Abstract
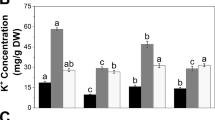
Growth, ionic responses, and expression of candidate genes to salinity stress were examined in two perennial ryegrass accessions differing in salinity tolerance. The salinity tolerant (PI265349) and sensitive accessions (PI231595) were subjected to 75-mM NaCl for 14 days in a growth chamber. Across two accessions, salinity stress increased shoot dry weight and concentrations of malondialdehyde (MDA) and Na+ in the shoots and roots, but decreased shoot Ca2+ and root K+ concentrations. Salinity stress also increased root expressions of SOS1, PIP1, and TIP1. Plant height and chlorophyll content were unaffected by salinity stress in the tolerant accession but significantly decreased in the sensitive accession. Shoot MDA content did not change in the tolerant accession but increased in the sensitive accession. A more dramatic increase in Na+ was found in the roots of the sensitive accession. Relative to the control, salinity stress reduced expression of SOS1, NHX1, PIP1, and TIP1 in the shoots but increased expression of these genes in the roots of the tolerant accession. Expression levels of SOS1 increased in the roots and expression of NHX1 increased in the shoots but decreased in the roots of the sensitive accession under salinity stress. A decline in PIP1 expression in the shoots and dramatic increases in TIP expression in both shoots and roots were found in the sensitive accession under salinity stress. The results suggested maintenance of plant growth and leaf chlorophyll content, lesser Na+ accumulation in the roots, and lower lipid peroxidation in the shoots which could be associated with salinity tolerance. The decreased expressions of SOS1, NHX1, and TIP1 in the shoots, and increased expressions of NHX1 and PIP1 in the roots might also be related to salinity tolerance in perennial ryegrass.



Similar content being viewed by others
References
Ahmad P, John R, Sarwat M, Umar S (2008) Responses of proline, lipid peroxidation and antioxidative enzymes in two varieties of Pisum sativum L. under salt stress. Intl J Plant Production 2:353–366
Azadi A, Hervan EM, Mohammadi SA, Moradi F, Nakhoda B, Vahabzade M (2011) Screening of recombinant inbred lines for salinity tolerance in bread wheat (Triticum aestivum L.). Afri J Biotech 10:12875–12881
Banjara M, Zhu L, Shen G, Payton P, Zhang H (2012) Expression of an Arabidopsis sodium/proton antiporter gene (AtNHX1) in peanut to improve salt tolerance. Plant Biotechnol Rep 6:59–67
Cabot C, Sibole JV, Barceló J, Poschenrieder C (2009) Sodiu–-calcium interactions with growth, water, and photosynthetic parameters in salt-treated beans. J Plant Nutr Soil Sci 172:637–643
Chen Y, Ye Y (2014) Effects of salinity and nutrient addition on mangrove Excoecaria agallocha. PLoS One 9:e93337
Dai J, Schlossberg MJ, Huff DR (2008) Salinity tolerance of 33 greens-type Poa annua experimental lines. Crop Sci 48:1187–1192
Dhindsa RS, Plumb-Dhindsa P, Thorpe TA (1981) Leaf senescence: correlated with increased leaves of membrane permeability and lipid peroxidation and decreased levels of superoxide dismutase and catalase. J Exp Bot 32:93–101
Diédhiou CJ, Popova OV, Golldack D (2009) Transcript profiling of the salt-tolerant Festuca rubra ssp. litoralis reveals a regulatory network controlling salt acclimatization. J Plant Physiol 166:697–711
Du Y, Hei Q, Liu Y, Zhang H, Xu K, Xia T (2010) Isolation and characterization of a putative vacuolar Na+/H+ antiporter gene from Zoysia japonica L. J Plant Biol 53:251–258
Gao J, Sun J, Cao P, Ren L, Liu C, Chen S, Chen F, Jiang J (2016) Variation in tissue Na+ content and the activity of SOS1 genes among two species and two related genera of Chrysanthemum. BMC Plant Biol 16:98
Grattan S, Grieve C (1998) Salinity–mineral nutrient relations in horticultural crops. Sci Hort 78:127–157
Health RL, Packer L (1968) Photoperoxidation in isolated chloroplasts. I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophys 125:189–198
Hove RM, Ziemann M, Bhave M (2015) Identification and expression analysis of the barley (Hordeum vulgare L.) aquaporin gene family. PLoS One 10:e0128025
Hu Y, Schmidhalter U (1997) Interactive effects of salinity and macronutrient level on wheat. II. Composition. J Plant Nutr 20:1169–1182
Hu T, Li H, Zhang X, Luo H, Fu J (2011) Toxic effect of NaCl on ion metabolism, antioxidative enzymes and gene expression of perennial ryegrass. Ecotoxicol Environ Saf 74:2050–2056
Hu L, Li H, Pang H, Fu J (2012a) Responses of antioxidant gene, protein and enzymes to salinity stress in two genotypes of perennial ryegrass (Lolium perenne) differing in salt tolerance. J Plant Physiol 169:146–156
Hu L, Huang Z, Liu S, Fu J (2012b) Growth response and gene expression in antioxidant-related enzymes in two bermudagrass genotypes differing in salt tolerance. J Amer Soc Hort Sci 137:134–143
Hu W, Yuan Q, Wang Y, Cai R, Deng X, Wang J, Zhou S, Chen M, Chen L, Huang C, Ma Z, Yang G, He G (2012c) Overexpression of a wheat aquaporin gene, TaAQP8, enhances salt stress tolerance in transgenic tobacco. Plant Cell Physiol 53:2127–2141
Jha D, Shirley N, Tester M, Roy J (2010) Variation in salinity tolerance and shoot sodium accumulation in Arabidopsis ecotypes linked to differences in the natural expression levels of transporters involved in sodium transport. Plant Cell Environ 33:793–804
Krishnan S, Brown RN (2009) Na+ and K+ accumulation in perennial ryegrass and red fescue accessions differing in salt tolerance. Intl Soc Turf Res J 11:817–827
Lalitha S (2004) Primer premier 5. Biotech Softw Internet Rep 1:270–272. https://doi.org/10.1089/152791600459894
Liu M, Jiang Y (2015) Genotypic variation in growth and metabolic responses of perennial ryegrass exposed to short-term waterlogging and submergence stress. Plant Physiol Biochem 95:57–64
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT C T method. Methods 25:402–408
Maurel C, Verdoucq L, Luu DT, Santoni V (2008) Plant aquaporins: membrane channels with multiple integrated functions. Ann Rev Plant Biol 59:595–624
Munns R, James RA (2003) Screening methods for salinity tolerance: a case study with tetraploid wheat. Plant Soil 253:201–218
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681
Qian Z-J, Song J-J, Chaumont F, Ye Q (2015) Differential responses of plasma membrane aquaporins in mediating water transport of cucumber seedlings under osmotic and salt stresses. Plant Cell Environ 38:461–473
Ruiz D, Martínez V, Cerdá A (1997) Citrus response to salinity: growth and nutrient uptake. Tree Physiol 17:141–150
Ruiz-Carrasco K, Antognoni F, Coulibaly AK, Lizardi S, Covarrubias A, Martínez EA, Molina-Montenegro MA, Biondi A, Zurita-Silva A (2011) Variation in salinity tolerance of four lowland genotypes of quinoa (Chenopodium quinoa Willd.) as assessed by growth, physiological traits, and sodium transporter gene expression. Plant Physiol Biochem 49:333–1341
Sahoo DP, Kumar S, Mishra S, Kobayashi Y, Panda SK, Sahoo L (2016) Enhanced salinity tolerance in transgenic mungbean overexpressing Arabidopsis antiporter (NHX1) gene. Mol Breed 36:144
Shi H, Quintero FJ, Pardo JM, Zhu J-K (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long distance Na+ transport in plants. Plant Cell 14:465–477
Silva EN, Silveira JAG, Rodrigues CRF, Viégas RA (2015) Physiological adjustment to salt stress in Jatropha curcas is associated with accumulation of salt ions, transport and selectivity of K+, osmotic adjustment and K+/Na+ homeostasis. Plant Biol 5:1023–1029
Tang J, Camberato JJ, Yu X, Luo N, Bian S, Jiang Y (2013a) Growth response, carbohydrate and ion accumulation of diverse perennial ryegrass accessions to increasing salinity. Sci Hort 154:73–81
Tang J, Yu X, Luo N, Xiao F, Camberato JJ, Jiang Y (2013b) Natural variation of salinity response, population structure and candidate genes associated with salinity tolerance in perennial ryegrass accessions. Plant Cell Environ 36:2021–2033
Thalji T, Shalaldeh G (2007) Screening wheat and barley genotype for salinity resistance. J Agron 6:75–78
Uddin MK, Juraimi AS, Ismail MR, Hossain MA, Othman R, Rahim AA (2012) Physiological and growth responses of six turfgrass species relative to salinity tolerance. Sci World J 2012:905468. https://doi.org/10.1100/2012/905468
Vysotskaya L, Hedley PE, Sharipova G, Veselov D, Kudoyarova G, Morris J, Jones HG (2010) Effect of salinity on water relations of wild barley plants differing in salt tolerance. AoB PLANTS. https://doi.org/10.1093/aobpla/plq006
Wang S, Zhang Q, Watkins E (2011) Evaluation of salinity tolerance of prairie junegrass, a potential low-maintenance turfgrass species. HortScience 46:1038–1043
Wang L, Li Q, Lei Q, Feng C, Gao Y, Zheng X, Zhao Y, Wang Z, Kong J (2015) MZPIP2;1: an aquaporin involved in radial water movement in both water uptake and transportation, altered the drought and salt tolerance of transgenic Arabidopsis. PLoS One 10:e0142446
Wellburn AR (1994) The spectral determination of chlorophylls a and b, as well as total carotenoids, using various solvents with spectrophotometers of different resolution. J Plant Physiol 144:307–313
Wu G, Wang S (2012) Calcium regulates K+/Na+ homeostasis in rice (Oryza sativa L.) under saline conditions. Plant Soil Environ 58:121–127
Wu Y, Chen Q, Chen M, Chen J, Wang X (2005) Salt tolerant transgenic perennial ryegrass (Lolium perenne L.) obtained by Agrobacterium tumefaciens-mediated transformation of the vacuolar Na+/ H+ antiporter gene. Plant Sci 169:65–73
Xu C, Wang M, Zhou L, Quan T, Xia G (2013) Heterologous expression of the wheat aquaporin gene TaTIP2;2 compromises the abiotic stress tolerance of Arabidopsis thaliana. PLoS One 8:e79618
Yang Q, Chen Z-Z, Zhou X-F, Yin H-B, Li X, Xin X-F, Hong X-H, Zhu J-K, Gong Z (2009) Overexpression of SOS (Salt Overly Sensitive) genes increases salt tolerance in transgenic Arabidopsis. Mol Plant 2:22–31
Yin X, Zhang C, Song X, Jiang Y (2017) Interactive short-term effects of waterlogging and salinity stress on growth and carbohydrate, lipid peroxidation, and nutrients in two perennial ryegrass cultivars. J Am Soc Hort Sci 142:110–118
Zhou L, Wang C, Liu RF, Han Q, Vandeleur RK, Du J, Tyerman S, Shou H (2014) Constitutive overexpression of soybean plasma membrane intrinsic protein GmPIP1;6 confers salt tolerance. BMC Plant Biol 14:181
Acknowledgements
This work was supported by the China Scholarship Council (No. 201708430007).
Author information
Authors and Affiliations
Contributions
ML conducted the experiments, performed the statistical analysis of data and prepared the manuscript. XS performed ion analysis. YJ supervised the overall work and contributed to data interpretation and discussion as well as paper writing.
Corresponding author
Additional information
Communicated by S Abe.
Rights and permissions
About this article
Cite this article
Liu, M., Song, X. & Jiang, Y. Growth, ionic response, and gene expression of shoots and roots of perennial ryegrass under salinity stress. Acta Physiol Plant 40, 112 (2018). https://doi.org/10.1007/s11738-018-2687-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-018-2687-7