22 (upper arrow) and CH261-105E16 (lower arrow). The same four BACs were also identified as positives for UBAP2L2 (or UBAP2W)
Out of these seven BACs, there were four clones, CH261-105E16, CH261-114G22, CH261-019I23 and CH261-053P09, that also hybridized to the UBAP2L2 (or UBAP2W) specific overgo probe. These appeared to overlap with both the UBE2R2L and UBAP2L2 genes that can be confirmed by their close localization in the chicken genome assembly GRCg6a (Fig. 3a; Online resource 2: Fig. S2-1a). Indeed, the distance between these two genes was less than 20 Kb, with that between their Z-linked counterparts being even smaller (~ 2.5 Kb), making possible for a BAC clone to encompass the two neighboring genes.
Ensembl genome browser views for the GGAW (a) and GGAZ (b) genomic regions showing close localization of two gametologous gene pairs in the chicken genome assembly GRCg6a. Gene positions: a UBAP2L2 at 2,472.0.48,083 bp (forward strand) vs UBE2R2L (or ENSGALG00000048542) at 67,938.0.141,552 bp (reverse strand; green highlight); b UBE2R2 at 7,304,045.0.7,356,140 bp (forward strand; green highlight) vs UBAP2 at 7,358,707.0.7,496,787 bp (reverse strand)
Overgo-based cross-species hybridization
Using the chicken UBE2R2L-specific overgo, we, in addition to the chicken BAC library, successfully screened four other avian libraries (Table 2). The number of positive clones ranged between 1 to 15. For instance, in the California condor BAC library CHORI-262, we identified one positive clone CH262-037I12 that was putatively mapped in silico to the W chromosome and was further used for FISH on the Z and W in this endangered species and other birds (Romanov et al. 2006, 2009; Modi et al. 2009).
BAC-based FISH
We used three out of the seven UBE2R2L-specific chicken BACs, i.e., CH261-019I23, CH261-105E16 and CH261-114G22, to identify precise cytogenetic location of UBE2R2L on GGAW (Table 1). Previously, these three clones also produced co-localized signals for UBAP2L2 (or UBAP2W) on GGAW but not for its Z-linked counterpart UBAP2 (or UBAP2Z; Sazanov et al. 2006; Blagoveschensky et al. 2011). The fine FISH mapping of the three BAC clones to GGAW proved unequivocally that they contained sequences of both UBE2R2L and UBAP2L2. Examples of mapping CH261-114G22 to GGAW using double-color FISH, in combination with different ATP5A1-positive BACs, are presented in Fig. 4. Remarkably, the ATP5A1-specific clone CH261-064F22, detected cytogenetically on GGAZ (Flpter, 0.14 ± 0.033; Blagoveschensky et al. 2011), also co-localized on GGAW, along with the CH261-114G22 BAC clone containing the GGAW-specific UBAP2L2 DNA sequence (Fig. 4e).
In situ hybridization of chicken mitotic chromosomes with BAC clone DNA. a, b A metaphase plate showing localization for CH261-033F10 (a, ATP5A1 on GGAZ and GGAW) and CH261-114G22 (b, UBE2R2L and UBAP2L2 on GGAW). c, d A metaphase plate representing TAM31-099N01 (c, ATP5A1 on GGAZ) and CH261-114G22 (d, UBE2R2L and UBAP2L2 on GGAW). e, f Signal location for CH261-064F22 (green, ATP5A1 on GGAZ and GGAW) and co-localized CH261-114G22 (red, UBE2R2L and UBAP2L2 on GGAW); and TAM31-100C09 (f, green, ATP5A1 on GGAZ) and CH261-114G22 (red, UBE2R2L and UBAP2L2 on GGAW); same metaphase plate as c, d. Signals of hybridization are shown by the arrows. Photographs a–d adopted from Blagoveschensky et al. (2011)
Discussion
The origin of sex gonosomes in birds, mammals and reptiles appear to be different. While mammals use the XX/XY system, the genetic sex determination independently evolved in birds, and utilizes the ZZ/ZW system (Matsubara et al. 2006). Effective study of the gametologous genes underlying the ZZ/ZW system, especially for cross-species comparison, requires the availability of appropriate molecular and cytogenetic tools. In this study, we demonstrated efficiency in using gene sequence-derived overgos to discover specific BACs that, in turn, are applicable to FISH map gametologs to one or both gonosomes.
Within-species use of overgos to scan BAC libraries is, as a rule, a straightforward and reliable technique. Overgo-based interspecies hybridization success rate may vary due to a number of reasons: (1) evolutionary distance from the chicken, a source for the used overgo probes, and respective sequence divergence, especially within non-coding regions (Modi et al. 2009); (2) genome coverage, i.e., genome representation in a BAC library; (3) variation in quality of BAC DNAs arrayed on the gridded filters; and (4) specific and changeable BAC filter hybridization conditions in an individual screen experiment and in a particular lab.
Comparative chromosome painting and FISH mapping of DNA clones, including BACs, have been broadly employed for studying chromosome rearrangements between chicken and other birds (e.g., Griffin et al. 1999; Shibusawa et al. 2002, 2004). By implementing BAC-based FISH and using zebra finch BACs for a number of Z chromosome genes, including UBE2R2, Itoh et al. (2006) were able to map them to the zebra finch Z and W chromosomes. We previously mapped pairs of the chicken gametologous genes located in the PAR: ATP5A1Z—ATP5A1W and UBAP2–UBAP2L2 (Sazanov et al. 2006; Blagoveschensky et al. 2011). Here, we cytogenetically localized in the chicken genome UBE2R2L, a previously unmapped W-linked homolog of another gametologous pair, UBE2R2–UBE2R2L, but were unable to assign UBE2R2L-containing BACs to the Z gametolog. On the other hand, BAC filter hybridization using a lambda phage clone containing the W-linked turkey AD012W gene, also known as LOC100303669 or UBAP2W (accession numbers AY188758; see for further details Harry et al. 2003; Sazanov et al. 2006; Romanov et al. 2019), resulted in discovery of five positive clones in the Texas A&M University genomic jungle fowl BAC library (code TAM31; Lee et al. 2003; Ren et al. 2003). Sazanov et al. (2006) and Blagoveschensky et al. (2011) cytogenetically assigned these BACs to Z but not to W in the chicken and Japanese quail. That could be because the turkey W-linked lambda phage clone cross-hybridized to the GGAZ-positive chicken BACs but not to GGAW-specific ones. Blagoveschensky et al. (2011) also mapped three other W-specific chicken clones to both Z and W in the chicken and Japanese quail. This ambiguity in applying successfully clones as FISH probes to another heterochromosome within and between species might depend on how different sequences of particular genomic regions on Z and W are, and, to some extent, how rigorous or relaxed DNA–DNA hybridization conditions are in a given experiment. In any event, it can be deduced that BAC clones cross-hybridizing to opposite gonosomes carry sequences on the Z and W chromosomes with a lesser degree of divergence, which conform to areas of the gonosomes that have recently stopped recombining and are located inside the PAR (Blagoveschensky et al. 2011).
Using the chicken UBE2R2 mRNA sequence as a query, we additionally performed a BLAST® search as can be viewed in Fig. 5. In this BLAST® tree comprising 25 birds, a basal group was represented by three galliform species, i.e., the chicken, turkey, and rock ptarmigan (Lagopus muta), suggesting that their UBE2R2 genes were most ancient by origin among the compared species. There was another gene cluster for a younger group of waterfowl including five species of geese, swans and ducks (order Anseriformes). A greater divergence relative to the chicken was characteristic of more recent evolutionary clades such as Accipitriformes, Apodiformes, Falconiformes, Psittaciformes and Passeriformes. Tinamou (Palaeognathae) got into this large group only due to availability of a partial mRNA sequence for this species that biased its divergence estimate, although in the respective protein tree (Online resource 1: Fig. S1-5b) it occupied a more basal position. Also, we noticed that some members of a younger order Passeriformes were scattered among various evolutionary tree branches (Fig. 5), which might reflect a perceptible diversification of genome organization within this clade. Similar relationships between various avian clades were also noticed based on UBE2R2/UBE2R2L gene/protein sequence alignments (Online resource 1: Figs. S1-4 and S1-5). The observed patterns of the UBE2R2/UBE2R2L-related molecular evolution was largely in agreement with the current general understanding of evolution and taxonomy in the class Aves.
A distance tree of BLAST® search results produced using BLAST® pairwise alignments between the chicken UBE2R2L gene sequence query (XM_025144594.3; 3185 bp) and 24 other sequences. The search was limited to records that include birds (Aves; taxid:8782). Database: nt; tree method: Fast Minimum Evolution; maximum sequence difference: 0.75. The query is highlighted in yellow; green nodes denote hawks and eagles
In a previous study (Modi et al. 2009), the California condor clone CHORI-262 37I12 positive for the chicken UBE2R2L overgo was cytogenetically mapped by FISH to the condor Z-linked homolog, with no signal being determined on the W chromosome. Similarly, use of the same condor W-linked clone on the black stork chromosomes revealed its localization on Zq near telomere (though the stork W was not looked at; Modi et al. 2009; Fig. 6). In the chicken, the UBE2R2 gene is located on Zp (Figs. 1 and 3) and demonstrates a pericentric inversion(s) in the zebra finch (Itoh et al. 2006). Similar pericentric inversion(s) of a much greater degree obviously occurred in the California condor and black stork, both species attributed to Ciconiiformes (see, however, a note on this controversial attribution for the condor in Romanov et al. 2009), with their UBE2R2 genes being on Zq, close to the telomere. The revealed inversion patterns are indicative of specific intrachromosomal rearrangements in the Z chromosome evolution in these four avian lineages. These observations imply that the Z chromosome gene content has been conserved during the 90–100 million years since the split between the chicken and other three lineages, while the chromosome has undergone significant reorganization (Itoh et al. 2006).
Comparisons of the Z chromosomes in the zebra finch, chicken, California condor, and black stork using ideograms. Maps for the stork and condor were created using the prior FISH results (Modi et al. 2009), whereas those for the zebra finch and chicken are based on genome sequences
Conclusions
In the present study, we established a reliable pipeline of bioinformatic, molecular and mapping tools used for genomic analysis and mapping to explore avian sex chromosomes (gonosomes) and genes they harbor. We bioinformatically evaluated in the chicken genome a previously understudied pair of gametologous genes (i.e., Z- and W-linked homologs), UBE2R2 and UBE2R2L, and, using FISH, mapped the W-linked counterpart, UBE2R2L. Three of the seven used BAC clones were also cytogenetically assigned to GGAW and co-localized with another sex chromosome-linked UBAP2L2 gene. Although the chicken UBE2R2L (i.e. W-linked) clones were FISH mapped to GGAW, they did not cross-hybridize to GGAZ, which was in accord with other similar observations on inconsistent cross-hybridization of specific clones between the two heterochromosomes within and among species. Overall, this and other relevant investigations have established that cross-heterochromosome and interspecies hybridization can be useful for the scanning of avian BAC libraries and the FISH mapping of individual genes (including gametologs), genomic regions (like PAR), and whole chromosomes. This integrative research approach is instrumental in comprehending the evolutionary history in birds that will be further illuminated by future cytogenomic studies of various Aves clades.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
Change history
26 May 2023
A Correction to this paper has been published: https://doi.org/10.1007/s11756-023-01446-y
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
BACPAC Genomics (2020) CHORI-261: Chicken BAC Library. BACPAC Genomics, Emeryville, CA, USA. https://bacpacresources.org/library.php?id=120. Accessed 14 Aug 2022
Blagoveschensky IY, Sazanova AL, Stekol’nikova VA, Fomichev KA, Barkova OY, Romanov MN, Sazanov AA (2011) Investigation of pseudoautosomal and bordering regions in avian Z and W chromosomes with the use of large insert genomic BAC clones. Russ J Genet 47:272–278. https://doi.org/10.1134/S1022795411020050
Cai WW, Reneker J, Chow CW, Vaishnav M, Bradley A (1998) An anchored framework BAC map of mouse chromosome 11 assembled using multiplex oligonucleotide hybridization. Genomics 54:387–397. https://doi.org/10.1006/geno.1998.5620
Cuff JA, Birney E, Clamp ME, Barton GJ (2000) ProtEST: protein multiple sequence alignments from expressed sequence tags. Bioinformatics 16:111–116. https://doi.org/10.1093/bioinformatics/16.2.111
Florijn RJ, Bonden LA, Vrolijk H, Wiegant J, Vaandrager JW, Baas F, den Dunnen JT, Tanke HJ, van Ommen GJ, Raap AK (1995) High-resolution DNA Fiber-FISH for genomic DNA mapping and colour bar-coding of large genes. Hum Mol Genet 4:831–836. https://doi.org/10.1093/hmg/4.5.831
GRCg6a (2018) Genome Reference Consortium Chicken Build 6a. National Library of Medicine, National Center for Biotechnology Information. https://www.ncbi.nlm.nih.gov/assembly/GCF_000002315.6. Accessed 14 Aug 2022
Griffin DK, Haberman F, Masabanda J, O’Brien P, Bagga M, Sazanov A, Smith J, Burt DW, Ferguson-Smith M, Wienberg J (1999) Micro- and macrochromosome paints generated by flow cytometry and microdissection: tools for mapping the chicken genome. Cytogenet Cell Genet 87:278–281. https://doi.org/10.1159/000015449
Harry DE, Zaitlin D, Marini PJ, Reed KM (2003) A first-generation map of the turkey genome. Genome 46:914–924. https://doi.org/10.1139/g03-042
Howe KL, Achuthan P, Allen J, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, Azov AG, Bennett R, Bhai J, Billis K, Boddu S, Charkhchi M, Cummins C, Da Rin Fioretto L, Davidson C, Dodiya K, El Houdaigui B, Fatima R, Gall A, Garcia Giron C, Grego T, Guijarro-Clarke C, Haggerty L, Hemrom A, Hourlier T, Izuogu OG, Juettemann T, Kaikala V, Kay M, Lavidas I, Le T, Lemos D, Gonzalez Martinez J, Marugán JC, Maurel T, McMahon AC, Mohanan S, Moore B, Muffato M, Oheh DN, Paraschas D, Parker A, Parton A, Prosovetskaia I, Sakthivel MP, Salam AIA, Schmitt BM, Schuilenburg H, Sheppard D, Steed E, Szpak M, Szuba M, Taylor K, Thormann A, Threadgold G, Walts B, Winterbottom A, Chakiachvili M, Chaubal A, De Silva N, Flint B, Frankish A, Hunt SE, IIsley GR, Langridge N, Loveland JE, Martin FJ, Mudge JM, Morales J, Perry E, Ruffier M, Tate J, Thybert D, Trevanion SJ, Cunningham F, Yates AD, Zerbino DR, Flicek P (2021) Ensembl 2021. Nucleic Acids Res 49(D1):D884–D891. https://doi.org/10.1093/nar/gkaa942
Itoh Y, Mizuno S (2002) Molecular and cytological characterization of SspI-family repetitive sequence on the chicken W chromosome. Chromosome Res 10:499–511. https://doi.org/10.1023/a:1020944414750
Itoh Y, Kampf K, Arnold AP (2006) Comparison of the chicken and zebra finch Z chromosomes shows evolutionary rearrangements. Chromosome Res 14:805–815. https://doi.org/10.1007/s10577-006-1082-1
Jin J, Li X, Gygi SP, Harper JW (2007) Dual E1 activation systems for ubiquitin differentially regulate E2 enzyme charging. Nature 447:1135–1138. https://doi.org/10.1038/nature05902
Kerstens H, Groenen M (2004–2008) Chicken FPC (ChickFPC). Generic genome browser version 1.67. Generic Model Organism Database Project. Genome Sequencing Centre at Washington University (St. Louis, MO, USA) and Wageningen University (The Netherlands). https://web.archive.org/web/20080118014911/http://www.animalsciences.nl/ChickFPC/. Accessed 14 Aug 2022
Lawson Handley LJ, Ceplitis H, Ellegren H (2004) Evolutionary strata on the chicken Z chromosome: implications for sex chromosome evolution. Genetics 167:367–376. https://doi.org/10.1534/genetics.167.1.367. PubMed: https://pubmed.ncbi.nlm.nih.gov/15166161/. PubMed Central: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1470863/
Lee MK, Ren CW, Yan B, Cox B, Zhang HB, Romanov MN, Sizemore FG, Suchyta SP, Peters E, Dodgson JB (2003) Construction and characterization of three BAC libraries for analysis of the chicken genome. Anim Genet 34:151–152. https://doi.org/10.1046/j.1365-2052.2003.00965_5.x
Maglott D, Ostell J, Pruitt KD, Tatusova T (2011) Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res 39(Database issue):D52–D57. https://doi.org/10.1093/nar/gkq1237
Matsubara K, Tarui H, Toriba M, Yamada K, Nishida-Umehara C, Agata K, Matsuda Y (2006) Evidence for different origin of sex chromosomes in snakes, birds, and mammals and step-wise differentiation of snake sex chromosomes. Proc Natl Acad Sci U S A 103:18190–18195. https://doi.org/10.1073/pnas.0605274103
Modi WS, Romanov M, Green ED, Ryder O (2009) Molecular cytogenetics of the California condor: evolutionary and conservation implications. Cytogenet Genome Res 127:26–32. https://doi.org/10.1159/000272458
Navarro Gonzalez J, Zweig AS, Speir ML, Schmelter D, Rosenbloom KR, Raney BJ, Powell CC, Nassar LR, Maulding ND, Lee CM, Lee BT, Hinrichs AS, Fyfe AC, Fernandes JD, Diekhans M, Clawson H, Casper J, Benet-Pagès A, Barber GP, Haussler D, Kuhn RM, Haeussler M, Kent WJ (2021) The UCSC Genome Browser database: 2021 update. Nucleic Acids Res 49(D1):D1046–D1057. https://doi.org/10.1093/nar/gkaa1070
Nefedov M, Zhu B, Thorsen J, Shu CL, Cao Q, Osoegawa K, de Jong P (2003) New chicken, turkey, salmon, bovine, porcine and sheep genomic BAC libraries to complement world wide effort to map farm animals genomes. In: Carollo V, Grant D, Lazo G (eds) Plant and Animal Genome XI Conference. Scherago International, New York, p 96
Papadopoulos JS, Agarwala R (2007) COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23:1073–1079. https://doi.org/10.1093/bioinformatics/btm076
Pigozzi MI, Solari AJ (2005) Meiotic recombination in the ZW pair of a tinamid bird shows a differential pattern compared with neognaths. Genome 48:286–290. https://doi.org/10.1139/g04-117
Ren C, Lee MK, Yan B, Ding K, Cox B, Romanov MN, Price JA, Dodgson JB, Zhang HB (2003) A BAC-based physical map of the chicken genome. Genome Res 13:2754–2758. https://doi.org/10.1101/gr.1499303
Resources (2000–2013) Database of BACs Assigned to Chicken Genes and Markers. US Poultry Genome Project. Michigan State University, East Lansing, MI, USA. https://web.archive.org/web/20131028030006/http://poultry.mph.msu.edu/resources/Resources.htm#bacdata. Accessed 14 Aug 2022
Romanov MN, Dodgson JB (2006) Cross-species overgo hybridization and comparative physical mapping within avian genomes. Anim Genet 37:397–399. https://doi.org/10.1111/j.1365-2052.2006.01463.x
Romanov MN, Price JA, Dodgson JB (2003) Integration of animal linkage and BAC contig maps using overgo hybridization. Cytogenet Genome Res 102:277–281. https://doi.org/10.1159/000075763
Romanov MN, Koriabine M, Nefedov M, de Jong PJ, Ryder OA (2006) Construction of a California condor BAC library and first-generation chicken-condor comparative physical map as an endangered species conservation genomics resource. Genomics 88:711–718. https://doi.org/10.1016/j.ygeno.2006.06.005
Romanov MN, Tuttle EM, Houck ML, Modi WS, Chemnick LG, Korody ML, Mork EM, Otten CA, Renner T, Jones KC, Dandekar S, Papp JC, Da Y, Comparative Sequencing Program NISC, Green ED, Magrini V, Hickenbotham MT, Glasscock J, McGrath S, Mardis ER, Ryder OA (2009) The value of avian genomics to the conservation of wildlife. BMC Genomics 10(Suppl 2):S10. https://doi.org/10.1186/1471-2164-10-S2-S10
Romanov MN, Dodgson JB, Gonser RA, Tuttle EM (2011) Comparative BAC-based mapping in the white-throated sparrow, a novel behavioral genomics model, using interspecies overgo hybridization. BMC Res Notes 4:211. https://doi.org/10.1186/1756-0500-4-211
Romanov MN, Betuel AM, Chemnick LG, Ryder OA, Kulibaba RO, Tereshchenko OV, Payne WS, Delekta PC, Dodgson JB, Tuttle EM, Gonser RA (2019) Widely applicable PCR markers for sex identification in birds. Russ J Genet 55:220–231. https://doi.org/10.1134/S1022795419020121
Sayers EW, Beck J, Bolton EE, Bourexis D, Brister JR, Canese K, Comeau DC, Funk K, Kim S, Klimke W, Marchler-Bauer A, Landrum M, Lathrop S, Lu Z, Madden TL, O’Leary N, Phan L, Rangwala SH, Schneider VA, Skripchenko Y, Wang J, Ye J, Trawick BW, Pruitt KD, Sherry ST (2021) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 49(D1):D10–D17. https://doi.org/10.1093/nar/gkaa892
Sazanov AA, Sazanova AL, Stekolnikova VA, Trukhina AV, Kozyreva AA, Smirnov AF, Romanov MN, Lawson Handley LJ, Malewski T, Dodgson JB (2006) Chromosomal localization of the UBAP2Z and UBAP2W genes in chicken. Anim Genet 37:72–73. https://doi.org/10.1111/j.1365-2052.2005.01392.x
Schmid M, Nanda I, Guttenbach M, Steinlein C, Hoehn M, Schartl M, Haaf T, Weigend S, Fries R, Buerstedde JM, Wimmers K, Burt DW, Smith J, A’Hara S, Law A, Griffin DK, Bumstead N, Kaufman J, Thomson PA, Burke T, Groenen MA, Crooijmans RP, Vignal A, Fillon V, Morisson M, Pitel F, Tixier-Boichard M, Ladjali-Mohammedi K, Hillel J, Mäki-Tanila A, Cheng HH, Delany ME, Burnside J, Mizuno S (2000) First report on chicken genes and chromosomes 2000. Cytogenet Cell Genet 90:169–218. https://doi.org/10.1159/000056772
Schmid M, Nanda I, Hoehn H, Schartl M, Haaf T, Buerstedde JM, Arakawa H, Caldwell RB, Weigend S, Burt DW, Smith J, Griffin DK, Masabanda JS, Groenen MA, Crooijmans RP, Vignal A, Fillon V, Morisson M, Pitel F, Vignoles M, Garrigues A, Gellin J, Rodionov AV, Galkina SA, Lukina NA, Ben-Ari G, Blum S, Hillel J, Twito T, Lavi U, David L, Feldman MW, Delany ME, Conley CA, Fowler VM, Hedges SB, Godbout R, Katyal S, Smith C, Hudson Q, Sinclair A, Mizuno S (2005) Second report on chicken genes and chromosomes 2005. Cytogenet Genome Res 109:415–479. https://doi.org/10.1159/000084205
Shibusawa M, Nishida-Umehara C, Masabanda J, Griffin DK, Isobe T, Matsuda Y (2002) Chromosome rearrangements between chicken and guinea fowl defined by comparative chromosome painting and FISH mapping of DNA clones. Cytogenet Genome Res 98:225–230. https://doi.org/10.1159/000069813
Shibusawa M, Nishida-Umehara C, Tsudzuki M, Masabanda J, Griffin DK, Matsuda Y (2004) A comparative karyological study of the blue-breasted quail (Coturnix chinensis, Phasianidae) and California quail (Callipepla californica, Odontophoridae). Cytogenet Genome Res 106:82–90. https://doi.org/10.1159/000078569
Smit AFA, Hubley R, Green P (1996–2010) RepeatMasker Open-3.0. Institute for Systems Biology. http://www.repeatmasker.org. Accessed 14 Aug 2022
Stiglec R, Ezaz T, Graves JAM (2007a) A new look at the evolution of avian sex chromosomes. Cytogenet Genome Res 117:103–109. https://doi.org/10.1159/000103170
Stiglec R, Ezaz T, Graves JAM (2007b) Reassignment of chicken W chromosome sequences to the Z chromosome by fluorescence in situ hybridization (FISH). Cytogenet Genome Res 116:132–134. https://doi.org/10.1159/000097432
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Wheeler DL, Chappey C, Lash AE, Leipe DD, Madden TL, Schuler GD, Tatusova TA, Rapp BA (2000) Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 28:10–14. https://doi.org/10.1093/nar/28.1.10
Acknowledgements
We are greatly in debt to Professors Jerry B. Dodgson, Pieter J. de Jong, Oliver A. Ryder, Elaina M. Tuttle, and Rusty A. Gonser for their kind support and provision with avian BAC library-related resources. We are also very grateful to Drs. Igor Yu. Blagoveschensky, Kirill A. Fomichev, and Victoria A. Stekol’nikova for their assistance in FISH experiments.
Elaina M. Tuttle died before this paper was submitted to the journal.
Funding
A.A. Sazanov and A.L. Sazanova were supported by the grants from the Russian Foundation for Basic Research (06–04-48031-a and 07–04-13500).
Author information
Authors and Affiliations
Contributions
Conceptualization: A.A. Sazanov and M.N. Romanov; Methodology: M.N. Romanov, A.A. Sazanov and A.L. Sazanova; Formal analysis and investigation: M.N. Romanov; Writing – original draft preparation: A.A. Sazanov and M.N. Romanov; Writing – review and editing: D.K. Griffin and M.N. Romanov; Funding acquisition: A.A. Sazanov; Resources: M.D. Nefedov; Supervision: A.A. Sazanov and D.K. Griffin.
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding authors state that there is no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sazanov, A.A., Sazanova, A.L., Nefedov, M.D. et al. A pair of gametologous genes provides further insights into avian comparative cytogenomics. Biologia 78, 2737–2746 (2023). https://doi.org/10.1007/s11756-023-01395-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01395-6