Abstract
With rapid urbanization and industrialization, rural rivers in China are facing deterioration in water quality and ecosystem health. Microorganisms living in river sediments are involved in biogeochemical processes, mineralization, and degradation of pollutants. Understanding bacterial community distribution in rural rivers could help evaluate the response of river ecosystems to environmental pollution and understand the river self-purification mechanism. In this study, the relationship between characteristics of sediment microbial communities and the surrounding environmental factors in a typical rural river was analyzed using 16S rRNA gene sequencing technology. The results showed that the dominant bacterial groups in the river sediment were Proteobacteria, Actinobacteria, Chloroflexi, Acidobacteria, Bacteroidetes, and Firmicutes, accounting for 83.61% of the total microbial load. Different areas have different sources of pollution which give rise to specific dominant bacteria. The upstream part of the river flows through an agricultural cultivation area where the dominant bacteria were norank_f_Gemmatimonadaceae, Haliangium, and Pseudolabrys, possessing obvious nitrogen- and phosphorus-metabolizing activities. The midstream section flows through an urban area where the dominant bacteria were Marmoricola, Nocardioides, Gaiella, Sphingomonas, norank_f_67-14, Subgroup_10, Agromyces, and Lysobacter, with strong metabolizing activity for toxic pollutants. The dominant bacteria in the downstream part were Clostridium_sensu_stricto_1, norank_f__Bacteroidetes_vadinHA17, Candidatus_Competibacter, and Methylocystis. Redundancy analysis and correlation heatmap analysis showed that environmental factors: ammonia nitrogen (NH4+–N) and total nitrogen (TN) in the sediment, and pH, temperature, TN, electrical conductivity (EC), and total dissolved solids (TDS) in the water, significantly affected the bacterial community in the sediment. The PICRUSt2 functional prediction analysis identified that the main function of bacteria in the sediment was metabolism (77.3%), specifically carbohydrate, amino acid, and energy metabolism. These activities are important for degrading organic matter and removing pollutants from the sediments. The study revealed the influence of organic pollutants derived from human activities on the bacterial community composition in the river sediments. It gave a new insight into the relationship between environmental factors and bacterial community distribution in rural watershed ecosystems, providing a theoretical basis for self-purification and bioremediation of rural rivers.










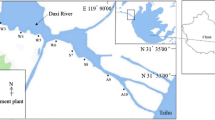
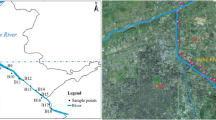
source regions in the upper, middle, and lower reaches of rivers. PICRUSt2 was used to identify the functional groups associated with level 3 functional categories
Similar content being viewed by others
Data availability
The datasets generated during and analyzed during the current study are available from the corresponding author on reasonable request.
References
Abhijit M, Sufia Z (2016) Marine ecosystem: an overview. Springer, India, pp 1–19. https://doi.org/10.1007/978-81-322-2707-6_1
Allgaier M, Grossart HP (2006) Diversity and seasonal dynamics of Actinobacteria populations in four lakes in northeastern Germany. Appl Environ Microbiol 72(5):3489–3497. https://doi.org/10.1128/AEM.72.5.3489-3497.2006
Beale DJ, Karpe AV, Ahmed W, Cook S, Morrison PD, Staley C, Sadowsky MJ, Palombo EA (2017) A community multi-omics approach towards the assessment of surface water quality in an urban river system. Int J Environ Res Public Health 14(3):303. https://doi.org/10.3390/ijerph14030303
Bordel S, Rojas A, Muñoz R (2019) Reconstruction of a genome scale metabolic model of the polyhydroxybutyrate producing methanotroph methylocystis parvus OBBP. Microb Cell Fact 18(1):104. https://doi.org/10.1186/s12934-019-1154-5
Boyer EW, Howarth RW, Galloway JN, Dentener FJ, Green PA, Vorosmarty CJ (2006) Riverine nitrogen export from the continents to the coasts. Glob Biogeochem Cycles 20(1):GB1S91
Cai W, Li Y, Wang P, Niu L, Zhang W, Wang C (2016) Revealing the relationship between microbial community structure in natural biofilms and the pollution level in urban rivers: a case study in the Qinhuai River basin, Yangtze River Delta. Water Sci Technol: A J Int Assoc Water Pollut Res 74(5):1163–1176. https://doi.org/10.2166/wst.2016.224
Chebbi A, Mnif S, Mhiri N, Jlaiel L, Sayadi S, Chamkha M (2014) A moderately thermophilic and mercaptan-degrading Bacillus licheniformis strain CAN55 isolated from gas-washing wastewaters of the phosphate industry, Tunisia. Int Biodeterior Biodegradation 94:207–213. https://doi.org/10.1016/j.ibiod.2014.08.001
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics (oxford, England) 34(17):i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Chu Z, Wang K, Li X, Zhu M, Yang L, Zhang J (2015) Microbial characterization of aggregates within a one-stage nitritation-anammox system using high-throughput amplicon sequencing. Chem Eng J 262:41–48. https://doi.org/10.1016/j.cej.2014.09.067
Cornejo-Granados F, Lopez-Zavala AA, Gallardo-Becerra L, Mendoza-Vargas A, Sánchez F, Vichido R, Brieba LG, Viana MT, Sotelo-Mundo RR, Ochoa-Leyva A (2017) Microbiome of Pacific whiteleg shrimp reveals differential bacterial community composition between wild, aquacultured and AHPND/EMS outbreak conditions. Sci Rep 7(1):11783. https://doi.org/10.1038/s41598-017-11805-w
Cottingham KL, Brown BL, Lennon JT (2001) Biodiversity may regulate the temporal variability of ecological systems. Ecol Lett 4(1):72–85. https://doi.org/10.1046/j.1461-0248.2001.00189.x
Dai T, Zhang Y, Tang Y, Bai Y, Tao Y, Huang B, Wen D (2016a) Identifying the key taxonomic categories that characterize microbial community diversity using full-scale classification: a case study of microbial communities in the sediments of Hangzhou Bay. FEMS Microbiol Ecol 92(10):fiw150. https://doi.org/10.1093/femsec/fiw150
Dai Y, Yang Y, Wu Z, Feng Q, Xie S, Liu Y (2016b) Spatiotemporal variation of planktonic and sediment bacterial assemblages in two plateau freshwater lakes at different trophic status. Appl Microbiol Biotechnol 100(9):4161–4175. https://doi.org/10.1007/s00253-015-7253-2
Drury B, Rosi-Marshall E, Kelly JJ (2013) Wastewater treatment effluent reduces the abundance and diversity of benthic bacterial communities in urban and suburban rivers. Appl Environ Microbiol 79(6):1897–1905. https://doi.org/10.1128/AEM.03527-12
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–998. https://doi.org/10.1038/nmeth.2604
Feng BW, Li XR, Wang JH, Hu ZY, Meng H, XiangLY QZX (2009) Bacterial diversity of water and sediment in the Changjiang estuary and coastal area of the East China Sea. FEMS Microbiol Ecol 70(2):80–92. https://doi.org/10.1111/j.1574-6941.2009.00772.x
Fu R, Cheng R, Wang S, Li J, Zhang J (2021) Succinoglycan Riclin reshaped the soil microbiota by accumulating plant probiotic species to improve the soil suppressiveness on Fusarium wilt of cucumber seedlings. Int J Biol Macromol 182:1883–1892. https://doi.org/10.1016/j.ijbiomac.2021.05.180
Fuhrman JA (2009) Microbial community structure and its functional implications. Nature 459(7244):193–199. https://doi.org/10.1038/nature08058
Goetghebuer L, Servais P, George IF (2017) Carbon utilization profiles of river bacterial strains facing sole carbon sources suggest metabolic interactions. FEMS Microbiol Lett 364(10): https://doi.org/10.1093/femsle/fnx098
Guo K, Wu N, Li W, Baattrup-Pedersen A, Riis T (2021) Microbial biofilm community dynamics in five lowland streams. Sci Total Environ 798:149169. https://doi.org/10.1016/j.scitotenv.2021.149169
Guo XP, Yang Y, Niu ZS, Lu DP, Zhu CH, Feng JN, Wu JY, Chen YR, Tou FY, Liu M, Hou L (2019) Characteristics of microbial community indicate anthropogenic impact on the sediments along the Yangtze Estuary and its coastal area, China. Sci Total Environ 648:306–314. https://doi.org/10.1016/j.scitotenv.2018.08.162
Gupta RS, Mok A (2007) Phylogenomics and signature proteins for the alpha Proteobacteria and its main groups. BMC Microbiol 7:106. https://doi.org/10.1186/1471-2180-7-106
Hahn MW, Lünsdorf H, Wu Q, Schauer M, Höfle MG, Boenigk J, Stadler P (2003) Isolation of novel ultramicrobacteria classified as actinobacteria from five freshwater habitats in Europe and Asia. Appl Environ Microbiol 69(3):1442–1451. https://doi.org/10.1128/AEM.69.3.1442-1451.2003
Hug LA, Castelle CJ, Wrighton KC, Thomas BC, Sharon I, Frischkorn KR, Williams KH, Tringe SG, Banfield JF (2013) Community genomic analyses constrain the distribution of metabolic traits across the Chloroflexi phylum and indicate roles in sediment carbon cycling. Microbiome 1(1):22. https://doi.org/10.1186/2049-2618-1-22
Jia W, Yang YC, Yang LY, Gao Y (2021) High-efficient nitrogen removal and its microbiological mechanism of a novel carbon self-sufficient constructed wetland. Sci Total Environ 775:145901. https://doi.org/10.1016/j.scitotenv.2021.145901
Kaiser K, Canedo-Oropeza M, McMahon R, Amon R (2017) Origins and transformations of dissolved organic matter in large Arctic rivers. Sci Rep 7(1):13064. https://doi.org/10.1038/s41598-017-12729-1
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, KnightR BeikoRG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814–821. https://doi.org/10.1038/nbt.2676
Li Q, Zhao Y, Zhang X, Wei YQ, Qiu LL, Wei ZM (2015) Spatial heterogeneity in a deep artificial lake plankton community revealed by PCR-DGGE fingerprinting. Chin J Oceanol Limnol 33(3):624–635. https://doi.org/10.1007/s00343-015-4184-9
Li Y, Yang Z (2020) Soil nutrients and soul microbial characteristics and their relationship under different land use types in Minjiang River Basin. Res Soil Water Conserv 27(1):33–46 ((In Chinese))
Liao H, Yu K, Duan Y, Ning Z, Li B, He L, Liu C (2019) Profiling microbial communities in a watershed undergoing intensive anthropogenic activities. Sci Total Environ 647:1137–1147. https://doi.org/10.1016/j.scitotenv.2018.08.103
Liu J, Chen X, Shu HY, Lin XR, Zhou QX, Bramryd T, Shu WS, Huang LN (2018) Microbial community structure and function in sediments from e-waste contaminated rivers at Guiyu area of China. Environ Pollut 235:171–179. https://doi.org/10.1016/j.envpol.2017.12.008
Liu L, Yang J, Yu X, Chen G, Yu Z (2013) Patterns in the composition of microbial communities from a subtropical river: effects of environmental, spatial and temporal factors. PLoS One 8(11):e81232. https://doi.org/10.1371/journal.pone.0081232
Liu S, Ren H, Shen L, Lou L, Tian G, Zheng P, Hu B (2015) pH levels drive bacterial community structure in sediments of the Qiantang River as determined by 454 pyrosequencing. Front Microbiol 6:285. https://doi.org/10.3389/fmicb.2015.00285
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics (oxford, England) 27(21):2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Matias MG, Combe M, Barbera C, Mouquet N (2013) Ecological strategies shape the insurance potential of biodiversity. Front Microbiol 3:432. https://doi.org/10.3389/fmicb.2012.00432
Nakkabi A, Sadiki M, Fahim M, Ittobane N, Koraichi SI, Barkai H, El Abed S (2015) Biodegradation of poly (ester urethane)s by Bacillus subtilis. Int J Environ Res 9(1):157–162. https://doi.org/10.4028/www.scientific.net/MSF.494.297
Ni M, Chen Y, Pan Y, Huang Y, Li DP, Li L, Huang B, Song Z (2022) Study on community structure and metabolic mechanism of dominant polyphosphate-accumulating organisms (PAOs) and glycogen-accumulating organisms (GAOs) in suspended biofilm based on phosphate recovery. Sci Total Environ 815:152678. https://doi.org/10.1016/j.scitotenv.2021.152678
Numberger D, Ganzert L, Zoccarato L, Mühldorfer K, Sauer S, Grossart HP, Greenwood AD (2019) Characterization of bacterial communities in wastewater with enhanced taxonomic resolution by full-length 16S rRNA sequencing. Sci Rep 9(1):9673. https://doi.org/10.1038/s41598-019-46015-z
Ouyang L, Chen H, Liu X, Wong MH, Xu F, Yang X, Xu W, Zeng Q, Wang W, Li S (2020) Characteristics of spatial and seasonal bacterial community structures in a river under anthropogenic disturbances. Environ Pollut 264:114818. https://doi.org/10.1016/j.envpol.2020.114818
Pascault N, Roux S, Artigas J, Pesce S, Leloup J, Tadonleke RD, Debroas D, Bouchez A, Humbert JF (2014) A high-throughput sequencing ecotoxicology study of freshwater bacterial communities and their responses to tebuconazole. FEMS Microbiol Ecol 90(3):563–574. https://doi.org/10.1111/1574-6941.12416
Patowary K, Saikia RR, Kalita MC et al (2015) Degradation of polyaromatic hydrocarbons employing biosurfactant-producing Bacillus pumilus KS2. Ann Microbiol 65:225–234. https://doi.org/10.1007/s13213-014-0854-7
Pearce DA (2005) The structure and stability of the bacterioplankton community in Antarctic freshwater lakes, subject to extremely rapid environmental change. FEMS Microbiol Ecol 53(1):61–72. https://doi.org/10.1016/j.femsec.2005.01.002
Puopolo G, Tomada S, Pertot I (2018) The impact of the omics era on the knowledge and use of Lysobacter species to control phytopathogenic micro-organisms. J Appl Microbiol 124(1):15–27. https://doi.org/10.1111/jam.13607
Puopolo G, Tomada S, Sonego P, Moretto M, Engelen K, Perazzolli M, Pertot I (2016) The Lysobacter capsici AZ78 genome has a gene pool enabling it to interact successfully with phytopathogenic microorganisms and environmental factors. Front Microbiol 7:96. https://doi.org/10.3389/fmicb.2016.00096
Qi M, Liang B, Zhang L, Ma X, Yan L, Dong W, Kong D, Zhang L, Zhu H, Gao SH, Jiang J, Liu SJ, Corvini PF, Wang A (2021) Microbial interactions drive the complete catabolism of the antibiotic sulfamethoxazole in activated sludge microbiomes. Environ Sci Technol 55(5):3270–3282. https://doi.org/10.1021/acs.est.0c06687
Qian-Qian Z, Sheng-Long J, Ke-Mao L, Zhen-Bing W, Hong-Tao G, Jin-Wen H, Shu-Yi W, Yao-Yao L, Guo-Jie W, Ai-Hua L (2021) Community structure of bacterioplankton and its relationship with environmental factors in the upper reaches of the Heihe River in Qinghai Plateau. Environ Microbiol 23(2):1210–1221. https://doi.org/10.1111/1462-2920.15358
Qiu C, Bao Y, Petropoulos E, Wang Y, Zhong Z, Jiang Y, Ye X, Lin X, Feng Y (2022) Organic and inorganic amendments shape bacterial indicator communities that can, in turn, promote rice yield. Microorganisms 10(2):482. https://doi.org/10.3390/microorganisms10020482
Qiu W, Sun J, Fang M, Luo S, Tian Y, Dong P, Xu B, Zheng C (2019) Occurrence of antibiotics in the main rivers of Shenzhen, China: association with antibiotic resistance genes and microbial community. Sci Total Environ 653:334–341. https://doi.org/10.1016/j.scitotenv.2018.10.398
Reddy GV, Reddy BR, Tlou MG (2014) Biodegradation of 2-hydroxyquinoxaline (2-HQ) by Bacillus sp. J Hazard Mater 278:100–107. https://doi.org/10.1016/j.jhazmat.2014.05.080
Ren C, Teng Y, Chen X, Shen Y, Xiao H, Wang H (2021) Impacts of earthworm introduction and cadmium on microbial communities composition and function in soil. Environ Toxicol Pharmacol 83:103606. https://doi.org/10.1016/j.etap.2021.103606
Roberto AA, Van Gray JB, Leff LG (2018) Sediment bacteria in an urban stream: spatiotemporal patterns in community composition. Water Res 134:353–369. https://doi.org/10.1016/j.watres.2018.01.045
Shah RM, Crosswell J, Metcalfe SS, Carlin G, Morrison PD, Karpe AV, Palombo EA, Steven ADL, Beale DJ (2019) Influence of human activities on broad-scale estuarine-marine habitats using omics-based approaches applied to marine sediments. Microorganisms 7(10):419. https://doi.org/10.3390/microorganisms7100419
Shah RM, Stephenson S, Crosswell J, Gorman D, Hillyer KE, Palombo EA, Jones O, Cook S, Bodrossy L, van de Kamp J, Walsh TK, Bissett A, Steven A, Beale DJ (2022) Omics-based ecosurveillance uncovers the influence of estuarine macrophytes on sediment microbial function and metabolic redundancy in a tropical ecosystem. Sci Total Environ 809:151175. https://doi.org/10.1016/j.scitotenv.2021.151175
Shen JF, Lin LJ (2017) State-sponsored and spontaneous urbanization in Fujian province of China, 1982–2010. Cities 60:420–427. https://doi.org/10.1016/j.cities.2016.05.021
Shen CC, Xiong JB, Zhang HY, Feng YZ, Lin X, Li XG, Li XY, Liang WJ, Chu HY (2013) Soil pH drives the spatial distribution of bacterial communities along elevation on Changbai Mountain. Soil Biol and Biochem 57:204–211. https://doi.org/10.1016/j.soilbio.2012.07.013
Sheng P, Yu Y, Zhang G, Huang J, He L, Ding J (2016) Bacterial diversity and distribution in seven different estuarine sediments of Poyang Lake, China. Environ Earth Sci 75(6):479. https://doi.org/10.1007/s12665-016-5346-6
Shi X, Ng KK, Li XR, Ng HY (2015) Investigation of intertidal wetland sediment as a novel inoculation source for anaerobic saline wastewater treatment. Environ Sci Technol 49(10):6231–6239. https://doi.org/10.1021/acs.est.5b00546
SIEPS (2017) Water standard scheme of Minjiang Yuelai ferry control unit in Meishan City. Sichuan Institute of Environmental Protection Science, Chengdu (In Chinese)
Singha LP, Pandey P (2021) Rhizosphere assisted bioengineering approaches for the mitigation of petroleum hydrocarbons contamination in soil. Crit Rev Biotechnol 41(5):749–766. https://doi.org/10.1080/07388551.2021.1888066
Song X, Yu D, Qiu Y, Qiu C, Xu L, Zhao J, Wang X (2022) Unexpected phosphorous removal in a Candidatus_Competibacter and Defluviicoccus dominated reactor. Biores Technol 345:126540. https://doi.org/10.1016/j.biortech.2021.126540
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44(4):846–849. https://doi.org/10.1099/00207713-44-4-846
Suzuki M, Tachibana Y, Takizawa R, Morikawa T, Takeno H, Kasuya K (2021) A novel poly(3-hydroxybutyrate)-degrading actinobacterium that was isolated from plastisphere formed on marine plastic debris. Polym Degrad Stab 183:109461. https://doi.org/10.1016/j.polymdegradstab.2020.109461
Vlassi A, Nesler A, Perazzolli M, Lazazzara V, Büschl C, Parich A, Puopolo G, Schuhmacher R (2020) Volatile organic compounds from Lysobacter capsici AZ78 as potential candidates for biological control of soilborne plant pathogens. Front Microbiol 11:1748. https://doi.org/10.3389/fmicb.2020.01748
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73(16):5261–5267. https://doi.org/10.1128/AEM.00062-07
Wu HC, Chen PC, Tsay TT (2010) Assessment of nematode community structure as a bioindicator in river monitoring. Environ Pollut (Baking, Essex: 1987) 158(5):1741–1747. https://doi.org/10.1016/j.envpol.2009.11.015
Wu Y, Wang YC, Hu MM, Liu DF, Huang YL, Li DH, Xu T (2015) Ecological distribution and its influencing factors of bacterioplankton in the typical tributaries of Three Gorges Reservoir. Chin J Ecol (04): pp 1060–1065. https://doi.org/10.13292/j.1000-4890.20150304.019
Xiao Y, Chen S, Gao Y, Hu W, Hu M, Zhong G (2015) Isolation of a novel beta-cypermethrin degrading strain Bacillus subtilis BSF01 and its biodegradation pathway. Appl Microbiol Biotechnol 99(6):2849–2859. https://doi.org/10.1007/s00253-014-6164-y
Xiong W, Sun Y, Zhang T, Ding X, Li Y, Wang M, Zeng Z (2015) Antibiotics, antibiotic resistance genes, and bacterial community composition in fresh water aquaculture environment in China. Microb Ecol 70(2):425–432. https://doi.org/10.1007/s00248-015-0583-x
Xue L, Ren H, Li S, Leng X, Yao X (2017) Soil bacterial community structure and co-occurrence pattern during vegetation restoration in karst rocky desertification area. Front Microbiol 8:2377. https://doi.org/10.3389/fmicb.2017.02377
Yamada T, Sekiguchi Y (2009) Cultivation of uncultured chloroflexi subphyla: significance and ecophysiology of formerly uncultured chloroflexi ‘subphylum i’ with natural and biotechnological relevance. Microbes Environ 24(3):205–216. https://doi.org/10.1264/jsme2.me09151s
Yang F, Zeng X, Ning K, Liu KL, Lo CC, Wang W, Chen J, Wang D, Huang R, Chang X, Chain PS, Xie G, Ling J, Xu J (2012) Saliva microbiomes distinguish caries-active from healthy human populations. ISME J 6(1):1–10. https://doi.org/10.1038/ismej.2011.71
Yang L, Shin HS, Hur J (2014) Estimating the concentration and biodegradability of organic matter in 22 wastewater treatment plants using fluorescence excitation emission matrices and parallel factor analysis. Sensors (basel, Switzerland) 14(1):1771–1786. https://doi.org/10.3390/s140101771
Yang Y, Gao Y, Huang X, Ni P, Wu Y, Deng Y, Zhan A (2019) Adaptive shifts of bacterioplankton communities in response to nitrogen enrichment in a highly polluted river. Environ Pollut 245:290–299. https://doi.org/10.1016/j.envpol.2018.11.002
Zhao L, Zhao Z, Zhang K, Zhang X, Xu S, Liu J, Liu B, Hong Q, Qiu J, He J (2021) Cotinine hydroxylase CotA initiates biodegradation of wastewater micropollutant cotinine in Nocardioides sp. strain JQ2195. Appl Environ Microbiol 87(18):e0092321. https://doi.org/10.1128/AEM.00923-21
Zhao M, Sun PF, Du LN, Wang G, Jia XM, Zhao YH (2014) Biodegradation of methyl red by Bacillus sp. strain UN2: decolorization capacity, metabolites characterization, and enzyme analysis. Environ Sci Pollut Res Int 21(9):6136–6145. https://doi.org/10.1007/s11356-014-2579-3
Zhuang M, Sanganyado E, Li P, Liu W (2019) Distribution of microbial communities in metal-contaminated nearshore sediment from Eastern Guangdong, China. Environ Pollut 250:482–492. https://doi.org/10.1016/j.envpol.2019.04.041
Acknowledgements
This study is supported by the National Natural Science Foundation of China (Grant no. 52170104; Grant no. 51979237), the Science and Technology Program of Sichuan Province (2021YJ0382), and the Fundamental Research Funds for the Central Universities (2682021CX066).
Author information
Authors and Affiliations
Contributions
Cheng Yang selected the topic, designed the research methodology, collected and sorted the data, and wrote the first draft. Han Zhang and Zhuo Zeng selected the topic, designed the research methodology, and reviewed and finalized the manuscript. Dongdong Gao collected and sorted the data. Guangyi He and Yuanyuan Wang designed the research methodology. Ying Liu, Yan Wang, and Xinyu Du analyzed the data. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yang, C., Zeng, Z., Zhang, H. et al. Distribution of sediment microbial communities and their relationship with surrounding environmental factors in a typical rural river, Southwest China. Environ Sci Pollut Res 29, 84206–84225 (2022). https://doi.org/10.1007/s11356-022-21627-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-21627-7