Abstract
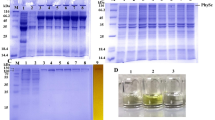
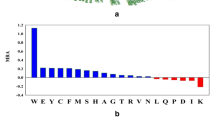
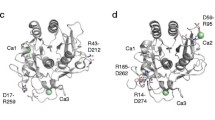
Protein thermostability is an inherent characteristic of proteins from thermophilic microorganisms, and therefore enables these organisms to survive at extreme temperatures. Although it is well-known that thermostable proteins are critical for the growth of thermophilic organisms, the structural basis of protein thermostability is not yet fully understood. The histidine-containing phosphocarrier (HPr) protein, a phosphate shuttle protein in the phosphoenolpyruvate-dependent sugar transport system (PTS) of bacterial species, is an ideal model for investigating protein thermostability with respect to its small size and deficiency in disulphide bonds or cofactors. In this study, the HPr protein from Thermoanaerobacter tengcongensis (TtHPr) is cloned and purified. Crystal structure with good quality has been determined at 2.3 Å resolution, which provides a firm foundation for exploring the thermostable mechanism. However, it shows that the crystal structure is conserved and no clue can be obtained from this single structure. Furthermore, detailed comparison of sequence and structure with the homologs from meso- or thermophilic bacteria shows no obvious rule for thermostability, but the extra salt-bridge existing only in thermophilic bacteria might be a better explanation for thermostability of HPr. Thus, mutations are performed to interrupt the salt-bridge in HPrs in thermophilic bacteria. Using site-directed mutations and the circular dichroism method, thermostability is evaluated, and the mutational variations are shown to have a faster denaturing rate than for wild-type viruses, indicating that mutations cause instability in the HPrs. Understanding the higher-temperature resistance of thermophilic and hyperthermophilic proteins is essential to studies on protein folding and stability, and is critical in engineering efficient enzymes that can work at a high temperature.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Xue Y, Xu Y, Liu Y, et al. Thermoanaerobacter tengcongensis sp. nov., a novel anaerobic, saccharolytic, thermophilic bacterium isolated from a hot spring in Tengcong, China. Int J Syst Evol Microbiol, 2001, 51: 1335–1341 11491330, 1:CAS:528:DC%2BD3MXmtVSmurs%3D
Bao Q, Tian Y, Li W, et al. A complete sequence of the T. tengcongensis genome. Genome Res, 2002, 12: 689–700 11997336, 10.1101/gr.219302, 1:CAS:528:DC%2BD38XjvFSgt7g%3D
Szilagyi A, Zavodszky P. Structural differences between mesophilic, moderately thermophilic and extremely thermophilic protein subunits: results of a comprehensive survey. Structure (London, England: 1993), 2000, 8: 493–504 10801491, 10.1016/S0969-2126(00)00133-7, 1:CAS:528:DC%2BD3cXjsleqtb4%3D
Knapp S, de Vos WM, Rice D, et al. Crystal structure of glutamate dehydrogenase from the hyperthermophilic eubacterium Thermotoga maritima at 3.0 Å resolution. J Mol Biol, 1997, 267: 916–932 9135121, 10.1006/jmbi.1996.0900, 1:CAS:528:DyaK2sXivFWks7w%3D
Karshikoff A, Ladenstein R. Ion pairs and the thermotolerance of proteins from hyperthermophiles: a “traffic rule” for hot roads. Trends Biochem Sci, 2001, 26: 550–556 11551792, 10.1016/S0968-0004(01)01918-1, 1:CAS:528:DC%2BD3MXms1OmtL8%3D
Russell R J, Ferguson J M, Hough D W, et al. The crystal structure of citrate synthase from the hyperthermophilic archaeon Pyrococcus furiosus at 1.9 Å resolution. Biochemistry, 1997, 36: 9983–9994 9254593, 10.1021/bi9705321, 1:CAS:528:DyaK2sXkvVSqsb8%3D
Sterner R, Liebl W. Thermophilic adaptation of proteins. Crit Rev Biochem Mol Biol, 2001, 36: 39–106 11256505, 10.1080/20014091074174, 1:CAS:528:DC%2BD3MXit1Cjur8%3D
Vogt G, Woell S, Argos P. Protein thermal stability, hydrogen bonds, and ion pairs. J Mol Biol, 1997, 269: 631–643 9217266, 10.1006/jmbi.1997.1042, 1:CAS:528:DyaK2sXktlejtbw%3D
Suvd D, Fujimoto Z, Takase K, et al. Crystal structure of Bacillus stearothermophilus alpha-amylase: possible factors determining the thermostability. J Biochem (Tokyo), 2001, 129: 461–468 1:CAS:528:DC%2BD3MXjtlSisr0%3D
Eijsink V G, Vriend G, Van der Zee J R, et al. Increasing the thermostability of the neutral proteinase of Bacillus stearothermophilus by improvement of internal hydrogen-bonding. Biochem J, 1992, 285: 625–628 1637352, 1:CAS:528:DyaK38XltVOhtrc%3D
Tanner J J, Hecht R M, Krause K L. Determinants of enzyme thermostability observed in the molecular structure of Thermus aquaticus D-glyceraldehyde-3-phosphate dehydrogenase at 25 angstroms resolution. Biochemistry, 1996, 35: 2597–2609 8611563, 10.1021/bi951988q, 1:CAS:528:DyaK28Xosl2jtQ%3D%3D
Matsui I, Harata K. Implication for buried polar contacts and ion pairs in hyperthermostable enzymes. FEBS J, 2007, 274: 4012–4022 17683331, 10.1111/j.1742-4658.2007.05956.x, 1:CAS:528:DC%2BD2sXhtVSmt7rO
Bae E, Phillips G N Jr. Structures and analysis of highly homologous psychrophilic, mesophilic, and thermophilic adenylate kinases. J Biol Chem, 2004, 279: 28202–28208 15100224, 10.1074/jbc.M401865200, 1:CAS:528:DC%2BD2cXlt1Cltbw%3D
Del Vecchio P, Elias M, Merone L, et al. Structural determinants of the high thermal stability of SsoPox from the hyperthermophilic archaeon Sulfolobus solfataricus. Extremophiles, 2009, 13: 461–470 19247785, 10.1007/s00792-009-0231-9, 1:CAS:528:DC%2BD1MXls1SltbY%3D
Zhou X X, Wang Y B, Pan Y J, et al. Differences in amino acids composition and coupling patterns between mesophilic and thermophilic proteins. Amino Acids, 2008, 34: 25–33 17710363, 10.1007/s00726-007-0589-x, 1:CAS:528:DC%2BD1cXlt1Clsw%3D%3D
Postma P W, Keizer H G, Koolwijk P. Transport of trehalose in Salmonella typhimurium. J Bacteriol, 1986, 168: 1107–1111 3023298, 1:CAS:528:DyaL2sXivFykug%3D%3D
Cornilescu G, Lee B R, Cornilescu C C, et al. Solution structure of the phosphoryl transfer complex between the cytoplasmic A domain of the mannitol transporter IIMannitol and HPr of the Escherichia coli phosphotransferase system. J Biol Chem, 2002, 277: 42289–42298 12202490, 10.1074/jbc.M207314200, 1:CAS:528:DC%2BD38XotF2gur0%3D
Weng Q P, Elder J, Jacobson G R. Site-specific mutagenesis of residues in the Escherichia coli mannitol permease that have been suggested to be important for its phosphorylation and chemoreception functions. J Biol Chem, 1992, 267: 19529–19535 1527071, 1:CAS:528:DyaK38Xlt1KksbY%3D
Postma P W, Lengeler J W, Jacobson G R. Phosphoenolpyruvate: carbohydrate phosphotransferase systems of bacteria. Microbiol Rev, 1993, 57: 543–594 8246840, 1:CAS:528:DyaK3sXmsVeqtbk%3D
Azuaga A I, Neira J L, van Nuland N A. HPr as a model protein in structure, interaction, folding and stability studies. Protein Pept Lett, 2005, 12: 123–137 15723638, 10.2174/0929866053005818, 1:CAS:528:DC%2BD2MXhvVCitr8%3D
Jia Z, Quail J W, Waygood E B, et al. The 2.0-Å resolution structure of Escherichia coli histidine-containing phosphocarrier protein HPr. A redetermination. J Biol Chem, 1993, 268: 22490–22501 8226757, 1:CAS:528:DyaK3sXlslKitb8%3D
Herzberg O, Reddy P, Sutrina S, et al. Structure of the histidine-containing phosphocarrier protein HPr from Bacillus subtilis 2.0-Å resolution. Proc Natl Acad Sci USA, 1992, 89: 2499–2503 1549615, 10.1073/pnas.89.6.2499, 1:CAS:528:DyaK38XltVOitbo%3D
Jia Z, Vandonselaar M, Hengstenberg W, et al. The 1.6 Å structure of histidine-containing phosphotransfer protein HPr from Streptococcus faecalis. J Mol Biol, 1994, 236: 1341–1355 8126724, 10.1016/0022-2836(94)90062-0, 1:CAS:528:DyaK2cXksFamsL8%3D
Pieper U, Kapadia G, Zhu P P, et al. Structural evidence for the evolutionary divergence of Mycoplasma from gram-positive bacteria—the histidine-containing phosphocarrier protein. Structure, 1995, 3: 781–790 7582895, 10.1016/S0969-2126(01)00213-1, 1:CAS:528:DyaK2MXns1Kktrs%3D
Wittekind M, Rajagopal P, Branchini B R, et al. Solution structure of the phosphocarrier protein HPr from Bacillus subtilis by two-dimensional NMR spectroscopy. Protein Sci, 1992, 1: 1363–1376 1303754, 10.1002/pro.5560011016, 1:CAS:528:DyaK3sXjsVKrug%3D%3D
Klevit R E, Waygood E B. Two-dimensional 1H NMR studies of histidine-containing protein from Escherichia coli. 3. Secondary and tertiary structure as determined by NMR. Biochemistry, 1986, 25: 7774–7781 3542036, 10.1021/bi00371a073, 1:CAS:528:DyaL28XmtVCqt7s%3D
Yin J, Liu Y, Li J, et al. Expression, crystallization and characterization of a novel 6-phospho-β-glucosidase from Thermoanaerobacter tengcongensis MB4. Chin J Biochem Mol Biol, 2008, 24: 916–924 1:CAS:528:DC%2BD1cXhtlOnsL3M
Otwinowski Z, Minor W, Carter C W, et al. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol, 1997, 276: 307–326 10.1016/S0076-6879(97)76066-X, 1:CAS:528:DyaK2sXivFehsbw%3D
Vagin A, Teplyakov A. MOLREP: an automated program for molecular replacement. J Appl Crystallogr, 1997, 30: 1022–1025 10.1107/S0021889897006766, 1:CAS:528:DyaK1cXhslehu7Y%3D
Murshudov G N, Vagin A A, Dodson E J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Cryst D, 1997, 53: 240–255 10.1107/S0907444996012255, 1:STN:280:DC%2BD2czpsFegsw%3D%3D
Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta Cryst D, 2004, 60: 2126–2132 10.1107/S0907444904019158
Adams P D, Grosse-Kunstleve R W, Hung L W, et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D Biol Crystallogr, 2002, 58: 1948–1954 12393927, 10.1107/S0907444902016657
Laskowski R A, MacArthur M W, Moss D S, et al. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr, 1993, 26: 283–291 10.1107/S0021889892009944, 1:CAS:528:DyaK3sXit12lurY%3D
Barlow D, Thornton J. Ion-pairs in proteins. J Mol Biol, 1983, 168: 867–885 6887253, 10.1016/S0022-2836(83)80079-5, 1:CAS:528:DyaL3sXltlGrsL0%3D
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Cryst D, 1994, 50: 760–763
Vriend G. WHAT IF: a molecular modeling and drug design program. J Mol Graph, 1990, 8: 52–56 2268628, 10.1016/0263-7855(90)80070-V, 1:CAS:528:DyaK3cXitFGksL8%3D
DeLano W L. The PyMOL molecular graphics system. Palo Alto: DeLano Scientific. 2002
Razvi A, Scholtz J M. A thermodynamic comparison of HPr proteins from extremophilic organisms. Biochemistry, 2006, 45: 4084–4092 16566582, 10.1021/bi060038+, 1:CAS:528:DC%2BD28Xit1eltb8%3D
Ramachandran G N, Sasisekharan V. Conformation of polypeptides and proteins. Adv Protein Chem, 1968, 23: 283–438 4882249, 10.1016/S0065-3233(08)60402-7, 1:CAS:528:DyaE3cXhtFOmsL8%3D
Homeyer N, Essigke T, Ullmann G M, et al. Effects of histidine protonation and phosphorylation on histidine-containing phosphocarrier protein structure, dynamics, and physicochemical properties. Biochemistry, 2007, 46: 12314–12326 17918862, 10.1021/bi701228g, 1:CAS:528:DC%2BD2sXhtFWktr%2FJ
Li W F, Zhou X X, Lu P. Structural features of thermozymes. Biotechnol Adv, 2005, 23: 271–281 15848038, 10.1016/j.biotechadv.2005.01.002, 1:CAS:528:DC%2BD2MXjsVehu7Y%3D
Ben M S, Aghajari N, Ben A M, et al. Enhancement of the thermostability of the maltogenic amylase MAUS149 by Gly312Ala and Lys436Arg substitutions. Bioresour Technol, 2011, 102: 1740–1746 10.1016/j.biortech.2010.08.082
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Feng, C., Gao, F., Liu, Y. et al. Crystal structure of histidine-containing phosphocarrier protein from Thermoanaerobacter tengcongensis MB4 and the implications for thermostability. Sci. China Life Sci. 54, 513–519 (2011). https://doi.org/10.1007/s11427-011-4182-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-011-4182-x