Abstract
Purpose
The interplay between microbial taxonomy, functional diversity, and land-use changes across diverse ecosystems must be better understood. This study explores, through a statistical modeling approach, the intricate relationships between microbial communities, soil properties (nutrient contents and physicochemical parameters), and land-use changes, using stable isotopes ratio (δ13C/δ15N) as ecosystem function indicator.
Methods
Using Structural Equation Modeling (SEM) this study aims to investigate how soil microbial communities, soil properties and ecosystem function are interconnected in three ecosystems undergoing land-use change in Nevado de Toluca, Mexico.
Results
Multivariate analyses of microbial communities and soil parameters for the different land-uses reveal significant variation in both microbial composition and soil properties (i.e. nutrient contents) mainly explained by the land-use history of the studied ecosystems. Through the SEM approach, it was possible to disentangle the relative contribution of land-use change, microbial community composition and nutrient contents to the ecosystem function defined as the δ13C/δ15N ratio.
Conclusions
Microbial communities are the main drivers of soil ecosystem functioning. However, through SEM approach it is possible to formally test direct and indirect relationships impacting microbial communities and the resulting functional consequences, contributing to a mechanistic understanding of the ecological implications of land-use change, aiding in sustainable land-use decision-making.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Microbial communities, as both the architects and operators of ecosystems, play a fundamental role in the mineralization of plant residues, thereby significantly influencing the cycling and storage of carbon and nitrogen (Wang et al. 2020). In recent years, the interplay between microbial taxonomy and functional diversity has been recognized as a pivotal factor steering soil nutrient cycling across various ecosystems, such as crop fields, grasslands, and forests (He et al. 2023; Singavarapu et al. 2023). Nevertheless, the nuanced responses of microbial community structure and function to the complex mosaics of land-use changes across varying temporal scales remain elusive (George et al. 2019). In this context, the utilization of stable isotopes in soil ecology emerges as a powerful instrument for exploring the historical activities of microbial communities. Stable isotopes, such as 13C and 15N, serve as robust indicators of previous environmental changing conditions in soil over time, through intricate biogeochemical processes (Craine et al. 2015; Zhang et al. 2018; Choi et al. 2023; Park et al. 2023). Changes in land-use from natural ecosystems to agriculture or forestry, can notably impact isotopic composition (Gerschlauer et al. 2019; Ngaba et al. 2019). For instance, the replacement of native C3 plants with C4 crops and shifts in organic matter inputs – in terms of quantity, quality, and composition – can exert direct effects on the microbial communities engaged in organic matter decomposition, altering δ13C values (Diels et al. 2004; Wynn and Bird 2008). In addition, altered nutrient inputs from fertilization practices and changes in primary productivity can shift δ15N values (Craine et al. 2015; Park et al. 2023). Thus, in principle, δ13C/δ15N ratio as isotopic tapestry of any ecosystem can be taken as an indicator of its historical functioning, offering invaluable insights into past alterations in vegetation, shifts in carbon and nitrogen sources, as well as, the microbial activity dynamics (Choi et al. 2023; Tang et al. 2023).
Microbial communities are pivotal not only in nutrient cycling and organic matter decomposition but also in the restoration of ecological interactions (Singh et al. 2019; Hu et al. 2022). However, understanding how these communities and ecosystem functioning recover after land-use change is complex and influenced by multiple factors such as severity and duration of the land-use alteration, ecosystem type, and microbial communities adaptability (Tosi et al. 2016; McKinley 2019; Pérez-Hernández and Gavilán 2021). These relationships involve ecological interactions and feedback mechanisms. In this context, Structural Equation Modeling (SEM) is a powerful tool to understand the complex processes and mechanisms such as trophic interactions (Shao et al. 2015; Jiang et al. 2020), community dynamics (Luan et al. 2020; Ma et al. 2022), and ecosystem stability (Chen et al. 2021; Xu et al. 2021; Zhang et al. 2021a, b) based in causal-effect relationships (Heck and Thomas 2020). SEM offers a valuable approach for testing multivariate hypotheses that integrate multiple variables and relationships in a quantitative and visual manner (Shao et al. 2015; Hair et al. 2021). Also, SEM allows evaluating different models and incorporate empirical data from diverse sources into a unified framework that considers direct and indirect cause-effect relationships, including measurement error, typically overlooked in traditional linear models, (Saborío‐Montero et al. 2020; Lai and Hsiao 2022;). Moreover, SEM enables the modeling of composite variables as latent variables, which are not individually quantified but are inferred from multiple observed or measured variables, allowing for the identification of the most plausible and parsimonious explanation for the observed patterns (Fan et al. 2016).
In this study, employing a SEM approach, we identified the relative significance of the microbial community composition, environmental variables, and metabolic diversity focused on the δ13C/δ15N isotopic signature as indicator of ecosystem functioning along an altitudinal gradient of land use conversion in Nevado de Toluca, Mexico. The bidirectional relationship between microbial composition and nutrient content has been extensively documented (Jing et al. 2020; Jiao et al. 2021; Kang et al. 2021; Philippot et al. 2023). Moreover, the relationship between soil microbial composition and environmental variables (including management practices) in different functional outcomes has been investigated in previous research (Sun et al. 2020; Liu et al. 2022; Xue et al. 2022). In addition, previous studies have shown that soil microbial composition and nutrient dynamics are also influenced by some environmental variables such as pH, soil organic matter (SOM) quality and quantity, nutrient content, etc. (Lauber et al. 2008; Trivedi et al. 2016; Fierer 2017; Philippot et al. 2023). Finally, it has been documented that land-use change negatively influences biodiversity and nutrient content, (Felipe-Lucia et al. 2020; Hasan et al. 2020). Based on these findings, we hypothesized the reciprocal (bidirectional) interaction between soil properties and microbial communities, and microbial communities acting as mediators of the effects of land-use change in soil functional response (δ13C/δ15N). Testing these hypothetical relationships is crucial for understanding the consequences of land-use changes in the underlying mechanisms of functional ecosystem processes and provide valuable insights for a sustainable land-management decision making.
2 Materials and methods
2.1 Experimental design and study sites
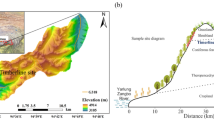
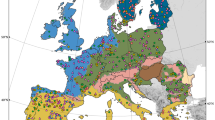
The study was conducted at Nevado de Toluca, Central Mexico (Fig. 1), which is the fourth highest formation in Mexico reaching 4645 m above sea level (masl). The climate in the area is characterized by semicol-subhumid C(E)wig and cold E(T)Hwig conditions (García 1990). The main ecosystems observed are temperate forests, which include species such as Abies, Pinus and Quercus, situated between 3000 to 4100 masl. Additionally, there are high mountain grasslands consisting of Festuca and Calamagrostis, found at altitudes ranging from 4100 to 4500 masl (Calderón de Rzedowski and Rzedowski 2005).
To statistically evaluate the relationship between land-use change, soil properties (nutrient contents and physicochemical parameters), microbial diversity, and their influence on stable isotopes measures as ecosystem functioning indicators, soil samples were collected from six sites with different land-use at different altitudinal floors (3200, 3400 and 4000 masl). The corresponding pairs of sites with contrasting land-use were: grassland-burned grassland, pine-reforested pine and fir-agriculture. As contextual background for the sites, we include the C and N stable isotopes of the dominant vegetation (González Contreras 2019) in each ecosystem studied (Table 1).
2.2 Sampling
At each site, a 30 × 30 m plot was established for sampling. Specifically, three equidistant rows (10 m apart) were drawn, and three soil cores (7.5 cm in diameter and 10 cm deep) were taken from three equidistant points (10 m apart) in each row. For the physicochemical and microbial diversity analyses, samples taken at each point were combined into one composite sample per row for a total of three composite samples per site (N = 18). Samples for microbial diversity analyses were added to 50 mL centrifuge tubes containing RNAlater™ (SIGMA) (1:1), RNAlater™ was removed before sample storage at -80 ºC until DNA extraction. Samples for the physicochemical analyses were stored in high density black plastic bags at room temperature (RT) until processed. Samples for C and N isotopic measurements were taken individually (N = 36) and stored at RT.
2.3 Physicochemical analyses
Soil samples were sieved to 2 mm and air-dried for physicochemical characterization. Soil organic carbon (SOC) was analyzed using a ground subsample of 5 g of soil that passed through a 100-mesh screen by dry digestion at 900 ºC in an automated C- analyzer (Shimadzu 5050A, Canada), concentration of total soil N (TN) was determined by acid digestion in H2SO4 with the Kjeldahl method (Bremner 1960), phosphorus content (TP) was determined using the Olsen Method (Sims 2000). Ca, Mg and Fe concentrations were determined by atomic absorption, K and Na by atomic emission. Particle size distribution (clay, silt, and sand) was analyzed using the modified hydrometer method (Gee and Bauder 1986). The pH was determined in water (1:2 w/v). Analysis of variance (ANOVA) and Tukey’s HSD mean comparisons test at P < 0.05 were used to show differences of the soil physicochemical parameters among different land-use ecosystems.
2.4 DNA extraction and 16S rRNA sequencing
Genomic DNA from each sample was extracted using DNAeasy® PowerSoil® Kit (Qiagen, Valencia CA, USA) according to manufacturer instructions with the following modifications: 0.5 g of sample instead of 0.25 g was used and incubation periods at 4 °C were increased from 5 to 20 min. Genomic DNA samples were submitted to Macrogen laboratory for 16S rRNA gene sequencing of V3-V4 region (341F/805R primer set) (Herlemann et al. 2011) using an Illumina MiSeq instrument.
2.5 16S rRNA sequence data processing
Illumina raw sequences (7,720,761) were processed, and quality filtered using Quantitative Insights into Microbial Ecology (QIIME 2). Chimera and amplicon sequence variants (ASVs) identification were performed using the DADA2 algorithm. Because of quality, only forward sequences were trimmed by 30 base pairs (bp) and truncated at 220 bp during the ASV clustering in DADA2. The filtered sequences (5,489,149) were assigned using SILVA database (v138.1). For the compositional nature of the metagenomic data, a center log ratio (CLR) normalization was performed with the Microbiome Analyst web-based tool (https://www.microbiomeanalyst.ca). The raw data were deposited in the NCBI sequence read archive (SRA) under the accession number PRJNA1037141.
2.6 Characterization C and N stable isotopes
Soil samples for carbon (δ13C) and nitrogen (δ15N) stable isotope analyses were collected in the southwest corner of each plot: 0–15 and 15–30 cm-deep soil cores were collected with a 3.2 cm-diameter auger (Dorme Soil Samplers; Australia) and placed in sealed plastic bags and transported to the laboratory where they were kept refrigerated at 4 ºC until they were processed. The processed samples were sent to the Environmental Isotope Laboratory, University of Arizona, USA for dC13 and dN15 detection with a continuous flow mass spectrometer (Finnigan Delta PlusXL; USA) coupled to an elemental analyzer (Costech, Analytical Technologies Inc; USA) using acetanilide as standard for C and N. The values of δ13C and δ15N were expressed in parts per thousand (‰) according to the international standard (δ13C, Vienna-PeeDee Belemnite; δ15N, atmospheric N2) in the conventional δ-notation, where the X refers to 13C or 15N and Rsample and Rstandard are the 13C/12C or 15N/14N ratios of sample and standard, respectively:
2.7 Microbial diversity analyses
Alpha diversity
Chao1 diversity index was calculated from the ASV tables derived from the bioinformatic sequence analysis using the phyloseq R package (McMurdie and Holmes 2013). Post-hoc T-student tests were applied to detect differences between treatments using the R package (Constantin and Patil 2021).
2.8 Beta diversity
Aitchinson distance matrices were calculated and Non-Metric Multidimensional Scaling (NMDS) at order level were performed using the vegdist function of the vegan package in R (Oksanen et al. 2022). Permutational Analysis of Variance analysis (PERMANOVA) and LDA Effect size (LEfSe) analysis were performed to test differences between land-use categories using the Microbiome Analyst web-based tool (Dhariwal et al. 2017).
2.9 Modeling approach
A structural equation modeling approach (SEM) was used to tests the hypothesis that soil properties and microbial communities in reciprocal interaction act as mediators of the effects of land-use over the soil functional response (δ13C/δ15N ratio). Based on the literature, we expected: (i) direct unidirectional relationships between land-use and nutrient content (negative), nutrient content and δ13C/δ15N (positive and negative), microbial community and δ13C/δ15N (positive and negative) (Qun and Geng-Xing 2010; Adugna and Abegaz 2016; Djodjic et al. 2021; Farooq et al. 2021; Jeong et al. 2023), (ii) direct, bidirectional and positive relationship between microbial community and nutrient content (Allison et al. 2007; Cui et al. 2018; Xu et al. 2022; Philippot et al. 2023), and (iii) indirect relationship, mediated by microbial community, between nutrient content and δ13C/δ15N.
The input variables for the model were selected and optimized in two steps. First, since the number of explanatory variables corresponding to microbial diversity was very large (> 50 Bacteria and Archaea orders), we performed an efficient branch-and-bound algorithm implemented on the regsubsets function in the leaps R package (Lumley 2020) to determine the ten best models that better explain δ13C/δ15N ratio variances according with the Akaike Information Criterion (AIC). Second, to explain the variance of the outcome δ13C/δ15N ratio as ecosystem function indicator, a SEM was constructed using the R package the lavaan (Latent Variable Analysis, v 0.6–17, (Rosseel 2012), where the following interactions were tested: 1) land-use as a latent variable (time of conversion and managed/not managed categorical variable), 2) nutrient content as a composite variable (N, NO3, SOM) and, 3) microbial community, as latent variable (only the three orders that exhibit significant differences with the response variable).
Statistical significance estimates were calculated by Maximum Likelihood and Non-Linear Minimization Subject to Box Constraints (NLMINB) was used as optimization method. Finally, the goodness of fit for the overall model was judged by the X2-test (p < 0.05) and RMSEA (< 0.05) (Root-Mean-Square Error Approximation) statistic for statistical significance (Gao et al. 2020) previous.
3 Results
3.1 Soil physicochemical analyses
Soil physicochemical properties within the pairs of land use changes (original versus transformed) indicate variations from fir forest to agricultural land and, from pine forest to reforested pine (Table 2). Notably, the conversion from fir forest to agriculture presents the most substantial alterations, with 11 out of the 15 measured variables experiencing modifications. This conversion is marked by a shift to increased soil acidity, higher electrical conductivity, and elevated levels of organic matter, as well as, nitrogen, and calcium. Simultaneously, iron content decreased, while phosphorus, potassium, and, notably, nitrate increased. Conversely, the transition from pine forest to reforested pine showed an increase in organic matter, nitrogen, potassium, calcium, magnesium, and iron content.
In the case of the land use change from grassland to burned grassland, no statistically significant differences were observed (P < 0.05). Finally, ammonium, total phosphorus, and total carbon content remained unaltered in all sampled sites.
3.2 Microbial diversity
Microbial diversity measured as alpha diversity using the Chao1 index along different land use sites (original versus transformed) showed significant variations. Specifically, marked variations were evident in the transitions from fir forest to agricultural land and, from pine forest to reforested pine (Fig. 2). The conversion from fir forest to agriculture resulted in a reduction in diversity, while the transition from pine forest to reforested pine was associated with a diversity increase. Conversely, there were no differences in diversity between grassland and burned grassland sites.
Bacterial community diversity of land-use change pairs (original vs transformed). A Comparison of Fir vs. Agriculture B Comparison of Pine vs. Forested Pine C Comparison of Grassland vs. Burned Grassland. Box plots of alpha diversity calculated as Chao1 index. Box represents the interquartile range (IQR) between the first and third quartiles (25th and 75th), middle line inside the box represents the median and whiskers represent the lowest and highest values within 1.5 times the IQR. t-tests for significant differences between pairs were conducted and the P values are indicated on the top right corner of each comparison and significance is indicated as asterisk (P < 0.05)
Furthermore, these disparities were further underscored at beta diversity by the multidimensional ordination analysis, we observe differences associated with land-use in which geographic proximity also played a pivotal role, leading to greater similarity among nearby sites (Fig. 3a, b). In this context, factors such as soil texture (including sand, clay, and silt content) and altitude also emerged as influential determinants (Fig. 3a).
Bacterial beta-diversity across land-use types. a Non-metric multidimensional scaling (NMDS) plot for bacterial community composition dissimilarity across samples. Dissimilarity was estimated as Aitchinson distance with soil physicochemical parameters vectors generated using envfit. Shapes correspond to pairwise land-use change comparisons, and colors correspond to original and transformed environments (solid color, original; open-color, transformed environment). b Heatmap of the most significant bacterial orders per land-use type obtained throughout LEfSe analysis. Abundance corresponds to the z-score values of each microbial group
Also, these diversity shifts were consistent with the heatmap analysis conducted on the statistically associated groups identified through LefSe (Fig. 3b). The sites showing the most similarity were those within the grassland and burned grassland categories, followed by pine forest to reforested pine sites, while the fir forestland and agricultural sites displayed the most dissimilarity.
3.3 Isotope analyses and SEM
Stable isotope and δ13C/δ15N ratio analyses across sampled sites revealed distinctive isotopic signatures per site (Fig. 4). Grassland and burned grassland sites exhibited the highest similarity, whereas pine trees (both natural and reforested) diverged in opposite directions, and the fir’s signature was closely resembling native pines. Conversely, the agricultural site showed a marked variation from the other sample sites, displaying the lowest intraspecific variation.
Biplot of N and C stable isotope ratios for all land-use change samples (original and transformed). For each land-use type standard ellipses are drawn with confidence intervals at 95%: Fir (solid circles), Agriculture (open circles), Pine (solid triangles), Reforested pine (open triangles), Grassland (solid squares) and Burned grassland (open squares)
A Structural Equation Modeling approach (SEM) was used to test the hypothesis that soil properties and microbial communities act as mediators of the effects of land-use over the soil functional response (δ13C/δ15N ratio). The model showed that land-use exerted a negative relationship over the functional response of δ13C/δ15N ratio through nutrient content (path coefficient = -0.041, p < 0.05). Specifically, nutrient content had a direct and positive effect on the isotope signature ratio response, where the variables that explain variations in δ13C/ δ15N ratio were SOM (62.02, p < 0.01), N (-43.93, p < 0.05) and NO3 (-25.04, p < 0.05). Moreover, microbial community was directly correlated with nutrient content and had a strong direct relationship to the functional response (δ13C/δ15N ratio), through the three associated microbial orders Chitinophagales, (-19.59, p < 0.05), Shingobacteriales (25.63, p < 0.01) and Tepidspherales (29.13, p < 0.05). Finally, the multiple squared correlations highly explained (92.82%) the variation in the δ13C/δ15N ratio as soil functional ecosystem indicator.
4 Discussion
4.1 Physicochemical soil properties changes are associated with land use change
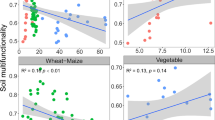
Land-use changes can influence soil nutrient dynamics and microbial community composition, thereby impacting crucial soil functions and physicochemical processes (Kooch et al. 2016). Our results of soil properties (i.e. nutrient content) showed changes associated with land-use, for instance, the transition from fir to agriculture exhibited the most pronounced alterations, in particular changes in SOM, N, P, K, Ca, Mg, Fe, NO3 (Table 2). This phenomenon is consistent with existing literature, highlighting that deforestation of entire ecosystems leads to nutrient depletion (Veldkamp et al. 2020; Bodo et al. 2021; Kumar et al. 2022). Specifically, the conversion of forests to agriculture has been shown to diminish organic matter (SOM), nitrogen, and calcium content—a trend reflected in our findings (Merloti et al. 2019; Peng et al. 2021). We also observe changes in agricultural field management, such as soil acidification (de Graaff et al. 2019) and elevation of nitrate and phosphorus levels, attributed to fertilizers commonly employed in potato cultivars (Solanum tuberosum L), including ammonium nitrate (AN), diammonium phosphate (DAP), and triple superphosphate (TSP) (Rosen et al. 2014).
Moreover, native forest conversion typically results in a net carbon loss in the soil (Yang et al. 2011). In contrast, conversion from pine to reforested pine showed an increase in soil organic matter and nutrient contents (N, K, Ca, Mg, and Fe). This increase could be attributed to the selection of frost-resistant Pinus montezumae for reforestation, exhibiting rapid growth, higher survival rates, and greater biomass compared to Pinus hartwegii, the native vegetation species in Nevado de Toluca (Viveros-Viveros et al. 2007; Rojas-García et al. 2019). Moreover, the transition from grassland (Festuca tolucensis) to burned grassland (due to firebreaks and controlled burning practices) did not result in significant soil properties alterations. This lack of statistical changes may be attributed to the lesser disturbance introduced to the environment where the burning is controlled and the soil structure is not affected (Neary et al. 1999), as opposed to the complete vegetation depletion observed in the fir-to-agriculture transition that affects soil structure, porosity, organic carbon, and nutrient availability or the plant species change observed in the native pine-to-reforested-pine scenario as reported in previous studies (Mueller et al. 2014; Goss-Souza et al. 2020; Hüblová and Frouz 2021).
4.2 Changes in microbial communities are associated with land-use change
Microbial diversity, or alpha diversity, is associated with land-use change and the specific management practices. Our study revealed that, land use change not only modified various soil properties but also resulted in changes in microbial diversity (alpha diversity) (Fig. 2). These changes in microbial diversity can be an increase or decrease in diversity, we observe both scenarios that also have been reported previously. In particular, reduction in microbial diversity was observed in the fir to agriculture land-use transition, potentially due to a reduction in soil organic carbon availability (Yu et al. 2012). Specifically in our case, the agricultural practices in the fir to agricultural transition are associated with potato agriculture that involves a complete depletion of any vegetation and natural input of organic carbon, being limited to intentional inputs that may not be sufficient to maintain microbial diversity. In contrast, an increase in microbial diversity was observed in the pine to reforested pine transition which could be attributed to the increased nutrient availability through intended nutrient inputs (Verchot 2010; Philippot et al. 2023). In particular, we attribute the increase in alpha diversity in the transition from pine to reforested pine to the enhanced nutrient supply from the leaf litter of Pinus montezumae compared to Pinus hartwegii (Lopez-Escobar et al. 2017).
Microbial community composition differences (beta diversity) are associated with specific management practices and the time elapsed since first intervention. We observe microbial composition differences associated with land-use and related factors such as soil properties (Fig. 3a, b). In this regard, it has been well documented that environmental perturbation (either natural or anthropogenic) can change microbial communities composition and if maintained, this change can be permanent (Hartmann et al. 2014; Yang et al. 2020). It is worth mentioning that the magnitude of differences in microbial communities can be associated with the time elapsed since the land-use change occurred. In particular, microbial communities that presented the greater differences are those in the transition from fir-dominated ecosystem-to-agricultural land, where land-use modification occured ∼ 50 years ago, while, intermediate compositional differences are observed in sites with a 30 year transition from pine to pine reforested. Finally the lowest compositional difference is observed in the grassland-to-burned grassland transition.
Land use transitions and associated management practices can be related to specific microbial groups. Bacterial orders whose abundances were statistically correlated with different land-uses and soil properties (Fig. 3b). For example, the highest abundance of Rhizobiales in fir and reforested pine where high nitrogen content is also observed. These orders have been reported as abundant in temperate forests and are key taxa contributing to the overall carbon decomposition process (Tláskal et al. 2017). Another example is Acidobacteriales, found in the pine forest site (P. hartwegii) and grasslands sites (natural and burned), this bacterial order has been reported to be abundant in coniferous forests and grasslands (Naether et al. 2012; Li et al. 2019). Previous studies have found that Acidobacteriales order is ubiquitous and abundant, mainly in soils with low carbon availability (constituting around 20% on average in soil microbiome surveys) (Janssen 2006), suggesting, adaptation to low substrate availabilities (oligotrophs) (Naether et al. 2012), and being consistent with the grasslands, and pine forest sites that exhibit the lowest total carbon contents (Table 2). Finally, Enterobacteriales and Corynebacteriales were abundant in the agriculturally managed site, these orders have been historically associated with fecal pollution associated with agricultural inputs (i.e. manure) and anthropogenic influence (Tkhakakhova et al. 2016; Devane et al. 2023).
4.3 Understanding the ecosystem functional response through isotopic signatures: soil properties and microbial community as mediators of land-use change
There is currently insufficient emphasis on utilizing δ13C and δ15N records as indicators of biochemical C and N cycling processes in microbial ecology studies (Park et al. 2023), despite their widespread application in plants and animals as indicators of trophic interactions (Wilkinson et al. 2022). While it is feasible to identify trophic pathways of C and N inputs into an ecosystem through stable isotope measures (De Clercq et al. 2015; Bieluczyk et al. 2023), the δ13C and δ15N ratio can offer an aggregate overview of all interactions and transformations involving C and N in an environment (Layman et al. 2007; Liu et al. 2018; Soldatova et al. 2024).
In this study, δ13C/δ15N was used as a reference of the C and N interactions and transformations in the environment (Layman et al. 2007; Liu et al. 2018; Soldatova et al. 2024), in order to show that each ecosystem and land-use possesses a unique isotopic signature (Fig. 4), reflecting different interactions among the components of the systems such as nutrient contents and microbial communities (Fig. 5). These relationships were formally tested using the Structural Equation Modeling (SEM) framework. Through this approach, we identified the effect of land-use over the soil functional response (δ13C/δ15N ratio), acting in cascade with the nutrient content and microbial community (Fig. 5). Furthermore, we found a negative relationship between land-use change and nutrient content, in particular Soil Organic Matter (SOM), total nitrogen and nitrates. This relationship has been previously documented on forest soils, where the concentration of these nutrients decreases when land-use changes (Merloti et al. 2019; Peng et al. 2021). Additionally, specific bacterial orders emerge as potential bioindicators, with their abundance being a factor linked to the functional ecosystem parameter δ13C/δ15N (Fig. 5). For instance, the abundance of Sphingobacteriales is positively related with the δ13C/δ15N ratio, previous studies associated the abundance of Sphingobacteriales with high soil organic carbon (SOC) mineralization (Whitman et al. 2016; Zhang et al. 2021a, b; Carrasco-Espinosa et al. 2022). Chitinophagales, recognized as chitinolytic and hydrocarbon degraders, also show a strong influence with δ13C/δ15N ratios (Kaoping et al. 2023) and exhibit reduced abundance in afforestation and land-use change processes (Lan et al. 2022). Tepidisphaerales, displaying good tolerance to low temperatures (4-15 ºC) (Ivanova et al. 2016), experience changes in abundance following restoration-fertilization processes (Cheng et al. 2021) and soil warming experiments (Parada-Pozo et al. 2022). Altogether the evidence here presented suggests potential roles for specific bacterial groups as biological indicators and soil nutrient drivers of C and N transformations, warranting further investigation through specific experiments such as mesocosms or complementary metagenomic analyses.
Structural equation model describing soil properties and microbial communities in reciprocal interaction as mediators of the effects of land-use over the soil functional response (δ13C/δ15N ratio). The model diagram depicts the different variables interactions (nutrient content, microbial communities and land-use) and their effect in the functional response (δC13/δN15). Latent variables (land-use and microbial communities) are represented as ellipses and composite variables (nutrient content) are represented as rectangular boxes; arrows depict the relationships (effects) between variables, their width is proportional to their coefficient path and its significance p-value (* P < 0.05, ** P < 0.01 and *** P < 0.001), which values are also included in the diagram. The model fit parameters correspond to the Root-Mean-Square Error Approximation statistic (RMSEA), Chi-square (X2) value and the significance probability of p-value
Finally, through SEM we tested the hypothesis of the role that nutrient content and microbial communities in reciprocal interaction play as mediators of the impact of land-use on ecosystem functioning. The resulting model shows that, in fact, land-use change (as environmental perturbation), directly impacts nutrient contents, and in turn nutrient contents interact reciprocally with microbial community composition that impact ecosystem function (Fig. 5). Although, in some sense, all these interactions have been previously documented (Chadwick et al. 2015; Adetunji et al. 2020; Thakur et al. 2020; Philippot et al. 2023), it has not been common to formally test not only the interactions but the strength and direction of such interactions. In particular, it is worth noticing, that although the effect of specific management practices or land-use transitions in ecosystem function is obscured by summarizing all sampled sites in a single model, we do demonstrate that interventions do change ecosystem function in idiosyncratic ways (Fig. 4), and that microbial communities act as strong mediators of such effects, also identifying specific microbial groups involved (Fig. 5), which represents a generalizable model for different ecosystems and/or environmental perturbations (Jorgensen 2021; Vispoel et al. 2023). Also, although the interaction between nutrient contents and microbial community is highly significant, its magnitude is not very high, may be due to the specific measurements included in this latent variable that can overlook functionally relevant nutrients in this interaction, including specific carbon substrates that could be better correlated with the abundance of specific microbial groups (Abraham et al. 1998; Boschker and Middelburg 2002; Trivedi et al. 2016).
5 Conclusions
Our findings underscore the intricate relationship between microbial community structure, soil properties, and environmental transformations, particularly land-use change. Specifically, we demonstrate the cause-effect relationships among land-use, nutrient contents and microbial communities, where soil properties and microbial communities interact reciprocally, acting as mediators of the effects of land-use change in soil functional responses (i.e. δ13C/δ15N ratio). This understanding highlights the potential use of bioindicators (microbial communities) by decision-makers regarding forest management in Mexico to facilitate the development of sustainable management strategies.
Data availability
Raw data and metadata were deposited in the NCBI Sequence Read Archive (SRA) under the accession number PRJNA1037141.
References
Abraham W-R, Hesse C, Pelz O (1998) Ratios of Carbon Isotopes in Microbial Lipids as an Indicator of Substrate Usage. Appl Environ Microbiol 64(11):4202–4209. https://doi.org/10.1128/AEM.64.11.4202-4209.1998
Adetunji AT, Ncube B, Mulidzi R, Lewu FB (2020) Management impact and benefit of cover crops on soil quality: A review. Soil till Res 204. https://doi.org/10.1016/J.STILL.2020.104717
Adugna A, Abegaz A (2016) Effects of land use changes on the dynamics of selected soil properties in northeast Wellega. Ethiopia Soil 2(1):63–70. https://doi.org/10.5194/soil-2-63-2016
Allison VJ, Condron LM, Peltzer DA, Richardson SJ, Turner BL (2007) Changes in enzyme activities and soil microbial community composition along carbon and nutrient gradients at the Franz Josef chronosequence. New Zealand Soil Biol Biochem 39:1770–1781. https://doi.org/10.1016/J.SOILBIO.2007.02.006
Bieluczyk W, Ometto Asselta F, Navroski D, Gontijo JB, Monteiro Venturini A, Mendes LW, Simon CP, Barbosa De Camargo P, Tadini AM, Martin-Neto L, Albertino Bendassolli J, Rodrigues RR, Van Der Putten WH, Tsai SM (2023) Linking above and belowground carbon sequestration, soil organic matter properties, and soil health in Brazilian Atlantic Forest restoration. J Environ Manag 344:301–4797. https://doi.org/10.1016/j.jenvman.2023.118573
Bodo T, Gimah BG, Seomoni KJ (2021) Deforestation and Habitat Loss: Human Causes, Consequences and Possible Solutions. J Geogr Res 4(2):22–30. https://doi.org/10.30564/jgr.v4i2.3059
Boschker HTS, Middelburg JJ (2002) Stable isotopes and biomarkers in microbial ecology. FEMS Microbiol Ecol 40(2):85–95. https://doi.org/10.1016/S0168-6496(02)00194-0
Bremner JM (1960) Determination of nitrogen in soil by the Kjeldahl method. J Agric Sci 55(1):11–33. https://doi.org/10.1017/S0021859600021572
Calderón de Rzedowski G, Rzedowski J (2005) Flora fanerogámica del Valle de México. INECOL-CONABIO, México
Carrasco-Espinosa K, Avitia M, Barrón-Sandoval A, Abbruzzini TF, Salazar Cabrera UI, Arroyo-Lambaer D, Uscanga A, Campo J, Benítez M, Wegier A, Rosell JA, Reverchon F, Hernández G, Boege K, Escalante AE (2022) Land-Use Change and Management Intensification Is Associated with Shifts in Composition of Soil Microbial Communities and Their Functional Diversity in Coffee Agroecosystems. Microorganisms 10:1763. https://doi.org/10.3390/microorganisms10091763
Chadwick D, Wei J, Yan’an T, Guanghui Y, Qirong S, Qing C (2015) Improving manure nutrient management towards sustainable agricultural intensification in China. Agric Ecosyst Environ 209:34–46. https://doi.org/10.1016/J.AGEE.2015.03.025
Chen L, Jiang L, Jing X, Wang J, Shi Y, Chu H, He J (2021) Above- and belowground biodiversity jointly drive ecosystem stability in natural alpine grasslands on the Tibetan Plateau. Glob Ecol Biogeogr 30(7):1418–1429. https://doi.org/10.1111/geb.13307
Cheng H, Wu B, Wei M, Wang S, Rong X, Du D, Wang C (2021) Changes in community structure and metabolic function of soil bacteria depending on the type restoration processing in the degraded alpine grassland ecosystems in Northern Tibet. Sci Total Environ 755. https://doi.org/10.1016/J.SCITOTENV.2020.142619
Choi W-J, Müller C, Zaman M, Nannipieri P (2023) Stable isotopes for the study of soil C and N under global change. Biol Fert Soils 59:485–486. https://doi.org/10.1007/s00374-023-01747-z
Constantin A, Patil I (2021) ggsignif: R Package for Displaying Significance Brackets for 'ggplot2'. PsyArxiv. https://doi.org/10.31234/osf.io/7awm6, https://psyarxiv.com/7awm6
Craine JM, Brookshire ENJ, Cramer MD, Hasselquist NJ, Koba K, Marin-Spiotta E, Wang L (2015) Ecological interpretations of nitrogen isotope ratios of terrestrial plants and soils. Plant Soil 396:1–26. https://doi.org/10.1007/s11104-015-2542-1
Cui Y, Fang L, Guo X, Wang X, Wang Y, Li P, Zhang Y, Zhang X (2018) Responses of soil microbial communities to nutrient limitation in the desert-grassland ecological transition zone. Sci Total Environ 642:45–55. https://doi.org/10.1016/J.SCITOTENV.2018.06.033
De Clercq T, Heiling M, Dercon G, Resch C, Aigner M, Mayer L, Mao Y, Elsen A, Steier P, Leifeld J, Merckx R (2015) Predicting soil organic matter stability in agricultural fields through carbon and nitrogen stable isotopes. Soil Biol Biochem 88:29–38. https://doi.org/10.1016/J.SOILBIO.2015.05.011
de Graaff MA, Hornslein N, Throop HL, Kardol P, van Diepen LTA (2019) Chapter One - Effects of agricultural intensification on soil biodiversity and implications for ecosystem functioning: a meta-analysis. Adv Agron 155:1–44. https://doi.org/10.1016/bs.agron.2019.01.001
Devane ML, Taylor W, Dupont P-Y, Armstrong B, Weaver L, Gilpin BJ (2023) Exploring the Bacterial Community in Aged Fecal Sources from Dairy Cows: Impacts on Fecal Source Tracking. Microorganisms. https://doi.org/10.3390/microorganisms11051161
Dhariwal A, Chong J, Habib S, King IL, Agellon LB, Xia J (2017) MicrobiomeAnalyst: A web-based tool for comprehensive statistical, visual and meta-analysis of microbiome data. Nucleic Acids Res 45:W180-188. https://doi.org/10.1093/nar/gkx295
Diels J, Vanlauwe B, Van der Meersch MK, Sanginga N, Merckx R (2004) Long-term soil organic carbon dynamics in a subhumid tropical climate: 13C data in mixed C3/C4 cropping and modeling with ROTHC. Soil Biol Biochem 36:1739–1750. https://doi.org/10.1016/J.SOILBIO.2004.04.031
Djodjic F, Bieroza M, Bergström L (2021) Land use, geology and soil properties control nutrient concentrations in headwater streams. Sci Total Environ 772. https://doi.org/10.1016/J.SCITOTENV.2021.145108
Fan Y, Chen J, Shirkey G, John R, Wu SR, Park H, Shao C (2016) Applications of structural equation modeling (SEM) in ecological studies: an updated review. Ecol Process. https://doi.org/10.1186/s13717-016-0063-3
Farooq T, Chen X, Shakoor A, Li Y, Wang J, Rashid M, Kumar U, Yan W (2021) Unraveling the Influence of Land-Use Change on δ13C, δ15N, and Soil Nutritional Status in Coniferous, Broadleaved, and Mixed Forests in Southern China: A Field Investigation. Plants 10:1499. https://doi.org/10.3390/plants10081499
Felipe-Lucia MR, Soliveres S, Penone C, Fischer M, Ammer C, Boch S, Boeddinghaus RS, Bonkowski M, Buscot F, Fiore-Donno AM, Frank K, Goldmann K, Gossner MM, Hölzel N, Jochum M, Kandeler E, Klaus VH, Kleinebecker T, Leimer S et al (2020) Land-use intensity alters networks between biodiversity, ecosystem functions, and services. PNAS 117(45):28140–28149. https://doi.org/10.1073/pnas.2016210117
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol. https://doi.org/10.1038/nrmicro.2017.87
Gao C, Shi D, Maydeu-Olivares A (2020) Estimating the Maximum Likelihood Root Mean Square Error of Approximation (RMSEA) with Non-normal Data: A Monte-Carlo Study. Struct Equ Model 27(2):192–201. https://doi.org/10.1080/10705511.2019.1637741
García E (1990) Carta de climas. Universidad Nacional Autónoma de México, Atlas Nacional de México
Gee GW, Bauder JW (1986) Particle-size analysis (2nd edition). In: Klute A (ed) Methods of soil analysis Part 1. Wiley, pp 383–411
George PBL, Lallias D, Creer S, Seaton FM, Kenny JG, Eccles RM, Griffiths RI, Lebron I, Emmett BA, Robinson DA, Jones DL (2019) Divergent national-scale trends of microbial and animal biodiversity revealed across diverse temperate soil ecosystems. Nat Commun 10:1107. https://doi.org/10.1038/s41467-019-09031-1
Gerschlauer F, Saiz G, Schellenberger Costa D, Kleyer M, Dannenmann M, Kiese R (2019) Stable carbon and nitrogen isotopic composition of leaves, litter, and soils of various ecosystems along an elevational and land-use gradient at Mount Kilimanjaro. Tanzania Biogeosciences 16(2):409–424. https://doi.org/10.5194/bg-16-409-2019
González Contreras A (2019) Abundancia de los isótopos estables d13C y d15N en suelos de bosques templados y pastizales alpinos naturales y transformados [Universidad Nacional Autónoma de México]. https://tesiunam.dgb.unam.mx/F/YYIY6B28B12E1KR6XY4GAE94BSPN5GR614BYNAP7XFESMMQ874-08625?func=full-set-set&set_number=046592&set_entry=000002&format=999
Goss-Souza D, Mendes LW, Rodrigues JLM, Tsai SM (2020) Ecological Processes Shaping Bulk Soil and Rhizosphere Microbiome Assembly in a Long-Term Amazon Forest-to-Agriculture Conversion. Microb Ecol 79:110–122. https://doi.org/10.1007/s00248-019-01401-y
Hair JF, Hult GTM, Ringle CM, Sarstedt M, Danks NP, Ray S (2021) An introduction to structural equation modeling. In Partial least squares structural equation modeling (PLS-SEM) using R, classroom companion: business. Springer, Cham, pp 1–29. https://doi.org/10.1007/978-3-030-80519-7_1
Hartmann M, Niklaus PA, Zimmermann S, Schmutz S, Kremer J, Abarenkov K, Lüscher P, Widmer F, Frey B (2014) Resistance and resilience of the forest soil microbiome to logging-associated compaction. ISME J 8:226–244. https://doi.org/10.1038/ismej.2013.141
Hasan SS, Zhen L, Miah MG, Ahamed T, Samie A (2020) Impact of land use change on ecosystem services: A review. Environ Dev 34. https://doi.org/10.1016/J.ENVDEV.2020.100527
He P, Ling N, Lü X-T, Zhang H-Y, Wang C, Wang R-Z, Wei C-Z, Yao J, Wang X-B, Han X-G, Nan Z (2023) Contributions of abundant and rare bacteria to soil multifunctionality depend on aridity and elevation. Appl Soil Ecol 188. https://doi.org/10.1016/J.APSOIL.2023.104881
Heck RH, Thomas SL (2020) An introduction to multilevel modeling techniques MLM and SEM approaches (Fourth edi). Routledge Taylor & Francis. https://doi.org/10.4324/9780429060274
Herlemann DP, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. https://doi.org/10.1038/ismej.2011.41
Hu L, Li Q, Yan J, Liu C, Zhong J (2022) Vegetation restoration facilitates belowground microbial network complexity and recalcitrant soil organic carbon storage in southwest China karst region. Sci Total Environ 820. https://doi.org/10.1016/J.SCITOTENV.2022.153137
Hüblová L, Frouz J (2021) Contrasting effect of coniferous and broadleaf trees on soil carbon storage during reforestation of forest soils and afforestation of agricultural and post-mining soils. J Environ Manag 290. https://doi.org/10.1016/J.JENVMAN.2021.112567
Ivanova AA, Kulichevskaya IS, Merkel AY, Toshchakov SV, Dedysh SN (2016) High Diversity of Planctomycetes in Soils of Two Lichen-Dominated Sub-Arctic Ecosystems of Northwestern Siberia. Front Microbiol 7. https://doi.org/10.3389/fmicb.2016.02065
Janssen PH (2006) Identifying the Dominant Soil Bacterial Taxa in Libraries of 16S rRNA and 16S rRNA Genes. Appl Environ Microbiol 72(3):1719–1728. https://doi.org/10.1128/AEM.72.3.1719-1728.2006
Jeong Y-J, Park H-J, Baek N, Seo B-S, Lee K-S, Kwak J-H, Choi S-K, Lee S-M, Yoon K-S, Lim S-S, Choi W-J (2023) Assessment of sources variability of riverine particulate organic matter with land use and rainfall changes using a three-indicator (δ13C, δ15N, and C/N) Bayesian mixing model. Environ Res J 216. https://doi.org/10.1016/J.ENVRES.2022.114653
Jiang Y, Luan L, Hu K, Liu M, Chen Z, Geisen S, Chen X, Li H, Xu Q, Bonkowski M, Sun B (2020) Trophic interactions as determinants of the arbuscular mycorrhizal fungal community with cascading plant-promoting consequences. Microbiome 8:142. https://doi.org/10.1186/s40168-020-00918-6
Jiao S, Peng Z, Qi J, Gao J, Wei G (2021) Linking bacterial-fungal relationships to microbial diversity and soil nutrient cycling. MSystems. https://doi.org/10.1128/mSystems.01052-20
Jing X, Chen X, Fang J, Ji C, Shen H, Zheng C, Zhu B (2020) Soil microbial carbon and nutrient constraints are driven more by climate and soil physicochemical properties than by nutrient addition in forest ecosystems. Soil Biol Biochem 141. https://doi.org/10.1016/J.SOILBIO.2019.107657
Jorgensen TD (2021) How to Estimate Absolute-Error Components in Structural Equation Models of Generalizability Theory. Psych 3:113–133. https://doi.org/10.3390/psych3020011
Kang E, Li Y, Zhang X, Yan Z, Wu H, Li M, Yan L, Zhang K, Wang J, Kang X (2021) Soil pH and nutrients shape the vertical distribution of microbial communities in an alpine wetland. Sci Total Environ 774. https://doi.org/10.1016/J.SCITOTENV.2021.145780
Kaoping Z, Song Y, Xu H, Fei C (2023) Soil C:N: P stoichiometry regulates the response of soil bacterial community composition and putative functions to coastal reclamation. J Soils Sediments. https://doi.org/10.1007/s11368-023-03657-x
Kooch Y, Moghimian N, Bayranvand M, Alberti G (2016) Changes of soil carbon dioxide, methane, and nitrous oxide fluxes in relation to land use/cover management. Environ Monit Assess 188:346. https://doi.org/10.1007/s10661-016-5342-z
Kumar R, Kumar A, Saikia P (2022) Deforestation and Forests Degradation Impacts on the Environment. In: Singh V (ed) Environmental Degradation: Challenges and Strategies for Mitigation. Springer, Cham, pp 19–46. https://doi.org/10.1007/978-3-030-95542-7_2
Lai MHC, Hsiao Y-Y (2022) Two-stage path analysis with definition variables: An alternative framework to account for measurement error. Psychol Methods 27(4):568–588. https://doi.org/10.1037/met0000410
Lan J, Wang S, Wang J, Qi X, Long Q, Huang M (2022) The Shift of Soil Bacterial Community After Afforestation Influence Soil Organic Carbon and Aggregate Stability in Karst Region. Front Microbiol 13. https://doi.org/10.3389/fmicb.2022.901126
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415. https://doi.org/10.1016/J.SOILBIO.2008.05.021
Layman CA, Arrington DA, Montaña CG, Post DM (2007) Can stable isotope ratios provide for community-wide measures of trophic structure? Ecology 88(1):42–48. https://doi.org/10.1890/0012-9658(2007)88[42:CSIRPF]2.0.CO;2
Li W, Liu X, Niu S (2019) Differential responses of the acidobacterial community in the topsoil and subsoil to fire disturbance in Pinus tabulaeformis stands. PeerJ 7. https://doi.org/10.7717/peerj.8047
Liu S, Sun Y, Shi F, Liu Y, Wang F, Dong S, Li M (2022) Composition and Diversity of Soil Microbial Community Associated With Land Use Types in the Agro-Pastoral Area in the Upper Yellow River Basin. Front Plant Sci 13. https://doi.org/10.3389/fpls.2022.819661
Liu Y, Hu C, Hu W, Wang L, Li Z, Pan J, Chen F (2018) Stable isotope fractionation provides information on carbon dynamics in soil aggregates subjected to different long-term fertilization practices. Soil till Res 177:54–60. https://doi.org/10.1016/J.STILL.2017.11.016
Lopez-Escobar NF, Gómez-Guerrero A, Velázquez-Martínez A, Fierros-González AM, Castruita-Esparza LU, Vera-Castillo JAG (2017) Reservoirs and nutrient dynamics in two stands of Pinus montezumae Lamb in Tlaxcala, Mexico. Rev Chapingo Ser Cienc For Ambiente 24(1):115–129. https://doi.org/10.5154/r.rchscfa.2017.09.055
Luan L, Liang C, Chen L, Wang H, Xu Q, Jiang Y, Sun B (2020) Coupling Bacterial Community Assembly to Microbial Metabolism across Soil Profiles. Msystems 5(3):e00298-e320. https://doi.org/10.1128/mSystems.00298-20
Lumley T (2020) Leaps: regression subset selection. Version 3.1. https://CRAN.Rproject.org/package=leaps
Ma T, Zhan Y, Chen W, Xu S, Wang Z, Tao Y, Shi X, Sun B, Ding G, Li J, Wei Y (2022) Impact of aeration rate on phosphorus conversion and bacterial community dynamics in phosphorus-enriched composting. Bioresour Technol 364. https://doi.org/10.1016/J.BIORTECH.2022.128016
McKinley VL (2019) Effects of Land Use and Restoration on Soil Microbial Communities. In: Hurst CJ (ed) Understanding Terrestrial Microbial Communities, Advances in Environmental Microbiology. Springer, Nature Switzerland, pp 173–242. https://doi.org/10.1007/978-3-030-10777-2_7
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8(4). https://doi.org/10.1371/journal.pone.0061217
Merloti LF, Mendes LW, Pedrinho A, Fonseca De Souza L, Ferrari BM, Tsai SM (2019) Forest-to-agriculture conversion in Amazon drives soil microbial communities and N-cycle. Soil Biol Biochem. https://doi.org/10.1016/j.soilbio.2019.107567
Mueller RC, Paula FS, Mirza BS, Rodrigues JLM, Nüsslein K, Bohannan BJM (2014) Links between plant and fungal communities across a deforestation chronosequence in the Amazon rainforest. ISME J 8:1548–1550. https://doi.org/10.1038/ismej.2013.253
Naether A, Foesel BU, Naegele V, Wüst PK, Weinert J, Bonkowski M, Alt F, Oelmann Y, Polle A, Lohaus G, Gockel S, Hemp A, Kalko EKV, Linsenmair KE, Pfeiffer S, Renner S, Schöning I, Weisser WW, Wells K et al (2012) Environmental Factors Affect Acidobacterial Communities below the Subgroup Level in Grassland and Forest Soils. Appl Environ Microbiol 78(20):7398–7406. https://doi.org/10.1128/AEM.01325-12
Neary DG, Klopatek CC, DeBano LF, Ffolliott PF (1999) Fire effects on belowground sustainability: a review and synthesis. For Ecol Manag 122:51–71. https://doi.org/10.1016/S0378-1127(99)00032-8
Ngaba MJY, Hu Y-L, Bol R, Ma X-Q, Jin S-F, Mgelwa AS (2019) Effects of land use change from natural forest to plantation on C, N and natural abundance of 13C and 15N along a climate gradient in eastern China. Sci Rep 9:16516. https://doi.org/10.1038/s41598-019-52959-z
Oksanen J, Simpson G, Blanchet F, Kindt R, Legendre P, Minchin P, O'Hara R, Solymos P, Stevens M, Szoecs E, Wagner H, Barbour M, Bedward M, Bolker B, Borcard D, Carvalho G, Chirico M, De Caceres M, Durand S, Evangelista H, FitzJohn R, Friendly M, Furneaux B, Hannigan G, Hill M, Lahti L, McGlinn D, Ouellette M, Ribeiro Cunha E, Smith T, Stier A, Ter Braak C, Weedon J (2022) _vegan: Community Ecology Package_. R package version 2.6-4
Parada-Pozo G, Bravo LA, Sáez PL, Cavieres LA, Reyes-Díaz M, Abades S, Alfaro FD, De la Iglesia R, Trefault N (2022) Vegetation drives the response of the active fraction of the rhizosphere microbial communities to soil warming in Antarctic vascular plants. FEMS Microbiol Ecol 98:1–12. https://doi.org/10.1093/femsec/fiac099
Park H-J, Baek N, Lim S-S, Jeong Y-J, Seo B-S, Kwak J-H, Lee S-M, Yun S-I, Kim H-Y, Arshad MA, Choi W-J (2023) Coupling of δ13C and δ15N to understand soil organic matter sources and C and N cycling under different land-uses and management: a review and data analysis. Biol Fert Soils 59:487–499. https://doi.org/10.1007/s00374-022-01668-3
Peng S, Liu W, Xu G, Pei X, Millerick K, Duan B (2021) A meta-analysis of soil microbial and physicochemical properties following native forest conversion. Catena. https://doi.org/10.1016/j.catena.2021.105447
Pérez-Hernández J, Gavilán RG (2021) Impacts of Land-Use Changes on Vegetation and Ecosystem Functioning: Old-Field Secondary Succession. Plants 10:990. https://doi.org/10.3390/plants10050990
Philippot L, Chenu C, Kappler A, Rillig MC, Fierer N (2023) The interplay between microbial communities and soil properties. Nat Rev Microbiol 10(1038):1–14. https://doi.org/10.1038/s41579-023-00980-5
Qun M, Geng-Xing Z (2010) Effects of Different Land Use Types on Soil Nutrients in Intensive Agricultural Region. J Nat Resour 25(11):1834–1844. https://doi.org/10.11849/ZRZYXB.2010.11.003
Rojas-García F, Fredericksen TS, Vazquez Lozada S, Endara Agramont AR (2019) Impact of timber harvesting on carbon storage in montane forests of central Mexico. New for 50:1043–1061. https://doi.org/10.1007/s11056-019-09714-z
Rosen CJ, Kelling KA, Stark JC, Porter GA (2014) Optimizing Phosphorus Fertilizer Management in Potato Production. Am J Potato Res 91:145–160. https://doi.org/10.1007/s12230-014-9371-2
Rosseel Y (2012) lavaan: An R Package for Structural Equation Modeling. J Stat Softw 48(2):1–21. https://doi.org/10.18637/jss.v048.i02
Saborío-Montero A, Gutiérrez-Rivas M, García-Rodríguez A, Atxaerandio R, Goiri I, López de Maturana E, Jiménez-Montero JA, Alenda R, González-Recio O (2020) Structural equation models to disentangle the biological relationship between microbiota and complex traits: Methane production in dairy cattle as a case of study. J Anim Breed Genet 137(1):36–48. https://doi.org/10.1111/jbg.12444
Shao Y, Bao W, Chen D, Eisenhauer N, Zhang W, Pang X, Xu G, Fu S (2015) Using structural equation modeling to test established theory and develop novel hypotheses for the structuring forces in soil food webs. Pedobiologia 58(4):137–145. https://doi.org/10.1016/J.PEDOBI.2015.06.001
Sims JT (2000) Soil test phosphorus: Olsen P. In: Pierzynski GM (ed) Methods of phosphorus analysis for soils, sediments, residuals, and waters. Southern Cooperative Series Bulletin No.396, SERA-IEG 17, Manhattan, KS, pp 20–22. http://www.soil.ncsu.edu/sera17/publications/sera17-2/pm_cover.htm
Singavarapu B, Du J, Beugnon R, Cesarz S, Eisenhauer N, Xue K, Wang Y, Bruelheide H, Wubet T (2023) Functional Potential of Soil Microbial Communities and Their Subcommunities Varies with Tree Mycorrhizal Type and Tree Diversity. Microbiol Spectr. https://doi.org/10.1128/spectrum.04578-22
Singh AK, Sisodia A, Sisodia V, Padhi M (2019) Role of Microbes in Restoration Ecology and Ecosystem Services. New and Future Developments in Microbial Biotechnology and Bioengineering. https://doi.org/10.1016/B978-0-444-64191-5.00004-3
Soldatova E, Krasilnikov S, Kuzyakov Y (2024) Soil organic matter turnover: Global implications from δ13C and δ15N signatures. Sci Total Environ 912. https://doi.org/10.1016/J.SCITOTENV.2023.169423
Sun Y, Luo C, Jiang L, Song M, Zhang D, Li J, Li Y, Ostle NJ, Zhang G (2020) Land-use changes alter soil bacterial composition and diversity in tropical forest soil in China. Sci Total Environ 712. https://doi.org/10.1016/J.SCITOTENV.2020.136526
Tang S, Ma Q, Marsden KA, Chadwick DR, Luo Y, Kuzyakov Y, Wu L, Jones DL (2023) Microbial community succession in soil is mainly driven by carbon and nitrogen contents rather than phosphorus and sulphur contents. Soil Biol Biochem 180. https://doi.org/10.1016/J.SOILBIO.2023.109019
Thakur AK, Mandal KG, Raychaudhuri S (2020) Impact of crop and nutrient management on crop growth and yield, nutrient uptake and content in rice. Paddy Water Environ 18:139–151. https://doi.org/10.1007/s10333-019-00770-x
Tkhakakhova AK, Chernov TI, Ivanova EA, Kutovaya OV, Kogut BM, Zavalin AA (2016) Changes in the metagenome of the prokaryotic community of chernozems under the influence of mineral fertilizers. Russ Agric Sci 42(1):62–65. https://doi.org/10.3103/s1068367416010201
Tláskal V, Zrůstová P, Vrška T, Baldrian P (2017) Bacteria associated with decomposing dead wood in a natural temperate forest. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fix157
Tosi M, Correa OS, Soria MA, Vogrig JA, Sydorenko O, Montecchia MS (2016) Land-use change affects the functionality of soil microbial communities: A chronosequence approach in the Argentinian Yungas. Appl Soil Ecol 108:118–127. https://doi.org/10.1016/J.APSOIL.2016.08.012
Trivedi P, Delgado-Baquerizo M, Trivedi C, Hu H, Anderson IC, Jeffries TC, Zhou J, Singh BK (2016) Microbial regulation of the soil carbon cycle: evidence from gene–enzyme relationships. ISME J 10:2593–2604. https://doi.org/10.1038/ismej.2016.65
Veldkamp E, Schmidt M, Powers JS, Corre MD (2020) Deforestation and reforestation impacts on soils in the tropics. Nat Rev Earth Environ 1:590–605. https://doi.org/10.1038/s43017-020-0091-5
Verchot LV (2010) Impacts of Forest Conversion to Agriculture on Microbial Communities and Microbial Function. In: Dion P (ed) Soil Biology and Agriculture in the Tropics. Springer, Berlin, Heidelberg, pp 45–63. https://doi.org/10.1007/978-3-642-05076-3_3
Vispoel WP, Hong H, Lee H, Jorgensen TD (2023) Analyzing Complete Generalizability Theory Designs Using Structural Equation Models. Appl Meas Educ 36(4):372–393. https://doi.org/10.1080/08957347.2023.2274573
Viveros-Viveros H, Sáenz-Romero C, López-Upton J, Vargas-Hernández JJ (2007) Growth and frost damage variation among Pinus pseudostrobus, P. montezumae and P. hartwegii tested in Michoacán. México for Ecol Manag 253:81–88. https://doi.org/10.1016/j.foreco.2007.07.005
Wang Y, Liu L, Yang J, Duan Y, Luo Y, Taherzadeh MJ, Li Y, Li H, Awasthi MK, Zhao Z (2020) The diversity of microbial community and function varied in response to different agricultural residues composting. Sci Total Environ 715. https://doi.org/10.1016/J.SCITOTENV.2020.136983
Whitman T, Pepe-Ranney C, Enders A, Koechli C, Campbell A, Buckley DH, Lehmann J (2016) Dynamics of microbial community composition and soil organic carbon mineralization in soil following addition of pyrogenic and fresh organic matter. ISME J 10:2918–2930. https://doi.org/10.1038/ismej.2016.68
Wilkinson C, Lim RBH, Liew JH, Kwik JTB, Tan CLY, Heok Hui T, Yeo DCJ (2022) Empirical food webs of 12 tropical reservoirs in Singapore. Biodivers Data J 10. https://doi.org/10.3897/BDJ.10.e86192
Wynn JG, Bird MI (2008) Environmental controls on the stable carbon isotopic composition of soil organic carbon: implications for modelling the distribution of C3 and C4 plants. Australia Tellus B Chem Phys Meteorol 60(4):604. https://doi.org/10.1111/j.1600-0889.2008.00361.x
Xu M, Li X, Kuyper TW, Xu M, Li X, Zhang J (2021) High microbial diversity stabilizes the responses of soil organic carbon decomposition to warming in the subsoil on the Tibetan Plateau. Glob Change Biol 27(10):2061–2075. https://doi.org/10.1111/gcb.15553
Xu M, Wang J, Zhu Y, Han X, Ren C, Yang G (2022) Plant Biomass and Soil Nutrients Mainly Explain the Variation of Soil Microbial Communities During Secondary Succession on the Loess Plateau. Microb Ecol 83(1):114–126. https://doi.org/10.1007/s00248-021-01740-9
Xue P, Minasny B, McBratney AB (2022) Land-use affects soil microbial co-occurrence networks and their putative functions. Appl Soil Ecol 169. https://doi.org/10.1016/J.APSOIL.2021.104184
Yang J, Li A, Yang Y, Li G, Zhang F (2020) Soil organic carbon stability under natural and anthropogenic-induced perturbations. Earth-Sci Rev 205. https://doi.org/10.1016/J.EARSCIREV.2020.103199
Yang Y, Luo Y, Finzi AC (2011) Carbon and nitrogen dynamics during forest stand development: a global synthesis. New Phytol 190(4):977–989. https://doi.org/10.1111/j.1469-8137.2011.03645.x
Yu Y, Shen W, Yin Y, Zhang J, Cai Z, Zhong W (2012) Response of soil microbial diversity to land-use conversion of natural forests to plantations in a subtropical mountainous area of southern China. Soil Sci Plant Nut 58(4):450–461. https://doi.org/10.1080/00380768.2012.708645
Zhang G, Yu X, Xu J, Duan H, Rafay L, Zhang Q, Li Y, Liu Y, Xia S (2018) Effects of environmental variation on stable isotope abundances during typical seasonal floodplain dry season litter decomposition. Sci Total Environ 630:1205–1215. https://doi.org/10.1016/J.SCITOTENV.2018.02.298
Zhang R, Wang Z, Niu S, Tian D, Wu Q, Gao X, Schellenberg MP, Han G (2021a) Diversity of plant and soil microbes mediates the response of ecosystem multifunctionality to grazing disturbance. Sci Total Environ 776. https://doi.org/10.1016/J.SCITOTENV.2021.145730
Zhang S, Fang Y, Luo Y, Li Y, Ge T, Wang Y, Wang H, Yu B, Song X, Chen J, Zhou J, Li Y, Chang SX (2021b) Linking soil carbon availability, microbial community composition and enzyme activities to organic carbon mineralization of a bamboo forest soil amended with pyrogenic and fresh organic matter. Sci Total Environ 801. https://doi.org/10.1016/J.SCITOTENV.2021.149717
Acknowledgements
This work is part of the PhD research of KCE, who received a scholarship from the Consejo Nacional de Humanidades, Ciencia y Tecnología (CONAHCyT, scholarship no. 443871) and extends thanks to the Doctorado en Ciencias Biomédicas, UNAM. Also, the authors acknowledge technical assistance of Rodrigo González Chauvet, Alberto Barrón-Sandoval, Gloria A. Sarmina, Alan Heres and Laurette Prince. We like to thank to the Santiago-Tlacotepc community members for their time and solidarity accompaniment.
Funding
This work was supported by UNAM-PAPIIT IN207619.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Research involving human participants and/or animals
No studies with human participants or animals were performed by any of the authors to obtain data for this paper.
Conflicts of interest
The authors hereby declare that they have no conflicts of interest.
Additional information
Responsible editor: Dulce Flores-Rentería
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Carrasco-Espinosa, K., Avitia, M., Santini, N.S. et al. Nutrient contents and microbial communities as mediators of the effects of land-use in ecosystem functioning in alpine ecosystems from Central Mexico. J Soils Sediments (2024). https://doi.org/10.1007/s11368-024-03804-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11368-024-03804-y