Abstract
Eight diketopiperazines including five previously unreported derivatives were isolated from an endophytic fungus cultured from the medicinal plant Globularia alypum collected in Algeria. The strain was characterised by means of morphological studies and molecular phylogenetic methods and was found to represent a species of a new genus in the Chaetomiaceae, for which we propose the name Batnamyces globulariicola. The taxonomic position of the new genus, which appears phylogenetically related to Stolonocarpus and Madurella, was evaluated by a multi-locus genealogy and by morphological studies in comparison to DNA sequence data reported in the recent monographs of the family. The culture remained sterile on several culture media despite repeated attempts to induce sporulation, and only some chlamydospores were formed. After fermentation in submerged culture and extraction of the cultures with organic solvents, the major secondary metabolites of B. globulariicola were isolated and their chemical structures were elucidated by extensive spectral analysis including nuclear magnetic resonance (NMR) spectroscopy, high-resolution electrospray ionisation mass spectrometry (HRESIMS), and electronic circular dichroism (ECD) measurements. The isolated compounds were tested for their biological activities against various bacteria, fungi, and two mammalian cell lines, but only three of them exhibited weak cytotoxicity against KB3.1 cells, but no antimicrobial effects were observed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, it has been demonstrated that fungal endophytes are playing an important role in most ecosystems of the world. These fungi, which colonise their host plants without causing any disease symptoms, have been shown to represent various known and new phylogenetic lineages (Blackwell and Vega 2018).
Moreover, several studies have led to the isolation and structure elucidation of important secondary metabolites from endophytic fungi, thus raising the prospect of using such organisms as alternative sources of lead compounds for development of new drugs or agrochemical pesticides (Suryanarayanan et al. 2009; Bills and Gloer 2016). Moreover, it is now also being attempted to use these organisms as biocontrol agents and biofertilisers (Hyde et al. 2019; White et al. 2019).
In our ongoing search for bioactive fungal metabolites from fungi associated with Algerian medicinal plants (Noumeur et al. 2017; Teponno et al. 2017), 12 secondary metabolites including five previously unreported diketopiperazines (1–5) were isolated from the culture of a fungus that was obtained from Globularia alypum (Plantaginaceae) using a well-established protocol that has been used for the isolation of endophytic fungi for several decades. The producer organism turned out to be new to Science and is formally described in the present paper. Moreover, the isolation of its secondary metabolites, and their structure elucidation and their preliminary biological characterisation are reported.
Material and methods
Origin and isolation of the strain
Fresh healthy roots of the medicinal plant Globularia alypum (Plantaginaceae) were collected in June 2015 from Ain Touta (Batna, Algeria). Along with other cultures, strain CBS 144474 was isolated according to established protocols involving surface disinfection (Noumeur et al. 2017; Teponno et al. 2017) and was maintained in liquid nitrogen at the HZI culture collection, Braunschweig, Germany. Morphology and macroscopic features of the culture were determined on several different culture media including yeast-malt-glucose (YMG; Fig. 1), see Richter et al. (2016), and all other the standard media that are currently in use for induction of sporulation of Sordariomycetes, i.e. potato dextrose agar (PDA), oatmeal agar (OA), synthetic nutrient-poor agar (SNA), and malt extract agar (MEA) (cf. Crous et al. 2009; Wang et al. 2019b). Cardinal temperature requirements for growth were only checked on YMG at temperatures ranging from 24 to 43 °C.
Molecular analysis and sequencing
Genomic DNA extraction from the fungal colonies growing on YMG was performed using an EZ-10 Spin Column Genomic DNA Miniprep kit (Bio Basic Canada Inc., Markham, Ontario, Canada) as specified by the manufacturer. The internal transcribed spacer 1 and 2 including the intervening 5.8S nrDNA (ITS), the 28S nrDNA (LSU) including the D1/D2 domains, a part of the DNA-directed RNA polymerase II second largest subunit gene (RPB2), and the β-tubulin gene (TUB2) were selected for phylogenetic inference. PCR and generation of DNA sequences followed the procedure outlined by Wendt et al. (2018).
After a BLAST result based on GenBank data, which allocated the new fungus to the Chaetomiaceae, the phylogenetic affinities were studied based on an analysis of a combined ITS, LSU, RPB2, and TUB2 dataset in comparison with DNA sequence data reported in the recent monographs of the family (Wang et al. 2016a, b, 2019a, b). Alignments were made using the web interface MAFFT v.7 (Katoh and Standley 2013), followed by manual adjustments with MEGA v. 6 (Tamura et al. 2013). Phylogenetic analysis was performed using maximum likelihood (ML) and Bayesian inference (BI) approaches under RAxML-HPC2 on XSEDE 8.2.10 (Stamatakis 2014) using the Cipres Science gateway portal (Miller et al. 2010) and MrBayes v. 3.2.6 (Ronquist et al. 2012), respectively. For BI, the best evolutionary model for each locus was determined using MrModeltest v. 2.0 (Nylander 2004). The maximum likelihood analysis used the GTRGAMMA model. Obtained trees were viewed in FigTree v. 1.1.2 (Rambaut 2009) and subsequently visually prepared and edited in Adobe® Illustrator® CS6. Confident branch support is defined as Bayesian posterior probabilities (PP) ≥ 0.95 and maximum likelihood bootstrap values (ML-BS) ≥ 70%.
Instrumentation
Optical rotations were determined with a Perkin Elmer (Überlingen, Germany) 241 MC polarimeter (using the sodium D line and a quartz cuvette with a 10-cm path length and 0.5-mL volume). Circular dichroism (CD) spectra were recorded on a JASCO spectropolarimeter, model J-815 (JASCO, Pfungstadt, Germany). Nuclear magnetic resonance (NMR) spectra were recorded on a Bruker (Bremen, Germany) 500 MHz Avance III spectrometer with a BBFO (plus) SmartProbe (1H 500 MHz, 13C 125 MHz) and a Bruker 700 MHz Avance III spectrometer with a 5-mm TCI cryoprobe (1H 700 MHz, 13C 175 MHz), locked to the deuterium signal of the solvent. Chemical shifts are given in parts per million (ppm) and coupling constants in Hertz (Hz). Spectra were measured at 24.8 °C in CD3OD, acetone-d6, and deuterated chloroform; chemical shifts were referenced to residual solvent signals with resonances at δH/C 3.31/49.15 for CD3OD, δH/C 2.05/29.92 for acetone-d6, and δH/C 7.24/77.23 for deuterated chloroform. High-performance liquid chromatography coupled with diode array detector and mass spectrometric detection (HPLC-DAD/MS) analysis was performed using an amaZon speed ETD ion trap mass spectrometer (Bruker Daltonics) in positive and negative ionisation modes. The mass spectrometer was coupled to an Agilent 1260 series HPLC-UV system (Agilent Technologies) (Santa Clara, CA, USA) (column 2.1 × 50 mm, 1.7 μm, C18 Acquity uPLC BEH (Waters); solvent A: H2O + 0.1% formic acid; solvent B: acetonitrile (ACN) + 0.1% formic acid; gradient: 5% B for 0.5 min, increasing to 100% B in 20 min, maintaining isocratic conditions at 100% B for 10 min, flow = 0.6 mL/min, UV−vis detection 200–600 nm). High-resolution electrospray (HR-ESI) MS spectra were recorded on a maXis ESI TOF mass spectrometer (Bruker Daltonics) (scan range m/z 100–2500, rate 2 Hz, capillary voltage 4500 V, dry temperature 200 °C), coupled to an Agilent 1200 series HPLC-UV system (column 2.1 × 50 mm, 1.7 μm, C18 Acquity uPLC BEH (Waters); solvent A: H2O + 0.1% formic acid; solvent B: ACN + 0.1% formic acid; gradient: 5% B for 0.5 min, increasing to 100% B in 19.5 min, maintaining 100% B for 5 min, FR= 0.6 mL/min, UV−vis detection 200–600 nm). The molecular formulas were calculated including the isotopic pattern (Smart Formula algorithm). Preparative HPLC purification was performed at room temperature on an Agilent 1100 series preparative HPLC system (ChemStation software (Rev. B.04.03 SP1); binary pump system; column: Kinetex 5u RP C18 100 Å, dimensions 250 × 21.20 mm; mobile phase: ACN + 0.05% trifluoroacetic acid (TFA) (solvent B) and water + 0.05% TFA (solvent A); flow rate 20 mL/min; diode-array UV detector; 226 fraction collector).
Fermentation, extraction, and isolation
Pieces of well-colonised agar of strain CBS 144474 from YMG plates were inoculated in Q61/2 medium (Chepkirui et al. 2019) in a 500-mL Erlenmeyer flask containing 200 mL of media and incubated at 23 °C for 7 days. This seed culture was used to inoculate 25 other flasks (5 L) of the same medium composition after homogenisation with a Heidolph Silent Crusher. The flasks were incubated at 23 °C under constant shaking at 140 rpm on a rotary shaker for 12 days. After separation of the mycelia by vacuum filtration, the supernatant was extracted with ethyl acetate (EtOAc). The EtOAc fraction was dried over anhydrous Na2SO4, filtered, and concentrated under vacuum to yield 590 mg of extract. The wet mycelia were extracted twice with acetone, then with methanol in an ultrasonic bath at 40 °C for 30 min. The resulting solution was evaporated to yield an aqueous phase, which was further extracted with EtOAc (3 × 500 mL). After drying over anhydrous Na2SO4, the EtOAc fraction was concentrated under vacuum to yield 416.7 mg of crude extract. In the meantime, pieces of a well-grown YMG agar plate of strain CBS 144474 were inoculated in a YMG medium (Richter et al. 2016), in a 500-mL Erlenmeyer flask containing 200 mL of media, and incubated at 23 °C for 8 days. The homogenised seed culture was used to inoculate 25 other flasks (5 L) of the same medium composition. The flasks were incubated at 23 °C under constant shaking at 140 rpm on a rotary shaker for 11 days. After separation, the supernatant and the mycelia were extracted as described above to give 408.5 and 512.8 mg of extracts, respectively.
The crude supernatant extract from the fermentation in Q61/2 medium (ca. 400 mg) was purified by preparative HPLC using a gradient of 25–50% solvent B for 40 min, 50–100% B for 10 min, and 100% B for 10 min. The fractions were combined according to UV absorption at 220, 280, and 325 nm and concurrent HPLC-MS analyses. Compounds 5 (0.9 mg; Rt = 11.12 min), 1 (1 mg; Rt = 13.80 min), 2 (0.8 mg; Rt = 15.40 min), 8 (0.8 mg; Rt = 21.34 min), 3 (1.2 mg; Rt = 25.42 min), 6 (1.4 mg; Rt = 29.84 min), 7 (0.7 mg; Rt = 32.16 min), 4 (0.7 mg; Rt = 36.12 min), and 9 (0.5 mg; Rt = 38.69 min) were eluted.
The supernatant extract from the fermentation in YMG medium (ca. 400 mg) was also purified by preparative HPLC. The gradient used was 5–35% solvent B for 40 min, 35–100% solvent B for 20 min, and 100% B for 10 min. The fractions were combined according to UV absorption at 220, 280, and 325 nm to yield compounds 10 (8.7 mg; Rt = 18.17 min), 11 (9.3 mg; Rt = 19.69 min), and 12 (3.7 mg; Rt = 23.11 min).
Summary of spectral data for the new compounds (1–5)
(3R,6Z)-3-Thiomethyl-6-[4-O-[(2E)-4-hydroxy-3-methylbut-2-enyl]benzylidene]piperazine-2,5-dione (1): yellowish gum; \( {\left[\alpha \right]}_D^{25} \)+ 22.0 (c 0.05, MeOH); UV (MeOH) λmax (log ε): 204 (4.37), 229 (4.26), 323 nm (4.34); CD (c 0.5 mg/mL, EtOH) λmax 309 (+), 235 nm (+); 1H NMR (CD3OD, 700 MHz) and 13C NMR (CD3OD, 175 MHz) data (see Table 1); HRESIMS: m/z 349.1217 [M + H]+ (calcd for C17H21N2O4S+, 349.1217).
(3R,6Z)-3-Thiomethyl-6-[4-O-[(2Z)-4-hydroxy-3-methylbut-2-enyl]benzylidene]piperazine-2,5-dione (2): yellowish gum; \( {\left[\alpha \right]}_D^{25} \) + 37.5 (c 0.04, MeOH); UV (MeOH) λmax (log ε): 203 (4.13), 229 (3.97), 323 nm (3.96); CD (c 0.5 mg/mL, EtOH) λmax 316 (+), 237 nm (+); 1H NMR (CD3OD, 700 MHz) and 13C NMR (CD3OD, 175 MHz) data (see Table 1); HRESIMS: m/z 349.1217 [M + H]+ (calcd for C17H21N2O4S+, 349.1217).
(3R,6Z)-3-Hydroxy-6-[4-O-(3-methylbut-2-enyl)benzylidene]piperazine-2,5-dione (3): yellowish gum; \( {\left[\alpha \right]}_D^{25} \)+ 26.7 (c 0.06, MeOH); UV (MeOH) λmax (log ε): 206 (4.22), 224 (4.26), 321 nm (4.15); CD (c 0.5 mg/mL, EtOH) λmax 309 (+), 228 nm (+); 1H NMR (CD3OD, 700 MHz) and 13C NMR (CD3OD, 175 MHz) data (see Table 1); HRESIMS: m/z 303.1334 [M + H]+ (calcd for C16H19N2O4+, 303.1339).
(3R,6Z)-3-Thiomethyl-6-[4-O-(3-methylbut-2-enyl)benzylidene]piperazine-2,5-dione (4): yellowish gum; \( {\left[\alpha \right]}_D^{25} \) + 48.6 (c 0.035, MeOH); UV (MeOH) λmax (log ε): 202 (4.19), 229 (4.02), 324 nm (4.00); CD (c 0.5 mg/mL, EtOH) λmax 310 (+), 230 nm (+); 1H NMR (CD3OD, 700 MHz) and 13C NMR (CD3OD, 175 MHz) data (see Table 2); HRESIMS: m/z 355.1088 [M + Na]+ (calcd for C17H20N2NaO3S+, 355.1092), 687.2282 [2 M + Na]+.
(3S,6R)-3,6-Bisthiomethyl-6-[4-O-[(2Z)-4-hydroxy-3-methylbut-2-enyl]phenylmethyl]piperazine-2,5-dione (5): yellowish gum; \( {\left[\alpha \right]}_D^{25} \)+ 33.3 (c 0.045, MeOH); UV (MeOH) λmax (log ε): 208 (4.16), 226 (4.06), 275 nm (3.38); CD (c 0.5 mg/mL, EtOH) λmax 315 (+), 248 (−), 220 nm (−); 1H NMR (CD3OD, 700 MHz) and 13C NMR (CD3OD, 175 MHz) data (see Table 2); HRESIMS: m/z 397.1249 [M + H]+ (calcd for C18H25N2O4S2+, 397.1250), 419.1068 [M + Na]+ (calcd for C18H24N2O4S2Na+, 419.1075).
Screening for biological activities
The antimicrobial activity and the in vitro cytotoxicity (IC50) were evaluated according to our previously reported procedures (Sandargo et al. 2018; Teponno et al. 2017). Briefly, minimum inhibitory concentrations (MICs) in μg/mL of the isolated compounds were determined by serial dilution assays against Schizosaccharomyces pombe DSM 70572, Pichia anomala DSM 6766, Mucor hiemalis DSM 2656, Candida albicans DSM 1665, Rhodotorula glutinis DSM 10134, Micrococcus luteus DSM 1790, Bacillus subtilis DSM 10, Escherichia coli DSM 1116, Staphylococcus aureus DSM 346, Mycobacterium smegmatis ATCC 700084, Chromobacterium violaceum DSM 30191, and Pseudomonas aeruginosa DSM PA14. The assays were carried out in 96-well microtiter plates in YMG media for filamentous fungi and yeast and EBS for bacteria. Gentamycin, kanamycin, nystatin, and oxytetracycline were used as positive control, and the negative control was methanol. The cytotoxicity against HeLa cells KB3.1 and mouse fibroblasts L929 cells was determined by using the MTT (2-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) method in 96-well microplates. The cell lines were cultured in DMEM (Gibco). Briefly, 60-μL aliquots of serial dilutions from an initial stock of 1 mg/mL in MeOH of the test compounds were added to 120-μL aliquots of a cell suspension (5 × 104 cells/mL) in 96-well microplates. After 5 days of incubation, an MTT assay was performed, and the absorbance was measured at 590 nm using an ELISA plate reader (Victor). The concentration at which the growth of cells was inhibited to 50% of the control (IC50) was obtained from the dose-response curves. Epothilone B was used as a positive control, while methanol was used as a negative control.
Results and discussion
Taxonomy
The fungus was isolated as sterile mycelia without any reproductive structures, and its identification by phenotypic characters was impossible by conventional methods. The morphology and macroscopic features of the culture on the plate were determined on YMG, PDA, and OA.
Molecular phylogeny
The concatenated alignment consisted of 65 taxa including representatives of 35 genera in the Chaetomiaceae (cf. Table 3). Five isolates representing four species of the family Podosporaceae were selected as outgroups. The alignment contained 3055 characters (including gaps) and is composed of four partitions: 858 characters for RPB2, 961 characters for TUB2, 664 characters for ITS, and 572 characters for LSU. Of the total characters, 1573 were constant, 1220 were parsimony-informative, and 262 were parsimony-uninformative. For the Bayesian inference, the GTR+I+G model was selected as optimal for RPB2, TUB2, ITS, and LSU based on the result of the MrModeltest. Isolate CBS 144474 was located in a separate clade along with representatives of the genera Stolonocarpus, Madurella, and Canariomyces (ML-BS = 100%, PP = 0.99), but could not be accommodated in either of these genera (Fig. 2). Therefore, a novel genus Batnamyces is proposed.
Batnamyces Noumeur, gen. nov. MB 832844
Etymology—in reference to the town in Algeria where the type was collected.
Diagnosis—Differs from the genera Canariomyces, Stolonocarpus, and Madurella, to which it appears phylogenetically most closely related, in the absence of sexual features and conidiogenous structures, except for producing terminal chains of hyphal chlamydospores.
Type species: Batnamyces globulariicolaNoumeur, sp. nov. MB 832845 (Fig. 1)
Morphological characteristics of the new genus Batnamyces. a Colonies obverse and reverse on PDA at 23 °C after 21 days. b Colonies obverse and reverse on OM at 23 °C after 21 days. c Colonies obverse and reverse on YMG at 23 °C after 21 days. d and e chlamydospore-like structures formed at hyphal tips. Scale bars d, e = 10 μm
Typus: Algeria, Batna, Ain Touta from roots of Globularia alypum Plantaginaceae, June 2015, S. R. Noumeur (holotype CBS H-23624), ex-type culture in CBS 144474; GenBank Acc nos of DNA sequences: MT075917 (ITS/LSU), MT075918 (RPB2), MT075919 (TUB2).
Colonies on YMG and OMA at 23 °C spread over the whole 9-cm Petri dish after 14 days while they attain a diameter of 53 mm on PDA, initially appearing dark brown, then becoming covered with white patches with age (Fig. 1b). Mycelium on SNA, with brown, thick-walled, smooth, branched, septate, 5–6 μm diam hyphae, giving rise to hyaline, thin-walled, smooth, branched, septate, 1.5–2 μm diam hyphae. Colonies remained sterile on SNA, PDA, OA, and MEA (see images in Supplementary information). After several transfers onto new OA agar plates, oval, chlamydospores (5–12 × 8 μm) were formed in short chains, arising from the hyphal tips (Fig. 1e). Even after several months of subcultivation, no other conidiogenous structures or sexual morph were observed either in the old or the newly inoculated plates. The growth maximum was determined to be 28–30 °C, and no growth was observed at 37 °C.
Notes—The new genus Batnamyces is primarily defined based on its molecular phylogeny, since we neither observed the characteristic structures of the asexual nor sexual morph of the species in this genus. Its classification in the family was inferred from the molecular phylogeny that was established on the basis of a multi-locus genealogy comprising representatives of all important genera of Chaetomiaceae. The genera Batnamyces, Canariomyces, Madurella, and Stolonocarpus formed a single lineage (Fig. 2). Canariomyces species typically produce non-ostiolate ascomata together with single-celled conidia arising from reduced conidiophores that are reduced to a hyphal cell (cf. figs. 19–22 in Wang et al. 2019b). Madurella species usually produce only sterile (non-sporulating) hyphae and sparse aerial mycelium, growing restrictedly in culture and often producing buff, cinnamon, sienna, or orange exudates diffusing into the agar (cf. fig. 18 in Wang et al. 2019b). On the other hand, Stolonocarpus is characterised by non-ostiolate ascomata arising from a stolon-like mycelium and covered by pigmented hypha-like hairs (fig. 41 in Wang et al. 2019b). The genus Batnamyces is more similar to Canariomyces and Stolonocarpus than to Madurella with respect to the morphology of the colonies and mycelia but can be easily distinguished from them by the lack of reproductive structures. Since the ex-type strain of Batnamyces was obtained by using an isolation procedure, which is well established for endophytes, from an endemic plant in an area that has never been studied intensively for the biodiversity of its mycobiota, it did not come as a surprise that no reproductive structures are produced. After all, it is pretty well known that endophytic fungi often do not produce any propagules. However, we isolated the fungus only one time and can therefore not be sure about its actual lifestyle. Poor statistic support (PP < 0.95; ML-BS = 85%) also implied that B. globulariicola was not a member of either Madurella or Stolonocarpus.
Phylogenetic tree resulting from a maximum likelihood analysis of the concatenated RPB2, TUB2, ITS, and LSU sequence data sets. The confidence values are indicated at the notes: bootstrap proportions from the ML analysis above branches, and the posterior probabilities from the Bayesian analysis below branches. “-” means lacking statistical support (< 70% for bootstrap proportions from ML or MP analyses; < 0.95 for posterior probabilities from Bayesian analyses). The branches with full statistical support (MP-BS = 100%; ML-BS = 100%; PP = 1.0) are indicated by thickened branches. Genus clades are discriminated with boxes of different colours. The scale bar shows the expected number of changes per site. The tree is rooted with members of the Podosporaceae. “T” after the respective DNA sequences are derived from ex-type (including ex-epitype and ex-neotype) specimens
Isolation and structure elucidation of compounds 1–5
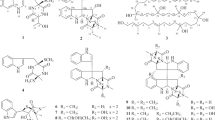
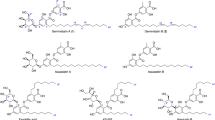
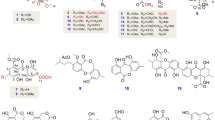
Fractionation of crude ethyl acetate extracts from the culture of Batnamyces globulariicola in Q61/2 and YMG media led to the isolation and structure elucidation of 5 new 2,5-diketopiperazines (1–5) together with seven known metabolites identified by spectroscopic analysis and comparison with literature data, such as Sch 54796 6 (Chu et al. 1993; Usami et al. 2002), Sch 54794 7 (Chu et al. 1993; Usami et al. 2002), cyclo-(glycyl-L-tyrosyl)-3,3-dimethylallyl ether 8 (Koolen et al. 2012), 4-O-(3-methylbut-2-enyl)benzoic acid 9 (Nozawa et al. 1989), L-Pro-L-Ile 10 (Ren et al. 2010), L-Pro-L-Leu 11 (Ren et al. 2010; Sansinenea et al. 2016), and L-Pro-L-Phe) 12 (Sansinenea et al. 2016) (Fig. 3).
Compound 1 was isolated as a yellowish gum. Its molecular formula C17H20N2O4S was deduced from the HRESIMS which exhibited the pseudomolecular ion peak at m/z 349.1217 [M + H]+ (calcd for C17H21N2O4S+, 349.1217). This was confirmed by the ESIMS ion cluster at m/z 371.11 [M + Na]+ and a prominent ion fragment and 301 [M + H–48]+ revealing the loss of a methanethiol (CH3SH) unit (Chu et al. 1993). Its 1H NMR spectrum displayed resonances for an AA′BB′ spin system at δH 7.45 (d, J = 8.6 Hz, H-9, and H-9′) and 7.00 (d, J = 8.6 Hz, H-10, and H-10′) suggesting the presence of a 1,4-disubstituted benzene ring in the molecule (Table 1). It also showed a signal assigned to a vinyl proton δH 6.87 (H-7) and a set of resonances depicted at δH 4.68 (brd, J = 6.4 Hz, H-12), 5.73 (m, H-12), 3.99 (s), 3.99 (s, H-15), and 1.76 (s, H-16) attributed to an O-isoprenol group. Other signals were those of a thiomethyl singlet at δH 2.26 (s, H-17) and a methine singlet at δH 4.97 (s, H-3). The downfield shift of the latter revealed its bis-heteroatom connectivity (Chu et al. 1997) (Table 1). The 13C NMR spectrum showed two amidocarbonyl carbon signals characteristic of a diketopiperazine core at δC 166.0 (C-2) and 163.5 (C-5) (Chu et al. 1993; Guimarães et al. 2010; Fu et al. 2011; Fan et al. 2017). It also displayed resonances at δC 125.3 (C-6), 119.3 (C-7), 126.8 (C-8), 132.1 (C-9, C-9′), 116.4 (C-10, C-10′), and 160.9 (C-11) evidencing the presence of an oxybenzylidene moiety. In addition, the signals of two methylenes at δC 66.8 (C-12) and 67.9 (C-15), a methine at δC 120.8 (C-13), a methyl at δC 14.2 (C-13), and a quaternary carbon at δC 141.5 (C-14) were assigned to the O-isoprenol group. The remaining signals were those of a methine at δC 59.6 (C-3) and the thiomethyl group at δC 13.1 (C-17) (Table 1). The location of the thiomethyl group at C-3 was further evidenced by the HMBC correlation observed between thiomethyl protons signal at δH 2.26 (s, H-17) and the carbon at δC 59.6 (C-3). Furthermore, the HMBC correlation from H-12 (δH 4.68) to carbon C-11 (δC 160.9) confirmed that the O-isoprenol group was linked at C-11 (Fig. 4). Careful examination of the 1H-1H COSY, HSQC, and HMBC spectra proved that 1 was related to Sch 56396, a metabolite produced by the fungus Tolypocladium sp. (Chu et al. 1997), the main difference was the hydroxylation of one methyl of the isopentenyl group to form 1. The E geometry for the Δ13,14 double bond was determined from the NOESY correlation depicted between the olefinic proton H-13 (δH 5.73) and the methylene protons H-15 (δH 3.99). To solve the stereochemistry of Δ6,7 double bond, a NOESY spectrum was measured in DMSO-d6. The lack of NOESY correlation between NH-1 depicted at δH 10.17 (s) and the vinyl proton H-7 (δH 6.87) was in favour of the Z configuration. This was further confirmed by the NOESY correlation depicted between NH-1 (δH 10.17 s) and H-9/H-9′ (δH 7.45, d, 8.6). The absolute configuration at C-3 was determined to be R by comparison of the experimental ECD of 1 (Fig. S9, Supporting information) with the calculated ECD spectra for the four stereoisomers (3R,6E; 3S,6E; 3R,6Z; 3S,6Z) of a related compound (Guimarães et al. 2010). Although the absorption on the experimental spectrum was not intense, the Cotton effects of 1 were in accordance with the experimental Cotton effects of 3R,6Z stereoisomer especially with positive absorption in the regions of 200–240 nm and 275–350 nm, respectively. The structure of compound 1 was unambiguously elucidated as (3R,6Z)-3-thiomethyl-6-[4-O-[(2E)-4-hydroxy-3-methylbut-2-enyl]benzylidene]piperazine-2,5-dione.
Metabolite 2 was obtained as a yellowish gum. It possessed the same molecular formula as 1 as evidenced by the HRESIMS which showed a protonated molecular ion peak at m/z 349.1217 [M + H]+ (calcd for C17H21N2O4S+, 349.1217) despite the fact that both compounds had different retention times. The 1H and 13C NMR spectra of 2 (Table 1) were closely related to those of 1, especially for signals of the diketopiperazine core and the oxybenzylidene moiety. The only difference was the downfield or upfield shifts of some 1H and 13C signals probably due to the change of configuration at the Δ13,14 double bond. This configuration was deduced to be Z from careful analysis of the NOESY spectrum which exhibited a cross-peak correlation between the methylene protons at δH 4.67 (brd, J = 6.4 Hz, H-12) and 4,16 (s, H-16). The NOESY correlation between the olefinic proton H-13 (δH, 5.57, m) and the methyl protons H-15 (δH 1.85) was also depicted. In view of determining the configuration of the Δ6,7 double bond, the NOESY spectrum of 2 was also measured in DMSO-d6. The NOESY correlation observed between the NH-1 proton signal (δH 10.17) and H-9/H-9′ (δH 7.45, d, J = 8.6) showed that Δ6,7 has the same geometry in metabolites 1 and 2. The experimental ECD spectrum of 2 (Fig. S18, Supporting information) was nearly identical to that of 1, leading to the deduction of the 3R absolute configuration for 2. The structure of metabolite 2 was thus concluded as (3R,6Z)-3-thiomethyl-6-[4-O-[(2Z)-4-hydroxy-3-methylbut-2-enyl]benzylidene]piperazine-2,5-dione.
Compound 3 was obtained as a yellowish gum. Its molecular formula was determined as C16H18N2O4 from the HRESIMS analysis which showed the pseudomolecular ion at m/z 303.1334 [M + H]+ (calcd for C16H19N2O4+, 303.1339). Its 1H NMR spectrum exhibited resonances for an AA′BB′ spin system at δH 7.46 (d, J = 8.7 Hz, H-9, and H-9′) and 6.98 (d, J = 8.7 Hz, H-10, and H-10′) suggesting a para-disubstituted aromatic ring in the structure. It also showed signals for a vinyl proton at δH 5.47 (m, H-13), an oxymethylene at δH 4.58 (brd, J = 6.6 Hz, H-12), and two vinyl connected methyl singlets at δH 1.79 (H-15) and 1.76 (H-16) characteristic of an O-isoprenyl moiety. Signals of the vinyl proton H-7 and the methine proton H-3 were observed at δH 6.80 (s) and 5.09 (s), respectively. The 13C NMR spectrum exhibited signals of two amidocarbonyl carbons at δC 167.0 (C-2) and 163.8 (C-5). Those characteristic of the oxybenzylidene moiety were observed at δC 125.4 (C-6), 119.5 (C-7), 126.8 (C-8), 132.1 (C-9, C-9′), 116.4 (C-10, C-10′), and 160.9 (C-11). The remaining resonances depicted at δC 66.1 (C-12), 121.1 (C-13), 139.1 (C-14), δC 26.0 (C-15), and 18.3 (C-16) confirmed the presence of the O-prenyl group in the molecule (Table 1). The downfield shift of C-3 in metabolite 3 (δC 76.1) with respect to compounds 1 and 2 (δC 59.6) suggested the presence of an OH group at C-3 in 3 instead of the thiomethyl moiety. This was further supported by mass analysis and the absence of the thiomethyl signal on the 1H and 13C NMR spectra. The structure was confirmed by a comprehensive analysis of the 2D NMR data, particularly 1H-1H COSY, HSQC, and HMBC spectra (Fig. 4). The configuration of the Δ6,7 double bond was determined to be Z by the NOESY spectrum measured in DMSO-d6 on which correlations were observed between NH-1 and H-9 (or H-9′) but not between NH-1 (δH 9.92, s) and H-7 (δH 6.80, s). The ECD spectrum of 3 (Fig. S27, Supporting information) indicated the same absolute configuration at C-3 (R) as for metabolites 1 and 2. The structure was finally concluded as (3R,6Z)-3-hydroxy-6-[4-O-(3-methylbut-2-enyl)benzylidene]piperazine-2,5-dione.
The molecular formula of 4 also obtained as a yellowish gum was deduced to be C17H20N2O3S from the HRESIMS which showed ion clusters [M + Na]+ at m/z 355.1088 (Calcd 355.1087) and [2M + Na]+ at m/z 687.2283 (Calcd 687.2281). The NMR data (Table 2) showed similarities with the previously described metabolites 1–3. Its 1H-NMR spectrum exhibited in addition to signals of the diketopiperazine and the oxybenzylidene moieties two olefinic methyl resonances at δH 1.76 and 1.79, an oxygenated methylene doublet at δH 4.58 (J = 6.6 Hz), and a vinyl proton signal at δH 5.46 (m) characteristic of a γ,γ-dimethylallyloxy moiety (Sritularak and Likhitwitayawuid 2006). Careful examination of the 1H-1H COSY, HSQC, and HMBC spectra proved that 4 was similar to Sch 56396 previously isolated from the fermentation broth of the fungus Tolypocladium sp. (Chu et al. 1997), but the only difference was on the sign of their optical rotations. Compound 4 showed a positive optical rotation, while the optical rotation of Sch 56396 was negative, confirming that they are stereoisomers. The Z configuration of the Δ6,7 was deduced from the chemical shift of H-7 (δH 6.87) in comparison with those of the same proton in compounds 1–3, since it was reported that the (Z)-vinyl proton of the 6-benzylidene-substituted piperazine-2,5-diones is farther downfield than the (E)-vinyl proton because of the deshielding effect of the 5-ketone (Fu et al. 2011). Since the configuration of the chiral centre C-3 of Sch 56396 was not determined, we measured the ECD spectrum of 4 (Fig. S36, Supporting information) and its comparison with those of compounds 1–3 allowed us to assign the 3R configuration. Compound 4 was then elucidated as (3R,6Z)-3-thiomethyl-6-[4-O-(3-methylbut-2-enyl)benzylidene]piperazine-2,5-dione, the stereoisomer of Sch 56396.
Compound 5 was obtained as a yellow gum from methanol. Its HRESIMS showed ion clusters at m/z 397.1249 [M + H]+ and 419.1068 [M + Na]+ consistent with the molecular formula of C18H24N2O4S2 (calcd for C18H25N2O4S2+, 397.1250; calcd for C18H24N2O4S2Na+, 419.1075). The presence of two thiomethyl groups was confirmed by the ion fragments depicted at m/z 349.1215 [M + H–48]+ and 301.1177 [M + H–2 × 48]+ revealing the loss of two methanethiol (CH3SH) units (Chu et al. 1993). Its 1H NMR spectrum showed in addition to the signals of the O-isoprenoltyrosine moiety at δH 7.26 (d, J = 9.0 Hz, H-9, and H-9′), 6.83 (d, J = 9.0 Hz, H-10, and H-10′), 4.61 (brd, J = 6.3 Hz, H-12), 5.47 (m, H-12), 4.15 (s, H-15), and 1.81 (s, H-16) those of two thiomethyl groups at δH 1.36 (s, H-18) and 2.23 (s, H-17) as well as a methylene group observed as an AX spin system at δH 2.94 (d, J = 13.5, H-7A) and 3.60 (d, J = 13.5, H-7X) (Table 2). Careful examination of the 13C, 1H-1H COSY, HSQC, and HMBC spectra proved metabolite 5 to have the same planar structure as meromutides A and B recently isolated after pleiotropic activation of natural products in Metarhizium robertsii by deletion of a histone acetyltransferase (Fan et al. 2017). The Z geometry for the Δ13,14 double bond was determined from the NOESY correlation depicted between the methylene protons at δH 4.61 (brd, J = 6.4 Hz, H-12) and 4.15 (s, H-16). Compound 5 with the Z geometry of the olefinic double bond can possess 4 stereoisomers (3S,6S), (3R,6S), (3S,6R), and (3R,6R). Up to now, only two of them, namely meromutide A (3S,6S) and meromutide B (3R,6S) were isolated and characterised. Its absolute configuration was determined by careful comparison of the chemical shifts for both protons and carbons of the thiomethyl groups linked at C-3 and C-6 of its known stereoisomers. For meromutide A, these chemical shifts were as follows: C-3 (δH 2.24; δC 13.9) and C-6 (δH 2.30; δC 13.4), while for meromutide B, they were C-3 (δH 2.21; δC 10.4) and C-6 (δH 1.20; δC 12.9) (Fan et al. 2017). The chemical shifts of the corresponding protons and carbons of the thiomethyl groups at C-3 and C-6 in compound 5 (also measured in methanol-d6) were different from those of the known stereoisomers ((δH 1.36; δC 10.6) and (δH 2.23; δC 13.4), respectively) indicating a change of configuration in one of the chiral centres. On the other hand, since metabolite 5 is the C-15 hydroxyl derivative of Sch54796 (6) and Sch54794 (7) possessing the known (3R,6S) and (3S,6S) configurations, its absolute configuration at C-6 must be R as supported by the positive optical rotation (+ 33.3, MeOH) in comparison with those of the known congeners (Sch54796 (− 25, DMSO) and Sch54794 (− 70, DMSO); see Chu et al. (1993). Furthermore, the NOESY correlation depicted between H-3 (δH 5.01, s) and CH3-18 (2.23, s) revealed the trans-orientation of the two thiomethyl groups on the diketopiperazine ring. To further confirm the 3S,6R configuration, we compared the ECD spectrum of metabolite 5 with that of fusaperazine, a related thiodiketopiperazine possessing a 3R,6R configuration obtained from a marine algae-derived fungus Penicillium sp. KMM 4672 (Yurchenko et al. 2019). Both compounds exhibited a negative Cotton effect between 220 and 240 nm, probably due to the common 6R configuration. However, a negative Cotton effect was observed on the ECD spectrum of compound 5 between 240 and 290 nm while a positive Cotton effect was depicted in the same zone of the ECD spectrum of fusaperazine. The structure of 5 was finally elucidated as (3S,6R)-3,6-bisthiomethyl-6-[4-O-[(2Z)-4-hydroxy-3-methylbut-2-enyl]phenylmethyl]piperazine-2,5-dione.
Diketopiperazines are cyclic peptides produced by bacteria and fungi arising from the cyclisation of two or more amino acids catalysed either by two-modular non-ribosomal peptide synthetases or by cyclodipeptide synthases (Huang et al. 2014; Brockmeyer and Li 2017). Since the isolated compounds are biogenetically related, their biosynthetic relationships were proposed. Compounds 10, 11, and 12 could be obtained from the cyclisation of L-proline and L-isoleucine, L-proline and L-leucine, and L-proline and L-phenylalanine, respectively. While working on the biosynthesis of the epidithiodiketopiperazine gliotoxin, Scharf et al. 2011 discovered that a specialised glutathione S-transferase (GliG) plays a key role in C-S bond formation (sulfurisation) and that bishydroxylation of the diketopiperazine by oxygenase (GliC) is a prerequisite for glutathione adduct formation. Cyclisation of glycine and L-tyrosine followed by O-prenylation affords cyclo-(glycyl-L-tyrosyl)-3,3-dimethylallyl ether 8 which could undergo bishydroxylation by oxygenase GliC to yield the intermediate 13 (not isolated). Thiolation of 13 in the presence of GliG could afford Sch 54796 (6) and Sch 54794 (7). The C-6 epimerisation of Sch 54794 (7) followed by hydroxylation at C-15 could lead compound 5. The intermediate 13 could also undergo dehydration to afford metabolite 3 which could give compound 9 after an oxidative cleavage of the C-6–C-7 double bond. Thiolation of metabolite 3 could also lead compound 4, which could undergo hydroxylation of one of the prenylmethyl groups to give 1 and 2 (Fig. 5).
Biological activities
Since some derivatives of diketopiperazines were reported to display significant antibiotic, antitumor, and immunosuppressant properties (Ameur et al. 2004), while others show a wide range of biological effects in cell cycle progression (Cui et al. 1996), the isolated compounds were tested for their antimicrobial and cytotoxic activities against various bacteria, fungi, and two mammalian cell lines, but only weak cytotoxic activity was observed for metabolites 1, 4, and 9 against KB3.1 cells. The evaluation of these compounds in additional bioassays is presently underway.
In general, Batnamyces globulariicola belongs to a group of Chaetomiaceae that has been poorly studied for secondary metabolites, suggesting that it will be worthwhile to examine further strains that appear phylogenetically related for the production of diketopiperazines and other secondary metabolites. The lack of suitable morphological features for the classification of these fungi using a polythetic approach could thus be compensated by chemotaxonomic methodology, as recently accomplished for some genera of the Xylariales (cf. Samarakoon et al. 2020; Wittstein et al. 2020).
Conclusion
Batnamyces globulariicola was only isolated once among several hundreds of cultures that inhabited the rhizosphere of the host plant during the course of a PhD thesis (Noumeur 2018). We selected this strain out of many others that were isolated concurrently because it turned out to represent a hitherto unknown phylogenetic lineage among the Sordariales. We were at first unsure whether to report the fungus as an “unknown member” of the Chaetomiaceae, along with the secondary metabolites, but finally decided to name the taxon, and place it in its phylogenetic context, as part of an ongoing taxonomic study (Wang et al. 2019b). It was rather surprising to see that it still did not fit in any of the known genera of the Chaetomiaceae even by comparison with the latest monographic work.
We are aware of the potential pitfalls regarding the classification of such a single strain as a member of a monotypic genus, and in particular the requests by some members of the mycological community that more than one culture needs to be deposited to justify the formal description of a new taxon. However, such arguments mostly come from scientists who are working with ubiquitous fungal genera that make up the bulk of new species in the large classes of Eurotiomycetes, Sordariomycetes, and Dothideomycetes, where ex-type strains as well as large numbers of congeneric isolates are readily available.
We cannot be absolutely certain whether our new fungus really represents an endophyte, because it was only isolated once. However, a sterile mycelium could hardly have survived the applied, rather harsh root disinfection procedure unless it forms persistent propagules which may have survived the chemical treatment. Nevertheless, we cannot exclude that it is heterothallic and that a sexual state may exist in nature. Even though we did not observe sporulation of the culture, aside from the production of chlamydospores, we cannot exclude that it forms persistent propagules in its natural habitat that we have not been able to induce in the laboratory. There remains a chance that a yet unknown stage of the fungus has persisted in a soil particle that was attached to the roots, and it should be attempted to re-isolate Batnamyces from other parts of the plant. The taxonomy of other fungi that were already frequently isolated from similar biotopes, such as the so-called dark septate root endophytes, which mostly belong to the Dothideomycetes (Knapp et al. 2015; Bonfim et al. 2016), poses a similar challenge. Their taxonomy cannot be resolved by using a morphocentric approach based on conidiogenous structures but must rely on molecular methods. Even here, a chemotaxonomic approach would be interesting to pursue to attain complementary phenotype-derived data.
References
Ameur RMB, Mellouli L, Chabchoub F, Fotso S, Bejar S (2004) Purification and structure elucidation of two biologically active molecules from a new isolated Streptomyces sp. US 24 strain. Chem Nat Compd 40:510–513
Bills GF, Gloer JB (2016) Biologically active secondary metabolites from the fungi. Microbiol Spectrum 4:6
Blackwell M, Vega FE (2018) Lives within lives: hidden fungal biodiversity and the importance of conservation. Fungal Ecol 35:127–134
Bonfim JA, Vasconcellos RL, Baldesin LF, Sieber TN, Cardoso EJ (2016) Dark septate endophytic fungi of native plants along an altitudinal gradient in the Brazilian Atlantic forest. Fungal Ecol 20:202–210
Brockmeyer K, Li SM (2017) Mutations of residues in pocket P1 of a cyclodipeptide synthase strongly increase product formation. J Nat Prod 80:2917–2922
Cai L, Jeewon R, Hyde KD (2006) Molecular systematics of Sordariaceae based on multiple gene sequences and morphology. Mycol Res 110:137–150
Chepkirui C, Cheng T, Sum WC, Matasyoh JC, Decock C, Praditya DF, Wittstein K, Steinmann E, Stadler M (2019) Skeletocutins A-L: antibacterial agents from the Kenyan wood-inhabiting basidiomycete, Skeletocutis sp. J Agric Food Chem 31:8468–8475
Chu M, Mierzwa R, Truumees I, Gentile F, Patel M, Gullo V, Chan TM, Puar MS (1993) Two novel diketopiperazines isolated from the fungus Tolypocladium sp. Tetrahedron Lett 34:7537–7540
Chu M, Truumees I, Mierzwa R, Patel M, Puar MS (1997) Sch 56396: a new c-fos proto-oncogene inhibitor produced by the fungus Tolypocladium sp. J Antibiot 50:1061–1063
Crous PW, Verkley GJ, Groenewald JZ, Samson RA (2009) Fungal biodiversity. CBS laboratory manual series 1. Centraalbureau voor Schimmelcultures, Utrecht
Cui CB, Kakeya H, Okada G, Onose R, Osada H (1996) Novel mammalian cell cycle inhibitors, tryprostatins A, B and other diketopiperazines produced by Aspergillus fumigatus. J Antibiot 49:527–533
De Hoog GS, Ahmed SA, Najafzadeh MJ, Sutton DA, Saradeghi Keisari M, Fahal AH, Eberhart U, Verkley GJ, Xin L, Stielow B, van de Sande WWJ (2013) Phylogenetic findings suggest possible new habitat and routes of infection of human eumyctoma. PLoS Negl Trop Dis 7:e2229
Fan A, Mi W, Liu Z, Zeng G, Zhang P, Hu Y, Fang W, Yin WB (2017) Deletion of a histone acetyltransferase leads to the pleiotropic activation of natural products in Metarhizium robertsii. Org Lett 19:1686–1689
Fu P, Liu P, Qu H, Wang Y, Chen D, Wang H, Li J, Zhu W (2011) α-Pyrones and diketopiperazine derivatives from the marine-derived actinomycete Nocardiopsis dassonvillei HR10-5. J Nat Prod 74:2219–2223
Guimarães DO, Borges WS, Vieira NJ, De Oliveira LF, Da Silva CHTP, Norberto P, Lopes NP, Dias LG, Durán-Patrón R, Collado IG, Pupo MT (2010) Diketopiperazines produced by endophytic fungi found in association with two Asteraceae species. Phytochemistry 71:1423–1429
Huang RM, Yi XX, Zhou Y, Su X, Peng Y, Gao CH (2014) An update on 2,5-diketopiperazines from marine organisms. Mar Drugs 12:6213–6235
Hyde KD, Xu JC, Rapior S, Jeewon R, Lumyong S et al (2019) The amazing potential of fungi, 50 ways we can exploit fungi industrially. Fungal Divers 97:1–136
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Knapp DG, Kovács GM, Zajta E, Groenewald JZ, Crous PW (2015) Dark septate endophytic pleosporalean genera from semiarid areas. Persoonia 35:87
Koolen HHF, Soares ER, Da Silva FMA, De Souza AQL, De Medeiros LS, Filho ER, De Almeida RA, Ribeiro IA, Pessoa C, De Morais MO, Da Costa PM, De Souza ADL (2012) An antimicrobial diketopiperazine alkaloid and co-metabolites from an endophytic strain of Gliocladium isolated from Strychnos cf. toxifera. Nat Prod Res 26:2013–2019
Miller MA, Pfeiffer W, Schwartz T (2010) Creating the CIPRES Science Gateway for inference of large phylogenetic trees. In: Proceedings of the Gateway Computing Environments Workshop (GCE), November 14, 2010, New Orleans, Louisiana: pp. 1–8
Noumeur SR (2018) Identification of bioactive natural products from endophytic fungi isolated from Globularia alypum. PhD dissertation, University of Setif, Algeria
Noumeur SR, Helaly SE, Jansen R, Gereke M, Stradal TEB, Harzallah D, Stadler M (2017) Preussilides A−F, bicyclic polyketides from the endophytic fungus Preussia similis with antiproliferative activity. J Nat Prod 80:1531–1540
Nozawa K, Takada M, Udagawa SL, Nakajima S, Kawai KI (1989) Three p-hydroxybenzoic acid derivatives from Talaromyces derxii. Phytochemistry 28:655–656
Nylander JAA (2004) MrModeltest v. 2. Programme distributed by the author. Evolutionary Biology Centre. Uppsala University
Rambaut A (2009) FigTree v. 1.3.1. Computer program and documentation distributed by the author at. http://tree.bio.ed.ac.uk/software/
Ren S, Ma W, Xu T, Lin X, Yin H, Yang B, Zhou XF, Yang XW, Long L, Lee KJ, Gao Q, Liu Y (2010) Two novel alkaloids from the South China Sea marine sponge Dysidea sp. J Antibiot 63:699–701
Richter C, Helaly SE, Thongbai B, Hyde KD, Stadler M (2016) Pyristriatins A and B: pyridino-cyathane antibiotics from the basidiomycete Cyathus cf. striatus. J Nat Prod 79:1684–1688
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Samarakoon MC, Thongbai B, Hyde KD, Brönstrup M, Beutling U, Lambert C, Miller AN, Liu JK, Promputtha I, Stadler M (2020) Elucidation of the life cycle of the endophytic genus Muscodor and its transfer into the genus Induratia in Induratiaceae fam. nov., based on a polyphasic taxonomic approach. Fungal Divers. https://doi.org/10.1007/s13225-020-00443-9
Sandargo B, Thongbai B, Praditya D, Steinmann E, Stadler M, Surup F (2018) Antiviral 4-hydroxypleurogrisein and antimicrobial pleurotin derivatives from cultures of the nematophagous basidiomycete Hohenbuehelia grisea. Molecules 23:2697
Sansinenea E, Salazar F, Jiménez J, Mendoza A, Ortiz A (2016) Diketopiperazines derivatives isolated from Bacillus thuringiensis and Bacillus endophyticus, establishment of their configuration by X-ray and their synthesis. Tetrahedron Lett 57:2604–2007
Scharf DH, Remme N, Habel A, Chankhamjon P, Scherlach K, Heinekamp T, Hortschansky P, Brakhage AA, Hertweck C (2011) A dedicated glutathione S-transferase mediates carbon-sulfur bond formation in gliotoxin biosynthesis. J Am Chem Soc 133:12322–12325
Sritularak B, Likhitwitayawuid K (2006) Flavonoids from the pods of Millettia erythrocalyx. Phytochemistry 67:812–817
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Suryanarayanan TS, Thirunavukkarasu N, Govindarajulu MB, Sasse F, Jansen R, Murali TS (2009) Fungal endophytes and bioprospecting. Fungal Biol Rev 23(1–2):9–19
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Teponno RB, Noumeur SR, Helaly SE, Hüttel S, Harzallah D, Stadler M (2017) Furanones and anthranilic acid derivatives from the endophytic fungus Dendrothyrium variisporum. Molecules 22:1674–1685
Usami Y, Aoki S, Hara T, Nu A (2002) New dioxopiperazine metabolites from a Fusarium species separated from a marine alga. J Antibiot 55:655–659
Van den Brink J, Samson RA, Hagen F, Boekhout T, de Vries RP (2012) Phylogeny of the industrial relevant, thermophilic genera Myceliophthora and Corynascus. Fungal Divers 52:197–207
Van den Brink J, Facun K, de Vries M et al (2015) Thermophilic growth and enzymatic thermostability are polyphyletic traits within Chaetomiaceae. Fungal Biol 119:1255–1266
Wang XW, Houbraken J, Groenewald JZ, Meijier M, Andersen B, Nielsen KF, Crous PW, Samson RA (2016a) Diversity and taxonomy of Chaetomium and chaetomium-like fungi from indoor environments. Stud Mycol 84:145–224
Wang XW, Lombard L, Groenewald JZ, Li J, Videira S, Samson RA, Liu XZ, Crous PW (2016b) Phylogenetic reassessment of the Chaetomium globosum species complex. Persoonia 36:83–133
Wang XW, Yang FY, Meijer M, Kraak B, Sun BD, Jiang YL, Wu YM, Bai FY, Seifert KA, Crous PW, Samson RA (2019a) Redefining Humicola sensu stricto and related genera in the Chaetomiaceae. Stud Mycol 93:65–153
Wang XW, Bai FY, Bensch K, Meijer M, Sun BD, Han YF, Crous PW, Samson RA, Yang FY, Houbraken J (2019b) Phylogenetic re-evaluation of Thielavia with the introduction of a new family Podosporaceae. Stud Mycol 93:155–252
Wendt L, Sir EB, Kuhnert E, Heitkämper S, Lambert C, Hladki AI, Romero AI, Luangsa-ard JJ, Srikitikulchai P, Peršoh D, Stadler M (2018) Resurrection and emendation of the Hypoxylaceae, recognised from a multi-gene genealogy of the Xylariales. Mycol Prog 17:115–154
White JF, Kingsley KL, Zhang Q, Verma R, Obi N, Dvinskikh S, Elmore MT, Verma SK, Gond SK, Kowalski KP (2019) Endophytic microbes and their potential applications in crop management. Pest Manag Sci 75(10):2558–2565
Wittstein K, Cordsmeier A, Lambert C, Wendt L, Sir EB, Weber J, Wurzler N, Petrini LE, Stadler M (2020) Identification of Rosellinia species as producers of cyclodepsipeptide PF1022 A and resurrection of the genus Dematophora as inferred from polythetic taxonomy. Stud Mycol 96:1–16
Yurchenko AN, Berdyshev DV, Smetanina OF, Ivanets EV, Zhuravleva OI, Rasin AB, Khudyakova YV, Popov RS, Sergey A, Dyshlovoy SA, Amsberg GV, Afiyatullov SS (2019) Citriperazines A-D produced by a marine algae-derived fungus Penicillium sp. KMM 4672. Nat Prod Res, in press. https://doi.org/10.1080/14786419.2018.1552696
Acknowledgements
The technical assistance of Nadine Wurzler and Vanessa Stiller is appreciated. We thank Wera Collisi for conducting the bioassays, Christel Kakoschke for recording NMR spectra and Sabrina Karwehl for HRESIMS measurements.
Funding
Open Access funding provided by Projekt DEAL. RBT and SEH are grateful for financial support from the Alexander von Humboldt Foundation. SRN acknowledges the Ministry of Higher Education and Scientific Research (MESRS) of Algeria for the financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Section Editor: Hans-Josef Schroers
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noumeur, S.R., Teponno, R.B., Helaly, S.E. et al. Diketopiperazines from Batnamyces globulariicola, gen. & sp. nov. (Chaetomiaceae), a fungus associated with roots of the medicinal plant Globularia alypum in Algeria. Mycol Progress 19, 589–603 (2020). https://doi.org/10.1007/s11557-020-01581-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-020-01581-9