Abstract
Forecasting invasive-pathogen dynamics is paramount to anticipate eradication and containment strategies. Such predictions can be obtained using a model grounded on partial differential equations (PDE; often exploited to model invasions) and fitted to surveillance data. This framework allows the construction of phenomenological but concise models relying on mechanistic hypotheses and real observations. However, it may lead to models with overly rigid behavior and possible data-model mismatches. Hence, to avoid drawing a forecast grounded on a single PDE-based model that would be prone to errors, we propose to apply Bayesian model averaging (BMA), which allows us to account for both parameter and model uncertainties. Thus, we propose a set of different competing PDE-based models for representing the pathogen dynamics, we use an adaptive multiple importance sampling algorithm (AMIS) to estimate parameters of each competing model from surveillance data in a mechanistic-statistical framework, we evaluate the posterior probabilities of models by comparing different approaches proposed in the literature, and we apply BMA to draw posterior distributions of parameters and a posterior forecast of the pathogen dynamics. This approach is applied to predict the extent of Xylella fastidiosa in South Corsica, France, a phytopathogenic bacterium detected in situ in Europe less than 10 years ago (Italy 2013, France 2015). Separating data into training and validation sets, we show that the BMA forecast outperforms competing forecast approaches.
Similar content being viewed by others
1 Introduction
The emergence of exogenous pathogens in new territories may induce severe sanitary and socio-economical crises. Mainly, such crises are reinforced by the eventual long delay between pathogen establishment in the new territory and its detection (Jones and Baker 2004; Faria et al. 2014; Soubeyrand et al. 2018). As this delay increases, the potential for pathogen expansion and the resulting cost for pathogen eradication or containment generally increases. Hence, reconstructing the past dynamics of the pathogen (Boys et al. 2008; Roques et al. 2016; Soubeyrand and Roques 2014) and predicting its future extent (Chapman et al. 2015; Peterson et al. 2003) are key steps for understanding the pathogen epidemiology, designing eradication or containment strategies and assessing their potential efficiency.
Partial differential equations (PDE) have been extensively used for modeling spatio-temporal population dynamics (Skellam 1951; Shigesada et al. 1995; Turchin 1998; Okubo and Levin 2002). PDE can precisely be used for past dynamics reconstruction and future extent prediction, by exploiting their ability (1) to represent dynamics in a phenomenological and concise way, and (2) to be fitted to data by attaching a probabilistic model of observations within a state-space modeling framework (the combination of (1) and (2) corresponds to the so-called physical-statistical or mechanistic-statistical approach; Berliner 2003; Wikle 2003; Roques et al. 2011; Soubeyrand and Roques 2014). To apply such an approach, the PDE is generally chosen to be parsimonious for identifiability reasons (i.e., the ability to estimate its parameters given the information contained in data). However, when left parsimonious, a PDE may not be proficient in describing all the processes and sources of variability involved in epidemiological dynamics. In addition, various structures of PDE are likely to be considered as candidate models for a given epidemic. When the goal of the study is to draw predictions, the use of a single model is prone to prediction error because this model may not have taken into account crucial drivers of the dynamics. This limitation can be circumvented by associating random terms to the PDE as proposed by Wikle (2003), or by considering a set of competing PDE-based models and applying either a model selection strategy (Burnham et al. 1995) or a model aggregation strategy (Hoeting et al. 1999).
As part of the model aggregation strategy, the Bayesian model averaging (BMA) approach has been proposed by Leamer (1978) to reduce and account for parameter and model uncertainties. This approach consists in performing a weighted average of candidate models in a Bayesian way and hence combining multiple predictions and multiple estimations of shared parameters (Raftery 1996; Madigan and Raftery 1994; Wintle et al. 2003). Theoretically, BMA provides better average predictive ability, as measured by a logarithmic scoring rule, than using any single model (Madigan and Raftery 1994). Axiomatically, this result depends on the assumption that the data are generated in the following stages (Fletcher 2018): (1) a model is selected at random from the set of candidate models using prior model probabilities, (2) the parameter values for this model are generated using the relevant prior distribution, and (3) the data are generated from the selected model and parameter values.
The BMA efficiency has been largely explored, in particular with respect to its theoretical properties (Rubin and Schenker 1986; Madigan and Raftery 1994), leave-one-out predictive performance (Madigan et al. 1995; Lamon and Clyde 2000) and numerical performance (George and McCulloch 1993; Viallefont et al. 2001). While BMA is an intuitively attractive solution to the problem of accounting for model uncertainty, it presents several difficulties related to its numerical implementation (Hoeting et al. 1999). By dint of some pioneering work implementing BMA (Madigan and Raftery 1994; Raftery 1996), this approach has then been applied in numerous study domains such as medicine (Oehler et al. 2009), ecology (Wintle et al. 2003), meteorology (Raftery et al. 2005), genetics (Yeung et al. 2005), economical and political sciences (Sidman et al. 2008), engineering and physical sciences (Parkinson and Liddle 2013) and epidemiology (Viallefont et al. 2001). Despite ample literature on BMA and its usefulness, it has been marginally applied in the context of predictive epidemiology.
In this article, we investigate the application of BMA in the context of pathogen-dynamics prediction using PDE-based models and we want to test its efficiency on a real case study. The models are grounded on a family of reaction-diffusion equations, some of which include spatially heterogeneous diffusion and reproduction terms. Our aim is to compute, from post-introduction data, the BMA posterior distribution of a certain quantity of interest \(\Delta \), which is typically the introduction time or location of the pathogen or its future spatial extent. Following Abboud et al. (2019), we apply to each model the Adaptive Multiple Importance Sampling algorithm (AMIS; Cornuet et al. 2012) for providing an empirical approximation, obtained via a weighted sample \({\{\Delta _n, w _n\}}_{n=1}^N\) of size N, of the posterior distribution of \(\Delta \) given the specified model. Then, for drawing BMA posterior samples of \(\Delta \), we compute posterior probabilities of models using different approximations of the integrated likelihood that have been proposed in the literature. Namely, we compare an estimator of the integrated likelihood, which is easily obtained by averaging AMIS un-normalized importance weights (Bugallo et al. 2015), to estimators of the integrated likelihood grounded on information criteria (McElreath 2018), as well as harmonic mean estimators (Raftery 1996; Gelfand and Dey 1994).
This approach is first tested on simulated data and then applied to make predictions concerning the dynamics of the phytopathogenic bacterium Xylella fastidiosa (Xf) in South Corsica, France. This quarantine pathogen in Europe has significantly impacted olive production in Puglia, Italy, and presents a drastic risk of environmental degradation due to its ability to reach a large variety of plant species. It is currently present in a large part of Corsica island and more marginally in Southern mainland France (Denancé et al. 2017a; Soubeyrand et al. 2018; Martinetti and Soubeyrand 2019). Xf might cause a major sanitary crisis in France, as the one caused in Italy since 2013 where the socio-economical impacts are considerable due to the death and felling of numerous olive trees in Puglia. In the case of South Corsica, spatio-temporal and presence–absence post-introduction surveillance data were collected from an intensive surveillance plan implemented by governmental agencies after the first in situ detection of Xf in 2015 in the city of Propriano. These data covering about three years and a half are separated into a training set (nearly 2 years) used for fitting the models and a validation set (nearly 1.5 years) used for comparing the forecasts obtained with BMA, the best PDE-based model (i.e., the model with the largest posterior model probability computed in the BMA procedure), an ensemble approach (i.e., the equiprobable average of all the PDE-based models parameterized by the maximum likelihood estimates) and two data-oriented prediction strategies not grounded on PDE.
The paper is organized as follows: Data are briefly described in Sect. 2. The competing models coupling a partial differential equation and a Bernoulli observation process are presented in Sect. 3. The Bayesian model averaging technique is described in Sect. 4. The simulation study is presented in Sect. 5. Results obtained from surveillance data for Xf in South Corsica are detailed in Sect. 6 where we specifically focus on model comparison, parameter inference and out-of sample predictive performance. Finally, Sect. 7 provides a conclusion and a discussion of perspectives.
2 Surveillance Data with Presence–Absence Records
In this article, we analyze spatio-temporal presence–absence data collected in South Corsica, France, and informing if sampled plants are positive or negative to Xf based on a molecular test. Data have been collected since the first detection of the bacterium in the study region in July 2015. Between July 2015 and January 2019, approximately 9500 plants were sampled, among which 900 have been diagnosed as infected with a real-time polymerase chain reaction (real-time PCR) technique (Denancé et al. 2017b). GPS coordinates, sampling dates and sanitary statuses (healthy/infected) are available for all the sampled plants. Spatial locations and sanitary statuses at the sampling times are shown in Fig. 1, left.
As for other bacteria, the growth and mortality of Xf are affected by various environmental variables such as habitability of the environment, nutrients, climatic conditions and availability of dissemination means (typically, insect vectors). In this study, to account for spatial heterogeneity in the diffusion and the reproduction regimes of the epidemics, we use temperature data to divide the spatial domain denoted by \(\Omega \) into two complementary and non-overlapping sub-domains, and different diffusion and growth terms are applied to the two sub-domains. We exploit a freely available database (PVGIS\(\copyright \) European Communities, 2001–2008) providing, in particular, monthly averages of the daily minimum temperature reconstructed over a grid with 1\(\times \)1 km spatial resolution (Huld et al. 2006); these monthly averages correspond to the period 1995–2003, which is between the probable introduction around 1960 of Xf in South Corsica (Soubeyrand et al. 2018) and the observation window starting in 2015. We use these data to build the average of the daily minimum temperature over January and February, say \(T({\textbf {x}})\) for any location \({\textbf {x}}\); see Fig. 1, right. Average daily minimum temperature in Winter is known to be a crucial factor for the presence or abundance of Xf (Anas et al. 2008; Martinetti and Soubeyrand 2019).
Remark. In this work we divide the spatial domain based on a static assessment of winter temperatures. This allows us to take into account a relatively large spatial heterogeneity of the environment since the winter temperature, as we defined it, ranges from \(-\) 0.7 \(^\circ \)C in the coldest area of South Corsica to 6.8 \(^\circ \)C in its warmest area. Ideally, we should also take into account the annual variability of winter temperatures and eventual other environmental variables in the dynamic process, as proposed by Botella et al. (2022) for instance. Such data may be obtained from a data base such as WorldClim (https://worldclim.org/) at the 1\(\times \)1 km resolution that we consider here.
3 Competing Models
Here, a set of models based on parabolic partial differential equations is used to describe pathogen dynamics at large spatial scales. As explained in the introduction, these models have been extensively used to represent population dynamics in a phenomenological and concise way, and can be fitted to data in a hierarchical modeling setting incorporating a probabilistic model of observations. In this section, we propose a family of mechanistic models and we present the model for the observation process.
3.1 Family of Mechanistic Models
We introduce a discrete family \({\varvec{\mathcal {M}}}=\{\mathcal {M}_i(\tilde{T}): 0\le i \le I; \tilde{T}\in \mathcal {T} \}\) of models governing the probability \(u(t,{\textbf {x}})\) of a host located at site \({{\textbf {x}}}=(x_1,y_1)\in \Omega \) to be infected (i.e., sick because of the pathogen under consideration) at time t, where \(I\in \mathbb {N}^*\) and \(\mathcal {T}\) is a finite collection of real values. The label i refers to a model structure, i.e. a specific form for the parabolic PDE. The label \(\tilde{T}\) refers in our application to a temperature threshold, which splits the spatial domain into two sub-domains where diffusion and growth terms may be different. The generic form of models in family \({\varvec{\mathcal {M}}}\) satisfies:
where the first line is the reaction-diffusion equation, the second line gives boundary conditions, the third line gives initial conditions, \(\varvec{\Delta }=\dfrac{\partial ^2}{\partial x_1^2}+\dfrac{\partial ^2}{\partial x_2^2}\) is the 2-dimensional diffusion operator of Laplace, and \(\varvec{\nabla }=\dfrac{\partial }{\partial x_1}+\dfrac{\partial }{\partial x_2}\) is the 2-dimensional gradient operator.
The diffusion coefficient \(D_{i\tilde{T}}({{\textbf {x}}})\) may be spatially heterogeneous and is defined as a spatial regularization of \( d_{i\tilde{T}}({{\textbf {x}}})=\sum _{k=1}^2 D_{i\tilde{T}k}{\mathbbm {1}}({{\textbf {x}}}\in \Omega _{\tilde{T}k}),\,\,\,\forall i\le I, \forall \tilde{T}\in \mathcal {T},\) where \({\textbf {x}}\mapsto {\mathbbm {1}} ({{\textbf {x}}}\in \Omega _{\tilde{T}k})\) is the indicator function taking the value 1 if \({\textbf {x}}\in \Omega _{\tilde{T}k}\) and 0 otherwise, and the sub-domains \(\Omega _{\tilde{T}1}\) and \(\Omega _{\tilde{T}2}\) are defined by thresholding the spatial function T, with the threshold value \(\tilde{T}\) such that: \(\Omega _{\tilde{T}1}=\{{{\textbf {x}}}\in \Omega :T({{\textbf {x}}})> \tilde{T}\}\) and \(\Omega _{\tilde{T}2}=\{{{\textbf {x}}}\in \Omega :T({{\textbf {x}}})\le \tilde{T}\}\). In our application, T is a measure of temperature in winter, \(\Omega _{\tilde{T}1}\) is the warm region of \(\Omega \), and \(\Omega _{\tilde{T}2}\) the cold one. If \(D_{i\tilde{T}1}=D_{i\tilde{T}2}\), then the diffusion coefficient is spatially homogeneous. The spatial regularization is required for the existence and the uniqueness of a classic solution \(u(t,{{\textbf {x}}})\) of Eq. (1); see Roques (2013). Thus \(D_{i\tilde{T}}\) is defined as:
where \(\phi \) is the normal regularization kernel: \(\phi ({{\textbf {x}}})=\dfrac{1}{2\pi \mathcal {V}}\exp \left( {-\dfrac{\Vert {\textbf {x}} \Vert ^2}{2\mathcal {V}}}\right) \), and the transition speed \(\mathcal {V}\) has to be tuned to approach more or less the piecewise constant function \(d_{i\tilde{T}}\). In other words, \(D_{i\tilde{T}}\) locally smoothes the step function \(d_{i\tilde{T}}\) around the transition border between the two subdomains with a bandwidth determined by \(\mathcal {V}\).
The reproduction term may also be either spatially heterogeneous or not. In the homogeneous case, we consider a logistic growth
where b is the intrinsic growth rate of the epidemic and \(K\in (0,1]\) is a plateau for the probability of infection (i.e., an analog to the carrying capacity of the environment). With such a reproduction term, the growth is nearly exponential when u is small, and the growth rate is around zero when u reaches the plateau K, which gives the maximum prevalence of the pathogen in the environment. In the heterogeneous case, the growth is logistic in \(\Omega _{\tilde{T}1}\) (i.e., warmest areas) and negative in \(\Omega _{\tilde{T}2}\) (i.e., coldest areas), mimicking a source-sink dynamics:
where \(\alpha \) is the decrease rate of the infection in \(\Omega _{\tilde{T}2}\). Thus, the pathogen is affected by two contradictory forces in the cold region: it is forced to decline with the rate \(\alpha \) but may be at the same time reinforced by the sources of infection located in the warm region.
In the application, we will consider three model structures:
-
\(\mathcal {M}_0(\tilde{T})\), under which \(D_{i\tilde{T}1}= D_{i\tilde{T}2}\) and \(f_{i\tilde{T}}\) satisfies Eq. (4), i.e. homogeneous diffusion but heterogeneous growth, like in Abboud et al. (2019);
-
\(\mathcal {M}_1(\tilde{T})\), under which \(D_{i\tilde{T}1}\ne D_{i\tilde{T}2}\) and \(f_{i\tilde{T}}\) satisfies Eq. (4), i.e. heterogeneous diffusion and growth;
-
\(\mathcal {M}_2(\tilde{T})\), under which \(D_{i\tilde{T}1}\ne D_{i\tilde{T}2}\) and \(f_{i\tilde{T}}\) satisfies Eq. (3), i.e. heterogeneous diffusion, but homogeneous growth.
The second equation in System (1) corresponds to the homogeneous Neumann condition on the boundary \(\partial \Omega \) of \(\Omega \) (i.e. with reflection on the boundary). This condition is formalized by setting that the gradient of the spatial function \(x\mapsto D_{i\tilde{T}}({{\textbf {x}}}) u(t,{{\textbf {x}}})\) is orthogonal to the outward normal vector n(x) at point x on the boundary \(\partial \Omega \), for all \(t\ge \tau _0\). Thus, physically, there is neither outward nor inward flux from and to \(\Omega \).
The spatial function \(u_0\) models the introduction of the pathogen in the study domain at time \(\tau _0\in {{\mathbb {R}}}\). Following Abboud et al. (2019), the introduction represents the initial phase of the outbreak corresponding to the arrival of the pathogen and its local establishment. Thus, \(u_0\) is not expressed as a Dirac function but as a kernel function centered around the central point of the introduction \(\tilde{{\textbf {x}}}_0=(\tilde{x}_0,\tilde{y}_0)\in \Omega \). More precisely, the probability of a host at \(\textbf{x}\) to be infected at \(\tau _0\) satisfies:
where \(P_0\) is the infection probability at \((\tau _0,\tilde{{\textbf {x}}}_0)\), \(\sigma ^2=\frac{r_0^2}{q}\), q is the 0.95-quantile of the \(\chi ^2\) distribution with two degrees of freedom, and \(r_0\) is the radius of the kernel. Thus, at \(\tau _0\), if we neglect border effects, 95% of the infected plants are located within the ball with center \(\tilde{{\textbf {x}}}_0\) and radius \( r _0\).
With these initial and boundary conditions, the equation system (1) is well-posed (Evans 1998) and, by constraining \(P_0\) in [0, K], the principle of parabolic comparison (Protter, MH and Weinberger, HF 1967) implies that the solution of (1) remains in the interval [0, K].
3.2 Probabilistic Model of the Observation Process
Let \(t_j\in {{\mathbb {R}}}\) denote the sampling time of host \(j\in \{1,\ldots ,J\}\), \(J\in {{\mathbb {N}}}^*\), \({\textbf {x}}_j\in \Omega \) its location and \(Y_j\in \{0,1\}\) its observed sanitary status at \(t_j\) (1 for infected, 0 for healthy). Given u, \(\mathcal {M}_i(\tilde{T})\) and \(\{(t_j,{\textbf {x}}_j):1\le j \le J\}\), the sanitary statuses \(Y_j\), \(j\in \{1,\ldots ,J\}\), are independent random variables following Bernoulli distributions with success probabilities \(u(t_j,{\textbf {x}}_j)\):
where u depends on the model \(\mathcal {M}_i(\tilde{T})\) and its vector of parameters \(\Theta _{i\tilde{T}}\).
Remark. This data model was proposed in Abboud et al. (2019) for its simplicity. It could be refined to account for sampling errors classically encountered in epidemiology, e.g. false-positive and false-negative observations, as well as spatial and temporal dependencies not accounted for in the process model.
4 Bayesian Model Averaging
4.1 Principle
Briefly, the BMA consists in estimating the expectation of the posterior distributions of the variable of interest \(\Delta \) provided under all the competing models and weighted by the posterior model probabilities (Raftery 1996; Hoeting et al. 1999). In the modeling setting introduced above, \(\Delta \) is typically a vector of shared parameters such as the introduction point (\(\tilde{{\textbf {x}}}_0\),\(\tau _0\)), the temperature threshold \(\tilde{T}\) or the spatial probability of infection u over a future period. Using Gelfand’s bracket notation for probability distributions (Gelfand and Smith 1990), the BMA posterior distribution of \(\Delta \) given training data Y satisfies:
The posterior model probability of \(\mathcal {M}_i(\tilde{T})\) is:
The integrated likelihood \([Y|\mathcal {M}_i(\tilde{T})]\) of \(\mathcal {M}_i(\tilde{T})\), which may be a complex integral depending on the dimension of the unknowns and eventual dependencies, satisfies:
where \(\Theta \) is the vector of parameters of \(\mathcal {M}_i(\tilde{T})\), \([Y|\Theta ,\mathcal {M}_i(\tilde{T})]\) is the likelihood under \(\mathcal {M}_i(\tilde{T})\), \([\Theta |\mathcal {M}_i(\tilde{T})]\) is the prior distribution of \(\Theta \) under \(\mathcal {M}_i(\tilde{T})\), and \([\mathcal {M}_i(\tilde{T})]\) is the prior probability of \(\mathcal {M}_i(\tilde{T})\). The posterior mean of \(\Delta \) is a weighted average of the posterior means under the competing models:
The posterior variance is expressed as follows:
4.2 Implementation
The BMA-posterior distribution of \(\Delta \) is computed from the following two-step process: First, we compute the posterior distribution of \(\Delta \) given model \(\mathcal {M}_i(\tilde{T})\) and training data Y for all \(i\in \{1,\ldots ,I\}\); Second, we compute the posterior model probabilities.
4.2.1 Approximation of the Posterior Distribution of \(\Delta \) Given a Model \(\mathcal {M}_i(\tilde{T})\)
For any mechanistic-statistical model defined in Sect. 3, neither the posterior distribution nor the likelihood can be expressed analytically, especially because they depend on the solution u of the PDE that cannot be written in a closed form. Thus, following Abboud et al. (2019), we use the adaptive multiple importance sampling algorithm (AMIS; Cornuet et al. 2012), that consists of successively generating parameter vectors under an adaptive proposal distribution and assigning/updating weights for the parameter vectors. To design efficient importance sampling algorithms, the auxiliary proposal distribution should be chosen as close as possible to the posterior distribution. However, the posterior distribution being unknown, the crucial choice of the proposal is a difficult task (Gelman et al. 1996). The main aim of AMIS is to overcome this difficulty by tuning the coefficients of the proposal distribution picked from a parametric family of distributions, generally the Gaussian one, at the end of each iteration. In this framework, at each iteration, new coefficient values for the proposal distribution are determined using the current weighted posterior sample, then the posterior sample is augmented by generating new replicates from the newly tuned proposal distribution and the weights of the cumulative posterior sample are updated. The AMIS algorithm provides a weighted posterior sample \(\{(\Delta _m^l,w_m^l):1\le m\le M,1\le l\le L\}\) of size ML, which forms an empirical approximation of the posterior distribution \([\Delta |Y,\mathcal {M}_i(\tilde{T})]\) (m stands for the iteration; l stands for the replicate generated at iteration m). Conditions leading to the convergence in probability of the posterior mean of any function (integrable with respect to the posterior distribution) of the parameters are described in Cornuet et al. (2012) and Marin et al. (2019), and are satisfied in our case (Abboud et al. 2019).
We implemented the AMIS algorithm in the R statistical software, with calls to the software Freefem++ for numerically solving the PDE in the mechanistic-statistical environment (MSE; https://informatique-mia.inrae.fr/mse/). We performed parallel computation to compensate for the non-negligible time of PDE resolution. With \((M,L)=(50,10^4)\) (which ensured the stabilization of the posterior samples using the deviation measure described by Abboud et al. 2019) and the use of 100 cluster cores (the cluster being composed of 40-cores nodes Xeon(R) 2.2 Ghz, 228 Go RAM), the estimation procedure for one model takes approximately 1.75 days in average. Unlike the MCMC algorithm that is often used in the mechanistic-statistical framework (Soubeyrand and Roques 2014; Lanzarone et al. 2017), AMIS, as a purely Monte Carlo algorithm, can be easily parallelized, its tuning parameters are automatically adapted at each iteration, and all the samples generated throughout the algorithm are recycled thanks to the update of weights at each iteration. The AMIS algorithm provides at each iteration an assessment of the posterior distribution of parameters, which is expected to be stable after a burn-in period and to converge to the true posterior distribution.
4.2.2 Computation of the Integrated Likelihood
Computing the posterior model probability requires the evaluation of the integrated likelihood as shown in Eq. (8). Various methods to estimate the integrated likelihood (that is not analytically tractable in our case) are encountered in the literature. In this article, we briefly assess several of these methods with respect to their impact on BMA predictions. Nine integrated likelihood estimators, described in Online Resource 1, Section S1, are considered: the un-normalized weight estimator denoted by UWE (Bugallo et al. 2015), the estimator based on an information criterion (McElreath 2018) of which we consider six different specifications denoted by WAIC\(_1\), WAIC\(_2\), BIC, DIC\(_1\), DIC\(_2\) and IC, and the harmonic mean estimator (Newton and Raftery 1994; Gelfand and Dey 1994) of which we consider two specifications denoted by HME\(_1\) and HME\(_2\).
4.2.3 Priors and Posterior Samples
For the applications, we assume as a prior knowledge that the models are equally weighted. Because several model structures with different sets of parameters are considered, the prior distribution of \(\Theta \) partly depends on the model structure. These distributions combine vague uniform and Dirac distributions (Dirac distributions are considered for \(r_0\) and \(p_0\) for identifiability issues) and are provided in Online Resource 1, Section S2. AMIS is then applied to obtain a weighted posterior sample of size \(ML=5\times 10^5\) for each candidate model; see Sect. 4.2.1. Posterior model probabilities, empirical approximations of BMA posterior distributions and other posterior quantities (including predicted infection maps) were approximated by sampling with replacement \(10^4\) models\(\times \)parameters with respect to model and parameter weights.
5 Application to Simulated Data
We perform a simulation study to illustrate the heterogeneity in the predictive performance of the inference approach (by focusing on the posterior model probabilities) when one considers the different methods presented in Sect. 4.2.2 for computing the integrated likelihood.
The simulation study is carried out by generating three different data sets \(\{\mathcal {O}^{(g)}:g=1,2,3\}=\{(t_j,{\textbf {x}}_j,Y_j^{(g)}):1\le j \le J\}\) from two different generative models. The simulations are performed by using characteristics of real data analyzed in the next section: we use the same spatial domain \(\Omega \), observations locations \({\textbf {x}}_j\) and observation times \(t_j\) and, for most of the parameters, we use values close to parameter estimates obtained in the next section. The two models that we consider are \(\mathcal {M}_0(5.5)\), in which the diffusion is homogeneous (\(D_1=D_2\)), and \(\mathcal {M}_1(5.5)\), in which the diffusion is heterogeneous. For the latter model, we consider two cases: \(D_2=0.9D_1\) and \(D_2=0.1D_1\). Table S1 in Online Resource 1 summarizes model specifications and provides parameter values. Figures S1–S3 in Online Resource 1 display the posterior map of the introduction location and the marginal posterior distributions of the other parameters when the true model is fitted to data. We can observe that the true parameter values are all in their respective \(95\%\) credible intervals.
The relevancy of each approach for computing posterior model probabilities is assessed by fitting the two models \(\mathcal {M}_0(5.5)\) and \(\mathcal {M}_1(5.5)\) to the three simulated data sets and by checking, for each data set, whether the true model has the largest posterior probability. Both candidate models \(\mathcal {M}_0(5.5)\) and \(\mathcal {M}_1(5.5)\) have equal prior weights. Posterior model probabilities provided in Table 1 are computed with each of the nine estimators of the integrated likelihood listed in Sect. 4.2.2. Only posterior model probabilities estimated from UWE, HME\(_2\), WAIC\(_1\) and WAIC\(_2\) are consistent in the sense that the true model has the largest posterior probability.
In the real case study presented below, we use the estimator grounded on un-normalized weights (UWE) because it is considered as one of the most efficient approaches for assessing the integrated likelihood, displaying desirable convergence properties (Ford and Gregory 2007), and because, from a practical viewpoint, it is easily obtained from AMIS output.
6 Application to Xylella fastidiosa Data
6.1 Model Comparison
In the analysis of Xylella fastidiosa (Xf) data in South Corsica, we first compare the 27 models under consideration (the three model shapes presented in Sect. 3 times nine temperature thresholds, from 4 \(^\circ \)C to 6 \(^\circ \)C every 0.25 \(^\circ \)C). The models are fitted to training data consisting of all the samples from July 2015 to April 2017 (subsequent data from May 2017 to January 2019 are used for assessing predictive performance of diverse forecast approaches in Sect. 6.3). The stabilization of AMIS assessed as in Abboud et al. (2019) is satisfactory for all models (see Online Resource 1, Figure S4 for an example). Posterior model probabilities are provided in Online Resource 1, Table S2, for the computation based on UWE as well as for the computations based on the other approximation approaches. Regarding the best model, UWE and HME\(_2\) lead to \(\mathcal {M}_0(4.75)\) (homogeneous diffusion but heterogeneous growth with a temperature threshold equal to 4.75 \(^\circ \)C), DIC\(_1\) and IC lead to \(\mathcal {M}_0(5.75)\) and the others lead to \(\mathcal {M}_1(5.75)\) (homogeneous diffusion and growth). Thereafter, we only analyze results obtained with UWE, as explained in Sect. 5. Overall, models with the largest posterior probabilities computed from UWE are \(\mathcal {M}_0\) and \(\mathcal {M}_1\) with temperature thresholds from 4.75 \(^\circ \)C to 5.5 \(^\circ \)C.
6.2 Inference from BMA
BMA posterior distributions of shared parameters (the temperature threshold, the introduction time and the introduction location) are displayed in Figs. 2 and 3. Figure 2 illustrates an advantage of BMA since one obtains a posterior distribution for the temperature threshold instead of the unique value obtained by Abboud et al. (2019) with a model selection approach. The introduction of Xf tends to be relatively ancient but uncertain (posterior mean: 1956; posterior median: 1954; posterior SD: 15 years; 95%-posterior interval: [1933;1988]). The inference of the introduction date provided by the best model is slightly less variable (posterior mean: 1958; posterior median: 1957; posterior SD: 14 years; 95%-posterior interval: [1934;1986]). A larger difference in estimation uncertainty arises for the introduction location, which is more uncertain in the BMA inference than, for instance, in the inference resulting from the model selected by (Abboud et al. 2019, Fig. 6). The BMA estimation possibly better reflects the uncertainty about the introduction location than a single-model estimation.
6.3 Forecast Performance
We assess forecast performance by considering different in-sample predictors trained over the period from July 2015 to April 2017 (8152 observations) and evaluated over the period from May 2017 to January 2019 (1523 observations). We consider five in-sample predictors:
-
Best model: the posterior mean of u computed and averaged over the period May 2017–January 2019, which is provided by the best model selected from and fitted to training data (the best model is the model with the highest posterior model probability given by Eq. (8));
-
BMA: the posterior mean of u computed and averaged over the period May 2017–January 2019, which is provided by BMA applied to training data;
-
Ensemble: the mean of the 27 maximum likelihood estimates (MLE) of u computed and averaged over the period May 2017–January 2019 obtained under the 27 models, where the MLE of u corresponds for each model to the parameter vector simulated in the AMIS algorithm leading to the largest likelihood value;
-
Climatology: the so-called climatology forecast (Mason 2004), which is simply the mean of \(\{Y_j:t_j\in [\text {July 2015--April 2017}]\}\);
-
Kernel smoother: the spatial kernel smoother of \(\{({\textbf {x}}_j,Y_j):t_j\in [\text {July 2015--April} \text {2017}]\}\), using the Epanechnikov kernel, which is proportional to \(d\mapsto (1-d^2)1(|d|\le 1)\), where d is the geographical distance scaled by a bandwidth value ranging from 2.5 to 25 km. We assume that the kernel smoother predicts 0 if no observations are available within the bandwidth.
The in-sample forecasts are computed over a regular square grid with 1 km\(\times \)1 km cells covering \(\Omega \) and are compared to several benchmark out-of-sample forecasts computed over the same grid and offering different visions of the true sanitary situation between May 2017 and January 2019. Two types of out-of-sample forecasts are considered:
-
Local infection proportions: the raw mean of \(Y_j\) observed in each grid cell from May 2017 to January 2019. Sampling over the validation period was made in a limited number of grid cells mostly distributed near the shoreline. In particular, areas with very low infection risk based on earlier observations were not sampled (typically high altitude areas). Hence, to better represent the sanitary situation across space in the validation map, we augmented the set of grid cells where local infection proportions could be computed as raw means of \(Y_j\) with a set of grid cells where local infection proportions were fixed at zero. This set, displayed in Figure S5 in Online Resource 1, was constructed by identifying grid cells far from past detections of Xf and far from observations during the validation period, that is to say grid cells satisfying two conditions: (1) no infection observed in training data up to 10 km; (2) no observation in validation data up to 10 km.
-
Smoothed infection proportions: the spatial kernel smoother of validation data, i.e. \(\{({\textbf {x}}_j,Y_j):t_j\in [\text {May 2017--January 2019}]\}\), computed at each grid cell center. Here also, the smoother is grounded on an Epanechnikov kernel with bandwidth values ranging from 2.5 to 25 km, and it predicts 0 if no observations are available within the bandwidth.
The motivation for considering as benchmarks the smoothed infection proportions with varying bandwidths (in addition to the raw local infection proportions) is that we want to assess the ability of the competing predictors to forecast infection probability at different spatial scales.
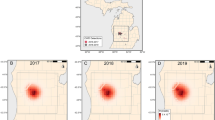
Thus, Fig. 4 shows the benchmark out-of-sample predictions consisting of the local infection proportions and the smoothed infection proportions obtained with the 15 km-kernel bandwidth (top right box), and the five in-sample predictions (as well as posterior standard deviations of u for the best model and the BMA; bottom box). Figures S6 and S7 in Online Resource 1 display the same plots for a kernel bandwidth equal to 5 and 25 km, respectively (only the in- and out-of-sample smoothers are different in these figures). Overall, the best model, the BMA, the ensemble and the kernel smoother provide spatially heterogeneous predictions that may be relevant but a quantitative assessment of the prediction quality is needed to finely compare the predictions (see the paragraph below). Furthermore, like for the estimation of the introduction location, BMA yields a clearly more uncertain inference of the infection probability u across space than the best model.
The root-mean-squared error (RMSE) with respect to each benchmark out-of-sample forecast is computed to measure the predictive performance of the BMA, the best model, the ensemble, the climatology and the kernel smoothing, i.e. to measure how close are the forecasts to the true sanitary situation between May 2017 and January 2019. This quantity is calculated over the regular square grid with 1 km\(\times \)1 km cells already introduced above:
where \(\hat{{\bar{u}}}_h\) is the average (in time and space) prediction of u in grid cell h over May 2017–January 2019 provided by one of the predictors; \({\bar{u}}_h\) is either the local infection proportion in grid cell h over the validation period or the average (in time and space) of u in grid cell h provided by the spatial kernel smoother with bandwidth \(b>0\) applied to validation data (where b ranges from 2.5 to 25 km); H is the number of grid cells.
Figure 5 shows the RMSE values for the five in-sample forecast approaches and a kernel bandwidth ranging from 2.5 to 25 km. Looking only at the comparison with the raw local infection proportions over the validation period (red symbols), the BMA outperforms the other approaches (best model, ensemble, climatology and kernel smoother whatever the bandwidth). The same holds true when one compares the competing in-sample forecasts to the out-of-sample predictions computed as smoothed infection proportions obtained with different bandwidths (black symbols). The climatology, which predicts the same infection probability everywhere, obviously does not account for the major spatially-structured effect of cold temperatures in winter on Xf reproduction or propagation. For small bandwidths, the high probability areas identified by the kernel smoother applied to training data and to validation data are spatially close but do not exactly coincide. In contrast, the quite smooth predictions based on the BMA, the best model and the ensemble do not predict peaks of infection as observed in the out-of-sample forecast but correctly delineate regions where these peaks can arise. When the bandwidth is large, the out-of-sample forecast tends to a very smooth function that even yields significant positive infection probabilities in regions where Xf reproduction or propagation is hampered (i.e. cold regions in winter). This bias incorporated in what we called a vision of the true sanitary situation is also encountered in the in-sample forecast based on kernel smoothing (and to some extent in the climatology that can be considered as a kernel smoothing with very large bandwidth). This bias partly explains the improvement of the prediction ability of the kernel smoothing and the climatology when the bandwidth increases. However, even for large bandwidths, the BMA prediction is better.
7 Discussion
We have presented how to use Bayesian model averaging (BMA) to infer and predict pathogen dynamics from multiple candidate PDE-based models, with an application to Xylella fastidiosa. This approach has a large potential of application in epidemiology where compartmental models based on ordinary or partial differential equations are widespread and often used to forecast epidemics as illustrated during the current COVID-19 pandemic (e.g., see Bertozzi et al. 2020; Kissler et al. 2020; Roques et al. 2020, among many other references). BMA is a valuable approach to compensate for the supposed lack of flexibility of such deterministic models. BMA can also be exploited for model selection, using the posterior model probabilities to identify the best model. However, we have tested nine methods proposed in the literature to compute these probabilities and we obtained rather heterogeneous results (even if only 3 models out of 27 were identified as the most probable models by the nine methods). We chose the computation based on un-normalized weights calculated within the importance sampling algorithm. This natural choice appears to be efficient in our application (it led to a BMA-forecast clearly better than the best-model-forecast and the ensemble-forecast), but further work is required to provide robust advises for computing model weights in BMA.
The model-averaging framework that we describe in this article may be used for a broad range of applications not limited to pathogen invasion dynamics. It may especially be applied to predict more general population dynamics (invasive or not) by adapting (1) the competing PDE-based models to the species dynamics and (2) the observation-process model to the available data. For instance, the model-averaging framework may be adapted to studies in which PDEs were fitted to data concerning the population dynamics of insects (Ovaskainen et al. 2008; Roques et al. 2011), birds (Wikle 2003), terrestrial mammals (Louvrier et al. 2020) and marine mammals (Williams et al. 2018). The framework is not limited to PDE-based phenomenological models: other relatively concise model formalisms may be used, such as those discussed for plant, animal and pathogen dynamics by Pyšek and Hulme (2005), Schurr et al. (2012), Leitner and Kühn (2018) and Roques and Soubeyrand (2023), even if the additional flexibility eventually offered by alternative models might be an issue depending on the information brought by data. If such a case occurs, the model averaging approach may however allow the analyst to reduce estimation difficulties by simplifying a comprehensive model (affected by identifiability problems) into multiple partial models (less prone to identifiability problems), which are fitted to data independently and then combined to draw probabilistic predictions. To illustrate this proposal, consider a situation where one aims to infer the dynamics of a species (typically a plant species) with multiple possible dispersal vectors (e.g., rivers, winds, animals and humans). If fitting a comprehensive model accounting for all the dispersal vectors leads to identifiability problems, one may fit partial models instead (each of them taking into account a single dispersal vector), and combine these models with BMA to draw predictions, to infer the relative contribution of each dispersal vector, and even to test hypotheses about them (Bartoš et al. 2021). Although appealing, this proposal should be tested with caution: considering a single dispersal vector in a model might lead to biased estimation of the effect of this vector, and might require an informative prior to avoid this bias.
Locations of plants (left), sampled from July 2015 to January 2019, that have been detected as positive (green dots) or negative (blue dots) to Xf in South Corsica, France, and map of the average of the daily minimum temperature (right) in Celsius degrees over January and February and a 1\(\times \)1 km-pixel grid (Color figure online)
BMA Posterior distributions of the introduction time \(\tau _0\) (histogram) and the introduction point \(\tilde{{\textbf {x}}}_0\) (color palette). The prior for \(\tau _0\) was uniform over [1931, 2015] (red line). The prior for \(\tilde{{\textbf {x}}}_0\) was uniform over \(\Omega _{\tilde{T}1}\) for model \(\mathcal {M}_{i}(\tilde{T}),\,0\le i\le I\) (Color figure online)
Training data (top left box), out-of-sample predictions computed either as a local infection proportion or as a kernel smoothing with a 15 km-bandwidth (top right box), and in-sample predictions (i.e. posterior means provided by the best model and the BMA, ensemble, climatology and kernel smoothing with a bandwidth of 15 km; bottom box) obtained from training data. Map of standard deviations are also provided in the bottom box for the best model and BMA. The color palette (bottom right) applies to all plots. The range of values taken in each map is provided below the map (Color figure online)
RMSE values showing the performance of the in-sample predictors (BMA; best model; ensemble; climatology; kernel smoothing with varying bandwidth) with respect to each of the out-of-sample predictions (local infection proportions in red; smoothed infection proportions with varying bandwidth in black). Each in-sample kernel smoother obtained with a given bandwidth value is compared both to the local infection proportions (red crosses) and to the smoothed infection proportion using the same bandwidth (black crosses), but not to the smoothed infection proportions using other bandwidth values (Color figure online)
Concerning the emergence and spread of Xylella fastidiosa, we obtained consistent results with previous studies. Thus, our estimation of the introduction date around 1956 [1933; 1988] is consistent with the phylogeny-based estimation (1971 [1924; 1994]) for the sequence type 7 of Xf provided by Dupas et al. (2023) and the estimation of Soubeyrand et al. (2018) based on a stochastic temporal model including a hidden host compartment (1985 [1978;1993]). Dupas et al. (2023) mention “a period of massive introduction of exotic plants, particularly in Corsica", starting around the 70’s, that supports their estimation, referring to Jeanmonod and Natali (1997)—this statement relies on the credible assumption that the introduction of Xf in an isolated disease-free territory can occur via the introduction of infected plants. Jeanmonod and Natali (1997) indeed estimated a rather constant and high rate of introduction of exotic plant taxa in Corsica from 1970 to 1997: around 50 taxa per decade. In contrast, from 1840 to 1970, this rate was estimated to be around 20 taxa per decade, which is clearly lower, but might be sufficient to favor the introduction of Xf in Corsica. A more in-depth analysis of the introduced plant species, with regard to those which are recognized as hosts of Xf, deserves to be carried out to evaluate the different estimates of the date of introduction. By the way, such an analysis could be exploited to propose a more informative prior of the introduction date in our Bayesian approach. The introduction location, which likely took place in the region of Ajaccio (West coast of Corsica), is consistent with the estimation provided by Abboud et al. (2019), but the application of BMA is expected to better reflect the uncertainty about this location. In addition, BMA provides a posterior distribution for the threshold in winter temperature that includes the threshold identified by Abboud et al. (2019) via a model selection procedure.
Based on the ample literature on model averaging and its benefits, we expected improved predictions and more realistic uncertainty estimation (Hoeting et al. 1999; Wintle et al. 2003). Precisely, in the Xf case study, BMA outperforms the best model in terms of prediction, and credibility intervals provided by BMA are larger than those provided by the best model. This observation could reflect the fact that BMA succeeds in better assessing model uncertainty and avoids overconfidence in the estimations and the predictions. However, for firmly confirming this result, complementary studies should be conducted to assess the calibration of credibility intervals.
As illustrated by the analysis of Xf data, the deterministic candidate models summarized by Eqs. (1–5) and describing the dynamics of the pathogen capture, overall, the discrepancies between regions of low- and high-probabilities of infection but do not capture the details, e.g., the spatio-temporal disease clusters that can be observed using kernel smoothing with small bandwidth. These details could be implicitly taken into account by coupling the partial differential equation to stochastic terms or observed covariates that would lead to more flexible realizations of the dynamics. Additional model components and covariates could be identified from the series of work grounded on mechanistic models and machine learning investigating the dynamics of Xylella fastidiosa and its environmental/biological drivers (Cendoya et al. 2020; Godefroid et al. 2019; Kottelenberg et al. 2021; Martinetti and Soubeyrand 2019; Raffini et al. 2020; Soubeyrand et al. 2018; White et al. 2017, 2020). An interesting perspective is also to include the models proposed in some of these references directly in the BMA analysis since BMA is expected to take advantage of different model structures.
Data availability
Xylella fastidiosa surveillance data are provided by the ESV Platform and accessible at https://doi.org/10.15454/RWBIWD (ESV 2022). Average of daily minimum temperature in January and February in Corsica are available at https://doi.org/10.5281/zenodo.6985147 (Soubeyrand and Abboud 2022).
References
Abboud C, Bonnefon O, Parent E, Soubeyrand S (2019) Dating and localizing an invasion from post-introduction data and a coupled reaction-diffusion-absorption model. J Math Biol 79:765–789
Anas O, Harrison UJ, Brannen PM, Sutton TB (2008) The effect of warming winter temperature on the severity of Pierce’s disease in the Appalachian mountains and Piedmont of the southeastern United States. Plant Health Progress 101094:450–459
Bartoš F, Gronau QF, Timmers B, Otte WM, Ly A, Wagenmakers E-J (2021) Bayesian model-averaged meta-analysis in medicine. Stat Med 40:6743–6761
Berliner LM (2003) Physical-statistical modeling in geophysics. J Geophys Res Atmos 108:8776
Bertozzi A, Franco E, Mohler G, Short M, Sledge D (2020) The challenges of modeling and forecasting the spread of COVID-19. Proc Natl Acad Sci 117:16732–16738
Botella C, Bonnet P, Hui C, Joly A, Richardson DM (2022) Dynamic species distribution modeling reveals the pivotal role of human-mediated long-distance dispersal in plant invasion. Biology 11:1293
Boys RJ, Wilkinson DJ, Kirkwood TBL (2008) Bayesian inference for a discretely observed stochastic kinetic model. Stat Comput 18:125–135
Bugallo MF, Martino L, Corander J (2015) Adaptive importance sampling in signal processing. Digit Signal Process 47:36–49
Burnham KP, White GC, Anderson DR (1995) Model selection strategy in the analysis of capture-recapture data. Biometrics 51:888–898
Cendoya M, Martínez-Minaya J, Dalmau V, Ferrer A, Saponari M, Conesa D, López-Quílez A, Vicent A (2020) Spatial Bayesian modeling applied to the surveys of Xylella fastidiosa in Alicante (Spain) and Apulia (Italy). Front Plant Sci 11:1204
Chapman DS, White SM, Hooftman DA, Bullock JM (2015) Inventory and review of quantitative models for spread of plant pests for use in pest risk assessment for the EU territory. EFSA Supporting Publications, 12:EN–795
Cornuet J, Marin J-M, Mira A, Robert CP (2012) Adaptive multiple importance sampling. Scand J Stat 39:798–812
Denancé N, Cesbron S, Briand M, Rieux A, Jacques M-A (2017) Is Xylella fastidiosa really emerging in France? In: Costa J, Koebnik R (eds) 1st Annual Conference of the EuroXanth - COST Action Integrating Science on Xanthomonadaceae for integrated plant disease management in Europe, vol 7. EuroXanth, Portugal
Denancé N, Legendre B, Briand M, Olivier V, Boisseson C, Poliakoff F, Jacques M-A (2017) Several subspecies and sequence types are associated with the emergence of Xylella fastidiosa in natural settings in France. Plant Pathol 66:1054–1064
Dupas E, Durand K, Rieux A, Briand M, Pruvost O, Cunty A, Denancé N, Donnadieu C, Legendre B, Lopez-Roques C, Cesbron S, Ravigné V, Jacques M-A (2023) Suspicions of two bridgehead invasions of xylella fastidiosa subsp. multiplex in France. Commun Biol 6:103
ESV P (2022) Données de surveillance sur végétaux de Xylella fastidiosa
Evans LC (1998) Partial differential equations, volume 19 of Graduate studies in mathematics. Am Math Soc
Faria NR, Rambaut A, Suchard MA, Baele G, Bedford T, Ward MJ, Tatem AJ, Sousa JD, Arinaminpathy N, Pépin J, Posada D, Peeters M, Pybus OG, Lemey P (2014) The early spread and epidemic ignition of HIV-1 in human populations. Science 346:56–61
Fletcher D (2018) Model averaging. Springer, Berlin
Ford EB, Gregory PC (2007) Bayesian model selection and extrasolar planet detection. In: Statistical Challenges in Modern Astronomy IV, ASP Conference Series, vol 371, p 189
Gelfand AE, Dey DK (1994) Bayesian model choice: asymptotics and exact calculations. J R Stat Soc B 56:501–514
Gelfand AE, Smith AFM (1990) Sampling-based approaches to calculating marginal densities. J Am Stat Assoc 85:398–409
Gelman A, Roberts GO, Gilks WR et al (1996) Efficient metropolis jumping rules. Bayesian Stat 5:599–608
George EI, McCulloch RE (1993) Variable selection via Gibbs sampling. J Am Stat Assoc 88:881–889
Godefroid M, Cruaud A, Streito J-C, Rasplus J-Y, Rossi J-P (2019) Xylella fastidiosa: climate suitability of European continent. Sci Rep 9:8844
Hoeting JA, Madigan D, Raftery AE, Volinsky CT (1999) Bayesian model-averaging: a tutorial. Stat Sci 14:382–417
Huld TA, Šúri M, Dunlop ED, Micale F (2006) Estimating average daytime and daily temperature profiles within Europe. Environ Modell Softw 21:1650–1661
Jeanmonod D, Natali A (1997) Les xénophytes de Corse: un danger pour la flore indigène. Lagascalia 19:783–792
Jones DR, Baker RHA (2004) Introductions of non-native plant pathogens into Great Britain, 1970–2004. Plant Pathol 56:891–910
Kissler SM, Tedijanto C, Goldstein E, Grad YH, Lipsitch M (2020) Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science 368:860–868
Kottelenberg D, Hemerik L, Saponari M, Van Der Werf W (2021) Shape and rate of movement of the invasion front of Xylella fastidiosa spp. pauca in Puglia. Sci Rep 11:1–14
Lamon EC, Clyde MA (2000) Accounting for model uncertainty in prediction of chlorophyll a in lake okeechobee. J Agric Biol Environ Stat 5:297–322
Lanzarone E, Pasquali S, Gilioli G, Marchesini E (2017) A Bayesian estimation approach for the mortality in a stage-structured demographic model. J Math Biol 75:759–779
Leamer E (1978) Specification searches: Ad hoc inference with nonexperimental data. vol 53. Wiley
Leitner M, Kühn I (2018) Dispersal in plants and animals. In: Bunde A, Caro J, Kärger J, Vogl G (eds) Diffusive spreading in nature, technology and society. Springer International Publishing, Cham, pp 29–47
Louvrier J, Papaix J, Duchamp C, Gimenez O (2020) A mechanistic-statistical species distribution model to explain and forecast wolf (Canis lupus) colonization in South-Eastern France. Spatial Stat 36:100428
Madigan D, Gavrin J, Raftery AE (1995) Enhancing the predictive performance of Bayesian graphical models. Commun Stat Theory Methods 24:2271–2292
Madigan D, Raftery AE (1994) Model selection and accounting for model uncertainty in graphical models using Occam’s window. J Am Stat Assoc 89:1535–1546
Marin J-M, Pudlo P, Sedki M (2019) Consistency of adaptive importance sampling and recycling schemes. Bernoulli 25:1977–1998
Martinetti D, Soubeyrand S (2019) Identifying lookouts for epidemio-surveillance: application to the emergence of Xylella fastidiosa in France. Phytopathology 109:265–276
Mason SJ (2004) On using “climatology’’ as a reference strategy in the Brier and ranked probability skill scores. Mon Weather Rev 132:1891–1895
McElreath R (2018) Overfitting, regularization, and information criteria. In: Statistical rethinking: a Bayesian course with examples in R and Stan. Chapman and Hall/CRC
Newton MA, Raftery AE (1994) Approximate Bayesian inference with the weighted likelihood bootstrap. J R Stat Soc B 56:3–26
Oehler VG, Yeung KY, Choi YE, Bumgarner RE, Raftery AE, Radich JP (2009) The derivation of diagnostic markers of chronic myeloid leukemia progression from microarray data. Blood 114:3292–3298
Okubo A, Levin S (2002) Diffusion and ecological problems: modern perspectives. Springer Science & Business Media, New York
Ovaskainen O, Rekola H, Meyke E, Arjas E (2008) Bayesian methods for analyzing movements in heterogeneous landscapes from mark-recapture data. Ecology 89:542–554
Parkinson D, Liddle AR (2013) Bayesian model averaging in astrophysics: a review. Stat Anal Data Min ASA Data Sci J 6:3–14
Peterson RO, Vucetich JA, Page RE, Chouinard A et al (2003) Temporal and spatial aspects of predator-prey dynamics. Alces 39:215–232
Protter MH, Weinberger HF (1967) Maximum principles in differential equations. Prentice-Hall, Englewood Cliffs, New Jersey
Pyšek P, Hulme PE (2005) Spatio-temporal dynamics of plant invasions: linking pattern to process. Ecoscience 12:302–315
Raffini F, Bertorelle G, Biello R, D’Urso G, Russo D, Bosso L (2020) From nucleotides to satellite imagery: approaches to identify and manage the invasive pathogen Xylella fastidiosa and its insect vectors in Europe. Sustainability 12:4508
Raftery AE (1996) Approximate Bayes factors and accounting for model uncertainty in generalised linear models. Biometrika 83:251–266
Raftery AE, Gneiting T, Balabdaoui F, Polakowski M (2005) Using Bayesian model averaging to calibrate forecast ensembles. Mon Weather Rev 133:1155–1174
Roques L (2013) Modèles de réaction-diffusion pour l’écologie spatiale. Editions Quae, Paris
Roques L, Bonnefon O, Baudrot V, Soubeyrand S, Berestycki H (2020) A parsimonious approach for spatial transmission and heterogeneity in the COVID-19 propagation. R Soc Open Sci 7:201382
Roques L, Soubeyrand S (2023) Les invasions biologiques à la lumière des modèles. In: Lannou C, Rasplus J-Y, Soubeyrand S, Gautier M, Rossi J-P (eds) Crises sanitaires en agriculture: Les espèces invasives sous surveillance. Editions Quae, Versailles, pp 167–185
Roques L, Soubeyrand S, Rousselet J (2011) A statistical-reaction-diffusion approach for analyzing expansion processes. J Theor Biol 274:43–51
Roques L, Walker E, Franck P, Soubeyrand S, Klein E (2016) Using genetic data to estimate diffusion rates in heterogeneous landscapes. J Math Biol 73:397–422
Rubin DB, Schenker N (1986) Efficiently simulating the coverage properties of interval estimates. J R Stat Soc C 35:159–167
Schurr FM, Pagel J, Cabral JS, Groeneveld J, Bykova O, O’Hara RB, Hartig F, Kissling WD, Linder HP, Midgley GF, Schröder B, Singer A, Zimmermann NE (2012) How to understand species’ niches and range dynamics: a demographic research agenda for biogeography. J Biogeogr 39:2146–2162
Shigesada N, Kawasaki K, Takeda Y (1995) Modeling stratified diffusion in biological invasions. Am Nat 146:229–251
Sidman AH, Mak M, Lebo MJ (2008) Forecasting non-incumbent presidential elections: Lessons learned from the 2000 election. Int J Forecast 24:237–258
Skellam JG (1951) Random dispersal in theoretical populations. Biometrika 38:196–218
Soubeyrand S, Abboud C (2022). Data set: average daily minimum temperature in January and February in Corsica
Soubeyrand S, de Jerphanion P, Martin O, Saussac M, Manceau C, Hendrikx P, Lannou C (2018) What dynamics underly temporal observations? Application to the emergence of Xylella fastidiosa in France: probably not a recent story. New Phytol 219:824–836
Soubeyrand S, Roques L (2014) Parameter estimation for reaction-diffusion models of biological invasions. Popul Ecol 56:427–434
Turchin P (1998) Quantitative analysis of movement: measuring and modeling population redistribution in plants and animals. Sinauer, Sunderland, Massachusetts
Viallefont V, Raftery AE, Richardson S (2001) Variable selection and bayesian model averaging in case-control studies. Stat Med 20:3215–3230
White SM, Bullock JM, Hooftman DAP, Chapman DS (2017) Modelling the spread and control of Xylella fastidiosa in the early stages of invasion in Apulia, Italy. Biol Invasions 19:1825–1837
White SM, Navas-Cortés JA, Bullock JM, Boscia D, Chapman DS (2020) Estimating the epidemiology of emerging Xylella fastidiosa outbreaks in olives. Plant Pathol 69:1403–1413
Wikle CK (2003) Hierarchical Bayesian models for predicting the spread of ecological processes. Ecology 84:1382–1394
Williams PJ, Hooten MB, Womble JN, Esslinger GG, Bower MR (2018) Monitoring dynamic spatio-temporal ecological processes optimally. Ecology 99:524–535
Wintle BA, McCarthy MA, Volinsky CT, Kavanagh RP (2003) The use of Bayesian model averaging to better represent uncertainty in ecological models. Conserv Biol 17:1579–1590
Yeung KY, Bumgarner RE, Raftery AE (2005) Bayesian model averaging: development of an improved multi-class, gene selection and classification tool for microarray data. Bioinformatics 21:2394–2402
Acknowledgements
This research was funded by a PhD grant INRAE-Région PACA (Emplois Jeunes Doctorants 2016-2019), the HORIZON XF-ACTORS Project (grant SFS-09-2016), the HORIZON BeXyl Project (grant 101060593) and the ANR BEYOND Project (grant 20-PCPA-0002). We thank DGAL, Anses, SRAL, FREDON, LNR-LSV, certified laboratories and the ESV Platform for data collection, centralization and sharing. We thank Afidol for their endorsement in the PhD grant.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that the research was conducted in the absence of any financial or non-financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abboud, C., Parent, E., Bonnefon, O. et al. Forecasting Pathogen Dynamics with Bayesian Model-Averaging: Application to Xylella fastidiosa. Bull Math Biol 85, 67 (2023). https://doi.org/10.1007/s11538-023-01169-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11538-023-01169-w