Abstract
The primate skull hosts a unique combination of anatomical features among mammals, such as a short face, wide orbits, and big braincase. Together with a trend to fuse bones in late development, these features define the anatomical organization of the skull of primates—which bones articulate to each other and the pattern this creates. Here, I quantified the anatomical organization of the skull of 17 primates and 15 non-primate mammals using anatomical network analysis to assess how the skulls of primates have diverged from those of other mammals, and whether their anatomical differences coevolved with brain size. Results show that primates have a greater anatomical integration of their skulls and a greater disparity among bones than other non-primate mammals. Brain size seems to contribute in part to this difference, but its true effect could not be conclusively proven. This supports the hypothesis that primates have a distinct anatomical organization of the skull, but whether this is related to their larger brains remains an open question.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The skull of primates shows a peculiar combination of anatomical features. Compared to other mammals, primates in general have shortened snouts and wide forward-facing eye orbits (Fleagle et al., 2010). Primates also tend toward a more vertical posture regardless of locomotion behavior, which affected their cranial base morphology (Fleagle, 1999). One of the most remarkable feature of primates’ evolution is the enlargement of the brain relative to body size, especially in some groups like anthropoids (Boddy et al., 2012). Evolving bigger brains imposes functional and anatomical trade-offs among the different parts of the skull. For example, the fact that the back of the face is physically integrated with the cranial base limits changes in the orientation of the cranial base and in encephalization (Lieberman, 2011; Lieberman et al., 2000; McCarthy & Lieberman, 2001).
The loss of ossification centers and the fusion of bones during development has also influenced the anatomy of the primate skull (Esteve-Altava et al., 2013; Goodrich, 1931; Kardong, 2005; Koyabu et al., 2012, 2014; Rager et al., 2014; Richtsmeier et al., 2006). Modern mammals have lost bones that were once present in their synapsid ancestors, such as the prefrontal, postfrontal, or postorbital (Esteve-Altava et al., 2014; Sidor, 2001). In turn, primates fuse bones that tend to remain unfused in other mammals, resulting in bones such as the occipital, temporal, and sphenoid (Smith et al., 2020); haplorhines also fuse the left and right frontals into a single frontal bone (Rosenberger & Pagano, 2008), and in humans the premaxilla fuses to the maxilla (Lang, 1995). Comparative studies in tetrapods (Esteve-Altava et al., 2013), synapsids (Sidor, 2001), and archosaurs (Lee et al., 2020; Plateau & Foth, 2020; Sookias et al., 2020; Werneburg et al., 2019) have shown that the loss and fusion of bones can modify the anatomical organization of the skull—which bones articulate to each other and the pattern of interactions this creates—in a way that increases the anatomical integration of the skull and the disparity among bones through evolution. In this context, integration and disparity refer to the aforementioned anatomical organization: integration refers to how strongly skull bones are interconnected (as in number of connections among elements and of 3-node loops), and disparity refers to how different or heterogeneous bones are in terms of number of connections. These general properties can be proxy-measured in different ways (see Box 1). In turn, we can understand modularity as a pattern of structured integration, with groups or clusters of bones more integrated (connected) among them than to bones in other clusters, which allows different regions of the skull to still evolve semi-independently (Esteve-Altava, 2017a). Indeed, studies using network models found that anatomical integration is strong in the skull of primates (Esteve-Altava et al., 2015; Powell et al., 2018). However, these studies did not compare primates with non-primate mammals. Curiously, changes in anatomical organization that increase integration and disparity among bones seem to be more common in lineages that evolved bigger brains and globular neurocrania (e.g., birds compared to archosaurs (Marugán‐Lobón et al., 2016; Plateau & Foth, 2020), which is probably a consequence of the strong ties between brain and skull development (Richtsmeier & Flaherty, 2013). This relationship is even stronger for the neurocranial bones which intramembranous growth is linked to brain growth (Opperman, 2000). Building on that, we can propose the working hypothesis that primates have an anatomical organization distinct from that of other mammals, which is related to bigger brains.
Here, I quantified the anatomical organization of the skull of primates and non-primate mammals to test: (i) whether primates show a distinctive organization of the skull that sets them apart from other mammals; if so, (ii) what specific anatomical features create this difference; and (iii) how such anatomical features relate to the increase in brain size relative to body size characteristic of primates. To this end, the skulls of 17 primates and 15 non-primate mammals were modelled as networks and their anatomical organization measured using six topological variables. See Fig. 1 for a visual schema of the topological variables and the list of species compared. Topological variables were compared in a phylogenetic context to identify differences between primates and other non-primate mammals and to assess the relationship between cranial anatomical organization and brain size.
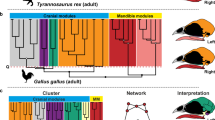
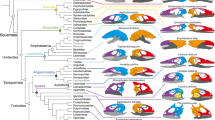
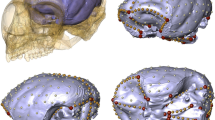
Modeling approach and phylogenetic context. A Network model of the human skull. Nodes represent bones and links represent physical joints (sutures and synchondroses). Topological variables help us to measure the structure and organization of network models. Their actual meaning depends on the nature of the relations that are being modelled as links (Butts, 2009). In skull networks links represent sutures and synchondroses, which embody growth and functional relations among bones (Herring & Teng, 2000; Opperman, 2000; Rafferty et al., 2003; Di Ieva et al., 2013), therefore, topological variables translate into morphological features that depend on growth and function, such as integration, modularity, and disparity (see Box 1). The six topological variables used here are illustrated with colors. D is the number of links (in gray) given the maximum number of links possible, which depends on the total number of nodes. C is the number of 3-node loops given the maximum number of loops possible (in blue, the 3-node loop between temporal, parietal, and occipital). L is the mean of all shortest paths (in green, shortest path of length two between the left maxilla and temporal). P is the even partition of nodes into many modules (dashed red lines indicate the three modules identified using an optimization algorithm, see Methods). H is the heterogeneity of connections among nodes and A is the correlation of connectivities: note how bones vary in their connections (the black circle size within nodes is proportional to their number of connections relative to the most connected node, the ethmoid with 13); in yellow, the connections of the four links of the lacrimal and of the 12 links of the sphenoid illustrate the two extremes of the variation. B Evolutionary relations of sampled mammals. The skulls of primates (including representatives of the Strepsirrhini and Haplorhini suborders) was compared with a diverse set of mammals spanning 12 orders, including marsupials and monotremes (Color figure online)
Materials and Methods
Anatomical Network Models
Anatomical network models of the typical young adult skull of 17 primates and nine non-primate mammals were gathered from existing literature (Esteve-Altava et al., 2013, 2015). The original studies defined age by dentition and ‘typical’ by the frequency of presenting or not an articulation, which tends to be a relatively stable feature. Six additional non-primates were coded to broaden the comparison: Antilope cervicapra, Dasypus novemcinctus, Equus caballus, Felis silvestris, Oryctolagus cuniculus, and Sus scrofa (see details in Supplementary Materials). Figure 1B shows the complete list of species. Following the modeling approach of previous studies (Rasskin-Gutman & Esteve-Altava, 2014): every bone was coded as a network node and every suture or synchondrosis between two bones was coded as a link connecting two nodes. Networks available at https://doi.org/10.6084/m9.figshare.14662257.v1.
Topological Variables
Six topological variables were measured: density of connections (D), mean clustering coefficient (C), mean shortest path length (L), heterogeneity (H), assortativity (A), and parcellation (P). D is the actual number of connections divided by the maximum theoretically possible given the number of nodes; D ranges from 0 to 1. C is the actual number of 3-node loops divided by the maximum theoretically possible given the number of nodes; C ranges from 0 to 1. L is the averaged minimum number of links separating all pairs of nodes divided by the number of pairs; L equals 1 when all nodes are directly connected and gets greater with longer distances between pairs of nodes. H is the coefficient of variation (σ/μ) of the number of connections per node; H ranges from 0 (all nodes have the same number of connections) to \(\sqrt{N-1}\) (maximum disparity among nodes’ connections). A is a preference for the nodes of a network to attach to other nodes that have a similar number of neighbors.; A ranges from -1 (dissimilar connectivity) to 1 (similar connectivity). Finally, P measures the extent to which nodes are evenly distributed into modules; P ranges from 0 (no modules) to 1 (as the number of modules increases). Networks have a higher P when they have more modules and also when nodes are similarly spread among them (e.g., a network with four modules of five nodes each has a higher parcellation than a network with one module of 17 nodes and three modules of two nodes). For a mathematical description of topological variables see (Esteve-Altava et al., 2019). Topological variables were measured using functions from the R package igraph (Csardi & Nepusz, 2006) and each variable rescaled for the whole sample using min–max normalization.
Calibrated Phylogeny and Brain Size Residuals
A consensus calibrated phylogeny for the 32 species was built from 100 trees pruned from the VertLife project (Upham et al., 2019) (Supplementary Data), using the mean edges method as implemented in the function consensus.edges of the package phytools (Revell, 2012). Brain size residuals were collected from a recent large-scale compilation by Burger et al. (2019), which includes all species in the study except Cynocephalus volans. This study reports an allometric relation of \(Brain=-1.26\times {Body}^{0.75}\), which I used to estimate the residual of C. volans using brain and body mass measurements from Lewitus et al. (2014). Raw data is available at https://doi.org/10.6084/m9.figshare.14662257.v1 (phylogeny: output2.nex; brain data: Burgeretal2019brainallometry.tsv).
Statistical Analysis
All statistical analyses and visualizations were performed in R (R Core Team, 2019) (available within Supplementary Results). A principal components analysis (PCA) was performed (without rotation) on the six topological variables with the function principal from the package psych (Revelle, 2019). The relationship between number of components, % of explained variance, and the load of variables on components were examined to decide the final number of components, as a compromise between capturing as much % of explained variance as possible but with each variable loading mostly onto only one component (Online Resource, Fig. S3). As a result, two obliquely rotated components were extracted (RC1 and RC2). Results were visualized using a phylomorphospace projection using the homonym function of the package phytools (Revell, 2012). Differences between primates and non-primates were tested with phylogenetic permutational multivariate analyses of variance (PERMANOVA) on raw topological variables and RCs, using the function aov.phylo of the package geiger (Harmon et al., 2008) with 10,000 permutations.
The association between RCs and brain size residuals was tested for the whole sample and for each group separately using phylogenetic methods under the Pagel’s λ model. Phylogenetic Pearson’s product moment correlations were calculated with a custom script, using the function phyl.vcv in phytools to measure the variance–covariance matrix of variables. Phylogenetic generalized least squares (PGLS) regressions were performed using the function gls from the package nlme (Pinheiro et al., 2019). To test whether a single regression for the whole sample fits the data better than separate regression models for primates and non-primates, a phylogenetic analysis of covariance (PANCOVA) was performed using the function gls.ancova of the package evomap (Smaers & Mongle, 2020).
The assumption of normality of the residual error for PGLS models was tested with a Lilliefors (Kolmogorov–Smirnov) test. Outliers were first visualized with boxplots (interquartile rule) and tested using PANCOVA, which has been shown to be the only correct way to directly test whether a single species is an outlier (Smaers & Rohlf, 2016).
Results
Anatomical Network Analysis
The 32 skull network models analyzed range in number of bones between 21 and 32 and in number of articulations between 47 and 99 (Online Resource, Table S1), producing a diverse set of networks, as captured by topological variables (Online Resource, Table S2). The phylogenetic PERMANOVA test supported a statistical difference in topological variables between primates and non-primates (Wilks’ lambda = 0.1596, F6,25 = 21.9322, p-value = 0.0097). A PCA of the topological variables was performed to visualize the overall pattern of variation across skulls and to select an adequate number of components. Two PCs were enough to explain 78.71% of the variance without increasing variable loads on a single PC above 1.5 (Online Resource, Fig. S3), which makes possible to interpret components anatomically because one variable is captured largely by only one component.
A PCA with oblique rotation was performed to extract the two main rotated components (RC). RC1 and RC2 explained 49.96% and 28.74% of the variance, respectively (Online Resource, Table S4). RC1 captured topological variables D, C, L, and P (Online Resource, Table S5), which are related to anatomical integration (Box 1). Greater values of RC1 align with greater values of D and C, and lower values of L and P; thus, RC1 ranks skull networks from less to more anatomically integrated. In turn, RC2 captured the variables H and A (Online Resource, Table S5), which are related to anatomical disparity (Box 1). Greater values of RC2 align with greater values of H and lower values of A; thus, RC2 ranks skull networks from less to more disparate. When analyzed in a phylogenetic context (Fig. 2), RCs also discriminate significatively between primates and non-primates (Wilks’ lambda = 0.3865, F2,29 = 23.012, p-value = 0.0298). These results support the working hypothesis that primates have an anatomical organization distinct from that of other mammals.
Phylomorphospace of skull network models defined by the two rotated components. Primates and non-primates occupy different regions of the space, with primates having larger values of RC1 and RC2 meaning that they have more anatomically integrated skulls and a greater disparity among bones than the other non-primate mammals sampled. Interestingly, the bat Pteropus directly occupies a position within the space occupied by primates (dashed line circle). The fact that bats share with primates a pervasive pattern of bone fusions during development may explain having a similar degree of anatomical disparity and disparity among bones
Network Anatomy and Brain Size
There was a significant positive phylogenetic correlation between RC1 and brain size residuals for the species in the sample (r = 0.5467, t = 3.5766, p-value = 0.0012). The PGLS of brain size residuals on RC1 confirmed bigger brains predict greater values of RC1 (slope = 1.3769 ± 0.6327, t = 2.1763, p-value = 0.0375; Online Resource, Fig. S6) in mammals. This baseline model fitting a single slope and intercept for the whole sample was compared with alternative models where primates and non-primates fit to different intercepts and/or slopes (Fig. 3). PANCOVA tests showed that none of the alternative models fitted the data significantly better than the baseline model (Online Resource, Tables S10, S11, S12). A closer examination of the relation between variables for primates and non-primates (Online Resource, Tables S13, S14, S15, S16) showed that only primates had a significant correlation between RC1 and brain size residuals (r = 0.6356, t = 3.1888, p-value = 0.0061), with a linear relation similar to that of the baseline model (slope = 2.2078 ± 0.8786, t = 2.5129, p-value = 0.0239). This may indicate that the correlation seen in the whole sample is mainly driven by primates alone. These results support an alternative working hypothesis, in which brain size affects anatomical integration overall, but brain size cannot explain differences of integration between primates and non-primates.
PGLS fits of brain size on anatomical integration. The baseline model (dashed black line) fitted the data better than the alternative models where primates (in green) and non-primates (in blue) have different slope and/or intercept. Note that the fit represented by the blue like is not significant and it has been included only for comparison and to show the alternative models compared (Color figure online)
PGLS residuals of brain size on RC2 failed the Lilliefors test of normality (D = 0.1788, p-value = 0.0107) and showed two outliers (Pan and Cercopithecus), visually identified using the interquartile rule and confirmed with a PANCOVA for differences in intercept between the outliers and the whole sample (F2,3 = 29.9505, p-value < 0.0001). After removing the confirmed outliers the PGLS residuals passed the normality test and showed no additional outliers (Online Resource, Table S19). However, no significant relationship was found between brain size residuals and RC2 for the whole sample (Online Resource, Tables S19 and S20) nor for primates and non-primates separately (Online Resource, Tables S21, S22, S23, S24). Thus, rejecting the working hypothesis that anatomical disparity of the skull is driven by brain size.
Discussion
These results support three points: (i) topological variables measured on skull network models can discriminate between primates and other non-primates mammals based on their anatomical organization; (ii) two features distinguish primates: a greater anatomical integration of the skull and a greater disparity among bones; and (iii) brain size seems to influence anatomical organization differently in primates and non-primates, but its true effect could not be conclusively demonstrated. Results support the working hypothesis that primates have a distinct anatomical organization of the skull compared to non-primate mammals, but whether this is related to primates evolving bigger brains remains an open question.
Primates have skulls with greater anatomical integration and disparity than the other mammals studied. This finding builds on the morphological interpretation of topological variables (Box 1) and their respective loading on each RC (Online Resource, Table S5). As introduced before, topological variables capture anatomical integration and disparity because connections in skull network models embody growth and functional relations between bones (Herring & Teng, 2000; Opperman, 2000; Rafferty et al., 2003; Di Ieva et al., 2013). Having a greater cranial anatomical integration means that, collectively, the bones of the skull interact more with one another, which affects form and function. Because a connection is a shared growth site between two bones (Opperman, 2000), changes in the shape of one bone, caused by genetic or epigenetic factors modifying growth, are more likely to produce a collateral change in an adjacent bone (Pearson & Woo, 1935; Woo, 1931), and indirectly to bones adjacent to that one (and so on). Therefore, the more structurally integrated (i.e., interconnected) is the whole skull, the stronger its shape integration and the weaker its modularity. This type of relationship between anatomy and shape would be a baseline expectation (actually, one that remains untested), with other factors also weighing in to determine the final phenotypic variation of the skull.
As related as they may seem, anatomical or structural integration and morphological integration are not interchangeable concepts, because the actual relation between organizational units (i.e., network anatomy) and variational units (i.e., shape co-variation) is not yet fully understood (Eble, 2005; Esteve-Altava, 2017b; Rasskin-Gutman & Buscalioni, 2001). Patterns of connectivity emerge from, and act on, growth and function: two of the main drivers of shape variation and morphological adaptation. The same is true for morphological integration (as shape co-variation), which is also characteristically strong in the skull of primates (Neaux et al., 2018, 2019).
From a more functional perspective, bone connections act also as sites of energy absorption and deformation (Herring, 2008), working together to diffuse strains across the skull (Moazen et al., 2009). Thus, we could speculate that the more interconnected is the skull, the more efficient its biomechanical functioning in terms of redistributing stress loads. Additionally, the manner in which topological variables capture anatomical integration (Box 1) is related to a kind of structural compactness or robustness (see Sidor, 2001 for a discussion of this argument in relation to bone losses in the skull of synapsids). Thus, as primates evolved bigger brains the need for more robust and protective skulls might have co-evolved.
On the other hand, having a greater disparity (in number of connections) among bones means that there are some bones that bear many interactions while others bear few. Results indicate this is what happens in primates, as it happens in modern mammals compared to stem mammals (Esteve-Altava et al., 2013).. In mammals, the number of connections seems to be related to the midline location of bones, which allows to articulate with bones from both sides of the skull; however, there are enough exceptions that prevent us to establish a constant relationship between location and connectivity. For example, in the human skull the ethmoid, frontal, and sphenoid have 13, 12, and 12 links, while most bones have 4–6 links (Fig. 1); the other two unpaired bones, vomer and occipital, have an average number of connections. Network studies have shown that systems with a few highly connected nodes and many poorly connected nodes are more robust to accidental damage (e.g., injury or loss), because accidents are more likely to happen to the most abundant type of node, which are the less connected ones. In contrast, the same systems are sensible to damage targeting the most connected nodes (Albert et al., 2000; Allesina et al., 2009), because they are more essential. This phenomenon has also been demonstrated in anatomical networks (Esteve-Altava et al., 2013, 2014; Murphy et al., 2018). Because more connected bones bear more growth and functional interactions, they are more constrained to change (Rasskin-Gutman & Esteve-Altava, 2018; Schoch, 2010). In contrast, bones with fewer connections can vary, specialize, or get lost more freely without compromising the integrity of the skull. However, when considering the potential effects of connections in constraining bones variation, it is important to also consider their relations with the surrounding soft tissues and functional matrices, in particular, the central neural and circulatory systems (Esteve-Altava & Rasskin-Gutman, 2014).
In mammals, a greater anatomical integration of the skull is related to bigger brains. The mechanisms by which bigger brains may produce more integrated skulls is unknown. A plausible explanation points to the role of the brain as a functional matrix in development (Moss & Young, 1960), and the way in which the bones and articulations of the neurocranium respond to brain growth (Lesciotto & Richtsmeier, 2019; Richtsmeier & Flaherty, 2013; Richtsmeier et al., 2006). It is also possible that bigger brains impose a selective pressure for more robust and protective skulls, which can be achieved by closing sutures and fusing bones together (Sidor, 2001), incidentally increasing anatomical integration and sometimes anisomerism (Esteve-Altava et al., 2014). Given that many lineages of primates also show trends toward bigger orbits and shorter faces (Smith et al., 2020), bigger brains may also produce a tighter packing of midfacial and basicranial bones (Bastir et al., 2010; Lesciotto & Richtsmeier, 2019). At the level of skull sutures and synchondroses, compression creates an environment that favors suture closure, for example, it enhances osteogenesis, narrows suture space, and immobilizes bones (Herring, 2008), which, as said before, is a process known to increase anatomical integration (Esteve-Altava et al., 2014). The correlation between anatomical integration and bigger brains is mostly driven by primates; non-primate mammals show no relationship between brain size and cranial integration. This is mostly because primates have a statistically significant linear fit between brain size and integration, while non-primates do not. To sustain this preliminary result, future studies will need to look at this relationship with similarly large samples in other orders of mammals.
In summary, network models and topological variables paint a picture of the skull that is not accessible to, and complements that of, other methods in morphology (Eble, 2005; Rasskin-Gutman & Esteve-Altava, 2014). Primates evolved a particular anatomical organization—in liaison with bigger brains?—that distinguish them from non-primate mammals. Future studies should address whether other lineages with relatively big brains, such as dolphins within cetaceans (Marino, 2009) or corvids within birds (Uomini et al., 2020), convergently evolved a similar relationship between cranial anatomy and brain size. On a more general note, the results presented here add up to a growing body of research that seeks to better understand morphological evolution, development, and function by using methods that allow us to studying the body’s anatomy as a complex biological system (Esteve-Altava et al., 2013, 2014, 2017, 2018, 2019; Saucède et al., 2015; Dos Santos et al., 2017; Kerkman et al., 2018; Murphy et al., 2018; Werneburg et al., 2019; Fernández et al., 2020; Plateau & Foth, 2020; Sookias et al., 2020).
Data Availability
All data and code are included in the submission.
References
Albert, R., Jeong, H., & Barabási, A.-L. (2000). Error and attack tolerance of complex networks. Nature, 406, 378–382.
Allesina, S., Bodini, A., & Pascual, M. (2009). Functional links and robustness in food webs. Philosophical Transactions of the Royal Society b: Biological Sciences, 364, 1701–1709.
Bastir, M., Rosas, A., Stringer, C., Manuel Cuétara, J., Kruszynski, R., Weber, G. W., et al. (2010). Effects of brain and facial size on basicranial form in human and primate evolution. Journal of Human Evolution, 58, 424–431.
Boddy, A. M., McGowen, M. R., Sherwood, C. C., Grossman, L. I., Goodman, M., & Wildman, D. E. (2012). Comparative analysis of encephalization in mammals reveals relaxed constraints on anthropoid primate and cetacean brain scaling. Journal of Evolutionary Biology, 25, 981–994.
Burger, J. R., George, M. A., Leadbetter, C., & Shaikh, F. (2019). The allometry of brain size in mammals. Journal of Mammalogy, 100, 276–283.
Butts, C. T. (2009). Revisiting the foundations of network analysis. Science, 325, 414–416.
Csardi, G., & Nepusz, T. (2006). The igraph software package for complex network research. InterJournal, 1695, 1–9.
Di Ieva, A., Bruner, E., Davidson, J., Pisano, P., Haider, T., Stone, S. S., et al. (2013). Cranial sutures: A multidisciplinary review. Child’s Nervous System, 29, 893–905.
Dos Santos, D. A., Fratani, J., Ponssa, M. L., & Abdala, V. (2017). Network architecture associated with the highly specialized hindlimb of frogs. PLoS ONE, 12, e0177819.
Eble, G. J. (2005). Morphological modularity and macroevolution: conceptual and empirical aspects. In D. Rasskin-Gutman & W. Callebaut (Eds.), Modularity: Understanding the development and evolution of natural complex systems (pp. 221–238). The MIT Press.
Esteve-Altava, B. (2017a). In search of morphological modules: A systematic review. Biological Reviews of the Cambridge Philosophical Society, 92, 1332–1347.
Esteve-Altava, B. (2017b). Challenges in identifying and interpreting organizational modules in morphology. Journal of Morphology, 278, 960–974.
Esteve-Altava, B., & Rasskin-Gutman, D. (2014). Beyond the functional matrix hypothesis: A network null model of human skull growth for the formation of bone articulations. Journal of Anatomy, 225, 306–316.
Esteve-Altava, B., Boughner, J. C., Diogo, R., Villmoare, B. A., & Rasskin-Gutman, D. (2015). Anatomical network analysis shows decoupling of modular lability and complexity in the evolution of the primate skull. PLoS ONE, 10, e0127653.
Esteve-Altava, B., Marugán-Lobón, J., Botella, H., & Rasskin-Gutman, D. (2013). Structural constraints in the evolution of the tetrapod skull complexity: Williston’s law revisited using network models. Evolutionary Biology, 40, 209–219.
Esteve-Altava, B., Marugán-Lobón, J., Botella, H., & Rasskin-Gutman, D. (2014). Random loss and selective fusion of bones originate morphological complexity trends in tetrapod skull networks. Evolutionary Biology, 41, 52–61.
Esteve-Altava, B., Molnar, J. L., Johnston, P., Hutchinson, J. R., & Diogo, R. (2018). Anatomical network analysis of the musculoskeletal system reveals integration loss and parcellation boost during the fins-to-limbs transition. Evolution, 72, 601–618.
Esteve-Altava, B., Pierce, S. E., Molnar, J. L., Johnston, P., Diogo, R., & Hutchinson, J. R. (2019). Evolutionary parallelisms of pectoral and pelvic network-anatomy from fins to limbs. Science Advances, 5, eaau7459.
Esteve-Altava, B., Vallès-Català, T., Guimerà, R., Sales-Pardo, M., & Rasskin-Gutman, D. (2017). Bone fusion in normal and pathological development is constrained by the network architecture of the human skull. Scientific Reports, 7, 3376.
Fernández, M. S., Vlachos, E., Buono, M. R., Alzugaray, L., Campos, L., Sterli, J., et al. (2020). Fingers zipped up or baby mittens? Two main tetrapod strategies to return to the sea. Biology Letters, 16, 20200281.
Fleagle, J. G. (1999). Primate adaptation and evolution. Academic Press.
Fleagle, J. G., Gilbert, C. C., & Baden, A. L. (2010). Primate cranial diversity. American Journal of Physical Anthropology, 142, 565–578.
Goodrich, E. S. (1931). Studies on the structure and development of vertebrates. Dover.
Harmon, L. J., Weir, J. T., Brock, C. D., Glor, R. E., & Challenger, W. (2008). GEIGER: Investigating evolutionary radiations. Bioinformatics (oxford, England), 24, 129–131.
Herring, S. W. (2008). Mechanical influences on suture development and patency. In D. P. Rice (Ed.), Craniofacial sutures. Development, disease and treatment, frontiers of oral biology (pp. 41–56). S. KARGER AG.
Herring, S. W., & Teng, S. (2000). Strain in the braincase and its sutures during function. American Journal of Physical Anthropology, 112, 575–593.
Kardong, K. V. (2005). Vertebrates: Comparative anatomy, function and evolution (4th ed.). MacGraw-Hill.
Kerkman, J. N., Daffertshofer, A., Gollo, L. L., Breakspear, M., & Boonstra, T. W. (2018). Network structure of the human musculoskeletal system shapes neural interactions on multiple time scales. Science Advances, 4, eaat0497.
Koyabu, D., Maier, W., & Sanchez-Villagra, M. R. (2012). Paleontological and developmental evidence resolve the homology and dual embryonic origin of a mammalian skull bone, the interparietal. Proceedings of the National Academy of Sciences USA, 109, 14075–14080.
Koyabu, D., Werneburg, I., Morimoto, N., Zollikofer, C. P. E., Forasiepi, A. M., Endo, H., et al. (2014). Mammalian skull heterochrony reveals modular evolution and a link between cranial development and brain size. Nature Communications, 5, 1–9.
Lang, J. (1995). Clinical anatomy of the masticatory apparatus and peripharyngeal spaces. Thieme.
Lee, H. W., Esteve-Altava, B., & Abzhanov, A. (2020). Evolutionary and ontogenetic changes of the anatomical organization and modularity in the skull of archosaurs. Scientific Reports, 10, 16138.
Lesciotto, K. M., & Richtsmeier, J. T. (2019). Craniofacial skeletal response to encephalization: How do we know what we think we know? American Journal of Physical Anthropology, 168, 27–46.
Lewitus, E., Kelava, I., Kalinka, A. T., Tomancak, P., & Huttner, W. B. (2014). An adaptive threshold in mammalian neocortical evolution. PLoS Biology, 12, e1002000.
Lieberman, D. E. (2011). The evolution of the human head. Belknap Press Harvard University Press.
Lieberman, D. E., Ross, C. F., & Ravosa, M. J. (2000). The primate cranial base: Ontogeny, function, and integration. American Journal of Physical Anthropology, 113, 117–169.
Marino, L. (2009). Brain size evolution. In W. F. Perrin, B. Würsig, & J. G. M. Thewissen (Eds.), Encyclopedia of Marine Mammals (pp. 149–152). Academic Press.
Marugán-Lobón, J., Watanabe, A., & Kawabe, S. (2016). Studying avian encephalization with geometric morphometrics. Journal of Anatomy, 229, 191–203.
McCarthy, R. C., & Lieberman, D. E. (2001). Posterior maxillary (PM) plane and anterior cranial architecture in primates. The Anatomical Record, 264, 247–260.
Moazen, M., Curtis, N., O’Higgins, P., Jones, M. E. H., Evans, S. E., & Fagan, M. J. (2009). Assessment of the role of sutures in a lizard skull: A computer modelling study. Proceedings of the Royal Society b: Biological Sciences, 276, 39–46.
Moss, M. L., & Young, R. W. (1960). A functional approach to craniology. American Journal of Physical Anthropology, 18, 281–292.
Murphy, A. C., Muldoon, S. F., Baker, D., Lastowka, A., Bennett, B., Yang, M., et al. (2018). Structure, function, and control of the human musculoskeletal network. PLOS Biology, 16, e2002811.
Neaux, D., Sansalone, G., Ledogar, J. A., Heins Ledogar, S., Luk, T. H. Y., & Wroe, S. (2018). Basicranium and face: Assessing the impact of morphological integration on primate evolution. Journal of Human Evolution, 118, 43–55.
Neaux, D., Wroe, S., Ledogar, J. A., Ledogar, S. H., & Sansalone, G. (2019). Morphological integration affects the evolution of midline cranial base, lateral basicranium, and face across primates. American Journal of Physical Anthropology, 170, 37–47.
Opperman, L. A. (2000). Cranial Sutures as intramembranous bone growth sites. Developmental Dynamics, 219, 472–485.
Pearson, K., & Woo, T. L. (1935). Further investigation of the morphometric characters of the individual bones of the human skull. Biometrika, 27, 424.
Pinheiro, J., Bates, D., DebRoy, S., Sarkar, D., & R Core Team. (2019). nlme: Linear and nonlinear mixed effects models. R package version 3.1-153. https://CRAN.R-project.org/package=nlme
Plateau, O., & Foth, C. (2020). Birds have peramorphic skulls, too: Anatomical network analyses reveal oppositional heterochronies in avian skull evolution. Communications Biology, 3, 1–12.
Powell, V., Esteve-Altava, B., Molnar, J., Villmoare, B., Pettit, A., & Diogo, R. (2018). Primate modularity and evolution: First anatomical network analysis of primate head and neck musculoskeletal system. Scientific Reports, 8, 2341.
R Core Team. (2019). R: A language and environment for statistical computing. R Foundation for Statistical Computing.
Rafferty, K. L., Herring, S. W., & Marshall, C. D. (2003). Biomechanics of the rostrum and the role of facial sutures. Journal of Morphology, 257, 33–44.
Rager, L., Hautier, L., Forasiepi, A., Goswami, A., & Sánchez-Villagra, M. R. (2014). Timing of cranial suture closure in placental mammals: Phylogenetic patterns, intraspecific variation, and comparison with marsupials. Journal of Morphology, 275, 125–140.
Rasskin-Gutman, D., & Buscalioni, A. D. (2001). Theoretical morphology of the Archosaur (Reptilia: Diapsida) pelvic girdle. Paleobiology, 27, 59–78.
Rasskin-Gutman, D., & Esteve-Altava, B. (2014). Connecting the dots: Anatomical network analysis in morphological evo-devo. Biological Theory, 9, 178–193.
Rasskin-Gutman, D., & Esteve-Altava, B. (2018). Concept of burden in evo-devo. In L. N. de la Rosa & G. Müller (Eds.), Evolutionary developmental biology: A reference guide (pp. 1–11). Cham: Springer.
Revell, L. J. (2012). phytools: An R package for phylogenetic comparative biology (and other things). Methods in Ecology and Evolution, 3, 217–223.
Revelle, W. R. (2019). psych: Procedures for personality and psychological research. Northwestern University.
Richtsmeier, J. T., & Flaherty, K. (2013). Hand in glove: Brain and skull in development and dysmorphogenesis. Acta Neuropathologica, 125, 469–489.
Richtsmeier, J. T., Aldridge, K., DeLeon, V. B., Panchal, J., Kane, A. A., Marsh, J. L., et al. (2006). Phenotypic integration of neurocranium and brain. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution, 306B, 360–378.
Rosenberger, A. L., & Pagano, A. S. (2008). Frontal fusion: Collapse of another anthropoid synapomorphy. The Anatomical Record, 291, 308–317.
Saucède, T., Laffont, R., Labruère, C., Jebrane, A., François, E., Eble, G. J., et al. (2015). Empirical and theoretical study of atelostomate (Echinoidea, Echinodermata) plate architecture: Using graph analysis to reveal structural constraints. Paleobiology, 41, 436–459.
Schoch, R. R. (2010). Riedl’s burden and the body plan: Selection, constraint, and deep time. Journal of Experimental Zoology Part b: Molecular and Developmental Evolution, 314B, 1–10.
Sidor, C. A. (2001). Simplification as a trend in synapsid cranial evolution. Evolution, 55, 1419–1442.
Smaers, J., & Mongle, C. (2020) Evomap: r package for the evolutionary mapping of continuous traits. Available from https://github.com/JeroenSmaers/evomap
Smaers, J. B., & Rohlf, F. J. (2016). Testing species’ deviation from allometric predictions using the phylogenetic regression. Evolution, 70, 1145–1149.
Smith, T. D., DeLeon, V. B., Vinyard, C. J., & Young, J. W. (2020). The skeletal anatomy of the newborn primate. Cambridge University Press.
Sookias, R. B., Dilkes, D., Sobral, G., Smith, R. M. H., Wolvaardt, F. P., Arcucci, A. B., et al. (2020). The craniomandibular anatomy of the early archosauriform Euparkeria capensis and the dawn of the archosaur skull. Royal Society Open Science, 7, 200116.
Uomini, N., Fairlie, J., Gray, R. D., & Griesser, M. (2020). Extended parenting and the evolution of cognition. Philosophical Transactions of the Royal Society B: Biological Sciences, 375, 20190495.
Upham, N. S., Esselstyn, J. A., & Jetz, W. (2019). Inferring the mammal tree: Species-level sets of phylogenies for questions in ecology, evolution, and conservation. PLoS Biology, 17, e3000494.
Werneburg, I., Esteve-Altava, B., Bruno, J., Torres Ladeira, M., & Diogo, R. (2019). Unique skull network complexity of Tyrannosaurus rex among land vertebrates. Scientific Reports, 9, 1520.
Woo, T. L. (1931). On the asymmetry of the human skull. Biometrika, 22, 324.
Acknowledgements
I thank Dr. Jeroen Smaers for his feedback on the phylogenetic analyses. Finally, I would like to thank Dr. Diego Rasskin-Gutman his contribution in developing anatomical network analysis and for his continuous support as a mentor and friend over the years. BE-A has received financial support through the Postdoctoral Junior Leader Fellowship Programme from “la Caixa” Banking Foundation (LCF/BQ/LI18/11630002).
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature.
Author information
Authors and Affiliations
Contributions
BE-A designed the study, collected and analyzed the data, interpreted the results, prepared the figures, and wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Esteve-Altava, B. Cranial Anatomical Integration and Disparity Among Bones Discriminate Between Primates and Non-primate Mammals. Evol Biol 49, 37–45 (2022). https://doi.org/10.1007/s11692-021-09555-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11692-021-09555-9