Abstract
The COVID-19 pandemic caused unprecedented damage to humanity, and while vaccines have been developed, they are not fully effective against the SARS-CoV-2 virus. Limited targeted drugs, such as Remdesivir and Paxlovid, are available against the virus. Hence, there is an urgent need to explore and develop new drugs to combat COVID-19. This study focuses on exploring microbial natural products from soil-isolated bacteria Streptomyces sp. strain 196 and RI.24 as a potential source of new targeted drugs against SARS-CoV-2. Molecular docking studies were performed on holoRdRp and nsp13, two key factors responsible for virus replication factor. Our in silico studies, K-252-C aglycone indolocarbazole alkaloid (K252C) and daunorubicin were found to have better binding affinities than the respective control drugs, with K252C exhibiting binding energy of − 9.1 kcal/mol with holoRdRp and − 9.2 kcal/mol with nsp13, and daunorubicin showing binding energy at − 8.1 kcal/mol with holoRdRp and − 9.3 kcal/mol with nsp13. ADMET analysis, MD simulation, and MM/GBSA studies indicated that K252C and daunorubicin have the potential to be developed as targeted drugs against SARS-CoV-2. The study concludes that K252C and daunorubicin are potential lead compounds that might suppress the inhibition of SARS-CoV-2 replication among the tested microbial compounds and could be developed as targeted drugs against COVID-19. In the future, further in vitro studies are required to validate these findings.
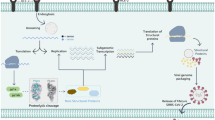
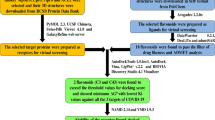
Graphical Abstract
















Similar content being viewed by others
Data availability
All data will be available on request.
References
Alam K, Mazumder A, Sikdar S et al (2022) Streptomyces: the biofactory of secondary metabolites. Front Microbiol 13:968053. https://doi.org/10.3389/fmicb.2022.968053
Al-Khodairy FM, Khan MK, Kunhi M et al (2013) In silico prediction of mechanism of erysolin-induced apoptosis in human breast cancer cell lines. A J Bioinform Res 3:62–71. https://doi.org/10.5923/j.bioinformatics.20130303.03
Angeletti S, Benvenuto D, Bianchi M et al (2020) COVID-2019: the role of the nsp2 and nsp3 in its pathogenesis. J Med Virol 92(6):584–588. https://doi.org/10.1002/jmv.25719
Asiedu SO, Kwofie SK, Broni E, Wilson MD (2021) Computational identification of potential anti-inflammatory natural compounds targeting the p38 mitogen-activated protein kinase (MAPK): implications for COVID-19-induced cytokine storm. Biomolecules 11(5):653. https://doi.org/10.3390/biom11050653
Atanasov AG, Zotchev SB, Dirsch VM et al (2021) Natural products in drug discovery: advances and opportunities. Nat Rev Drug Discov 20:200–216. https://doi.org/10.1038/s41573-020-00114-z
Chan JFW, Yuan S, Kok KH et al (2020) A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395(10223):514–523. https://doi.org/10.1016/S0140-6736(20)30154-9
Chen J, Malone B, Llewellyn E et al (2020) Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication-transcription complex. Cell 182(6):1560–1573.e13. https://doi.org/10.1016/j.cell.2020.07.033
Cobre AF, Maia Neto M, de Melo EB et al (2023) Naringenin-4′-glucuronide as a new drug candidate against the COVID-19 Omicron variant: a study based on molecular docking, molecular dynamics, MM/PBSA and MM/GBSA. J Biomol Struct Dyn 1–14. https://doi.org/10.1080/07391102.2023.2229446
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:42717. https://doi.org/10.1038/srep42717
Deshmukh N, Talkal R, Lakshmi B (2023) In silico screening of potential inhibitors from Cordyceps species against SARS-CoV-2 main protease. J Biomol Struct Dyn 1–17. https://doi.org/10.1080/07391102.2023.2225110
Ferdinands JM, Rao S, Dixon BE (2022) Waning of vaccine effectiveness against moderate and severe covid-19 among adults in the US from the VISION network: test negative, case-control study. BMJ 379:e072141. https://doi.org/10.1136/bmj-2022-072141
Hejazi II, Beg MA, Imam MA et al (2021) Glossary of phytoconstituents: can these be repurposed against SARS CoV-2? A quick in silico screening of various phytoconstituents from plant Glycyrrhiza glabra with SARS CoV-2 main protease. Food Chem Toxicol 150:112057. https://doi.org/10.1016/j.fct.2021.112057
Irshad R, Raj N, Gabr GA et al (2022) Integrated network pharmacology and experimental analysis unveil multi-targeted effect of 18α-glycyrrhetinic acid against non-small cell lung cancer. Front Pharmacol 13:1018974. https://doi.org/10.3389/fphar.2022.1018974
Khan A, Mohammad T, Shamsi A et al (2021) Identification of plant-based hexokinase 2 inhibitors: combined molecular docking and dynamics simulation studies. J Biomol Struct Dyn 40(20):10319–10331. https://doi.org/10.1080/07391102.2021.1942217
Krieger E, Vriend G (2015) New ways to boost molecular dynamics simulations. J Comput Chem 36(13):996–1007. https://doi.org/10.1002/jcc.23899
Kumar P, Kundu A, Kumar M et al (2019) Exploitation of potential bioactive compounds from two soil derived actinomycetes, Streptomyces sp. strain 196 and RI.24. Microbiol Res 229:126312. https://doi.org/10.1016/j.micres.2019.126312
Kwofie SK, Broni E, Asiedu SO et al (2021) Cheminformatics-based identification of potential novel anti-SARS-CoV-2 natural compounds of African origin. Molecules 26(2):406. https://doi.org/10.3390/molecules26020406
Liu DX, Liang JQ, Fung TS (2021) Human coronavirus-229E,-OC43,-NL63, and-HKU1 (Coronaviridae). Encyclopedia of Virology 428–440. https://doi.org/10.1016/B978-0-12-809633-8.21501-X
Markov PV, Ghafari M, Beer M et al (2023) The evolution of SARS-CoV-2. Nat Rev Microbiol 21:361–379. https://doi.org/10.1038/s41579-023-00878-2
Mathur S, Hoskins C (2017) Drug development: lessons from nature (review). Biomed Rep 6(6):612–614. https://doi.org/10.3892/br.2017.909
Mirza MU, Froeyen M (2020) Structural elucidation of SARS-CoV-2 vital proteins: computational methods reveal potential drug candidates against main protease, Nsp12 polymerase and Nsp13 helicase. J Pharm Anal 10(4):320–328. https://doi.org/10.1016/j.jpha.2020.04.008
Mohammad T, Siddiqui S, Shamsi A et al (2020) Virtual screening approach to identify high-affinity inhibitors of serum and glucocorticoid-regulated kinase 1 among bioactive natural products: combined molecular docking and simulation studies. Molecules 25(4):823. https://doi.org/10.3390/molecules25040823
Pascolutti M, Quinn RJ (2014) Natural products as lead structures: chemical transformations to create lead-like libraries. Drug Discov Today 19(3):215–221. https://doi.org/10.1016/j.drudis.2013.10.013
Patel CN, Goswami D, Jaiswal DG et al (2021) Pinpointing the potential hits for hindering interaction of SARS-CoV-2 S-protein with ACE2 from the pool of antiviral phytochemicals utilizing molecular docking and molecular dynamics (MD) simulations. J Mol Graph Model 105:107874. https://doi.org/10.1016/j.jmgm.2021.107874
Pires DE, Blundell TL, Ascher DB (2015) pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 58(9):4066–4072. https://doi.org/10.1021/acs.jmedchem.5b00104
Pitsillou E, Liang J, Hung A, Karagiannis TC (2022) The SARS-CoV-2 helicase as a target for antiviral therapy: identification of potential small molecule inhibitors by in silico modelling. J Mol Graph Model 114:108193. https://doi.org/10.1016/j.jmgm.2022.108193
Pollard AJ, Bijker EM (2021) A guide to vaccinology: from basic principles to new developments. Nat Rev Immunol 21:83–100. https://doi.org/10.1038/s41577-020-00479-7
Prasad N, Gopalakrishnan N, Sahay M, Gupta A, Agarwal SK et al (2020) Epidemiology, genomic structure, the molecular mechanism of injury, diagnosis and clinical manifestations of coronavirus infection: an overview. Indian J Nephrol 30(3):143–154. https://doi.org/10.4103/ijn.IJN_191_20
Rahimi A, Mirzazadeh A, Tavakolpour S (2021) Genetics and genomics of SARS-CoV-2: a review of the literature with the special focus on genetic diversity and SARS-CoV-2 genome detection. Genomics 113:1221–1232. https://doi.org/10.1016/j.ygeno.2020.09.059
Rasul HO, Thomas NV, Ghafour DD et al (2023) Searching possible SARS-CoV-2 main protease inhibitors in constituents from herbal medicines using in silico studies. J Biomol Struct Dyn 1–15. https://doi.org/10.1080/07391102.2023.2220040
Raubenolt BA, Islam NN, Summa CM et al (2022) Molecular dynamics simulations of the flexibility and inhibition of SARS-CoV-2 NSP 13 helicase. J Mol Graph Model 112:108122. https://doi.org/10.1016/j.jmgm.2022.108122
Rossi GA, Sacco O, Mancino E, Cristiani L, Midulla F (2020) Differences and similarities between SARS-CoV and SARS-CoV-2: spike receptor-binding domain recognition and host cell infection with support of cellular serine proteases. Infection 48(5):665–669. https://doi.org/10.1007/s15010-020-01486-5
Sargsyan K, Grauffel C, Lim C (2017) How molecular size impacts RMSD applications in molecular dynamics simulations. J Chem Theory Comput 13(4):1518–1524. https://doi.org/10.1021/acs.jctc.7b00028
Singh H, Dahiya N, Yadav M, Sehrawat N (2022) Emergence of SARS-CoV-2 new variants and their clinical significance. Can J Infect Dis Med Microbiol 2022:7336309. https://doi.org/10.1155/2022/7336309
Song Z, Xu Y, Bao L et al (2019) From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses 11(1):59. https://doi.org/10.3390/v11010059
Vardhan S, Sahoo SK (2022) Exploring the therapeutic nature of limonoids and triterpenoids against SARS-CoV-2 by targeting nsp13, nsp14, and nsp15 through molecular docking and dynamics simulations. J Tradit Complement Med 12(1):44–54. https://doi.org/10.1016/j.jtcme.2021.12.002
Vazquez C, Swanson SE, Negatu SG et al (2021) SARS-CoV-2 viral proteins NSP1 and NSP13 inhibit interferon activation through distinct mechanisms. PLoS ONE 16(6):e0253089. https://doi.org/10.1371/journal.pone.0253089
Vlasova AN, Saif LJ (2021) Bovine coronavirus and the associated diseases. Front Vet Sci 8:643220. https://doi.org/10.3389/fvets.2021.643220
Wang E, Sun H, Wang J et al (2019) End-point binding free energy calculation with MM/PBSA and MM/GBSA: strategies and applications in drug design. Design Chem Rev 119(16):9478–9508. https://doi.org/10.1021/acs.chemrev.9b00055
Wang B, Svetlov V, Wolf YI et al (2021) Allosteric activation of SARS-CoV-2 RdRp by remdesivir triphosphate and other phosphorylated nucleotides. bioRxiv
Wu F, Zhao S, Yu B, Chen YM, Wang W et al (2020) A new coronavirus associated with human respiratory disease in China. Nature 579(7798):265–269. https://doi.org/10.1038/s41586-020-2008-3
Xie S, Cao S, Wu J et al (2023) In silico-based screening of natural products as potential inhibitors of SARS-CoV-2 macrodomain 1. J Biomol Struct Dyn 1–9. https://doi.org/10.1080/07391102.2023.2226745
Xiong G, Wu Z, Yi J et al (2021) ADMETlab 2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties. Nucleic Acids Res 49(W1):W5–W14. https://doi.org/10.1093/nar/gkab255
Yin Y, Wunderink RG (2018) MERS, SARS and other coronaviruses as causes of pneumonia. Respirology 23(2):130–137. https://doi.org/10.1111/resp.13196
Yue K, Yao B, Shi Y et al (2022) The stalk domain of SARS-CoV-2 NSP13 is essential for its helicase activity. Biochem Biophys Res Commun 601:129–136. https://doi.org/10.1016/j.bbrc.2022.02.068
Zhu N, Zhang D, Wang W, Li X, Yang B et al (2020) A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382(8):727–733. https://doi.org/10.1056/NEJMoa2001017
Acknowledgements
Authors are thankful to Department of Zoology, University of Allahabad and Microbial Technology Lab, Acharya Narendra Dev College, University of Delhi.
Funding
The authors received no funding for this research.
Author information
Authors and Affiliations
Contributions
Formal analysis: PK, P, NR, MK and SK; investigation: PK, P and NR; methodology: PK, P, KUF and SK; supervision: PK, NM and MKK; visualization: PK and AMA; writing: original draft: PK, P, NR, MK, SK, KUF, AAK, RS and H.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
This declaration is “not applicable”.
Additional information
Communicated by Ran Wang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Note: Dr. Prateek Kumar is the first author as well as co-corresponding author in this manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumar, P., Parveen, Raj, N. et al. Natural products from Streptomyces spp. as potential inhibitors of the major factors (holoRdRp and nsp13) for SARS-CoV-2 replication: an in silico approach. Arch Microbiol 206, 88 (2024). https://doi.org/10.1007/s00203-023-03820-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03820-5