Abstract
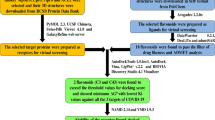
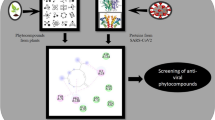
The current study reveals the inhibitory potential of novel bioactive compounds of mangrove actinomycetes against nsp10 of SARS-CoV-2. A total of fifty (50) novel bioactive (antibacterial, antitumor, antiviral, antioxidant, and anti-inflammatory) compounds of mangrove actinomycetes from different chemical classes such as alkaloids, dilactones, sesquiterpenes, macrolides, and benzene derivatives are used for interaction analysis against nsp10 of SARS-CoV-2. The six antiviral agents sespenine, xiamycin c, xiamycin d, xiamycin e, xiamycin methyl ester, and xiamycin A (obeyed RO5 rule) are selected based on higher binding energy, low inhibition constant values, and better-docked positions. The effective hydrogen and hydrophobic (alkyl, \(\pi\)–sigma, \(\pi\)–\(\pi\) T shaped and \(\pi\)-alkyl) interaction analysis reveals the four antivirals sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A are supposed to be the most auspicious inhibitors against nsp10 of SARS-CoV-2. Quantum chemistry methods such as frontier molecular orbitals and molecular electrostatic potential are used to explain the thermal stability and chemical reactivity of ligands. The toxicity profile shows that selected ligands are safe by absorption, distribution, metabolism, excretion, and toxicity profiling and also effective for inhibition of nsp10 protein of SARS-CoV-2. The molecular dynamic simulation investigation of apo and halo forms of nsp10 done by RMSD of C\(\alpha\) atoms of nsp10, all amino acid residues RMSF, count total number of hydrogen bonds and radius of gyration (Rg). MD simulations reveal the complexes are stable and increase the structural compactness of nsp10 in the binding pocket. The lead antiviral compounds sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A are recommended as the most promising inhibitors against nsp10 of SARS-CoV-2 pathogenicity.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The outbreak of COVID-19 around the world is caused by coronavirus named severe- acute-respiratory-syndrome-coronavirus 2 (SARS-CoV-2). The World Health Organization acknowledged this global outbreak a pandemic on March 12, 2020 (Jebril 2019). This infectious pneumonia-like disease trapped the 220 countries, and the number of confirmed deaths is 5,112,461 reported till November 17, 2021 (Meo et al. 2020). The common symptoms of COVID-19 include cold, fatigue, mucous production, lymphopenia, fever, hemoptysis, headache, breathing illness, and in some cases eventually caused death (Graham et al. 2020). The drug repurposing approach of reported drugs bemcentinib, sonidegib, antrafenine, Tegobuvir, itacitinib, siramesine, and phthalocyanine was previously used to inhibit the SARS-CoV-2 activity (Encinar and Menendez 2020). In current situation, though the vaccines have been approved and emerged as an effective tool to minimize the spread of COVID-19, a therapeutics drug is very imperative to put this pandemic under complete control.
Coronavirus belongs to family Coronaviridae, order Nidovirales, and subfamily Orthcoronavirinea. Coronavirus genome encoded 16 non-structural proteins (nsp1 to nsp16) by ORF 1a/1b, which are involved in RNA replication and translation activity (Rothan and Byrareddy 2020). In the present investigation, we considered the nsp10 protein of SARS-CoV-2, and their chain B contains 148 amino acid residues which are consist of two anti-parallel \(\beta\) sheets and two zinc-binding fingers that are used to interact with nsp16 and nsp14 for replication (Decroly 2011; Bouvet et al. 2014; Ma et al. 2015). In the nsp10/nsp16 methyltransferase complex, the nsp10 core plays a critical function in the activation of nsp16 that is essential for SARS-CoV-2 replication. The nsp10 activates the 2’-O-methyltransferase of nsp16 for its proper working in RNA replication (Chen 2011). SARS-CoV-2 2′-O-methyltransferase is an RNA cap-modifying enzyme with little enzymatic activity that is triggered by nsp10, which interacts with nsp16 and selects N7-methyl guanine RNA caps with 2′-O-MTase activity (Chen 2011). Essentially, lack of 2′-O-MTase activities leads to a considerable reduction in SARS-CoV infection, which is marked by lower viral reproduction and restricted respiration problems in animal models (Menachery et al. 2014). Now, the nsp10 core is an attractive target for development of antiviral drugs against coronavirus pathogenesis.
Recently novel bioactive compounds from mangrove actinomycetes gained importance due to their antibiotics (Sangkanu et al. 2017), antiviral (Xu et al. 2014), antitumor (Usha et al. 2010), and antioxidant activity to use as potential inhibitors against COVID-19. Fifty novel bioactive compounds from mangrove actinomycetes including different chemical classes such as alkaloids, dilactones, sesquiterpenes, macrolides, and benzene derivatives are used for molecular docking study. In the past, people acknowledged the mangrove as a medicinal tree and the secondary metabolites of mangrove actinomycetes are previously used in different medications (Xu et al. 2014). Mangrove plant is enriched with many microorganisms and their novel bioactive compounds are reported as an anti-inflammatory, antibacterial, antifungal, and antipyretic activity against animals, humans, and as well plants pathogens (Shilpi et al. 2012).
In recent years, mangrove actinomycetes have produced compounds with novel structures and potential medical use. Salinosporamide A is the well-known compound formed by Salinispora strain separated from mangrove samples, which is the primary and innovative marine actinomycete secondary metabolite used for cancer medication (Feling et al. 2003). The xiamycin compounds belonging to the indolosesquiterpenes class was first time isolated from prokaryotes, and these compounds showed anti-HIV activity (Ding et al. 2010). Streptomyces sp. HKI0595, obtained from the mangrove plant Kandelia candel, yielded five new kandenols A-E compounds that showed antimicrobial activities (Ding et al. 2012). The mangrove ecosystem is an almost unexplored source of innovative and effective natural products that need significantly more attention.
Computer-aided drug design (CADD) techniques are used to screen a wide range of natural and synthetic libraries for the development of successful lead active compounds against the targeted protein (Marshall 1987). The early-stage prediction of pharmacokinetic properties used in computer-aided drug design reduced the chance of failure at the experimental stage, save time and money where pharmacokinetic properties were analyzed by using in silico analysis. Quantum chemical methods in conjugation with molecular docking and dynamics are used to evaluate their chemical reactivity and suitability against SARS-CoV-2 (Zia 2021; Muhammad 2021; Haroon et al. 2021). In the current study, we aimed to find out potential inhibitors from novel bioactive compounds of mangrove actinomycetes against nsp10 of SARS-CoV-2 by using an in silico molecular docking approach and performed the DFT calculations of lead compounds to estimate some quantum parameters that predict inhibitors reactivity and stability toward targeted protein.
Computational methodology
Protein preparation
The high-resolution structure of non-structural protein nsp10/nsp16 methyltransferase (see Fig. 1) of COVID-19 (PDB ID 6WVN) was downloaded from protein data bank (Bank 1971) (https://www.rcsb.org/). The resolution of the crystal structure is 2.0 Å. 6WVN is a heterodimer and, contains nsp10 and nsp16 in complex form (Rosas-Lemus et al. 2020). The nsp10 consists of single-chain B and, contains 142 amino acid residues is used for docking analysis. The nsp10 protein preparation was done by Biovia Discovery Studio visualizer 2020 Client (Bhaskar et al. 2019) and MGL tools (Morris et al. 2009). Firstly, the water molecules, heteroatoms, and ligand moieties were removed from chain B of nsp10. Then the protonation of polar hydrogens and addition of Kollman charges were added to prepare for docking. The final files were saved as pdbqt formats along with assigned grid values in separate files.
Ligand preparation
Fifty novel bioactive compounds were isolated from mangrove actinomycetes and were used for molecular docking against nsp10 of SARS-CoV-2. These fifty bioactive compounds were downloaded from PubChem database (Kim et al. 2021) (https://pubchem.ncbi.nlm.nih.gov/) in sdf format. The 3D structure of ligands was previously obtained in sdf format and then changed to Pdb format. The prepared ligands were applied as input files in the AutoDock Vina 1.5.6 program (Ding, et al. 2012) for docking evaluation.
Molecular docking
The molecular docking of fifty novel bioactive compounds was performed by AutoDock vina (Trott and Olson 2010). During the docking analysis, the nsp10 of SARS-CoV-2 acted as a rigid receptor, and the ligands were flexible. The grid box covered the whole binding site of the protein and provided enough space for ligands to react with protein. The grid parameters that was generated in configuration file are as center_x = 64.6772, center_y = 15.5947, center_z = 9.4511 and x = 40 Å, y = 57 Å, z = 45 Å. Thereafter, Biova Discovery Studio visualizer 2020 Client was used for the interaction analysis of ligand–protein complexes (Samant and Javle 2020). The replica of molecular docking was prepared by using IGEMDOCK software (Balavignesh et al. 2013).
Quantum chemistry
Quantum chemistry, in terms of Frontier molecular orbital analysis (HOMO and LUMO) (Wei et al. 2003) and molecular electrostatic potential (MEPs) (Chidangil et al. 1998) of ligands, was used to evaluate their reactivity and thermal stability. The structure of the chosen lead compounds with the DFT method B3LYP with 6-311G* has been optimized with the Gauss View software. At the theoretical level B3LYP/6-311G*, the electrostatic molecular diagrams of antivirals were built.
Molecular dynamics (MD) simulation
Lead compounds (mangrove actinomycetes) as inhibitors of nsp10 of SARS-CoV-2 were identified from the docking analysis and were subjected to molecular dynamics simulation to evaluate inhibitory potential and conformational stability. Molecular dynamics simulation was performed by NAMD (Nelson et al. 1996) and VMD (Humphrey et al. 1996) programs. The molecular dynamics simulation was carried out at 60 ns (60 \(\times\) 106 fs), and boundary conditions were applied to run the simulations. Water molecules were added, and the system was neutralized with Na+ ions at 310 K temperature and 1 atm pressure, respectively. Further details are provided in supporting information of the article. The RMSD, RMSF, a radius of gyration, and the number of hydrogen bond values were used for further analysis of ligand–protein complexes.
ADMET study and drug likeness
The drug-likeness properties of drug candidates were examined from the online web tool SwissADME (Daina et al. 2017) (http://www.swissadme.ch/). AMDET (absorption, distribution, metabolism, excretion, and toxicity) properties are evaluated by pkCSM (Pires et al. 2015), an open database server that gives information about drug pharmacokinetics. The ligands smiles were provided to the server for running different predictions related to AMDET.
Results and discussion
Ligand selection and structure chemistry
Fifty novel bioactive compounds from mangrove actinomycetes are used for docking analysis against the nsp10 of SARS-CoV-2 to find out their inhibitory potential. The bioactive compounds of mangrove actinomycetes are isolated from different chemical classes alkaloids, dilactones, sesquiterpenes, macrolides, and benzene derivatives. The six antiviral compounds (see Fig. 2) were selected from fifty compounds of mangrove actinomycetes which belong to class indolosesquiterpene. The selected lead compounds are summarized in it with their molecular weight, compound ID, and smiles shown in Table S4. The antiviral compounds (a) L1(sespenine), (b) L2 (xiamycin C), (c) L3 (xiamycin D), (d) L4 (xiamycin E), (e) L5 (xiamycin methyl ester), (f) L6 (xiamycin A) are obtained from Streptomyces sp. mangrove endophyte (Xu et al. 2014). Xiamycin and its derivatives were reported as anti-HIV (Meng et al. 2015), antimicrobial (Christina et al. 2013) and antifungal against a broad range of fungi (Pfaffenbach et al. 2019), and antiviral activity (Ding et al. 2010; Kim et al. 2016).
Binding energy and inhibition constant of protein–ligand interactions
We docked fifty (50) novel ligands of mangrove actinomycetes against nsp10 of SARS-CoV-2, and thirty (30) ligands (see Table S1) are selected based on the RO5 rule for further analysis. After interaction analysis of selected (30) ligands it revealed, the top six lead compounds showed good interaction with nsp10 binding pocket and have high binding energy values. Table1 reveals the molecular docking analysis of the selected lead compounds of mangrove actinomycetes in terms of binding energy and inhibition constant Ki, which is docked in the binding pocket of nsp10 of SARS-CoV-2. The binding energy ranged from − 7.8 to − 8.8 kcal/mol of the lead compounds. The inhibition constant (Ki) defined the amount of the drug, needed for the 50% inhibition of enzymatic reactions (Yung-Chi and Prusoff 1973). The binding affinity of a ligand in the binding pocket of targeted protein is explained by its binding energy.
Another word used to assess the efficacy of the ligand targeted complex is binding affinity by measuring the interactions between the complex utilizing score functions, force field, and statistical approaches. The replica of selected lead compounds against nsp10 of SARS-CoV-2 by using IGEMDOCK software to check the reliability of docking results. The detail of replica was added in the supporting file (Table S2 and Figure S1). The six lead compounds of mangrove actinomycetes were selected due to their high binding energy and low inhibition constant values ranging from 0.335 to 1.827 μM. It is worth mentioning that after binding energy and inhibition constant investigations showed that all selected lead compounds of the mangrove actinomycetes are supposed to be promising inhibitors of the nsp10 of SARS-CoV-2. Figure 3 shows the graphical illustration of binding energy verse inhibition constant Ki of the selected lead compounds.
Binding interactions analysis
The interaction analysis of the lead compounds of mangrove actinomycetes toward nsp10 of SARS-CoV-2 is visualized by free software Biova Discovery Studio visualizer 2020 Client (Das et al. 2020). The binding energy range of selected antivirals of mangrove actinomycetes is − 7.8 to − 8.8 kcal/mol. The increasing inhibitory potential order of antivirals depending upon inhibition constant value is: sespenine > xiamycin A > xiamycin C > xiamycin D = xiamycin methyl ester > xiamycin E as shown in Table 1. On the base of higher binding energy and low inhibition constant values of four antivirals (sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A), interactions (hydrophilic and hydrophobic) in the binding pocket of nsp10 are visualized for further study. The interactions of compounds xiamycin D (L3) and xiamycin E (L4) are shown in Figure S2 (supporting file).
Sespenine (L1) antiviral of indolosesquiterpene class has higher binding energy -8.8 kcal/mol. The docking pose of L1 (sespenine) in the binding pocket of nsp10 in terms of total density surface and 2D interaction is shown in Fig. 4a, b. The compound L1 is formed four H-bonds through ALA4276, TYR4329, ALA4277, ASP4275, and VAL4274 amino acid residues of nsp10. The hydrophobic (alkyl and \(\pi\)-alkyl) interactions were shown by CYS4270, PRO4337, and ALA4271 amino acid residues of nsp10. The van der Waals interactions were displayed by amino acid residues ASP4335, ILE4334, HIS4336, and LEU4267.
The binding energy shown by xiamycin C (L2) is − 8.2 kcal/mol in the binding pocket of nsp10 Fig. 4c, d. The THR4292 amino acid residue formed four hydrogen bonds, and two H-bonds are formed by ARG4331 and TYR4283 amino acid residues of protease. The six hydrophobic interactions of four different types such as alkyl, \(\pi\)–sigma, \(\pi\)–\(\pi\) T-shaped, and \(\pi\)-alkyl are demonstrated by amino acid residues ILE4291, TYR4283, TYR4280, TYR4283, and ILE4291, respectively. The CYS4332 amino acid residue formed three \(\pi\)-sulfur bonds. The amino acid residues ALA4276, ASN4293, and GLN4289 are linked through van der Waals forces.
The docking pose of xiamycin methyl ester (L5) towards nsp10 of SARS-CoV-2 having binding energy − 7.9 kcal/mol as shown in Fig. 5a, b. The amino acid residue THR4292 is linked with two hydrogen bonds, and ARG4331 with one H-bond. The eleven hydrophobic (alkyl, \(\pi\)–sigma, \(\pi\)–\(\pi\) T-shaped, and \(\pi\)-alkyl) contacts are exhibited by TYR4280 with two bonds, four bonds with TYR4283 and ILE4291, ALA4276, ALA4279, and LEU42322 formed one bond. The three \(\pi\)-sulfur bonds are formed by CYS4332 amino acid residue. The van der Waals interactions are presented by ALA4276, ASN4293, and GLN4289 amino acid residues.
The docking pose of L5(Xiamycin methyl ester) (a) and L6(Xiamycin A) (b) in binding pocket of nsp10 showed as total density surface, the H-bond representation in purple and green mesh shows total density surface, 2D interactions of docked ligands L5(Xiamycin methyl ester) (c) and L6(Xiamycin A) (d) respectivel
The antiviral xiamycin A (L6) that was reported for its anti-HIV activity (Ding et al. 2010) was docked in the binding pocket of nsp10 with − 8.3 kcal/mol binding energy (Fig. 5c, d). The amino acid residue THR4292 is linked through two hydrogen bonds towards the binding pocket. The ten hydrophobic (alkyl, \(\pi\)–sigma, \(\pi\)–\(\pi\) T-shaped, and \(\pi\)-alkyl) contacts are revealed by TYR4283 with three bonds, three bonds with ILE4291 and TYR4280, ALA4276, ALA4279, and LEU42322 formed one bond. The three \(\pi\)-sulfur bonds are formed by CYS4332 amino acid residue. The van der Waals interactions are displayed by ALA4276, ASN4293, and GLN4289 amino acid residues.
The amino acid residues of nsp10 in its binding pocket interacting with six antivirals of mangrove actinomycetes along with bond distance in Å are mentioned in Table 2. It is valuable to mention after the study of the interaction of dock complexes, all are docked in the correct binding pose and show tight binding with amino acid residues. The antivirals sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A are supposed to be the most promising inhibitors of nsp10 of SARS-CoV-2. Further, the stability of ligand-nsp10 complexes was analyzed by molecular dynamics.
Frontier molecular orbital (FMO)
The chemical structures and reactivities of ligand molecules are immensely dependent on the charge distribution patterns and energetics of their frontier molecular orbitals (FMOs). When it comes to docking interactions between ligand and protein molecules, the structural chemistry of the ligand as well as partial atomic charge leads the interactions between ligand and protein macromolecule. So, the MOs and orbital energy gaps (\(\Delta E\)LUMO–HOMO) are a crucial property of ligand that demonstrates the ability of ligand how it interacts with protein (Abd El-Kareem et al. 2020). It's also worth noting that HOMO and LUMO are highly significant quantum chemical parameters that are exploited to compute many critical characteristics, such as chemical reactivity parameters. Secondly, the presence of partial charges on ligand and protein molecules plays a crucial role in leading the docking of ligand with protein. The ligands sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A are selected for FMO analysis on the base of high binding energy and effective interactions with nsp10 of SARS-CoV-2 as reported earlier in the molecular docking (Sect. 3.3). The energy bandgap (\(\Delta E\)LUMO–HOMO) and values of HOMO and LUMO orbitals are calculated in water as solvent as shown in Fig. 6. All the selected ligands showed no prominent difference in their energy gap values, and the xiamycin C from all shows a higher energy band gap \(\Delta E\) = 5.26 eV. The decreasing order of energy bandgap of selected ligands is: sespenine = xiamycin A < xiamycin methyl ester < xiamycin C. The selected ligands have a high value of energy bandgap and high thermal stability as shown in Fig. 7. The atomic coordinates of sespenine, xiamycin a, xiamycin methyl ester, and xiamycin C are given in the supporting file of article.
Molecular electrostatic potential (MEPs)
Molecular electrostatic potentials (MEPs) are maps of ground state electrostatic potentials on total electron density surfaces which were calculated for actinomycetes ligands. Molecular electrostatic potentials (MEPs) are valuable to insight into the three-dimensional structures of ligands having electrophilic and nucleophilic sites. MEPs also provide information about the size and shape of ligands' 3-D structures. The MEP diagrams of antivirals ligands including sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A are computed at B3LYP/6-311G* level of theory as shown in Fig. 7. The positive electrostatic potential maxima are indicated by the blue color, and the red area shows negative electrostatic potential maxima, while the green area demonstrates zero electrostatic potential on ligand’s total density surfaces. The red color indicates that minimum electrostatic potential or excess of unbound electrons act as electrophilic attack, and on the other hand, blue color shows maximum electrostatic or deficiency of electrons acts as a nucleophilic attack (Mehmood et al. 2018; Guégan et al. 2014). The heteroatoms are important to generate different potentials over ligand surfaces where nitrogen (N) atoms and oxygen (O) atoms create a positive and negative electrostatic potential in ligands (see Fig. 7). The electrostatic potential mapping shows that overall some negative and positive potentials are crucial to make ligand favorable for interacting with protein molecules (Rzęsikowska et al. 2019).
Drug likeness
A drug candidate must have good intestinal absorption and be easily excreted from the body to be a useful lead compound against target protein. The drug-likeness of selected lead compounds was predicted by Lipinski’s rule of 5. According to this rule, drug candidates should fulfil these following rules: molecular weight < 500D, hydrogen donor < 5, hydrogen acceptor < 10, and LogP < 5 (Chen et al. 2020). All the selected novel lead compounds from the indolosesquiterpene class of mangrove actinomycetes are obeyed the RO5 rule (see Table S5), and it proposed that these compounds have strong drug-likeness and potency to be a promising drug candidate.
ADMET profiling
For the advancement of an effective drug, the lead molecule must show maximum potency against targeted protein and have appropriate pharmacokinetic properties. ADMET profiling is used to find out the pharmacokinetic properties of the lead compound to be a promising inhibitor of COVID-19 (Fatima et al. 2020). The detail ADMET profiling of selected lead compounds is given in supporting file (see Table S6).
Molecular dynamic (MD) simulations
MD simulations were used to validate the docking results and obtain more insight into the ligand–protein complex's stability (Abdelrheem et al. 2020). The physical changes of atoms along with whole macromolecules are studied using molecular dynamics, which are efficiently used to investigate the structure-to-protein significance of a given system (Amera et al. 2020). Furthermore, it can also be used to show the properties, pattern, and strength of ligand–protein interactions as well as macromolecule dynamic conformational changes (Khan et al. 2021). The docked moiety nsp10 of SARS-CoV-2 along with L1(sespenine), L2 (xiamycin C), L5 (xiamycin methyl ester), and L6 (xiamycin A) was simulated to insight the structural deviations and stability of protein over 60 ns dynamic simulation. MD simulation of apo-form nsp10 and complexes was run to investigate, root mean square of deviation (RMSD) of C\(\alpha\) atoms of protease, all amino acid residues root mean square of fluctuation (RMSF), count total number of hydrogen bonds, and radius of gyration (Rg).
The root mean square of deviation (RMSD) determines the average distance between atoms of protein. It also explained the conformational change of C\(\alpha\) atoms of apo-form nsp10 over the 60 ns timescale simulation. The complexes and apo-form nsp10 RMSD values in graph form over 60 ns time trajectory are shown in Fig. 8A. The apo-form of nsp10 average RMSD value is 0.95 Å, which is a low RMSD value and shows less conformational changes and stability of protein (Bagaria et al. 2012; Carugo and Pongor 2001). The complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 show unique behavior over 60 ns simulation, and all complexes’ average RMSD values are in the range of 0.85 Å to 1.5 Å. The lower RMSD values of all complexes exemplify the correct binding pose of ligands and stability of complexes in the binding pocket of the protein.
Analysis of RMSD, RMSF, total number of hydrogen bonds, and radius of gyration (Rg) of apo-nsp10 protease of SARS-CoV-2 and four selected ligand–protein complexes over 60 ns time trajectory. A Root mean square deviation (RMSD, Å) of nsp10 C\(\alpha\) atoms during 60 ns simulation, B root mean square fluctuation (RMSF), values of RMSF are given at ordinate in Å and time (ns) at abscissa, C radius of gyration (Å), D No. of hydrogen bonds at 60 ns simulation
The root mean square of fluctuation (RMSF) is used to explain the fluctuation of all amino acid residues when the ligand interacted with protein. Figure 8B reveals the root mean square fluctuation of ligand-nsp10 complexes and apo-form nsp10. The average RMSF value of protein is 1.5 Å which is in the acceptable RMSF range. The low value of RMSF shows the protein exhibits less flexibility even when not interacted with ligands. The average RMSF values of complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 are 1.75, 1.15, 0.85, and 1.15 Å, respectively. All the complexes show no prominent fluctuations over the whole 60 ns simulation. The amino acid residues that show minor fluctuation peaks in all complexes are ARG4331, HIS4336, PRO4337, and ASN4338. The L5-nsp10 complex shows a more stable RMSF value over 60 ns time trajectory. The complexes L1-nsp10, L2-nsp10, and L6-nsp10 show similar RMSF at all residues.
Radius of gyration is a valuable indicator to check the compactness of protein which deals with the stability of protein over the whole time of simulations. The radius of gyration (Rg) is the root mean square distance from each atom of protein from its centroid, and it describes structural compactness. The high value of Rg shows that all amino acid residues in nsp10 chain B are loosely packed and have less structural compactness. The nsp10 protease is \(\beta\)-type protein that has a value of Rg in the range of 14.6 Å to 16.1 Å (Lobanov et al. 2008). Rg value of apo-form nsp10 increases after 10 ns and goes its maximum value 16.8 Å at 19 ns. The Rg value decreases after 33 ns in the range of 16.0 Å to 16.5 Å up to 60 ns simulation. The major reason for that unusual behavior of nsp10 is it is \(\beta\)-type protein that has less folding and amino acid residues. The average value of Rg of apo-nsp10 is 15.8 Å. The complexes show a low value of Rg as interacting amino acid residues in the binding pocket become more rigid after docking and increase the protein structural compactness. The complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 have an average value of Rg are 15.3, 15.2, 15.1, and 15.3 Å, individually. All complexes exhibit the same pattern of the radius of gyration during the whole 60 ns simulation. Figure 8C shows the radius of gyration of apo-nsp10 and complexes.
In a ligand–protein complex, the hydrogen bonding plays a very significant role in estimating the binding energy of the complex (Chowdhury 2020). The count of hydrogen bonds between ligand and protein complexes plays important role in the conformational stability of a protein. The total number of bonds calculated between ligand-nsp10 complexes and apo-nsp10 is revealed in Fig. 8D. The average of hydrogen bonds in apo-nsp10 is 12. All the complexes show a higher number of hydrogen bonds than apo-nsp10 and more stability of docked ligands in binding pocket. The hydrogen count of all complexes (L1, L2, L5, and L6) lies in the range of 20–60 H-bonds. Notably, all the complexes show similar behavior during the 60 ns simulation. After molecular dynamic analysis, it was proved that all the complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 are docked in the best binding pose and stabilize the confirmation of nsp10. The novel antivirals sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A from the indolosesquiterpene class of mangrove actinomycetes are the most promising inhibitors against nsp10 of SARS-CoV-2.
Conclusion
In this study, different computational methods such as molecular docking, quantum chemistry, molecular dynamics, and ADMET profiling were used to analyze the best possible inhibitors from fifty (50) novel bioactive compounds of mangrove actinomycetes against nsp10 of SARS-CoV-2. The six antivirals compounds L1(sespenine), L2 (xiamycin C), L3 (xiamycin D), L4 (xiamycin E), L5 (xiamycin methyl ester), and L6 (xiamycin A) were selected on the base of high binding energy (− 7.8 to − 8.8 kcal/mol) and low inhibition constant value (0.335 to 1.827 μM) through molecular docking analysis. The interaction (hydrophilic and hydrophobic) analysis discovered the four antivirals sespenine, xiamycin C, xiamycin methyl ester, and xiamycin A which showed effective hydrogen and hydrophobic interactions in the binding pocket of nsp10. The thermal stability and chemical reactivity of the above-mentioned four antiviral compounds were revealed by FMOs analysis and MEPs. ADMET profiling of selected antiviral compounds shows they have a low toxicity profile and are easily absorbed and extracted from the body. The multiple short replicate MD simulations of apo-form of nsp10 and its complexes are performed at 60 ns. The RMSD and RMSF of C\(\alpha\) atoms of nsp10, all amino acid residues RMSF, count total number of hydrogen bonds, and radius of gyration (Rg) were obtained and discussed. The complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 show unique behavior over 60 ns simulation, and all complexes’ average RMSD values are in the range of 0.85 Å to 1.5 Å. The average RMSF values of complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 are 1.75, 1.15, 0.85, and 1.15 Å, respectively. The radius of gyration analysis shows that all complexes L1-nsp10, L2-nsp10, L5-nsp10, and L6-nsp10 have an average value of Rg are 15.3, 15.2, 15.1, and 15.3 Å, individually. The hydrogen count of all complexes (L1, L2, L5, and L6) lies in the range of 20 to 60 H-bonds. The lead antiviral compounds L1(sespenine), L2 (xiamycin C), L5 (xiamycin methyl ester), and L6 (xiamycin A) were the most promising inhibitors against nsp10 of SARS-CoV-2 pathogenicity. Thus, we concluded that the current investigation would bring new scientific information for entitled compounds to be tested as therapeutic agent against COVID-19 disease through in vitro and in vivo experimental investigations.
References
Abd El-Kareem MSM et al (2020) Mass spectral fragmentation of Pelargonium graveolens essential oil using GC–MS semi-empirical calculations and biological potential. Processes 8(2):128
Abdelrheem DA et al (2020) The inhibitory effect of some natural bioactive compounds against SARS-CoV-2 main protease: insights from molecular docking analysis and molecular dynamic simulation. J Environ Sci Health Part A 55(11):1373–1386
Amera GM et al (2020) Screening of promising molecules against MurG as drug target in multi-drug-resistant-Acinetobacter baumannii-insights from comparative protein modeling, molecular docking and molecular dynamics simulation. J Biomol Struct Dyn 38(17):5230–5252
Bagaria A et al (2012) Protein structure validation by generalized linear model root-mean-square deviation prediction. Protein Sci 21(2):229–238
Balavignesh V et al (2013) Molecular docking study ON NS5B polymerase of hepatitis c virus by screening of volatile compounds from Acacia concinna and ADMET prediction. Int J Pharm Life Sci 4:2548–2558
Bank PD (1971) Protein data bank
Bhaskar V, Namboori K, Pappachen LK (2019) In silico discovery of novel ligands for anti-tubercular targets using computer aided drug design. Res J Pharm Technol 12(11):5646–5650
Bouvet M et al (2014) Coronavirus Nsp10, a critical co-factor for activation of multiple replicative enzymes. J Biol Chem 289(37):25783–25796
Carugo O, Pongor S (2001) A normalized root-mean-spuare distance for comparing protein three-dimensional structures. Protein Sci 10(7):1470–1473
Chen Y et al (2011a) Biochemical and structural insights into the mechanisms of SARS coronavirus RNA ribose 2′-O-methylation by nsp16/nsp10 protein complex. PLoS Pathog 7(10):e1002294
Chen X et al (2020) Analysis of the physicochemical properties of acaricides based on lipinski’s rule of five. J Comput Biol 27(9):1397–1406
Chidangil S, Shukla MK, Mishra PC (1998) A molecular electrostatic potential mapping study of some fluoroquinolone anti-bacterial agents. Mol Model Ann 4(8):250–258
Chowdhury P (2020) In silico investigation of phytoconstituents from Indian medicinal herb ‘Tinospora cordifolia (giloy)’against SARS-CoV-2 (COVID-19) by molecular dynamics approach. J Biomol Struct Dyn: 1–18
Christina A, Christapher V, Bhore SJ (2013) Endophytic bacteria as a source of novel antibiotics: an overview. Pharmacogn Rev 7(13):11
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7(1):1–13
Das D et al. (2020) In silico analysis of phytochemicals from Mucuna pruriens (L.) DC against Mycobacterium tuberculosis causing tuberculosis. Eur J Med Plants: 19–24
Decroly E et al (2011) Crystal structure and functional analysis of the SARS-coronavirus RNA cap 2′-O-methyltransferase nsp10/nsp16 complex. PLoS Pathog 7(5):e1002059
Ding L et al (2010) Xiamycin, a pentacyclic indolosesquiterpene with selective anti-HIV activity from a bacterial mangrove endophyte. Bioorg Med Chem Lett 20(22):6685–6687
Ding L et al (2012) Kandenols A-E, eudesmenes from an endophytic Streptomyces sp of the mangrove tree Kandelia candel. J Natl Prod 75(12):2223–2227
Encinar JA, Menendez JA (2020) Potential drugs targeting early innate immune evasion of SARS-coronavirus 2 via 2’-O-methylation of viral RNA. Viruses 12(5):525
Fatima S et al (2020) ADMET profiling of geographically diverse phytochemical using chemoinformatic tools. Fut Med Chem 12(1):69–87
Feling RH et al (2003) Salinosporamide A: a highly cytotoxic proteasome inhibitor from a novel microbial source, a marine bacterium of the new genus Salinospora. Angew Chem Int Ed 42(3):355–357
Graham CW et al (2020) Novel Wuhan (2019-NCoV) coronavirus. Am J Respir Crit Care Med 201(4):7–8
Guégan F et al (2014) Dual descriptor and molecular electrostatic potential: complementary tools for the study of the coordination chemistry of ambiphilic ligands. Phys Chem Chem Phys 16(29):15558–15569
Haroon M et al. (2021) Synthesis, antioxidant, antimicrobial and antiviral docking studies of ethyl 2-(2-(arylidene) hydrazinyl) thiazole-4-carboxylates. Zeitschrift für Naturforschung Cs
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38
Jebril N (2020) World Health Organization declared a pandemic public health menace: a systematic review of the coronavirus disease 2019 “COVID-19”, up to 26th March 2020. Available at SSRN 3566298
Khan RJ et al (2021) Targeting SARS-CoV-2: A systematic drug repurposing approach to identify promising inhibitors against 3C-like proteinase and 2′-O-ribose methyltransferase. J Biomol Struct Dyn 39(8):2679–2692
Kim S-H et al (2016) Antiviral indolosesquiterpenoid xiamycins C-E from a halophilic actinomycete. J Nat Prod 79(1):51–58
Kim S et al (2021) PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Res 49(D1):D1388–D1395
Lobanov MY, Bogatyreva N, Galzitskaya O (2008) Radius of gyration as an indicator of protein structure compactness. Mol Biol 42(4):623–628
Ma Y et al (2015) Structural basis and functional analysis of the SARS coronavirus nsp14–nsp10 complex. Proc Natl Acad Sci 112(30):9436–9441
Marshall GR (1987) Computer-aided drug design. Annu Rev Pharmacol Toxicol 27(1):193–213
Mehmood A et al (2018) An orbital-overlap complement to ligand and binding site electrostatic potential maps. J Chem Inf Model 58(9):1836–1846
Menachery VD et al (2014) Attenuation and restoration of severe acute respiratory syndrome coronavirus mutant lacking 2′-O-methyltransferase activity. J Virol 88(8):4251–4264
Meng Z et al (2015) Total synthesis and antiviral activity of indolosesquiterpenoids from the xiamycin and oridamycin families. Nat Commun 6(1):1–8
Meo SA et al (2020) Biological and epidemiological trends in the prevalence and mortality due to outbreaks of novel coronavirus COVID-19. J King Saud Univ-Sci 32(4):2495–2499
Morris G, Huey R, Lindstrom W (2009) Sanner Mf.; Belew Rk.; Goodsell Ds.; Olson AJ AutoDock4 and AutoDockTools4: Automated Docking with Selective Receptor Flexibility. J Comput Chem 30(16): 2785–279110.1002
Muhammad S et al (2021) Exploring the new potential antiviral constituents of Moringa oliefera for SARS-COV-2 pathogenesis: an in silico molecular docking and dynamic studies. Chem Phys Lett 767:138379
Nelson MT et al (1996) NAMD: a parallel, object-oriented molecular dynamics program. Int J Supercomput Appl High Perf Comput 10(4):251–268
Pfaffenbach M et al (2019) Total syntheses of xiamycins A, C, F, H and oridamycin A and preliminary evaluation of their anti-fungal properties. Angew Chem 131(43):15448–15452
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58(9):4066–4072
Rosas-Lemus M et al. (2020) High-resolution structures of the SARS-CoV-2 2′-O-methyltransferase reveal strategies for structure-based inhibitor design. Sci Signal 13(651)
Rothan HA, Byrareddy SN (2020) The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J Autoimmunity 109:102433
Rzęsikowska K, Krawczuk A, Kalinowska-Tłuścik J (2019) Electrostatic potential and non-covalent interactions analysis for the design of selective 5-HT7 ligands. J Mol Graph Model 91:130–139
Samant L, Javle V (2020) Comparative docking analysis of rational drugs against COVID-19 main protease
Sangkanu S et al (2017) Evaluation of antibacterial potential of mangrove sediment-derived actinomycetes. Microb Pathog 112:303–312
Shilpi J et al. (2012) Antinociceptive, anti-inflammatory, and antipyretic activity of mangrove plants: a mini review. Adv Pharmacol Sci 2012
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31(2):455–461
Usha R et al (2010) Antimicrobial and antiangiogenesis activity of Streptomyces parvulus KUAP106 from mangrove soil. Eur J Biol Sci 2(4):77–83
Wei Z et al (2003) Study of Pt/C and Pt–Fe/C catalysts for oxygen reduction in the light of quantum chemistry. J Electroanal Chem 541:185–191
Xu D-B et al (2014) Natural products from mangrove actinomycetes. Mar Drugs 12(5):2590–2613
Yung-Chi C, Prusoff WH (1973) Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem Pharmacol 22(23):3099–3108
Zia M et al (2021) Exploring the potential of novel phenolic compounds as potential therapeutic candidates against SARS-CoV-2, using quantum chemistry, molecular docking and dynamic studies. Bioorg Med Chem Lett 43:128079
Acknowledgements
The authors extend their appreciation to the Deanship of Scientific Research in King Khalid University for supporting the work through Project RGP.1/168/42.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Muhammad, S., Qaisar, M., Iqbal, J. et al. Exploring the inhibitory potential of novel bioactive compounds from mangrove actinomycetes against nsp10 the major activator of SARS-CoV-2 replication. Chem. Pap. 76, 3051–3064 (2022). https://doi.org/10.1007/s11696-021-01997-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11696-021-01997-x