Abstract
The identity of Achroomyces effusus is re-established with the use of morphological and DNA methods, and a neotype is selected. The species is conspecific with Colacogloea peniophorae, the generic type of Colacogloea, and has a priority over it. A new combination, C. effusa, is proposed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Pucciniomycotina (rust fungi and allies) are a subphylum of Basidiomycota heavily dominated by parasites, mainly inhabiting plants and other fungi. Only a few species, mostly with crust-like or pustulate basidiocarps, occur on plant remnants and are therefore considered as saprotrophs. A majority of these saprotrophic taxa belong to the Atractiellomycetes (Helicogloea Pat. and related genera) (Bauer et al. 2006; Aime et al. 2018; Spirin et al. 2018; Malysheva et al. 2020) and two species were assigned to the genus Achroomyces Bonord. (= Platygloea J. Schröt.). The generic type of the latter, A. disciformis (Fr.) Donk, produces pustulate or cushion-shaped, erumpent basidiocarps on still corticated branches of deciduous trees (almost exclusively on Tilia cordata). This species has been shown to be a member of the class Pucciniomycetes based on nrLSU sequence data (Bauer et al. 2006). Another species in the same genus, A. effusus (J. Schröt.) Mig., is a crust-like fungus appearing on wood of deciduous trees late in the season (Cohn 1889; Bourdot and Galzin 1909). Bandoni (1956) and Pilát (1957) treated it as a wood-inhabiting member of the genus, and this viewpoint has never been questioned. Only Krieglsteiner (1999) suggested A. effusus may be conspecific with Colacogloea peniophorae (Bourdot & Galzin) Oberw. & Bandoni (Microbotryomycetes, Pucciniomycotina), an intrahymenial fungus parasitizing corticioid fungi from the genus Peniophorella P. Karst. (Hymenochaetales, Agaricomycetes). However, Krieglsteiner’s idea was based on anatomical similarity of the two species and has not yet been verified with the use of DNA methods. In the present paper, we select a type for A. effusus and investigate phylogenetic relationships of this species.
Material and methods
Collections from herbaria H, C, GENT, PC were studied. Herbarium acronyms are given according to Thiers (2020). Microscopic routine and terminology follow Spirin et al. (2018). The following abbreviations are used in taxonomic section: L – mean basidiospore length, W – mean basidiospore width, Q’ – L/W ratio, Q – mean L/W ratio, n – number of measurements per specimens measured.
DNA study. We performed PCR directly from small fragments of dried basidiocarps without prior DNA extraction using the Phire Plant Direct PCR Kit (Thermo Scientific, USA) according to the manufacturer’s instructions.
The following primers were used for amplification and sequencing: ITS1-F (Gardes and Bruns 1993) and ITS4 (White et al. 1990) for the nrITS1-5.8S-ITS2 region, EF1-983F and EF1-1567R (Rehner and Buckley 2005) for a part of the TEF1 region, JS1 (Landvik 1996) and LR5 (Vilgalys and Hester 1990) for D1-D3 domains of nrLSU region. PCR products were purified applying the GeneJET Gel Extraction and DNA Cleanup Micro Kit (Thermo Scientific, USA). There were two bands for the nrITS region, which were successfully separated during electrophoresis and then sequenced as separate templates. Sequencing was performed with an ABI model 3500 Genetic Analyzer (Applied Biosystems, USA). Raw data were edited and assembled in MEGA X (Kumar et al. 2018). Molecular studies were mostly carried out at the Center for collective use of scientific equipment “Cellular and molecular technology of studying plants and fungi” (Komarov Botanical Institute, Russian Academy of Sciences, St. Petersburg, Russia).
Phylogenetic analyses. For this study, five nrITS, one TEF1 and three nrLSU sequences were generated. Additionally, 34 nrITS and 36 nrLSU sequences were retrieved from GenBank (www.ncbi.nlm.nih.gov/genbank/). Sequences were aligned with the MAFFT version 7 web tool (http://mafft.cbrc.jp/alignment/server/) with subsequent manual processing.
nrITS + nrLSU dataset comprised sequences from 15 genera accepted in Microbotryomycetes. The final alignment contained 1313 characters (including gaps). Phylogenetic reconstructions were performed with Maximum likelihood (ML) and Bayesian (BI) analyses. Before analyses, the best-fit substitution model for the alignment was estimated based on the Akaike Information Criterion (AIC) using FindModel web server (http://www.hiv.lanl.gov/content/sequence/findmodel/findmodel.html). The “GTR plus Gamma” model was chosen for the nrITS + nrLSU dataset. ML analysis was run on RAxML servers, v.0.9.0 (Kozlov et al. 2019) with one thousand rapid bootstrap replicates. BI analyses were performed with MrBayes 3.2.5 software (Ronquist et al. 2012), for two independent runs, each with 10 million generations, under described model and four chains with sampling every 100 generations. To check for convergence of MCMC analyses and to get estimates of the posterior distribution of parameter values Tracer v1.6 was used (Rambaut et al. 2014). We accepted the result where the ESS (Effective Sample Size) was above 5700 and the PSRF (Potential Scale Reduction Factor) was 1.
Newly generated sequences have been deposited in GenBank with corresponding accession numbers (see below in the text). Sequenced specimens are marked by asterisk.
Results
Platygloea effusa J. Schröt. was described from Silesia (that time Germany, now Poland) as growing on rotten stumps (Cohn 1889). No original material of this species survived up to the present moment. Later it was discovered in some other European countries although considered rare everywhere (Bourdot and Galzin 1909; Wakefield and Pearson 1917; Bandoni 1956; Pilát 1957). Donk (1966) argued that Achroomyces has a priority over Platygloea J. Schröt., and therefore the species was treated as A. effusus since then (Wojewoda 1981; Hansen and Knudsen 1997; Krieglsteiner 1999). Fortunately, the protologue as well as the subsequent reintroduction of A. effusus by Bourdot and Galzin (1909, 1927) are explicit enough to allow recognition of the species, describing a crust-like fungus with four-celled, transversally septate basidia and hyaline, ellipsoid basidiospores. Two recent specimens of A. effusus were selected for morphological and DNA-based study.
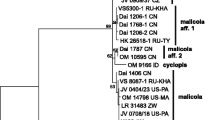
The nrITS, nrLSU and TEF1 sequences obtained from the two collections turned out identical to those ones of Colacogloea peniophorae in GenBank, thus confirming the aforementioned idea of Krieglsteiner (1999) about the identity of these two taxa. Additionally, two other ITS sequences generated from the same specimens ended up in Peniophorella praetermissa (P. Karst.) K.H. Larss. s. lato (GenBank MW293724, MW293725), the host species of C. peniophorae, although no signs of P. praetermissa basidiocarps (except a few hyphae and remains of cystidia) were detected in them. To verify these results, we studied and sequenced a typical specimen of C. peniophorae, which formed pustular, slimy fructifications (Bruggeman 6343) appeared directly on basidiocarps of P. praetermissa. ITS and LSU sequences of this specimen were found to be identical with those of A. effusus (Fig. 1). Since A. effusus has a priority over C. peniophorae, we combine it in the genus Colacogloea and treat the latter species as its synonym.
Combined phylogenetic midpoint-rooted nrITS + nrLSU topology from Bayesian analysis showing main lineages within the Microbotryomycetes. All sequences generated for this study are indicated in bold and provided with collection numbers. GenBank accession numbers are given for all other sequences. Support values (BS/PP) are given above the branches. Scale bar shows expected changes per site
Colacogloea effusa (J. Schröt.) V. Malysheva, Schoutteten & Spirin, comb. nov.
MB838144
≡ Platygloea effusa J. Schröt. in Cohn, Kryptogamen-Flora von Schlesien 3 (1): 384, 1889. Neotype (here designated, MBT395027). Denmark. Midtjylland: Norddjurs, Løvenholm Skov, rotten wood, 26.VIII.2009 Heilmann-Clausen 09–304* (C, isoneotype H) (GenBank ITS MW293723, LSU MW293727).
= Platygloea peniophorae Bourdot & Galzin, Bull. Soc. Mycol. France 25: 17, 1909.
Basidiocarps first hyaline, gelatinous, appearing as hardly visible patches or pustules on hymenial surface of Peniophorella praetermissa, then fusing together and finally forming perceptible (up to 5 cm wide and 1–2 mm thick), opalescent or yellowish, crustaceous fructifications with tuberculate hymenial surface, turning reddish or brownish in herbarium specimens. Monomitic; hyphae hyaline, clamped, thin-walled, mostly ascending, 1.5–2.5 µm in diam. Cystidia absent; hyphal endings abundant in hymenium, simple or sparsely branched, 1–2 µm in diam. Basidia four-celled, transversally septate, narrowly clavate, usually somewhat winding or moderately curved at the apex, (44)48–80(83) × (4.0)4.1–5.3(5.8) µm (n = 30/2), probasidia absent; sterigmata up to 40 × 1.5–2 µm. Basidiospores hyaline, thin-walled, ellipsoid to bean-shaped, (6.2)6.8–9.2(–9.3) × (4.1)4.3–5.8(5.9) µm (n = 60/2), L = 7.83–8.03, W = 5.13–5.18, Q’ = (1.3)1.4–1.8(1.9), Q = 1.53–1.56, with guttulate contents. Conidia present in some collections, hyaline, thick-walled, narrowly ovoid to subfusiform, often asymmetric or somewhat angular, 6–8 × 3–4(4.5) µm, distinctly cyanophilous.
Specimens examined. Finland. Varsinaissuomi: Raasepori, Framnäs, unidentified deciduous tree, 21.XI.2019 Pennanen 4226* (H, as A. effusus) (GenBank ITS MW293722, LSU MW293726, TEF1 MW298152). Netherlands. Utrecht: Zeist, Beerschoten, living Peniophorella praetermissa on decayed wood, 10.X.2019 Bruggeman 6343* (GENT) (GenBank ITS MW303962, LSU MW310243).
Discussion
While introducing P. peniophorae, Bourdot and Galzin (1909) explicitly compared it with Platygloea effusa J. Schröt. (= A. effusus). They differentiated P. peniophorae from the latter species based on thinner and smaller basidiocarps, shorter basidia, presence of conidia, and its growth on corticioid fungi (Peniophorella pubera (Fr.) P. Karst. and P. praetermissa were listed as host species). According to our study, none of these features is reliable for separating these two taxa. In particular, the length of basidia is highly variable even in one specimen and seemingly correlates to the basidiocarp thickness. The presence of conidia is also unstable: of two collections of A. effusus mentioned above, one possesses abundant conidia (Pennanen 4226) while the neotype (Heilmann-Clausen 09–304) is completely devoid of them.
Of 23 authentic specimens of P. peniophorae in Bourdot’s herbarium (PC), sixteen were collected from P. praetermissa, one from P. pubera and six from unidentified corticioid fungi. Specimens from the first host species were evidently the main source for the Bourdot’s original idea of P. peniophorae, and therefore this species should be treated as a synonym of C. effusa. Microscopic structures of C. effusa were illustrated by Bourdot and Galzin (1927), Wojewoda (1981) (as A. effusus) and Oberwinkler et al. (1990, as C. peniophorae). Macroscopic photographs of C. effusa can be found on Danmarks Svampeatlas web page (https://svampe.databasen.org, as A. effusus). Seemingly, C. effusa is able to consume its host species so aggressively that no visible signs of the latter are finally detectable. Certainly, this was a reason why A. effusus was believed to be a wood-inhabiting fungus. Nevertheless, a careful microscopic investigation and the use of molecular methods can re-confirm the presence of P. praetermissa.
Bandoni (1973) pointed out morphological differences of C. effusa specimens from different geographic areas. It may well be it represents a species complex. However, proving this hypothesis requires much more extensive sampling and incorporation of additional genetic markers. Colacogloea effusa is a generic type of Colacogloea Oberw. & Bandoni, a genus of which the majority of species is known only from asexual yeast stages (Wang et al. 2015; Li et al. 2020). Re-establishing its identity is important for future studies of species diversity and relations between sexual and asexual morphs in the Microbotryomycetes.
Data Availability
DNA sequences used in the present study are available in GenBank. Alignments were deposited in TreeBase. Fungal specimens are stored in public herbaria (as indicated under Specimens examined).
References
Aime MC, Urbina H, Liber JA, Bonito G, Oono R (2018) Two new endophytic Atractiellomycetes, Atractidochium hillariae and Proceropycnis hameedii. Mycologia 110:136–146
Bandoni RJ (1956) A preliminary survey of the genus Platygloea. Mycologia 48:821–840
Bandoni RJ (1973) Epistolae mycologicae II. Species of Platygloea from British Columbia. Syesis 6:229–232
Bauer R, Begerow D, Sampaio JP, Weiss M, Oberwinkler F (2006) The simple-septate basidiomycetes: a synopsis. Mycol Prog 5:41–66
Bourdot H, Galzin A (1909) Hyménomycètes de France. (I Hetérobasidiés). Bull Soc Mycol France 25:14–36
Bourdot H, Galzin A (1927) Hyménomycètes de France. Hetérobasidiés – Homobasidiés gymnocarpes. Sceaux
Cohn F (1889) Kryptogamen-Flora von Schlesien. Breslau
Donk MA (1966) Check list of European hymenomycetous heterobasidiae. Persoonia 4:145–335
Hansen L, Knudsen H (1997) Nordic macromycetes, vol 3. Nordsvamp, Copenhagen
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes – applications to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: A fast, scalable, and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics, btz305 https://doi.org/10.1093/bioinformatics/btz305
Krieglsteiner GJ (1999) Kritische Basidiomycetes-Arten aus Baden-Württemberg (Deutschland, Mitteleuropa) und taxonomische Konsequenzen. Beiträge Kenntnis Pilze Mitteleuropas 12:33–56
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Landvik S (1996) Neolecta, a fruit-body producing genus of the basal ascomycetes, as shown by SSU and LSU rDNA sequences. Mycol Res 100:199–202
Li AH, Yuan FX, Groenewald M, Bensch K, Yurkov A, Li K, Han PJ, Guo LD, Aime MC, Sampaio JP, Jindamorakot S, Turchetti B, Inacio J, Fungsin B, Wang QM, Bai FY (2020) Diversity and phylogeny of basidiomycetous yeasts from plant leaves and soil: Proposal of two new orders, three new families, eight new genera and one hundred and seven new species. Stud Mycol 96:17–140. https://doi.org/10.1016/j.simyco.2020.01.002
Malysheva V, Spirin V, Schoutteten N, De Lange R, Pennanen J, Larsson KH (2020) New and noteworthy species of Helicogloea (Atractiellomycetes, Basidiomycota) from Europe. Ann Bot Fenn 57:1–7
Oberwinkler F, Bauer R, Bandoni RJ (1990) Colacogloea: a new genus in the auricularioid Heterobasidiomycetes. Can J Bot 69:2531–2536
Pilát A (1957) Übersicht der europäischen Auriculariales und Tremellales unter besonderer Berücksichtigung der tschechoslowakishen Arten. Acta Mus Nat Prag 13B:115–210
Rambaut A, Suchard MA, Xie D, Drummond AJ (2014) Tracer v1.6. http://tree.bio.ed.ac.uk/software/tracer/ (Accessed on 20 Nov 2020)
Rehner SA, Buckley E (2005) A Beauveria phylogeny inferred from nuclear ITS and EF1-alpha sequences: evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 97:84–98
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Spirin V, Malysheva V, Trichies G, Savchenko A, Põldmaa K, Nordén J, Miettinen O, Larsson KH (2018) A preliminary overview of the corticioid Atractiellomycetes (Pucciniomycotina, Basidiomycota). Fung Syst Evol 2:311–340
Thiers B (2020) Index Herbariorum: a global directory of public herbaria and associated stuff [continuosly updated]. New York Botanical Garden’s Virtual Herbarium. http://sweetgum.nybg.org/ih. Accessed 20 Nov 2020
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriology 172:4238–4246
Wakefield EM, Pearson AA (1917) Additional resupinate basidiomycetes from the Weybridge District. Trans British Mycol Soc 6:136–143
Wang QM, Yurkov AM, Göker M, Lumbsch HT, Leavitt SD, Groenewald M, Theelen B, Liu XZ, Boekhout T, Bai FY (2015) Phylogenetic classification of yeasts and related taxa within Pucciniomycotina. Stud Mycol 81:149–189. https://doi.org/10.1016/j.simyco.2015.12.002
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA et al (eds) PCR protocols. A guide to methods and applications. Academic Press, San Diego, pp 315–322
Wojewoda W (1981) Mała flora grzybów. Tom 2. Krakow, Państwowe Wydawnictwo Naukowe Warsaw
Acknowledgements
Jacob Heilmann-Clausen (Denmark) and Jorma Pennanen (Finland) kindly provided us with their specimens of A. effusus.
Funding
Open Access funding provided by University of Helsinki including Helsinki University Central Hospital The Russian Foundation for Basic Researches, project No. 19–04-00024 and partly by the project of the Komarov Botanical Institute of the Russian Academy of Sciences (the author VM), as well as by the Research Foundation Flanders (FWO) under grant No. 11E0420N (Fellowship fundamental research of the author NS).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and data collection were performed by Vera Malysheva, Nathan Schoutteten and Viacheslav Spirin. Lab work and phylogenetic analyses were carried out by Vera Malysheva, Nathan Schoutteten and Annemieke Verbeken. The first draft of the manuscript was written by Viacheslav Spirin and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Conflict of interests
None.
Additional information
Section Editor: Meike Piepenbring.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Malysheva, V., Schoutteten, N., Verbeken, A. et al. Identity and typification of Achroomyces effusus (Pucciniomycotina, Basidiomycota). Mycol Progress 20, 413–417 (2021). https://doi.org/10.1007/s11557-021-01671-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-021-01671-2