Abstract
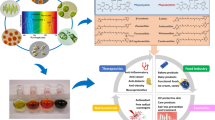
Cyanobacteria are ubiquitous photosynthetic prokaryotes responsible for the oxygenation of the earth’s reducing atmosphere. Apart from oxygen they are producers of a myriad of bioactive metabolites with diverse complex chemical structures and robust biological activities. These secondary metabolites are known to have a variety of medicinal and therapeutic applications ranging from anti-microbial, anti-viral, anti-inflammatory, anti-cancer, and immunomodulating properties. The present review discusses various aspects of secondary metabolites viz. biosynthesis, types and applications, which highlights the repertoire of bioactive constituents they harbor. Majority of these products have been produced from only a handful of genera. Moreover, with the onset of various OMICS approaches, cyanobacteria have become an attractive chassis for improved secondary metabolites production. Also the intervention of synthetic biology tools such as gene editing technologies and a variety of metabolomics and fluxomics approaches, used for engineering cyanobacteria, have significantly enhanced the production of secondary metabolites.

Data source CyanoMetDB



Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
References
Acuña UM, Zi J, Orjala J et al (2015) Ambiguine I isonitrile from Fischerella ambigua induces caspase-independent cell death in MCF-7 hormone dependent breast cancer cells. Int J Cancer Res 49:1655–1662
Acuña UM, Mo S, Zi J et al (2018) Hapalindole H induces apoptosis as an inhibitor of NF-KB and affects the intrinsic mitochondrial pathway in PC-3 androgen-insensitive prostate cancer cells. Anticancer Res 38:3299–3307. https://doi.org/10.21873/anticanres.12595
Anand N, Thajuddin N, Dadheech PK (2019) Cyanobacterial taxonomy: morphometry to molecular studies. In: Mishra AK, Tiwari DN, Rai AN (eds) Cyanobacteria: from basic science to applications. Academic Press, Cambridge. https://doi.org/10.1016/B978-0-12-814667-5.00003-9
Anzalone AV, Randolph PB, Davis JR et al (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157. https://doi.org/10.1038/s41586-019-1711-4
Babić O, Kovač D, Rašeta M et al (2016) Evaluation of antioxidant activity and phenolic profile of filamentous terrestrial cyanobacterial strains isolated from forest ecosystem. J Appl Phycol 28:2333–2342. https://doi.org/10.1007/s10811-015-0773-4
Bansal R, Pachauri S, Gururajaiah D et al (2021) Dual role of a dedicated GAPDH in the biosynthesis of volatile and non-volatile metabolites-novel insights into the regulation of secondary metabolism in Trichoderma virens. Microbiol Res 253:126862. https://doi.org/10.1016/j.micres.2021.126862
Behler J, Vijay D, Hess WR et al (2018) CRISPR-based technologies for metabolic engineering in cyanobacteria. Trends Biotechnol 36:996–1010. https://doi.org/10.1016/j.tibtech.2018.05.011
Bilal M, Rasheed T, Ahmed I et al (2017) High value compounds from microalgae with industrial exploitability—a review. Front Biosci 9:319–342
Blagojević D, Babić O, Rašeta M et al (2018) Antioxidant activity and phenolic profile in filamentous cyanobacteria: the impact of nitrogen. J Appl Phycol 30:2337–2346. https://doi.org/10.1007/s10811-018-1476-4
Burragoni SG, Jeon J (2021) Applications of endophytic microbes in agriculture, biotechnology, medicine, and beyond. Microbiol Res 245:126691. https://doi.org/10.1016/j.micres.2020.126691
Cabanillas AH, Pérez VT, Corral SM et al (2018) Cybastacines A and B: antibiotic sesterterpenes from a Nostoc sp. cyanobacterium. J Nat Prod 81:410–413. https://doi.org/10.1021/acs.jnatprod.7b00638
Cagide E, Becher PG, Louzao MC et al (2014) Hapalindoles from the cyanobacterium Fischerella: potential sodium channel modulators. Chem Res Toxicol 27:1696–1706. https://doi.org/10.1021/tx500188a
Calteau A, Fewer DP, Latifi A et al (2014) Phylum-wide comparative genomics unravel the diversity of secondary metabolism in Cyanobacteria. BMC Genomics 15:977–990. https://doi.org/10.1186/1471-2164-15-977
Cardllina JH II, Moore RE, Arnold EV et al (1979) Structure and absolute configuration of malyngolide, an antibiotic from the marine blue–green alga Lyngbya majuscula Gomont. J Org Chem 44:4039–4042. https://doi.org/10.1021/jo01337a003
Carpine R, Sieber S (2021) Antibacterial and antiviral metabolites from cyanobacteria: their application and their impact on human health. Cur Res Biotechnol 3:65–81. https://doi.org/10.1016/j.crbiot.2021.03.001
Carroll AL, Case AE, Zhang A, Atsumi S (2018) Metabolic engineering tools in model cyanobacteria. Metab Eng 50:47–56. https://doi.org/10.1016/j.ymben.2018.03.014
Castro I, Costa H, Turgeman-Grott I et al (2021) The lanthipeptide biosynthetic clusters of the domain Archaea. Microbiol Res 253:126884. https://doi.org/10.1016/j.micres.2021.126884
Cegłowska M, Szubert K, Wieczerzak E et al (2020) Eighteen new Aeruginosamide variants produced by the Baltic cyanobacterium Limnoraphis CCNP1324. Mar Drugs 18:1–13. https://doi.org/10.3390/md18090446
Chakdar H, Hasan M, Pabbi S et al (2021) High-throughput proteomics and metabolomic studies guide re-engineering of metabolic pathways in eukaryotic microalgae: a review. Bioresour Technol 321:124495. https://doi.org/10.1016/j.biortech.2020.124495
Chakdar H, Pabbi S (2016) Cyanobacterial phycobilins: production, purification, and regulation. In: Shukla P (ed) Frontier discoveries and innovations in interdisciplinary microbiology. Springer, New Delhi. https://doi.org/10.1007/978-81-322-2610-9_4
Chakdar H, Pabbi S (2017) Algal pigments for human health and cosmeceuticals. In: Rastogi RP et al (eds) Algal green chemistry: recent progress in biotechnology. Elsevier, Amsterdam, pp 171–188. https://doi.org/10.1016/B978-0-444-63784-0.00009-6
Chandra R, Pons-Faudoa FP, Saldívar RP, Rittmann BE (2020) Effect of ultra-violet exposure on production of mycosporine-like amino acids and lipids by Lyngbya purpurem. Biomass Bioenergy 134:1–7. https://doi.org/10.1016/j.biombioe.2020.105475
Chatterjee S, Ghosh R, Mandal NC (2020) Inhibition of biofilm-and hyphal-development, two virulent features of candida albicans by secondary metabolites of an endophytic fungus alternaria tenuissima having broad spectrum antifungal potential. Microbiol Res 232:126386. https://doi.org/10.1016/j.micres.2019.126386
Cheah YE, Xu Y, Sacco SA et al (2020) Systematic identification and elimination of flux bottlenecks in the aldehyde production pathway of Synechococcus elongatus PCC 7942. Metab Eng 60:56–65. https://doi.org/10.1016/j.ymben.2020.03.007
Chekan JR, Fallon TR, Moore BS (2020) Biosynthesis of marine toxins. Curr Opin Chem Biol 59:119–129. https://doi.org/10.1016/j.cbpa.2020.06.009
Chen L, Deng H, Cui H et al (2018) Inflammatory responses and inflammation-associated diseases in organs. Oncotarget 9:7204–7218. https://doi.org/10.18632/oncotarget.23208
Choi SY, Woo HM (2020) CRISPRi-dCas12a: a dCas12a-mediated CRISPR interference for repression of multiple genes and metabolic engineering in cyanobacteria. ACS Synth Biol 9:2351–2361. https://doi.org/10.1021/acssynbio.0c00091
Choi H, Engene N, Smith JE et al (2010) Crossbyanols A−D, toxic brominated polyphenyl ethers from the hawai’ian bloom-forming cyanobacterium leptolyngbya crossbyana. J Nat Prod 73:517–522. https://doi.org/10.1021/np900661g
Choi YH, Yang DJ, Kulkarni A et al (2015) Mycosporine-like amino acids promote wound healing through focal adhesion kinase (FAK) and mitogen-activated protein kinases (MAP Kinases) signaling pathway in keratinocytes. Mar Drugs 13:7055–7066. https://doi.org/10.3390/md13127056
Christiansen G, Fastner J, Erhard M et al (2003) Microcystin biosynthesis in planktothrix: genes, evolution, and manipulation. J Bacteriol 185:564–572. https://doi.org/10.1128/JB.185.2.564-572.2003
Daboussi F, Leduc S, Marechal A et al (2014) Genome engineering empowers the diatom phaeodactylum tricornutum for biotechnology. Nat Commun 5:1–7. https://doi.org/10.1038/ncomms4831
Demay J, Bernard C, Reinhardt A, Marie B (2019) Natural products from cyanobacteria: focus on beneficial activities. Mar Drugs 17:320–368. https://doi.org/10.3390/md17060320
Dittmann E, Gugger M, Sivonen K, Fewer DP (2015) Natural product biosynthetic diversity and comparative genomics of the cyanobacteria. Trends Microbiol 23:642–652. https://doi.org/10.1016/j.tim.2015.07.008
Dixit RB, Suseela MR (2013) Cyanobacteria: potential candidates for drug discovery. Antonie Van Leeuwenhoek 103:947–961. https://doi.org/10.1007/s10482-013-9898-0
Dos Santos DS, Rosa ME, Zanatta AP et al (2019) Neurotoxic effects of sublethal concentrations of cyanobacterial extract containing anatoxin-a(s) on nauphoeta cinerea cockroaches. Ecotoxicol Environ Saf 171:138–145. https://doi.org/10.1016/j.ecoenv.2018.12.068
Edwards DJ, Gerwick WH (2004) Lyngbyatoxin biosynthesis: sequence of biosynthetic gene cluster and identification of a novel aromatic prenyltransferase. J Am Chem Soc 126:11432–11433. https://doi.org/10.1021/ja047876g
Engene N, Gunasekera SP, Gerwick WH et al (2013) Phylogenetic inferences reveal a large extent of novel biodiversity in chemically rich tropical marine cyanobacteria. Appl Environ Microbiol 79:1882–1888. https://doi.org/10.1128/AEM.03793-12
Fagundes MB, Alvarez-Rivera G, Mendiola JA et al (2021) Phytosterol-rich compressed fluids extracts from phormidium autumnale cyanobacteria with neuroprotective potential. Algal Res 55:1–13. https://doi.org/10.1016/j.algal.2021.102264
Férir G, Huskens D, Noppen S et al (2014) Broad anti-HIV activity of the oscillatoria agardhii agglutinin homologue lectin family. J Antimicrob Chemother 69:2746–2758. https://doi.org/10.1093/jac/dku220
Fuentes-Tristan S, Parra-Saldivar R, Iqbal HMN, Carrillo-Nieves D (2019) Bioinspired biomolecules: mycosporine-like amino acids and scytonemin from lyngbya sp. with UV-protection potentialities. J Photochem Photobiol B Biol 201:1–32. https://doi.org/10.1016/j.jphotobiol.2019.111684
Furmaniak MA, Misztak AE, Franczuk MD et al (2017) Edible cyanobacterial genus arthrospira: actual state of the art in cultivation methods, genetics, and application in medicine. Front Microbiol 8:1–21. https://doi.org/10.3389/fmicb.2017.02541
Gaj T, Gersbach CA, Barbas CF III (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
Gao X, Xu H, Zhu Z et al (2020) Improved production of echinenone and canthaxanthin in transgenic nostoc sp PCC 7120 overexpressing a heterologous crtO gene from Nostoc flagelliforme. Microbiol Res. https://doi.org/10.1016/j.micres.2020.126455
Garrison AR, Giomarelli BG, Lear-Rooney CM et al (2014) The cyanobacterial lectin scytovirin displays potent in vitro and in vivo activity against zaire ebola virus. Antivir Res 112:1–7. https://doi.org/10.1016/j.antiviral.2014.09.012
Gordon GC, Korosh TC, Cameron JC et al (2016) CRISPR interference as a titratable, trans-acting regulatory tool for metabolic engineering in the cyanobacterium Synechococcus sp. strain PCC 7002. Metab Eng 38:170–179. https://doi.org/10.1016/j.ymben.2016.07.007
Guerreiro A, Andrade MA, Menezes C et al (2020) Antioxidant and cytoprotective properties of cyanobacteria: potential for biotechnological applications. Toxins 12:548–565. https://doi.org/10.3390/toxins12090548
Höckelmann C, Becher PG, von Reuss SH, Jüttner F (2009) Sesquiterpenes of the geosmin-producing cyanobacterium calothrix PCC 7507 and their toxicity to invertebrates. Z Naturforsch C 64:49–55. https://doi.org/10.1515/znc-2009-1-209
Hohlman RM, Sherman DH (2021) Recent advances in hapalindole-type cyanobacterial alkaloids: biosynthesis, synthesis, and biological activity. Nat Prod Rep. https://doi.org/10.1039/D1NP00007A
Jones MR, Pinto E, Torres MA et al (2021) CyanoMetDB, a comprehensive public database of secondary metabolites from cyanobacteria. Water Res 196:1–12. https://doi.org/10.1016/j.watres.2021.117017
Jose PA, Maharshi A, Jha B (2021) Actinobacteria in natural products research: progress and prospects. Microbiol Res 246:126708. https://doi.org/10.1016/j.micres.2021.126708
Kaczmarzyk D, Cengic I, Yao L, Hudson EP (2018) Diversion of the long-chain acyl-ACP pool in Synechocystis to fatty alcohols through CRISPRi repression of the essential phosphate acyltransferase PlsX. Metab Eng 45:59–66. https://doi.org/10.1016/j.ymben.2017.11.014
Kaloyannidis P, Hertzberg M, Webb K et al (2020) Brentuximab vedotin for the treatment of patients with relapsed or refractory hodgkin lymphoma after autologous stem cell transplantation. Br J Haematol 188:540–549. https://doi.org/10.1111/bjh.16201
Kamennaya NA, Ahn S, Park H et al (2015) Installing extra bicarbonate transporters in the cyanobacterium synechocystis sp. PCC6803 enhances biomass production. Metab Eng 29:76–85. https://doi.org/10.1016/j.ymben.2015.03.002
Kanno M, Carroll AL, Atsumi S (2017) Global metabolic rewiring for improved CO2 fixation and chemical production in cyanobacteria. Nat Commun 8:1–11. https://doi.org/10.1038/ncomms14724
Kaplon H, Muralidharan M, Schneider Z, Reichert JM (2020) Antibodies to watch in 2020. Mabs 12:1703531. https://doi.org/10.1080/19420862.2019.1703531
Kapuścik A, Hrouzek P, Kuzma M et al (2013) Novel aeruginosin-865 from nostoc sp. as a potent anti-infammatory agent. ChemBioChem 14:2329–2337. https://doi.org/10.1002/cbic.201300246
Kawaguchi M, Satake M, Zhang BT et al (2020) Neo-aplysiatoxin a isolated from okinawan cyanobacterium moorea producens. Molecules 25:457–463. https://doi.org/10.3390/molecules25030457
Kehr J-C, GattePicchi D, Ditmann E (2011) Natural product biosyntheses in cyanobacteria: a treasure trove of unique enzymes. Beilstein J Org Chem 7:1622–1635. https://doi.org/10.3762/bjoc.7.191
Keller L, Canuto KM, Liu C et al (2020) Tutuilamides A–C: vinyl-chloride-containing cyclodepsipeptides from marine cyanobacteria with potent elastase inhibitory properties. ACS Chem Biol 15:751–757. https://doi.org/10.1021/acschembio.9b00992
Khatri Y, Hohlman RM, Mendoza J et al (2020) Multicomponent microscale biosynthesis of unnatural cyanobacterial indole alkaloids. ACS Synth Biol 9:1349–1360. https://doi.org/10.1021/acssynbio.0c00038
Kim Tiam S, Gugger M, Demay J et al (2019) Insights into the diversity of secondary metabolites of planktothrix using a biphasic approach combining global genomics and metabolomics. Toxins 11:498. https://doi.org/10.3390/toxins11090498
Kitagaki S, Murata S, Asaoka K et al (2018) Planar chiral [2.2] paracyclophane-based bisoxazoline ligands: design, synthesis, and use in Cu-catalyzed inter and intramolecular asymmetric O–H insertion reactions. Chem Pharm Bull 66:1006–1014. https://doi.org/10.1248/cpb.c18-00519
Kiuru P, D’Auria MV, Muller CD et al (2014) Exploring marine resources for bioactive compounds. Planta Med 80:1234–1246. https://doi.org/10.1055/s-0034-1383001
Kleigrewe K, Almaliti J, Tian IY et al (2015) Combining mass spectrometric metabolic profiling with genomic analysis: a powerful approach for discovering natural products from cyanobacteria. J Nat Prod 78:1671–1682. https://doi.org/10.1021/acs.jnatprod.5b00301
Kłodawska K, Bujas A, Turos-Cabal M et al (2019) Effect of growth temperature on biosynthesis and accumulation of carotenoids in cyanobacterium anabaena sp. PCC 7120 under diazotrophic conditions. Microbiol Res 226:34–40. https://doi.org/10.1016/j.micres.2019.05.003
Komor AC, Kim YB, Packer MS et al (2016) Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533:420–424. https://doi.org/10.1038/nature17946
Kurmayer R, Deng L, Entfellner E (2016) Role of toxic and bioactive secondary metabolites in colonization and bloom formation by filamentous cyanobacteria planktothrix. Harmful Algae 54:69–86. https://doi.org/10.1016/j.hal.2016.01.004
Li S, Lowell AN, Yu F et al (2015) Hapalindole/ambiguine biogenesis is mediated by a cope rearrangement, C–C bond forming cascade. J Am Chem Soc 137:15366–15369. https://doi.org/10.1021/jacs.5b10136
Li L, Maclntyre LW, Brady SF (2021a) Refactoring biosynthetic gene clusters for heterologous production of microbial natural products. Curr Opin Biotechnol 69:145–152. https://doi.org/10.1016/j.copbio.2020.12.011
Li X, Huang L, Pan L et al (2021b) CRISPR/dCas9-mediated epigenetic modification reveals differential regulation of histone acetylation on aspergillus niger secondary metabolite. Microbiol Res 245:126694. https://doi.org/10.1016/j.micres.2020.126694
Liang Y, Hou J, Liu Y et al (2018) Textile dye decolorizing synechococcus PCC7942 engineered with CotA laccase. Front Bioeng Biotechnol 6:00095. https://doi.org/10.3389/fbioe.2018.00095
Lipinska-Zubrycka L, Klewicki R, Sojka M et al (2020) Anticandidal activity of Lactobacillus spp. in the presence of galactosyl polyols. Microbiol Res. https://doi.org/10.1016/j.micres.2020.126540
Liu D, Pakrasi HB (2018) Exploring native genetic elements as plug-in tools for synthetic biology in the cyanobacterium synechocystis sp. PCC 6803. Microb Cell Fact 17:1–8. https://doi.org/10.1186/s12934-018-0897-8
Liu L, Jokela J, Herfindal L (2014) 4-Methylproline guided natural product discovery: co-occurrence of 4-Hydroxy and 4-methylprolines in nostoweipeptins and nostopeptolides. ACS Chem Biol 9:2646–2655. https://doi.org/10.1021/cb500436p
Liu X, Hillwig ML, Koharudin LMI, Gronenborn AM (2016) Unified biogenesis of ambiguine, fischerindole, hapalindole and welwitindolinone: identification of a monogeranylated indolenine as a cryptic common biosynthetic intermediate by an unusual magnesium-dependent aromatic prenyltransferase. Chem Commun 52:1737–1740. https://doi.org/10.1039/C5CC10060G
Lopes G, Clarinha D, Vasconcelos V (2020) Carotenoids from cyanobacteria: a biotechnological approach for the topical treatment of psoriasis. Microorganisms 8:302–320. https://doi.org/10.3390/microorganisms8020302
Ma AT, Schmidt CM, Golden JW (2014) Regulation of gene expression in diverse cyanobacterial species by using theophylline-responsive riboswitches. Appl Environ Microbiol 80:6704–6713. https://doi.org/10.1128/AEM.01697-14
Manogar P, Vijayakumar S, Praseetha PK (2020) Evaluation of antioxidant and neuroprotective activities of lyngbya majuscula on human neural tissues. Gene Rep 19:1–5. https://doi.org/10.1016/j.genrep.2020.100661
Manzo E, Cutignano A, Pagano D et al (2017) A new marine-derived sulfoglycolipid triggers dendritic cell activation and immune adjuvant response. Sci Rep 7:1–10. https://doi.org/10.1038/s41598-017-05969-8
Martins J, Leikoski N, Wahlsten M et al (2018) Sphaerocyclamide, a prenylated cyanobactin from the cyanobacterium sphaerospermopsis sp. LEGE 00249. Sci Rep 8:1–9. https://doi.org/10.1038/s41598-018-32618-5
May DS, Crnkovic CM, Krunic A et al (2020) 15N stable isotope labeling and comparative metabolomics facilitates genome mining in cultured cyanobacteria. ACS Chem Biol 15:758–765. https://doi.org/10.1021/acschembio.9b00993
Méjean A, Mann S, Maldiney T et al (2009) Evidence that biosynthesis of the neurotoxic alkaloids anatoxin-a and homoanatoxin-a in the cyanobacterium oscillatoria PCC 6506 occurs on a modular polyketide synthase initiated by L-proline. J Am Chem Soc 131:7512–7513. https://doi.org/10.1021/ja9024353
Méjean A, Paci G, Gautier V, Ploux O (2014) Biosynthesis of anatoxin-a and analogues (anatoxins) in cyanobacteria. Toxicon 91:15–22. https://doi.org/10.1016/j.toxicon.2014.07.016
Mironenka J, Różalska S, Soboń A, Bernat P (2021) Trichoderma harzianum metabolites disturb fusarium culmorum metabolism: metabolomic and proteomic studies. Microbiol Res 249:126770. https://doi.org/10.1016/j.micres.2021.126770
Montaser R, Abboud KA, Paul VJ, Luesch H (2011) Pitiprolamide, a proline-rich dolastatin 16 analogue from the marine cyanobacterium lyngbya majuscula from guam. J Nat Prod 74:109–112. https://doi.org/10.1021/np1006839
Morone J, Alfeus A, Vasconcelos V, Martins R (2019) Revealing the potential of cyanobacteria in cosmetics and cosmeceuticals—a new bioactive approach. Algal Res 41:781–789. https://doi.org/10.1016/j.algal.2019.101541
Motoyama K, Tanida Y, Hata K et al (2016) Anti-inflammatory effects of novel polysaccharide sacran extracted from cyanobacterium aphanothece sacrum in various inflammatory animal models. Biol Pharm Bull 39:1172–1178. https://doi.org/10.1248/bpb.b16-00208
Myronovskyi M, Luzhetskyy A (2016) Native and engineered promoters in natural product discovery. Nat Prod Rep 33:1006–1019. https://doi.org/10.1039/C6NP00002A
Na D, Lee D (2010) RBSDesigner: software for designing synthetic ribosome binding sites that yields a desired level of protein expression. Bioinf 26:2633–2634. https://doi.org/10.1093/bioinformatics/btq458
Nainangu P, Antonyraj AP, Subramanian K et al (2020) In vitro screening of antimicrobial, antioxidant, cytotoxic activities, and characterization of bioactive substances from freshwater cyanobacteria oscillatoria sp. SSCM01 and phormidium sp. SSCM02. Biocatal Agric Biotechnol 29:1–11. https://doi.org/10.1016/j.bcab.2020.101772
Nakahira Y, Ogawa A, Asano H et al (2013) Theophylline-dependent riboswitch as a novel genetic tool for strict regulation of protein expression in cyanobacterium synechococcus elongatus PCC 7942. Plant Cell Physiol 5:1724–1735. https://doi.org/10.1093/pcp/pct115
Nazar R, Mohamed MI, Yousuff TN, Dharumadurai D (2018) Cyanobacterial toxins as biolarvicides for blood-sucking vectors. In: Tayagi BK, Dhanasekaran D (eds) Microbial control of vector-borne diseases. CRC Press, Boca Raton
Neuhof T, Schmieder P, Seibold M et al (2006) Hassallidin B-second antifungal member of the hassallidin family. Bioorg Med Chem Lett 16:4220–4222. https://doi.org/10.1016/j.bmcl.2006.05.094
Nguyen CP (2010) Synthetic studies toward scytophycin C. Ph.D. Thesis, The Scripps Research Institute, Jupiter, Florida
Niu TC, Lin GM, Xie LR et al (2018) Expanding the potential of CRISPR-Cpf1-based genome editing technology in the cyanobacterium anabaena PCC 7120. ACS Synth Biol 8:170–180. https://doi.org/10.1021/acssynbio.8b00437
Nowruzi B, Haghighat S, Fahimi H, Mohammadi E (2018) Nostoc cyanobacteria species: a new and rich source of novel bioactive compounds with pharmaceutical potential. J Pharm Health Serv Res 9:5–12. https://doi.org/10.1111/jphs.12202
Nowruzi B, Sarvari G, Blanco S (2020) Applications of cyanobacteria in biomedicine. In: Konur O (ed) Handbook of algal science, technology and medicine. Academic Press, London, pp 441–455. https://doi.org/10.1016/B978-0-12-818305-2.00028-0
Ogawa H, Iwasaki A, Sumimoto S et al (2017) Isolation and total synthesis of hoshinolactam, an antitrypanosomal lactam from a marine cyanobacterium. Org Lett 19:890–893. https://doi.org/10.1021/acs.orglett.7b00047
Oliver JWK, Atsumi S (2014) Metabolic design for cyanobacterial chemical synthesis. Photosynth Res 120:249–261. https://doi.org/10.1007/s11120-014-9997-4
Ongley SE, Bian X, Zhang Y et al (2013) High-titer heterologous production in E. coli of lyngbyatoxin, a protein kinase C activator from an uncultured marine cyanobacterium. ACS Chem Biol 8:1888–1893. https://doi.org/10.1021/cb400189j
Pagarete A, Ramos AS, Puntervoll P et al (2021) Antiviral potential of algal metabolites—a comprehensive review. Mar Drugs 19:1–23. https://doi.org/10.3390/md19020094
Parajuli A, Kwak DH, Dalponte L et al (2016) A unique tryptophan C-prenyltransferase from the kawaguchipeptin biosynthetic pathway. Angew Chem Int Ed 55:3596–3599. https://doi.org/10.1002/anie.201509920
Park SH, Lee K, Jang JW, Hahn JS (2019) Metabolic engineering of saccharomyces cerevisiae for production of shinorine, a sunscreen material, from xylose. ACS Synth Biol 8:346–357. https://doi.org/10.1021/acssynbio.8b00388
Patanaik B, Lindberg P (2015) Terpenoids and their biosynthesis in cyanobacteria. Life 5:269–293. https://doi.org/10.3390/life5010269
Patel VK, Sundaram S, Patel AK, Kalra A (2018) Characterization of seven species of cyanobacteria for high-quality biomass production. Arab J Sci Eng 43:109–121. https://doi.org/10.1007/s13369-017-2666-0
Pathak J, Sonker AS, Richa R et al (2017) Screening and partial purification of photoprotective pigment, scytonemin from cyanobacteria dominated biological crusts dwelling on the historical monuments in and around Varanasi, India. Microbiol Res 8:4–12. https://doi.org/10.4081/mr.2017.6559
Pattharaprachayakul N, Lee M, Incharoensakdi A, Woo HM (2020) Current understanding of the cyanobacterial CRISPR-Cas systems and development of the synthetic CRISPR-Cas systems for cyanobacteria. Enzyme Microb Technol 140:1–10. https://doi.org/10.1016/j.enzmictec.2020.109619
Pearson LA, Dittmann E, Mazmouz R et al (2016) The genetics, biosynthesis and regulation of toxic specialized metabolites of cyanobacteria. Harmful Algae 54:98–111. https://doi.org/10.1016/j.hal.2015.11.002
Pereira AR, Cao ZY, Engene N et al (2010) Palmyrolide A, an unusually stabilized neuroactive macrolide from Palmyra Atoll yanobacteria. Org Lett 12:4490–4493. https://doi.org/10.1021/ol101752n
Pérez VT, Ticona LA, Cabanillas AH et al (2020) Anti-inflammatory activity of glycolipids isolated from cyanobacterium nodularia harveyana. Nat Prod Res 27:1–6. https://doi.org/10.1080/14786419.2020.1837808
Pettit GR, Kamano Y, Herald CL et al (1987) The isolation and structure of a remarkable marine animal antineoplastic constituent: dolastatin 10. J Am Chem Soc 109:6883–6885. https://doi.org/10.1021/ja00256a070
Pipal M, Priebojova J, Koci T et al (2020) Field cyanobacterial blooms producing retinoid compounds cause teratogenicity in zebrafish embryos. Chemosphere 241:1–14. https://doi.org/10.1016/j.chemosphere.2019.125061
Ratnayake R, Gunasekera SP, Ma JJ et al (2020) Dolastatin 15 from a marine cyanobacterium suppresses HIF-1α mediated cancer cell viability and vascularization. ChemBioChem 21:2356–2366. https://doi.org/10.1002/cbic.202000180
Righini H, Francioso O, Martel Quintana A, Roberti R (2022) Cyanobacteria: a natural source for controlling agricultural plant diseases caused by fungi and oomycetes and improving plant growth. Hortic 8:58. https://doi.org/10.3390/horticulturae8010058
Rippka R, Deruelles J, Waterbury JB et al (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol 111:1–61. https://doi.org/10.1099/00221287-111-1-1
Rutkowska M, Płotka-Wasylka J, Majchrzak T et al (2019) Recent trends in determination of neurotoxins in aquatic environmental samples. Trends Anal Chem 112:112–122. https://doi.org/10.1016/j.trac.2019.01.001
Saad MH, El-Fakharany EM, Salem MS, Sidkey NM (2020) The use of cyanobacterial metabolites as natural medical and biotechnological tools. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2020.1838948
Saber AA, Hamed SM, Abdel-Rahim EF, Cantonati M (2018) Insecticidal prospects of algal and cyanobacterial extracts against the cotton leafworm spodoptera littoralis. Life Environ 68:199–212
Saha S, Esposito G, Urajová P et al (2020) Discovery of unusual cyanobacterial tryptophan-containing anabaenopeptins by MS/MS-based molecular networking. Molecules 25:3786. https://doi.org/10.3390/molecules25173786
Saini DK, Chakdar H, Pabbi S, Shukla P (2020) Enhancing production of microalgal biopigments through metabolic and genetic engineering. Crit Rev Food Sci Nutr 60:391–405. https://doi.org/10.1080/10408398.2018.1533518
Salis HM, Mirsky EA, Voigt CA (2009) Automated design of synthetic ribosome binding sites to control protein expression. Nat Biotechnol 27:946–950. https://doi.org/10.1038/nbt.1568
Salwan R, Sharma V (2020) Molecular and biotechnological aspects of secondary metabolites in actinobacteria. Microbiol Res 231:126374. https://doi.org/10.1016/j.micres.2019.126374
Sathasivam R, Radhakrishnan R, Hashem A, Abdallah EF (2019) Microalgae metabolites: a rich source for food and medicine. Saudi J Biol Sci 26:709–722. https://doi.org/10.1016/j.sjbs.2017.11.003
Scaife MA, Smith AG (2016) Towards developing algal synthetic biology. Biochem Soc Trans 44:716–722. https://doi.org/10.1042/BST20160061
Sebesta J, Peebles CAM (2020) Improving heterologous protein expression in Synechocystis sp. PCC 6803 for alpha-bisabolene production. Metab Eng Commun. https://doi.org/10.1016/j.mec.2019.e00117
Sengupta A, Madhu S, Wangikar PP (2020) A Library of tunable, portable, and inducer-free promoters derived from cyanobacteria. ACS Synth Biol 9:1790–1801. https://doi.org/10.1021/acssynbio.0c00152
Seo SW, Yang JS, Cho HS et al (2014) Predictive combinatorial design of mRNA translation initiation regions for systematic optimization of gene expression levels. Sci Rep 4:1–7. https://doi.org/10.1038/srep04515
Shah SAA, Akhter N, Auckloo BN et al (2017) Structural diversity, biological properties and applications of natural products from cyanobacteria. A review. Mar Drugs 15:1–30. https://doi.org/10.3390/md15110354
Shao J, Peng L, Luo S et al (2013) First report on the allelopathic effect of tychonema bourrellyi (cyanobacteria) against microcystis aeruginosa (cyanobacteria). J Appl Phycol 25:1567–1573. https://doi.org/10.1007/s10811-012-9969-z
Shih PM, Wu D, Latifi A, Axen SD et al (2013) Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. Proc Natl Acad Sci USA 110:1053–1058. https://doi.org/10.1073/pnas.1217107110
Sigamani S, Ramamurthy D, Natarajan H (2016) A review on potential biotechnological applications of microalgae. J App Pharm Sci 6:179–184. https://doi.org/10.7324/JAPS.2016.60829
Simpkin VL, Renwick MJ, Kelly R, Mossialos E (2017) Incentivising innovation in antibiotic drug discovery and development: progress, challenges and next steps. J Antibiot 70:1087–1096. https://doi.org/10.1038/ja.2017.124
Singh U, Singh P, Singh AK (2021) Identification of antifungal and antibacterial biomolecules from a cyanobacterium, arthrospira platensis. Algal Res 54:1–7. https://doi.org/10.1016/j.algal.2021.102215
Singh DP, Prabha R, Verma S et al (2017) Antioxidant properties and polyphenolic content in terrestrial cyanobacteria. 3 Biotech 7:1–14. https://doi.org/10.1007/s13205-017-0786-6
Singh SK, Kaur R, Bansal A et al (2020) Biotechnological exploitation of cyanobacteria and microalgae for bioactive compounds. In: Verma M, Chandel A (eds) Biotechnological production of bioactive compounds. Elsevier, Amsterdam, pp 221–258. https://doi.org/10.1016/B978-0-444-64323-0.00008-4
Siqueira AS, Lima AR, Aguiar DC et al (2021) Genomic screening and molecular dynamics simulations of cyanovirin-N homologs from cyanobacteria phylum. Proteins Struct Funct Bioinf 89:322–329. https://doi.org/10.1002/prot.26017
Sivonen K, Leikoski N, Fewer DP, Jokela J (2010) Cyanobactins-ribosomal cyclic peptides produced by cyanobacteria. Appl Microbiol Biotechnol 86:1213–1225. https://doi.org/10.1007/s00253-010-2482-x
Sizova I, Greiner A, Awasthi M et al (2013) Nuclear gene targeting in Chlamydomonas using engineered zinc-finger nucleases. Plant J 73:873–882. https://doi.org/10.1111/tpj.12066
Spolaore P, Joannis-Cassan C, Duran E, Isambert A (2006) Commercial applications of microalgae. J Biosci Bioeng 101:87–96. https://doi.org/10.1263/jbb.101.87
Stincone P, Brandelli A (2020) Marine bacteria as source of antimicrobial compounds. Crit Rev Biotechnol 40:306–319. https://doi.org/10.1080/07388551.2019.1710457
Storme J-Y, Golubic S, Wilmotte A et al (2015) Raman characterization of the UV-protective pigment gloeocapsin and its role in the survival of cyanobacteria. Astrobiol 15:843–857. https://doi.org/10.1089/ast.2015.1292
Sun L, Liu LP, Wang YZ (2022) Effect of ultrasonication on the metabolome and transcriptome profile changes in the fermentation of Ganoderma lucidum. Microbiol Res 254:126916. https://doi.org/10.1016/j.micres.2021.126916
Sweeney-Jones AM, Gagaring K, Antonova-Koch J et al (2020) Antimalarial peptide and polyketide natural products from the Fijian marine cyanobacterium Moorea producens. Mar Drugs 18:167–180. https://doi.org/10.3390/md18030167
Szwajgier D (2013) Anticholinesterase activity of phenolic acids and their derivatives. Z Naturforsch C 68:125–132. https://doi.org/10.1515/znc-2013-3-408
Tabarzad M, Atabaki V, Hosseinabadi T (2020) Anti-inflammatory activity of bioactive compounds from microalgae and cyanobacteria by focusing on the mechanisms of action. Mol Biol Rep 47:6193–6205. https://doi.org/10.1007/s11033-020-05562-9
Taiz L, Zeiger E (2006) Plant physiology, 4th edn. Sinauer Associate, Sunderland
Taton A, Ecker A, Diaz B et al (2020) Heterologous expression of cryptomaldamide in a cyanobacterial host. ACS Synth Biol 9:3364–3376. https://doi.org/10.1021/acssynbio.0c00431
Tsai SQ, Wyvekens N, Khayter C et al (2014) Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat Biotechnol 32:569–576. https://doi.org/10.1038/nbt.2908
Ungerer J, Pakrasi HB (2016) Cpf1 is a versatile tool for CRISPR genome editing across diverse species of cyanobacteria. Sci Rep 6:39681. https://doi.org/10.1038/srep39681
Vasudevan R, Gale GA, Schiavon AA et al (2019) CyanoGate: A modular cloning suite for engineering cyanobacteria based on the plant MoClo syntax. Plant Physiol 180:39–55. https://doi.org/10.1104/pp.18.01401
Vestola J, Shishido TK, Jokela J et al (2014) Hassallidins, antifungal glycolipopeptides, are widespread among cyanobacteria and are the end-product of a nonribosomal pathway. Proc Natl Acad Sci USA 111:E1909–E1917. https://doi.org/10.1073/pnas.1320913111
Videau P, Wells KN, Singh AJ et al (2016) Assessment of anabaena sp. strain PCC 7120 as a heterologous expression host for cyanobacterial natural products: production of lyngbyatoxin A. ACS Syn Biol 5:978–988. https://doi.org/10.1021/acssynbio.6b00038
Vining OB, Medina RA, Mitchell EA et al (2015) Depsipeptide companeramides from a panamanian marine cyanobacterium associated with the coibamide producer. J Nat Prod 78:413–420. https://doi.org/10.1021/np5007907
Walton KE (2012) Identification, isolation, and characterization of developmental toxins from the cyanobacterium Fischerella 52-1 using the Zebrafish (Danio rerio) embryo model. Dissertation, Florida International University, FIU Electronic Theses and Dissertations, 645. https://doi.org/10.25148/etd.FI12050207
Walton K, Berry JP (2016) Indole alkaloids of the stigonematales (cyanophyta): chemical diversity, biosynthesis and biological activity. Mar Drugs 14:1–28. https://doi.org/10.3390/md14040073
Wang X, Liu W, Xin C et al (2016) Enhanced limonene production in cyanobacteria reveals photosynthesis limitations. Proc Natl Acad Sci USA 113:14225–14230. https://doi.org/10.1073/pnas.1613340113
Wang M, Zhang J, He S, Yan X (2017) A review study on macrolides isolated from cyanobacteria. Mar Drugs 15:126–144. https://doi.org/10.3390/md15050126
Wang B, Li X, Tabudravu J et al (2021a) The chemical profile of activated secondary metabolites by overexpressing LaeA in Aspergillus niger. Microbiol Res 248:126735. https://doi.org/10.1016/j.micres.2021.126735
Wang H, Liang J, Yue Q et al (2021b) Engineering the acyltransferase domain of epothilone polyketide synthase to alter the substrate specificity. Microb Cell Fact 20:1–14. https://doi.org/10.1186/s12934-021-01578-3
Wendt KE, Ungerer J, Cobb RE et al (2016) CRISPR/Cas9 mediated targeted mutagenesis of the fast growing cyanobacterium Synechococcus elongatus UTEX 2973. Microb Cell Fact 15:115. https://doi.org/10.1186/s12934-016-0514-7
Wu C, Chen W, Chen J et al (2015) Preparation of monoPEGylated cyanovirin-N’s derivative and its anti-influenza A virus bioactivity in vitro and in vivo. J Biochem 157:539–548. https://doi.org/10.1093/jb/mvv013
Xu X, Liu Y, Du G, Liu L (2020) Systems biology, synthetic biology, and metabolic engineering. In: Liu Y et al (eds) Systems and synthetic metabolic engineering. Elsevier, Amnsterdam, pp 1–31. https://doi.org/10.1016/B978-0-12-821753-5.00001-0
Yang G, Cozad MA, Holland DA et al (2018) Photosynthetic production of sunscreen shinorine using an engineered cyanobacterium. ACS Synth Biol 7:664–671. https://doi.org/10.1021/acssynbio.7b00397
Zeng C, Wen B, Hou G et al (2017) Lipidomics profiling reveals the role of glycerophospholipid metabolism in psoriasis. Gigascience 6:1–11. https://doi.org/10.1093/gigascience/gix087
Acknowledgements
The authors are grateful to Director, ICAR-NBAIM, Mau for providing the infrastructural facilities and financial assistance to carry out the Ph.D. programme. This study is a part of the Ph.D. work of the first author under supervision of the corresponding author as co-supervisor. The authors are thankful to Dr. Pratyoosh Shukla, Professor, School of Biotechnology, Banaras Hindu University, India for his constructive comments during manuscript preparation.
Funding
No funds, grants or other support were received for this study.
Author information
Authors and Affiliations
Contributions
HC Conceptualization, Supervision, Visualization, Writing—Review & Editing. SV Investigation, Preparation of original draft, Writing—Review & Editing. ST Investigation, Preparation of original draft, Writing—Review & Editing. NS Writing—Review & Editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors’ declare that they have no known competing interest, financial or proprietary interest that could have appeared to influence the work discussed in this article.
Ethical approval
The authors’ declare that ethical approval is not applicable in this study.
Informed consent
The author’s declare informed consents are not applicable to this study.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Research involved in human and/or animal participants
The author’s declare human or animal rights are not applicable to this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Verma, S., Thapa, S., Siddiqui, N. et al. Cyanobacterial secondary metabolites towards improved commercial significance through multiomics approaches. World J Microbiol Biotechnol 38, 100 (2022). https://doi.org/10.1007/s11274-022-03285-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03285-6