Abstract
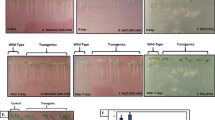
Glutaredoxins (Grxs) are glutathione (GSH)-mediated, small (10–15 kDa) oxidoreductases. In the present investigation, transgenic Arabidopsis thaliana over-expressing a glutaredoxin gene [LOC101493651 (CaGrx)] of chickpea (Cicer arietinum L.) has been evaluated against drought, and further transcriptomic studies were performed to know the effect of the CaGrx gene in transgenic Arabidopsis in drought tolerance along with wild-type (WT) and control © plants. A total of 1070 (transgenic) and 1423 (wild-type) significant differentially expressed genes (DEGs) were identified from the comparative analysis with control. Out of significant DEGs, 530 and 567 were up-regulated, and 540 and 856 were down-regulated in transgenic and wild-type (WT) plants, respectively. The Kyoto Encyclopedia of Genes and Genomes database (KEGG) identified 32 (Transgenic) and 20 (WT) metabolic pathways from the obtained unigenes. Tolerance against drought in the transgenics over-expressing CaGrx gene was by elevating the expression of genes involved in the plant defense system. This CaGrx administered tolerance in transgenics against drought by elevating glutaredoxin enzyme activity that induced various antioxidant enzymes antioxidants and reduced the level of stress markers. Leaf staining showed less H2O2 accumulation in transgenics as compared to higher accumulation in WT which shows a less stressful environment in transgenics under drought stress. Vast numbers of drought-responsive genes were retrieved, and among them, 28 differentially expressed genes (DEGs) were selected for quantitative real-time PCR that affirmed the reproducibility of the RNA-seq data.










Similar content being viewed by others
References
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Babraham Bioinformatics, Babraham Institute, Cambridge. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Ansari MA, Bano N, Kumar A, Dubey AK, Asif MH, Sanyal I, Pande V, Pandey V (2022) Comparative transcriptomic analysis and antioxidant defense mechanisms in cluster bean (Cyamopsis tetragonoloba (L.) Taub.) genotypes with contrasting drought tolerance. Funct Integr Genom 22:625–642. https://doi.org/10.1007/s10142-022-00860-w
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bouzid M, He F, Schmitz G, Häusler RE, Weber APM, Mettler-Altmann T, De Meaux J (2019) Arabidopsis species deploy distinct strategies to cope with drought stress. Ann Bot 124:27–40. https://doi.org/10.1093/aob/mcy237
Cao ZH, Zhang SZ, Wang RK, Zhang RF, Hao YJ (2013) Genome wide analysis of the apple MYB transcription factor family allows the identification of MdoMYB121 gene confering abiotic stress tolerance in plants. PLoS One 8:69955. https://doi.org/10.1371/journal.pone.0069955
Carvalho MD (2008) Drought stress and reactive oxygen species. Plant Signal Behav 3:156–165. https://doi.org/10.4161/psb.3.3.5536
Chen Z, Liu Y, Yin Y, Liu Q, Li N, Li X, He W, Hao D, Liu X, Guo C (2019) Expression of AtGA2ox1 enhances drought tolerance in maize. Plant Growth Regul 89:203–215. https://doi.org/10.1007/s10725-019-00526-x
Cheng NH, Liu JZ, Brock A, Nelson RS, Hirschi KD (2006) AtGRXcp, an Arabidopsis chloroplastic glutaredoxin, is critical for protection against protein oxidative damage. J Biol Chem 281:26280–26288. https://doi.org/10.1074/jbc.M601354200
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Couturier J, Jacquot JP, Rouhier N (2009) Evolution and diversity of glutaredoxins in photosynthetic organisms. Cell Mol Life Sci 66:2539–2557. https://doi.org/10.1007/s00018-009-0054-y
Das K, Roychoudhury A (2014) Reactive oxygen species (ROS) and response of antioxidants as ROS-scavengers during environmental stress in plants. Front Environ Sci 2:53. https://doi.org/10.3389/fenvs.2014.00053
Daudi A, O’Brien JA (2012) Detection of hydrogen peroxide by DAB staining in Arabidopsis leaves. Bio-Protoc 2:e263–e263
Ding S, He F, Tang W, Du H, Wang H (2019) Identification of maize CC-Type glutaredoxins that are associated with response to drought stress. Gene 10:610. https://doi.org/10.3390/genes10080610
Dubey AK, Kumar N, Kumar A, Ansari MA, Ranjan R, Gautam A, Meenakshi SN, Pandey V, Behera SK, Mallick S, Pande V, Sanyal I (2019) Over-expression of CarMT gene modulates the physiological performance and antioxidant defense system to provide tolerance against drought stress in Arabidopsis thaliana L. Ecotoxicol Environ Saf 171:54–65. https://doi.org/10.1016/j.ecoenv.2018.12.050
Fernandes AP, Holmgren A (2004) Glutaredoxins: glutathione-dependent redox enzymes with functions far beyond a simple thioredoxin backup system. Antioxid Redox Signal 6:63–74. https://doi.org/10.1089/152308604771978354
Foyer CH, Noctor G (2005) Redox homeostasis and antioxidant signaling: a metabolic interface between stress perception and physiological responses. Plant Cell 17:1866–1875. https://doi.org/10.1105/tpc.105.033589
Geng D, Chen P, Shen X, Zhang Y, Li X, Jiang L, Xie Y, Niu C, Zhang J, Huang X, Ma F (2018) MdMYB88 and MdMYB124 enhance drought tolerance by modulating root vessels and cell walls in apple. Plant Physiol 178:1296–1309. https://doi.org/10.1104/pp.18.00502
Gong X, Su Q, Lin D, Jiang Q, Xu J, Zhang J, Teng S, Dong Y (2014) The rice OsV4 encoding a novel pentatricopeptide repeat protein is required for chloroplast development during the early leaf stage under cold stress. J Integr Plant Biol 56:400–410. https://doi.org/10.1111/jipb.12138
Guo H, Wang Y, Wang L, Hu P, Wang Y, Jia Y, Zhang C, Zhang Y, Zhang Y, Wang C, Yang C (2017) Expression of the MYB transcription factor gene Bpl MYB 46 affects abiotic stress tolerance and secondary cell wall deposition in Betula platyphylla. Plant Biotechnol J 15:107–121. https://doi.org/10.1111/pbi.12595
Guo Y, Jiang Q, Hu Z, Sun X, Fan S, Zhang H (2018) Function of the auxin-responsive gene TaSAUR75 under salt and drought stress. Crop J 6:181–190. https://doi.org/10.1016/j.cj.2017.08.005
Jiang SC, Mei C, Liang S, Yu YT, Lu K, Wu Z, Wang XF, Zhang DP (2015) Crucial roles of the pentatricopeptide repeat protein SOAR1 in Arabidopsis response to drought, salt and cold stresses. Plant Mol Biol 88:369–385. https://doi.org/10.1007/s11103-015-0327-9
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360. https://doi.org/10.1038/nmeth.3317
Kolde R (2012) Pheatmap: pretty heatmaps. R package version, 1:747. https://CRAN.R-project.org/package=pheatmap
Kumar A, Dubey AK, Kumar V, Ansari MA, Narayan S, Meenakshi KS, Pandey V, Shirke PA, Pande V, Sanyal I (2020a) Over-expression of chickpea glutaredoxin (CaGrx) provides tolerance to heavy metals by reducing metal accumulation and improved physiological and antioxidant defence system. Ecotoxicol Environ Saf 192:110252. https://doi.org/10.1016/j.ecoenv.2020.110252
Kumar A, Dubey AK, Kumar V, Ansari MA, Narayan S, Meenakshi KS, Pandey V, Shirke PA, Pande V, Sanyal I (2020b) Overexpression of rice glutaredoxin genes LOC_Os02g40500 and LOC_Os01g27140 regulate plant responses to drought stress. Ecotoxicol Environ Saf 200:110721. https://doi.org/10.1016/j.ecoenv.2020.110721
Kumar A, Kumar V, Dubey AK, Ansari MA, Narayan S, Meenakshi KS, Pandey V, Pande V, Sanyal I (2021) Chickpea glutaredoxin (CaGrx) gene mitigates drought and salinity stress by modulating the physiological performance and antioxidant defense mechanisms. Physiol Mol Biol Plants 27:923–944. https://doi.org/10.1016/j.ecoenv.2020.110252
Laluk K, AbuQamar S, Mengiste T (2011) The Arabidopsis mitochondria-localized pentatricopeptide repeat protein PGN functions in defense against necrotrophic fungi and abiotic stress tolerance. Plant Physiol 156:2053–2068. https://doi.org/10.1104/pp.111.177501
Laporte D, Olate E, Salinas P, Salazar M, Jordana X, Holuigue L (2011) Glutaredoxin GRXS13 plays a key role in protection against photo-oxidative stress in Arabidopsis. J Exp Bot 63:503–515. https://doi.org/10.1093/jxb/err301
Laxa M, Liebthal M, Telman W, Chibani K, Dietz KJ (2019) The role of the plant antioxidant system in drought tolerance. Antioxidants 8:94. https://doi.org/10.3390/antiox8040094
Leng G, Hall J (2019) Crop yield sensitivity of global major agricultural countries to droughts and the projected changes in the future. Sci Total Environ 654:811–821. https://doi.org/10.1016/j.scitotenv.2018.10.434
Liu Y, Huang W, Xian Z, Hu N, Lin D, Ren H, Chen J, Su D, Li Z (2017) Overexpression of SlGRAS40 in tomato enhances tolerance to abiotic stresses and influences auxin and gibberellin signaling. Front Plant Sci 8:1659. https://doi.org/10.3389/fpls.2017.01659
Liu X, Lan J, Huang Y, Cao P, Zhou C, Ren Y, He N, Liu S, Tian Y, Nguyen T, Jiang L (2018) WSL5, a pentatricopeptide repeat protein, is essential for chloroplast biogenesis in rice under cold stress. J Exp Bot 69:3949–3961. https://doi.org/10.1093/jxb/ery214
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Mishra S, Jha S, Singh R, Chaudhary S, Sanyal I, Amla DV (2013) Transgenic chickpea expressing a recombinant human α1-proteinase inhibitor (α1-PI) driven by a seed-specific promoter from the common bean Phaseolus vulgaris (L.). Plant Cell Tissue Organ Cult 115:23–33. https://doi.org/10.1007/s11240-013-0336-9
O’Toole N, Hattori M, Andres C, Iida K, Lurin C, Schmitz-Linneweber C, Sugita M, Small I (2008) On the expansion of the pentatricopeptide repeat gene family in plants. Mol Biol Evol 25:1120–1128. https://doi.org/10.1093/molbev/msn057
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Rouhier N, Gelhaye E, Jacquot JP (2004) Plant glutaredoxins: still mysterious reducing systems. Cell Mol Life Sci 61:1266–1277. https://doi.org/10.1007/s00018-004-3410-y
Rouhier N, Lemaire SD, Jacquot JP (2008a) The role of glutathione in photosynthetic organisms: emerging functions for glutaredoxins and glutathionylation. Annu Rev Plant Biol 59:143–166. https://doi.org/10.1146/annurev.arplant.59.032607.092811
Rouhier N, San Koh C, Gelhaye E, Corbier C, Favier F, Didierjean C, Jacquot JP (2008b) Redox based antioxidant systems in plants: biochemical and structural analyses. Biochim Biophys Acta (BBA) Gen. Subj. 1780:1249–1260. https://doi.org/10.1016/j.bbagen.2007.12.007
Rubio-Pina JA, Zapata-Perez O (2011) Isolation of total RNA from tissues rich in polyphenols and polysaccharides of mangrove plants. Electron J Biotechnol 14:5. https://doi.org/10.2225/vol14-issue5-fulltext-8
Shi JB, Wang N, Zhou H, Xu QH, Yan GT (2019) The role of gibberellin synthase gene GhGA2ox1 in upland cotton (Gossypium hirsutum L.) responses to drought and salt stress. Biotechnol Appl Biochem 66:298–308. https://doi.org/10.1002/bab.1725
Singh R, Yadav R, Amla DV, Sanyal I (2016) Characterization and functional validation of two scaffold attachment regions (SARs) from Cicer arietinum (L.). Plant Cell Tissue Org Cult 125:135–148. https://doi.org/10.1007/s11240-015-0935-8
Sundaram S, Wu S, Ma LQ, Rathinasabapathi B (2009) Expression of a Pteris vittata glutaredoxin PvGRX5 in transgenic Arabidopsis thaliana increases plant arsenic tolerance and decreases arsenic accumulation in the leaves. Plant Cell Environ 32:851–858. https://doi.org/10.1111/j.1365-3040.2009.01963.x
Swift ML (1997) GraphPad prism, data analysis, and scientific graphing. J Chem Inf Comput Sci 37:411–412. https://doi.org/10.1021/ci960402j
Tan J, Tan Z, Wu F, Sheng P, Heng Y, Wang X, Ren Y, Wang J, Guo X, Zhang X, Cheng Z (2014) A novel chloroplast-localized pentatricopeptide repeat protein involved in splicing affects chloroplast development and abiotic stress response in rice. Mol Plant 7:1329–1349. https://doi.org/10.1093/mp/ssu054
Tang Y, Bao X, Zhi Y, Wu Q, Guo Y, Yin X, Zeng L, Li J, Zhang J, He W, Liu W (2019) Overexpression of a MYB family gene, OsMYB6, increases drought and salinity stress tolerance in transgenic rice. Front Plant Sci 10:168. https://doi.org/10.3389/fpls.2019.00168
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, Van Baren MJ, Salzberg SL, Wold BJ, Pachter L (2010) Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28:511–515. https://doi.org/10.1038/nbt.1621
Usadel B, Nagel A, Steinhauser D, Gibon Y, Bläsing OE, Redestig H, Sreenivasulu N, Krall L, Hannah MA, Poree F, Fernie AR (2006) PageMan: an interactive ontology tool to generate, display, and annotate overview graphs for profiling experiments. BMC Bioinform 7:1–8. https://doi.org/10.1186/1471-2105-7-535
Wickham H (2011) ggplot2. Wiley Interdiscip Rev Comput Stat 3:180–185. https://doi.org/10.1002/wics.147
Wu Q, Hu Y, Sprague SA, Kakeshpour T, Park J, Nakata PA, Cheng N, Hirschi KD, White FF, Park S (2017) Expression of a monothiol glutaredoxin, AtGRXS17, in tomato (Solanum lycopersicum) enhances drought tolerance. Biochem Biophys Res Commun 491:1034–1039. https://doi.org/10.1016/j.bbrc.2017.08.006
Wu J, Jiang Y, Liang Y, Chen L, Chen W, Cheng B (2019) Expression of the maize MYB transcription factor ZmMYB3R enhances drought and salt stress tolerance in transgenic plants. Plant Physiol Biochem 137:179–188. https://doi.org/10.1016/j.plaphy.2019.02.010
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2:100141. https://doi.org/10.1016/j.xinn.2021.100141
Yu G (2021) enrichplot: visualization of functional enrichment result. R package version 1.8. 1, 2020. https://github.com/GuangchuangYu/enrichplot
Yuan H, Liu D (2012) Functional disruption of the pentatricopeptide protein SLG1 affects mitochondrial RNA editing, plant development, and responses to abiotic stresses in Arabidopsis. Plant J 70:432–444. https://doi.org/10.1111/j.1365-313X.2011.04883.x
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61:49–64. https://doi.org/10.1146/annurev-arplant-042809-112308
Acknowledgements
The authors are grateful to the Director, CSIR-NBRI, Lucknow, for infrastructural support. We express gratitude towards C.S.I.R., New Delhi, for funds under the project MLP0026. The authors are also thankful to C.S.I.R., New Delhi (AK), and U.G.C., New Delhi (VK, NB) for providing research fellowships. Institutional manuscript number: CSIR-NBRI_MS/2021/09/03.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they do not have any conflict of interest.
Additional information
Communicated by V.P. Singh.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumar, A., Bano, N., Ansari, M.A. et al. Comparative transcriptomic analysis of the chickpea glutaredoxin (CaGrx) gene over-expressed in Arabidopsis thaliana is associated with drought tolerance by modulating the plant defense system. Acta Physiol Plant 45, 119 (2023). https://doi.org/10.1007/s11738-023-03612-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-023-03612-w