Abstract
The native vs. exotic status of reed canarygrass (RCG), a major invasive species of Minnesota wetlands, is unknown. The aim of this study was to investigate this native vs. exotic status to enhance its management. Genetic comparison of wild RCG populations from six Minnesota and six Czech Republic rivers was performed. A total of 2521 polymorphic SNP markers (single nucleotide polymorphisms) were used to evaluate 478 RCG samples across all collections. In the PCoA, all (n = 256) tested extant wild, riparian RCG genotypes from six Minnesota Rivers and six Czech Republic Rivers were genetically distinct, although some SNPs were common in both populations since they are the same species. DAPC analysis also resulted in the formation of two primary clusters separating the Minnesota Rivers and Czech Republic Rivers riparian samples, with little overlap; STRUCTURE analysis also supported this clustering with k = 4 groups as it separated the Czech Republic Rivers populations into three groups, along with Minnesota Rivers. The uniformity of PCoA, DAPC, STRUCTURE, and Evanno results indicates the distinct separation of Minnesota Rivers and Czech Republic Rivers populations. Portions of the genome (specific SNPs) are preserved or in common across continents, as indicated by STRUCTURE similarities. Nonetheless, overall significant SNP differences between the continents indicate that the Minnesota riparian populations are distinct enough from the European (Czech) collections to be delineated as native N. American RCG. PCoA of all the Minnesota RCG collections clustered Minnesota Rivers, Herbarium, Extant Herbarium, Research Field and Native Field collections together. STRUCTURE analysis (k = 2; Evanno) divided these Minnesota collections from the Commercial Field and Cultivars collections. There are two genetically distinct groups of RCG in Minnesota and since the Minnesota Rivers, the Research Field, the Native Field and pre-1930 herbaria collections clustered together, they are most likely native N. American types. Analysis of molecular variance (AMOVA) indicated that the genetic variation was more significant within, rather than among, the RCG populations. Native, historic herbaria types cluster together with all wild RCG river populations in Minnesota, all of which were distinct from those in Central Europe, suggesting native RCG type persistence in N. America. Also, cultivated forage types of RCG are distinct from wild RCG Minnesota river populations. The SNP genetic data shows that riparian Minnesota RCG populations are native. These data will facilitate future management strategies to control RCG as a native, but invasive, species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The challenges of establishing reed canarygrass (RCG; Phalaris arundinacea L.) as a native vs. exotic in North America are paramount as management priorities would change once the status were known (Anderson et al. 2021). Differential shifts in land management would vary, depending on where the species occurs, e.g. Tribal Land Managers would be interested in controlling exotic genotypes while preserving natives (if this is economically feasible), whereas State or Provincial Departments of Natural Resources and private agencies may choose to control aggressive, invasive populations regardless of their native/exotic status (Anderson et al. 2021). RCG is a perennial, wind-pollinated, cool-season wetland grass that is cultivated in temperate regions around the globe as a forage, bedding straw for livestock, ornamental crop, as well as for soil remediation, waste water treatments and biofuels (Olsen and Chong 1991; Galatowitsch et al. 1999; Samecka-Cymerman and Kempers 2001; Chekol et al. 2002; Lewandowski et al. 2003; Lavergne and Molofsky 2004; Sheaffer et al. 2008). RCG is an invasive species of wetlands and, in restoration efforts, it commonly prevents colonization for revegetation (Reinhardt and Galatowitsch 2004). In Europe, RCG is a common species in riparian habitats (Ambros and Štykar 1999) and wet meadows (Hroudová et al. 2009) but is classified as an archaeophyte (pre-Roman Empire era) rather than invasive (Anderson 2019). RCG is considered to be native to N. America in some instances (Piper 1914; Schoth 1929) while others postulate that it is an exotic, invasive varietas from Eurasia (Lavergne and Molofsky 2004). More recently, extant (current) N. American populations are considered to be a mixture of native N. American and European types or varietas (Lavergne and Molofsky 2007) or the products of recent RCG breeding programs, with cultivars outcompeting native populations (Merigliano and Lesica 1998). Likewise, RCG's exotic origin is considered to be a cause of its invasiveness in the State of Minnesota (USA) and wetlands elsewhere (Lavergne and Molofsky 2004).
The earliest indication (pre-1800s) that RCG is native to N. America is the use of this grass by Native Americans to weave traditional basketry items, fishing weirs, and thatch wigwam roofs (Turner v 1980; Densmore 2012). Additional evidence to support the native status of RCG comes from early plant collections (1825 to 1911) from the inland Northwest, USA (Merigliano and Lesica 1998). Schoth (1929) reports that RCG "was found from the New England States westward to the Pacific coast and as far as Tennessee" while others state that all RCG were native (Apfelbaum and Sams 1987). Herbarium RCG specimens indicated the presence of this species before and during Euro-American settlement across the N. American continent (Merigliano and Lesica 1998). Herbarium specimens were used as a tool to study past and current plant invasions (Wu et al. 2005) and proposed as a way to determine native vs. exotic status (Crawford et al. 2009). Molecular markers (SSRs or single sequence repeats) were used by Jakubowski et al. (2013) to assess the native vs. exotic status of RCG herbaria collections from N. America. Their RCG herbaria collection was compared to extant European RCG samples. Native N. American RCG populations were also discovered in Ontario, Canada, and remote areas elsewhere (Dore and McNeill 1980; LaVoie et al. 2005). Early American maps by Verendrye in 1737, Thompson in 1814, Long in 1823, and Pope in 1849 used the term Roseau (French for "reed") for the northern Minnesota river by that name (Prud’homme 1916; Northwest Regional Development Commission 2014). Two "reed-like" grasses, RCG and Phragmites communis, co-occur along the Roseau River, MN which is called "Ga-shashagunushkokawi-sibi" or "the-place-of-rushes-river" in Ojibwe (Northwest Regional Development Commission 2014). It was recently learned that an unplowed, pristine field in Roseau, MN was used to harvest hay during the Dust Bowl era (1930s) which was transported on the existing Constitutional Highways and sold throughout the Midwest (Magnusson 2012; Anderson 2019). This field, a segment of the ancient Lake Agassiz lakebed, has never been plowed, and remains 100% RCG to date (Anderson 2019).
The first reports of cultivated RCG in Europe are 1749 (Sweden), 1824 (England), and 1850 (Germany; Alway 1931). While some early cultivation in America could have been from European seed stocks, production on the west coast in the Coquille Valley (Coos Co., Oregon) in 1885, was from varietas most likely of native origin (Schoth 1929). The first commercial, low alkaloid cultivars, e.g. 'Venture' and 'Palaton' (Alderson and Sharp 1994), are the result of crosses among 'Flare', 'Vantage', 'Rise', and other (mostly wild) germplasm collections from Minnesota and Iowa (Fig. 1).
The pedigree of modern reed canarygrass forage cultivars, ‘Palaton’, ‘Venture’ and ‘Vantage’. Development of these cultivars in the USA spans the period from early 1900 to 1976. ‘Vantage’ was created from seed collected in Iowa and Minnesota and was released in 1972 (Alderson and Sharp 1994). ‘Palaton’ and ‘Venture’ are results of crosses of early cultivars (such as ‘Flare’, ‘Vantage’, ‘Rise’) and other germplasm accessions). Commercial forage cultivation of both cultivars started in 1985, with very limited release before this date (Alderson and Sharp 1994). ‘Palaton’ and ‘Venture’ are considered the most advanced forage types with very low levels of alkaloids
Genetic tests were inconclusive in showing whether invasive P. arundinacea were prevalent with European cultivars that escaped cultivation (Gifford et al. 2002) and neutral allozyme markers unique to French and Czech Republic P. arundinacea co-occurred with invasive N. American populations (87% shared diversity; Lavergne and Molofsky 2007). More precise genetic markers established genetic relationships among diverse sets of RCG populations. One of the first attempts to establish native vs. exotic status of RCG used amplified fragment length polymorphism (AFLP) markers to analyze landraces and improved cultivars from Europe and N. America (Casler et al. 2009). Jakubowski et al. (2013) is one of the first to report on genetically distinct native N. American populations of RCG, using herbaria samples (collection dates of 1875–2000) for comparison with those that were pre-1800, which were used as a benchmark for native N. American types. Later, 373 RCG accessions analyzed with 15 microsatellite markers showed that native populations of RCG were still present in N. America (Jakubowski et al. 2014). However, the majority of extant RCG samples from N. America clustered with those from Europe and Asia (Jakubowski et al. 2014). STRUCTURE analysis of inter-simple sequence repeat (ISSR) markers from N. American and European samples showed the best fit to two clusters (k = 2), separating most N. American and European samples (Nelson et al. 2014). The wild varietas, forage, ornamental exotic varietas and North American RCG populations harbored a high amount of genetic diversity within, as opposed to among populations. Thus, range expansion of reed canarygrass in North America was a not result of hybridization among exotic, forage, and native varietas (Nelson et al. 2014) despite previous theories to that effect (Lavergne and Molofsky 2007). More recent research using ISSRs on N. America (Minnesota) wild and cultivated samples showed two slightly overlapping groups separating cultivated samples from wild (Kávová et al. 2017). Anderson et al. (2016) showed that, based on ISSR markers, all European ornamental cultivars were distinct from wild Czech Republic populations (varietas) while the Czech forage 'Chrastava' was similar to wild types.
The State of Minnesota has the highest concentration of RCG in continental N. America, due partly to the large number of freshwater lakes (11,842 which are > 4.05 ha or > 10 acres; 21,871 which are > 1.01 ha or > 2.5 acres in size), natural rivers and streams (6564; Minnesota's waters flow outward in three directions: North to Hudson Bay in Canada, East to the Atlantic Ocean, South to the Gulf of Mexico; many feed into the Mississippi River which originates within the state), and wetlands (many of which were drained and tiled for commercial agriculture; Minnesota. Dept. of Natural Resources, n.d.; 1968). Roseau, MN is the site for production of seed of RCG forage cultivars and birdseed (P. canariensis; Anonymous 2012; Kávová et al. 2017; Anderson 2019). In the last century, RCG has spread widely throughout the state and land managers have been continuously confronted with its control as an invasive exotic species (Galatowitsch et al. 1999). However, given the growing body of evidence that many populations of N. American RCG are native varietas and not exotic, it is important to examine the status of this species in Minnesota. For comparative purposes, a large central European set of genotypes were included from the Czech Republic to determine whether Minnesota populations were genetically distinct.
The objectives of this study were to: (1) establish the native vs. exotic status of RCG in riparian populations of Minnesota by comparing germplasm collection from six dominant Minnesota and Czech Republic rivers (wild, riparian populations); (2) use herbaria specimens of RCG as a benchmark of its native presence in Minnesota; (3) evaluate Minnesota riparian (river) collections in comparison to commercial Minnesota forage cultivars and other wild (non-riparian) RCG collections to determine their native or exotic status. Associated null hypotheses tested were, respectively: (1) Ho = There is no difference among Minnesota and Czech Republic wild riparian (river) populations of RCG; (2) Ho = There is no difference among historic (herbaria) RCG accessions and Minnesota riparian populations; (3) Ho = There is no difference among Minnesota riparian populations, forage cultivars and wild (non-riparian) populations.
Material and methods
Plant collections
Herbaria RCG leaf samples, many of which were previously sampled for AFLP or ISSR/SSR analyses (Jakubowski et al. 2013; Nelson et al. 2014; Kávová et al. 2017), were collected from the University of Minnesota Herbarium (Bell Museum of Natural History, St. Paul, MN). This herbarium collection represents early North American RCG germplasm that is highly likely to be native to Minnesota (Merigliano and Lesica 1998; LaVoie et al. 2005). A sampling of the RCG herbarium collection (n = 17; Table 1) was selected to represent the earliest possible, native collections in Minnesota, along with additional earlier and more recent samples (Table 1). The historic cutoff for these native collections was the dust bowl era (< 1940), based on the shipment of RCG throughout the Midwest from Roseau, MN during this era (Anderson 2019). Herbaria samples fell into three categories, based on the time period of specimen collection: earliest possible (< 1940)—likely native (based on Jakubowski et al. 2013; Nelson et al. 2014; Kávová et al. 2017; Anderson 2019), after shipments started (> 1940; of questionable origin), and recent (of questionable origin, likely containing some exotics). Destructive sampling of about 2.5 cm × 0.5 cm of preserved (dry) mature leaf tip tissue using the least visible leaf in order to preserve the original specimen’s visual integrity and maintain their historic value as type specimens. Samples are termed hereafter the “Herbarium” collection.
Herbarium samples were supplemented with extant RCG collections that were in the same locations of each historic herbarium sample (Tables 1, 2), called hereafter the “Extant Herbarium” collection. Extant genotypes were found at n = 5 sites and a total of n = 60 genotypes collected for comparative analyses with the Minnesota Herbarium and Minnesota Rivers populations.
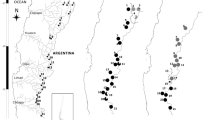
Leaf tissue samples of riparian RCG from six major Minnesota rivers (Mississippi, Minnesota, St. Croix, Red, Des Moines, Roseau; Fig. 2a) were collected every 30 km along each river, similarly to previous RCG collection methodology (Anderson et al. 2016). The headwaters of the Mississippi river originate in Minnesota and empty to the Gulf Basin; the Des Moines and Minnesota Rivers originate in Minnesota while the St. Croix River has headwaters in the adjacent State of Wisconsin (Fig. 2a). The Roseau and Red Rivers, also originating in Minnesota, flow north into Manitoba, Canada, and empty into the Lake Winnipeg / Hudson Bay watershed. Managed collected sites were avoided to minimize the potential impact of disturbance on the integrity of extant RCG populations. A total of 180 samples (n = 70 locations) were used (Table 3). This set of RCG is termed the “Minnesota Rivers” collection.
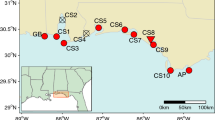
Locations of riparian, wet meadow or field reed canarygrass sample collections used in this study. All riparian collection sites were 30 km apart at undisturbed or unmanaged locations. a Minnesota, USA riparian populations from six rivers as well as extant herbarium populations (cf. Tables 2, 3); a research field at the Horticultural Research Center; and Roseau, MN populations and their distance apart (km), enlarged to show the precise locations of additional sympatric RCG collections sites (see inset). Three separate collections were made at this location: Native Field—this location was used to harvest hay for cattle feed during the Dust Bowl era (< 1930) which was shipped across the Midwest to dairy and beef cattle farms; this field is a segment of the Lake Agassiz lakebed (currently the Roseau Lake Wild Management Area, WMA), has never been plowed and remains 100% RCG as a pristine field; the adjacent and sympatric Roseau River RCG collection site; a commercial field for forage cultivar seed production, where ‘Palaton’, ‘Venture’ and ‘Vantage’ were grown for seed and hay (Anderson 2019). b Czech Republic
Leaf tissue of RCG collected from this location at the Horticultural Research Center, Chanhassen, MN, represents a mixed stand of the most likely RCG grass types similar to older forage cultivars such as ‘Rise’, but not the newer low-alkaloid types (Schaeffer 2019). This germplasm collection should be most likely similar to the wild RCG germplasm collection due to relatively minor germplasm advancement when compared to newer, low alkaloid ‘Palaton’ and ‘Venture’. Two perpendicular transects were run through the center of the population, with collection of three plant samples every 10 m along each transect with each of the samples being 10 m apart starting on the transect line and extending to the right and left thereof, respectively. A total of 56 samples were collected for this study. Samples from this location are termed the “Research Field” collection (Table 4).
The commercial field for forage seed production was located to the southwest of the set of transects of old Lake Agassiz lakebed in Roseau, MN USA (an unplowed circular field; Fig. 2a inset and Table 4). This location should include RCG commercially grown genotypes such as ‘Vantage’. Possibly numerous, old, native RCG genotypes have germinated at this location (Christenson 2012) (Anderson 2019). A total of 61 samples were extracted for this study. Samples from this location are termed the “Commercial Field” collection (Fig. 2a inset; Table 4).
Three advanced and closely related RCG forage cultivars were grown from the USDA GRIN (U.S. Dept. of Agriculture, Germplasm Resources Information Network; https://npgsweb.ars-grin.gov/gringlobal/search.aspx?): ‘Palaton’ (PI 531088), ‘Venture’ (PI 531089), and ‘Vantage’ (PI 578794; Table 4). Seeds were sown in 10 cm square pots filled with Sungro Professional Growing Mix (Sun Gro Horticulture; SKU:5105, Agawam, MA) and placed in a mist house for germination. Germinated seedlings were grown in a greenhouse with a 24.4 ± 3.0/18.3 ± 1.5 °C day/night daily interval and a 16 h long day photoperiod (0600–2200 HR) lighting (400 W high pressure sodium-high intensity discharge lamps, HPS-HID) set at a minimum of 150 μmol m−2 s−1 (Abbey et al. 2019). Both greenhouses were located in the St. Paul campus Plant Growth Facilities (University of Minnesota, St. Paul, MN USA) as previously described (Abbey et al. 2019). Plants were fertilized 2×/day, between 0700 and 0800 HR and 1600–1700 HR, using a constant liquid feed (CLF) of 125 ppm N from water-soluble 20N–4.4P–16.6K (Scotts, Marysville, OH). Fungicide drenches were applied in monthly rotations. Leaf sampling occurred ~4 wks. afterwards; true leaves were harvested and stored at – 20 °C until gDNA extraction was performed. These n = 3 USDA GRIN samples are termed the “Cultivars” collection (Table 4).
An unplowed field in Roseau, MN was used to harvest hay during the Dust Bowl era (1930s) which was transported and sold throughout the Midwest (Magnusson 2012) and may have led to the spread of RGC throughout the Midwest (Anderson 2019; Anderson et al. 2016). This pristine field, a segment of the ancient Lake Agassiz lakebed in the Roseau Lake Wild Management Area (WMA), remains unplowed and still contains 100% P. arundinacea (Fig. 2a inset; Anderson et al. 2016). A total of 30 specimens from a transect were collected for analysis (Table 4). This location most likely represents an extant, native RCG germplasm source and is termed the “Native Field” collection.
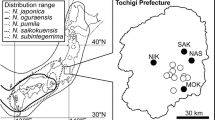
To compare the six MN rivers collections of RCG to central European genotypes, we conducted a sampling of six Czech Republic rivers: Labe, Vltava, Berunka, Lužnice, Orlice, and Tichá Orlice. The number of samples collected for analysis was n = 76 (n = 27 major locations) (Fig. 2b). Comparatively, the Czech Republic in Central Europe is at a similar latitude to Minnesota, has a large number of freshwater rivers that outflow both north and south, many historic fish ponds, canal systems, and wet meadows dominated by RCG (Anderson et al. 2016; Kávová et al. 2017; Anderson 2019). This collection represents wild Czech Republic RCG types (Table 5 and Fig. 2b) and its collection is called the “Czech Republic Rivers” collection.
DNA purification and genotyping
Approximately 15 cm of fresh, mature reed canarygrass leaf tissue per extant specimen was collected, frozen (− 4 °C), and stored in − 80 °C. Total gDNA was extracted from about 50 mg of leaf tissue using the gDNA extraction kit (96 Well Synergy™ Plant DNA Extraction Kit; OPS Diagnostics Laboratory, Lebanon, NJ) with small modifications to the manufacturer’s protocol. Due to the limited tissue supply of herbarium RCG samples, the Synergy™ 2.0 Plant DNA Extraction Kit (single tube DNA purification kit) was used to purify those samples individually. Both kits have the same extraction buffers and purification methodology; the only difference between them is a single tube extraction vs. a 96-well plate format, respectively. Fresh leaf tissue was kept on ice before processing the samples. Scissors and forceps used for tissue handling were cleaned in soapy water, rinsed twice in deionized water, and dried before cutting each sample. Tissue was ground for 15 min at 1500 rpm using a homogenizer (Geno/Grinder; SPEX SamplePrep, Metuchen, NJ). Purified gDNA was suspended in molecular grade water and kept at − 20 °C. DNA quality and quantity were checked with the use of a spectrophotometer (NanoDrop 2000; Thermo Scientific, Waltham, MA). Czech Republic RCG samples were purified following the protocol used by Kávová et al. (2013). The final purified herbarium and extant RCG samples (20µl of 20 ng·µL−1/sample) were submitted to Diversity Arrays Technologies (Bruce, ACT, Australia) for DNA polymorphism identification (SNPs). Genetic variability among RCG populations was assessed with the use of DArTseqLD, that offers a lower density of molecular markers while allowing SNP analysis with full genome representation. This method is highly suitable for non-model species, such as RCG, and has had utility in the analysis of genetic population structure analysis of herbaria RCG (Noyszewski et al. 2019).
Genomic DNA (gDNA) degradation among various RCG herbarium samples and its utility for SNP polymorphism analysis was previously assessed (Noyszewski et al. 2019). Collected herbarium DNA samples had varying levels of missing SNPs, higher than that of fresh tissue, with a mean of 4233 missing SNPs/genotype. To avoid biasing the analysis of herbarium samples with a large number of missing SNPs, these were filtered out based on the percent of missing data (gl.filter.callrate, threshold = 0.6). As a result, only n = 2 of the oldest (pre-1900) herbarium samples, No. 71158 (an undated specimen from the Otto Lugger private herbarium; extrapolated to be ≤ 1891 collection date) and No. 71175 (1891), were retained (Table 1). The rest of the early herbarium samples had a larger than average number of missing alleles or completely failed DArTseqLD genotyping. Additional herbarium samples (n = 15) were then selected with collection dates in the post-dust bowl period (1940–1985). Thus, after filtering, the final number of herbarium genotypes analyzed in the complete data set was n = 17 (from 1891 to 1985; Table 1).
Use of DNA-based approaches are necessary, particularly in species such as RCG where exotic and native forms are morphologically indistinguishable. Sequencing and subsequent detection of SNPs and genome-based methods have become well suited for non-model organisms (Ekblom and Galindo 2011; Fitzpatrick et al. 2016). Utilization of next-generation sequencing (NGS) as a means of generating multi-locus data for non-model organisms is now highly cost-effective and efficient. In particular, development of technologies such as DArTseqLD (Diversity Array Technology Pty Ltd., Canberra, ACT, Australia; https://www.diversityarrays.com/) can be used to delineate species' polymorphic data that can be used directly for genetic diversity data analysis. Bioinformatic packages such as dartR (Gruber et al. 2018) were developed to assist in DArTseqLDTM data analysis. DArTseq markers are co-dominant and based on SNP data. The development of DArTseq markers does not require a species’ genomic sequence and relies on genomic information complexity reduction to successfully analyze genetic diversity and population structure (Abu Zaitoun et al. 2018; Garot et al. 2019; Robbana et al. 2019) as well as genetic mapping (Barilli et al. 2018; Sánchez-Sevilla et al. 2015).
Data analysis
RCG were divided into three primary data sets (a) Minnesota Rivers and Czech Republic Rivers collections (Fig. 3), followed by (b) Minnesota Rivers, Herbarium, Extant Herbarium, Research Field, Native Field, Commercial Field, and Cultivars collections (Fig. 4) and (c) all analyzed samples (collections) together (Fig. 5), to assess overall within and among genotype and population differentiation. In total, after filtering by DArTseqLD and elimination of samples greater than the threshold of 4233 missing SNPs, a total of 2521 polymorphic DArTseqLD SNP markers were used to describe 478 RCG samples across all collections.
a PCoA plot of all tested extant, wild rivers reed canarygrass genotypes (n = 256) from Minnesota, USA and the Czech Republic, based on principal coordinates analysis (PCoA) of 2521 DArTseqLD SNP markers. Individuals are represented as dots and the groups as inertia ellipses with sampling locations connected with lines to the center of each ellipse; differing inertia ellipse sizes are due to sampling size of populations as well as genetic differences. Distinct clustering is due to SNP variation and distinctiveness of each continental grouping. b Graphical plot of all genotypes from Minnesota, USA and the Czech Republic, based on discriminant principle component analysis (DAPC) which resulted in the formation of two primary clusters with almost no overlap; c STRUCTURE bar plots showing the assignment of genotypes into two distinct genetic clusters (K = 2, Evanno method) separating “Minnesota Rivers” and “Czech Rivers” collections with each genotype represented in a vertical line, based on STRUCURE 2.3.4. of reed canarygrass and supported by PCoA and DAPC clusterings. Similar colors (randomly assigned) denote grouping of genotypes, based on shared SNPs (see text). Samples represent two distinct collections of genotypes from Minnesota (USA) and Czech Republic.
All Minnesota USA reed canarygrass genotypes (Herbarium, Extant Herbarium, Minnesota Rivers, Research Field, Commercial Field, Cultivars, and Native Field collections). a Scatter plot of all genotypes, based on principle coordinates analysis (PCoA) of SNP data for the first two principle components (PCoA1, 2); genotypes are represented as dots and the groups as inertia ellipses with sampling locations connected with lines to the center of each ellipse. b Scatter plot of genotypes in these Minnesota collections, based on discriminant principle component analysis (DAPC); Eigenvalues of the DAPC analysis are displayed in the bar plot inset; genotypes are represented as dots and the groups as inertia ellipses with sampling locations connected with lines to the center of each ellipse. c STRUCTURE bar plots showing the assignment of genotypes into two distinct genetic clusters (K = 2, Evanno method) based on STRUCURE 2.3.4. of reed canarygrass with each genotype represented in a vertical line. Similar colors (randomly assigned) denote grouping of genotypes, based on shared SNPs (see text). Analysis separated RCG genotypes into Commercial Field with ‘Venture’ and ‘Palaton’ and all other Minnesota collections. In addition, K = 3 suggest three clusters a) Minnesota Rivers, with two oldest herbarium specimens (71158 and 71175) and two other clusters Commercial Field and Minnesota Rivers and other MN collections. Approximate locations of selected individuals are plotted above STRUCTRUE output. Overall, n=399 individuals were analyzed with 2415 DArTseqLD SNP markers.
All Minnesota samples and Czech Rivers. a Scatter plot of all tested Minnesota USA reed canarygrass genotypes (Herbarium, Extant Herbarium, Minnesota Rivers, Research Field, Commercial Field, Cultivars, and Native Field collections) and Czech Republic Rivers, based on principle coordinates analysis (PCoA) of SNP data for the first two principle components (PCoA1, 2); Genotypes are represented as dots and the groups as inertia ellipses with sampling locations connected with lines to the center of each ellipse. b Scatter plot of genotypes in these Minnesota and Czech Republic Rivers collections, based on discriminant principle component analysis (DAPC); Eigenvalues of the DAPC analysis are displayed in the bar plot inset; Genotypes are represented as dots and the groups as inertia ellipses with sampling locations connected with lines to the center of each ellipse. c Structure bar plot showing the assignment of genotypes into two distinct genetic clusters (K = 2, Evanno method) based on STRUCURE 2.3.4. of reed canarygrass. Analysis separated RCG genotypes into a) Commercial Filed (with Venture and Palaton) b) other Minnesota collections and c) Czech Republic Rivers. In addition, K = 3 suggest three clusters a) primarily Commercial Filed, Minnesota Rivers (with other collections) and Czech Republic Rivers, with two oldest herbarium specimens (71158 and 71175) and few other (~1940) herbarium specimens. Approximate locations of selected individuals are plotted above STRUCTRUE output. Overall, n = 478 individuals were analyzed with 2521 DArTseqLD SNP markers
RStudio (RStudio Team 2015) was used to perform DArTseqLD data analyses. The dartR, adegenet, ade4 packages were used to analyze these data (Bougeard and Dray 2018; Gruber et al. 2018; Jombart and Ahmed 2011). Unlike many analyses where only one type of statistical program is used, we used several in this paper since they rely on different algorithms (PCoA, DAPC, STRUCTURE) and provide unique interpretations. PCoA was performed with the use of the dartR package (Jombart and Ahmed 2011). To visualize RCG data groupings, we used the discriminant analysis of principal components (DAPC) method (Jombart et al. 2010), a multivariate method that estimates the possible number of clusters of genetically related individuals. Clustering of the RCG populations was performed using the Bayesian clustering algorithm implemented in the STRUCTURE v 2.3.4 (Pritchard et al 2000; Falush et al 2003; Evanno et al 2005; Falush et al 2007; Hubisz et al 2009). For STRUCTURE analysis we used an admixture model with 10,000 burn-in iterations and 10,000 Markov Chain Monte Carlo iterations with six independent replicates for the assumed number of K clusters (values used from 2 to 17) for both “Minnesota rivers” vs. “Czech Republic Rivers” RCG collection, “Minnesota Rivers” vs. all other RCG collections and for all RCG samples and populations combined. The most likely number of assumed populations (K) was estimated by the Evanno method (Evanno et al. 2005) using web interface STRUCTURE harvester (Earl and vonHoldt, 2011) for each of these data sets.
To perform the Analysis of Molecular Variance (AMOVA; Excoffier et al. 1992) within and among RCG populations, collections, and to describe the degree of population differentiation, the fixation index (Fst) was used with the package GenAlex (Peakall and Smouse 2012). To determine the significance values for FST, a null distribution was calculated based on 999 permutations of the binary data matrix.
Results and discussion
The PCoA of 2,521 DArTseqLD SNP markers shows that all (n = 256) tested extant wild, riparian RCG genotypes from six Minnesota Rivers and six Czech Republic Rivers are genetically distinct (Fig. 3A). Since both PCoA-1 and PCoA-2 axes explain a relatively small percent of SNP genetic variation observed in both populations, 2.9% and 1.5%, respectively, neither a single nor or a few SNPs could be identified that differentiated RCG on the two continents. Rather, collectively, numerous SNPs are responsible for these differences. The additional DAPC analysis also resulted in the formation of two primary clusters separating the Minnesota Rivers and Czech Republic Rivers riparian samples, with almost no overlap (Fig. 3b). Similarly, STRUCTURE analysis supported this clustering of the PCoA and the DAPC, separating “Minnesota Rivers” and “Czech Rivers” collections (Fig. 3c). The most likely number of populations selected by the Evanno method was not k = 2, however, as that peak was significantly smaller than k = 4 (Fig. 6a), due to the occurrence of multiple samples in primarily the Czech Republic Rivers populations which share a significant number of SNPs with the Minnesota Rivers populations (Fig. 3c k = 4 plot). Thus, four is the most likely number of populations using the Evanno method (k = 4; ΔK = 762.8; Evanno et al. 2005), due to the separation of the “Czech Republic Rivers” into three separate, genetically diverse clusters (Fig. 3c k = 4 plot). DArTseqLD SNP markers are powerful enough to genetically differentiate the “Czech Republic Rivers” populations into three separate clusters within this collection.
Line graphs showing distribution of K value, indicating the most probable number of clusters (Evanno et al. 2005). a Line graph showing k = 4 (ΔK = 762.8) as the most likely number of populations for Minnesota River and Czech Rivers (Fig. 3c). Note the smaller peaks at K = 2 and K = 5. b Line graph showing k = 2 (ΔK = 67.53) as the most likely number of populations for Minnesota RCG collections (Fig. 4c). Note the smaller peaks at K = 3. c. Line graph showing k = 2 (ΔK = 80.83) as the most likely number of populations for all RCG collections (Fig. 5c). Note the smaller peaks at K = 3
The Minnesota Rivers populations, however, remain in one distinct cluster (Fig. 3). Together, the uniformity of PCoA, DAPC, and STRUCTURE (Evanno) statistical analyses of these 2521 SNPs indicate a distinct separation of “Minnesota Rivers” and “Czech Republic Rivers” populations. The only exception would be PCoA (Fig. 5) where overlap occurred among these two sets. Portions of the genome (specific SNPs) are preserved or in common across continents—as would be expected when comparing members of the same species, as indicated by STRUCTURE similarities within each of the groupings (Fig. 3c) as found in the overlap of the two in the PCoA (Fig. 5). Nonetheless, overall significant SNP differences between the continents indicate that the Minnesota riparian populations are distinct enough from the European (Czech) collections to be delineated as most likely native N. American RCG. This native status would refute the postulation by (Lavergne and Molofsky 2004) that RCG's exotic origin is a cause of its invasiveness. These findings would complement previous research with internal transcribed sequences (ITS) of P. arundinacea wherein the species appeared in two ITS clades with one embedded in Europe (with the center of origin in the Mediterranean Basin) while the other was distinctly N. American (Voshell and Hilu 2014; Voshell et al. 2015; Graper et al. 2021). Historic dispersal routes from the center of origin in the Mediterranean Basin into the Americas occurred via the Bering land route during the mid-Miocene epoch (Voshell and Hilu 2014). Subsequently evolutionary diversification led to the distinct N. American types, e.g. most likely the Minnesota populations tested herein (Graper et al. 2021). Introduction of European types into N. America via European settlers also occurred, although the current populations in Minnesota are genetically distinct enough to warrant separation from the tested central European populations. Whether this SNP differentiation holds for the remainder of Europe awaits discovery.
Analysis of molecular variance (AMOVA) showed that the majority of the total genetic variance was found within rather than among populations (Fig. 7), with a similar level of genetic variation found within Minnesota collections and the Minnesota Rivers and Czech Republic Rivers, 99% and 98%, respectively (Fig. 5). Similarly, high levels of genetic variation within populations were obtained in past RCG research with 84% (Jakubowski et al. 2011) ranging from 80 to 90%, depending on comparison scale (Nelson et al. 2014). Our SNP results, in conjunction with previous studies, confirm that RCG grass is genetically diverse within populations. However, Fst values (ranging from Fst = 0.003 to Fst = 0.026; Fig. 7) indicate that there is no significant genetic differentiation among all populations or groups (Fig. 5). The highest differentiation was observed between Cultivars and the Commercial Field collections and all other MN collections, in particular the wild Minnesota Rivers collection.
a Analysis of molecular variance (AMOVA) and the fixation index (Fst) among and within the tested reed canarygrass populations for all Minnesota collections (including Herbarium, Extant Herbarium, Minnesota Rivers, Research Center, Commercial Field, Cultivars, and the Native Field); Fst highlighted in red indicate significant fixation (p < 0.05), based on a null distribution calculated on 999 permutations of the binary data matrix; b Percentages of molecular variance among the Minnesota Rivers vs. c the Czech Republic Rivers collections
These SNP findings complement previous research using more precise molecular tools than gene products used for fingerprinting (Lavergne and Molofsky 2007). For example, too many missing SNPs due to DNA degradation were found in most of the identical herbaria specimens deemed as “native” in previous studies (Jakubowski et al 2013); as a result only two herbaria specimens < 1930 remained in the present study. As reviewed by Anderson (Anderson 2019), neutral isozyme markers proved to be inconclusive that European RCG cultivars had moved from cultivated fields into invasive N. American populations (Gifford et al. 2002). In contrast, Czech Republic and French RCG populations shared a high (87%) proportion of allozymes markers (Lavergne and Molofsky 2007). A smaller proportion of SNP markers were shared among Czech Republic Rivers and Minnesota Rivers populations (Fig. 3), due to the increased precision realized by comparing actual DNA sequence polymorphisms common to the species. Subsequent use by multiple labs of more precise molecular methods, such as AFLPs, SSRs and ISSRs, in historic as well as extant N. American genotypes confirmed the existence of native continental North American populations (Jakubowski et al. 2011, 2014). Nelson et al. (2014) determined that the population genetic structure of wild, forage, and ornamental European and N. American RCG harbored a high amount of genetic diversity within, as opposed to among, populations. Subsequent research has reconfirmed this in additional populations (Anderson et al. 2016; Nelson and Anderson 2016). Earlier SNP research by Noyszewski et al. (2019) furthers comparative molecular work from historic and extant RCG specimens. Thus, range expansion of P. arundinacea in N. America is not necessarily a result of hybridization among European, forage, and North American individuals (Nelson et al. 2014). Future research will be devoted to including a more comprehensive sampling across Europe, even though our sampling in central Europe is extensive. Previously, chloroplast sequencing differences were found in RCG from northwestern Europe (Perdereau et al 2017) although it is unknown whether or not nuclear DNA SNPs (as used herein) would also differ. In contrast, ITS (internal transcribed sequences) were the same for P. arundinacea samples across Europe but differed from N. American types. Thus, it may also be useful to incorporate ITS as an additional distinguishing molecular marker for RCG between the continents (Voshell and Hilu 2014).
PCoA (Fig. 4a) and DAPC (Fig. 4b) results from analysis of all the Minnesota RCG collections clustered the Minnesota Rivers, Herbarium, Extant Herbarium, Research Field and the Native Field collections together. The STRUCTURE analysis (k = 2, ΔK = 67.53; Fig. 4c) divided these Minnesota collections from the Commercial Field and Cultivars collections. Thus, there are two genetically distinct groups of RCG in Minnesota, as indicated by STRUCTURE (Evanno et al. 2005). Since the Minnesota Rivers, the Research Field, the Native Field and pre-1930 herbaria collections clustered together they are native N. American types. It remains to be determined, however, why the Commercial Field and Cultivars cluster separately from the N. American types but are unlikely to be European in origin (Fig. 5). This could indicate that the Commercial Field and low-alkaloid ‘Palaton’ and ‘Venture’ may represent other races of N. American or other European types. It will be critical to analyze additional extant and herbaria samples from across the United States and Canada to determine whether or not they align more closely with these collections, the native Minnesota (N. American) types or create other SNP clusters. Likewise, it is unknown whether roadside RCG populations along existing roads or “Constitutional Routes” emanating from Roseau, MN during the Dust Bowl era are genetically similar to those RCG descendants in our extant Native Field collection. We anticipate researching this question in the near future. Additionally, since (Voler and Smith 1965) suspected that RCG was absent in 1915 in the adjacent State of Iowa, its subsequent dispersal and spread therein may explain why RCG herbarium specimens don’t surface in Iowa herbaria until after the Dust Bowl era.
Our PCoA results explained 5.6 % of the total variance of the genetic composition of “Minnesota Rivers” and other collections for the first two principle components, PCoA1, 2 (Fig. 4a). PCoA (Fig. 4a) analysis of SNP data for most of the tested Minnesota USA reed canarygrass genotypes for this experiment (Herbarium, Extant Herbarium, Minnesota Rivers, Research Field, and Native Field collections) clustered together with few differences. Similar findings occurred with the DAPC analysis (Fig. 4b), although the Minnesota Rivers were distributed a bit more widely than the PCoA (Fig. 4a). In both analyses, the remarkable similarity of SNPs from such diverse geographic areas within MN (Table 3) was unexpected.
The first PCoA and STRUCTURE analyses comparing the Minnesota Rivers and Czech Republic Rivers (Fig. 4a, c, respectively) had indicated the Minnesota Rivers (Table 3) were in one grouping whereas the Czech Republic Rivers (Table 5) consisted of three populations (k = 4; Fig. 6b). This showed greater diversity in the European riparian populations than found in the N. American (Minnesota Rivers) and was also not expected. Thus, the Minnesota Rivers RCG genotypes act as one large, panmictic population by nature with a random mating strategy because grasses are wind-pollinated (anemophilous). The lack of particular structure within Minnesota Rivers populations may be attributable to human-mediated dispersal and self incompatibility (Carlson v 1996, 2015). Since RCG seeds are buoyant, genotype(s) could also easily spread downriver (Casler 2010). Likewise, prior to the arrival of European settlers, rivers were the main long-distance transportation corridors in Minnesota, the Midwest, and Canada. Since Native American tribes (U.S.A.) and First Nations (Canada) in temperate regions across N. America used RCG for a variety of purposes (Turner et al. 1980; Kindscher and Noguera 2002; Densmore 2012), it is likely that Native American tribes spread RCG seeds or vegetative propagules, which would also contribute to the overall lack of genetic differentiation among Minnesota Rivers. RCG does not have similar cultural significance in Europe and was used mainly as a forage crop, contained within pastures and wet meadows, or as an ornamental around dwellings and commercial buildings. Future research will be devoted to assessing larger populations of RCG genotypes along each river in both Minnesota and the Czech Republic to determine potential causes for the SNP configuration differences in RCG across the continents and whether SNPs are spread downstream in the flow of each river.
Herbaria specimens serve as a repository of historic plant biodiversity to allow for long-term scientific study (Besnard et al. 2018). However, early specimen preservation methods can negatively impact DNA quality and lead to various degrees of DNA degradation (Noyszewski et al. 2019). Additionally, allele dropout (non-specific loss of DNA sequences) and misincorporations (artificial nucleotide substitutions caused by DNA deamination during PCR amplification) could lead to false representation of allelic content or recognition of false polymorphism of herbaria specimens (Wandeler et al. 2003; Stiller et al. 2006;Sawyer et al. 2012 Burrell et al. 2015). Many traditional molecular markers analyses (SSRs, AFLPs, etc.) and current next generation-based methods, such as DArTseqLD, rely on amplification of DNA fragments with restriction digestion. Most likely these methods will not be greatly affected by the level of DNA degradation, since they are based on short DNA fragments (< 200 bp). When low quality gDNA is analyzed, next generation sequencing methods produce overrepresentation of “missing data”' compared to high quality DNA obtained from fresh tissue. The process of filtering against missing data points is one of quality control steps for the DArTseqLD method (Noyszewski et al. 2019).
Herbarium specimens in all three categories of specimen types (earliest possible—likely native, after shipments started, and recent) with relatively non-degraded DNA and sufficient numbers of SNPs (Table 1) clustered with the Minnesota Rivers accessions in both the PCoA (Fig. 4a), DAPC (Fig. 4b) and STRUCTURE (Fig. 4c) analyses, providing additional support that all of these riparian RCG are native N. American genotypes. The two earliest possible Herbarium samples with non-degraded DNA, Nos. 71158 and 71175, clustered together with the wild Minnesota Rivers and Extant Herbarium samples. Also, the more recently collected historic RCG herbarium samples in the categories “after shipments started” and “recent clustered” clustered within the wild Minnesota Rivers collection. Thus, regardless of herbarium specimen age, all categories had the same SNPs. Similarly, the DAPC method placed both pre-1900 samples in very close proximity to most of the extant Minnesota RCG rivers collections. Thus, regardless of the collection year, all tested RCG herbarium samples (pre 1900 to 1985) in the University of Minnesota Bell Museum Herbarium are genetically similar to extant Minnesota Rivers and Extant Herbarium collections. However, when Czech Republic Rivers were added (Fig. 5), PCoA showed that two earliest herbarium specimens (Nos. 71158 and 71175) with a few other earlier herbarium specimens (pre-1940) clustered within Czech Republic Rivers and few Minnesota Rivers samples that overlapped with Czech Republic Rivers. However, the majority of remaining herbarium samples (n = 12) clustered with Minnesota collections. Similarly as before PCoA results explained 5.6 % (PCoA-1 3.8% and PCoA 2%) of the total variance of the genetic composition. Similar grouping of collections and specific herbarium specimens was also observed with analysis by STRUCTURE (Fig. 6c). The most likely number of predicted clusters was k = 2 (ΔK = 80.83) with similar grouping of collections and individuals in PCoA. The DAPC however, clustered herbarium specimens much closer to Minnesota collections (Fig. 6b).
Jakubowski et al. (2013) evaluated herbarium samples (late 1800s and early 1900s) and found out that those samples were different from those of European and Asian origin, indicating that herbarium specimens were most likely native to the North American continent. Subsequent studies used the same herbarium samples and showed that only two, early herbarium samples clustered with extant North American samples (Jakubowski et al. 2014). However, STRUCTURE classification divided herbarium samples and all North American and Eurasian samples into three clusters (k = 4, Fig. 3b, Jakubowski et al. 2014): one with most of herbarium samples and the two North American and Eurasian samples. However allelic composition did not separate North American and Eurasian samples, those shared high levels of admixture. It is possible that herbarium samples that created a separate genetic cluster in Jakubowski’s et al. (2014) paper were biased due to the high level of degradation of gDNA that could impact genotyping results (Noyszewski et al. 2019).
As a secondary benchmark to the native RCG Herbarium collection (Table 1; Fig. 4; Jakubowski et al. 2014, 2013) those specimens with demographic information specific enough to approximate their current sites (all predate the existence of global positioning system or GPS technology) were collected (Tables 2, 3). These Extant Herbarium genotypes were sampled in 2012 across the State of Minnesota (Tables 1, 3). In some instances, it was not possible to find an exact match in location between the Herbarium and current day sites, e.g. one site present in the 1800s is now in the middle of a lake, landscape modifications had occurred over the past century at other sites, etc. Nonetheless, sufficient Extant Herbarium genotypes (n = 60) were located and collected at five sites (Table 3) to provide genotyping of descendants from their respective Herbarium ancestors. The SNP distribution of the Extant Herbarium genotypes, for both PCoA and DAPC analyses, clustered less tightly around the Minnesota Rivers and Herbarium collections, although their SNP range often surpassed those distributions creating a larger elipse (Fig. 4a, b). However, this distance is only slightly different when compared to other MN locations (Fig. 4b). Thus, some level of genetic variation and evolution may have occurred leading to the present-day Extant Herbarium genotypes, although the original corollary Herbarium specimens had high levels of DNA degradation, limiting SNPs generation.
The RCG Research Field collection (Table 4) represents a mixed stand of the most likely RCG grass types similar to older RCG forage cultivars such as ‘Rise’, but not the newer low-alkaloid cultivars currently in forage seed production in Roseau, MN (Schaeffer 2019). As posited earlier, this germplasm collection should most likely be similar to the Minnesota Rivers wild riparian collection due to relatively minor germplasm advancement when compared to newer, low alkaloid ‘Palaton’ and ‘Venture’. Indeed, this was the case in the PCoA (Fig. 4a) and DAPC (Fig. 4b) analyses, although in the PCoA, several RCG genotypes from this collection were genetically similar to ‘Vantage’ as well as extending towards alignment with the Commercial Field and Cultivars cluster. In the DAPC analysis, however, the Research Field genotypes did not extend as close to these collections. Nonetheless, the majority of the RCG genotypes in the Research Field are similar to the Minnesota Rivers, Herbarium, Extant Herbarium, and Native Field collections and do not appear to be introducing exotic RCG genes from European collections into the surrounding putatively native stands.
To evaluate the genetic composition and distance of the forage cultivars grown in Minnesota, we tested ‘Palaton’ and ‘Venture’ which are the most developed low alkaloid RCG cultivars planted in Minnesota for forage seed production (Alderson and Sharp 1994). ‘Vantage’ was also selected since it is one of the parents for ‘Palaton’ and ‘Venture’ and is less comparatively advanced and older (released in 1972; Table 1), although occasionally still in cultivation. The distinct clustering of the Commercial Field and Cultivars collections away from all other Minnesota RCG collections (Fig. 4a–c) is curious and unexpected. Both ‘Venture’ and ‘Palaton’ clustered with the Commercial Field genotypes in the PCoA as did ‘Vantage’ in the DAPC analysis (Fig. 4b), although ‘Vantage’ clustered toward the Minnesota Rivers, etc. in PCoA (Fig. 4a). The tighter clustering of ‘Vantage’ with the wild Minnesota Rivers collection is expected since it is the least developed (mainly seed collections in Iowa and southern Minnesota) of the three RCG forage cultivars tested. Further studies should analyze where additional historic forage cultivars (Table 1) from the University of Minnesota breeding program, as well as other N. American breeding efforts in adjacent U.S. states and across the provinces of Canada, cluster in relation to ‘Palaton’, ‘Venture’, the Commercial Field, and ‘Vantage’.
While the genotypes collected along the perpendicular transects through the Commercial Field in the 3642 hectare (9000 A) forage seed farm in Roseau, MN were reported by the farmer to originally be ‘Vantage’, they were not closely clustered in the PCoA with the known ‘Vantage’ seed obtained from the USDA GRIN collection (Fig. 4a). In fact, they clustered instead with ‘Palaton’ and ‘Venture’ which are more recent introductions and cultivated widely throughout Roseau, MN. It is important to note that ‘Vantage’ is one of the parents that contributed to both ‘Palaton’ and ‘Venture’ cultivars (Fig. 1). However, the DAPC analysis showed a much closer clustering of ‘Vantage’, but not overlapping, with the Commercial Field (Fig. 4b). The farmer producing RCG forage seed in this field conjectured that numerous old, native genotypes have germinated in this field, either from the extensive seed bank or from the adjacent Native Field (Christenson 2012, personal communication). While this could, indeed, be the case, none of the genotypes in the Commercial Field clustered with the Native Field (Fig. 4a, b). When all RCG samples and populations (including those from Czech Republic) and Commercial Field were compared, the Commercial Field consistently created separate cluster from all other RCG samples based on PCoA (Fig. 5a), DAPC (Fig. 5b) and STRUCTURE (Fig. 5c). However, PCoA (Fig. 5a) and STRUCTURE (Fig. 5c) indicate that relative genetic distance between Czech Rivers and Minnesota Rivers is relatively low. In overall comparison of RCG samples with use of STRUCTURE the k = 2 was determined to be the most likely number of clusters, indicating genetic distance to both wild RCG populations from Minnesota Rivers and Czech Republic Rivers to those from Commercial Field, with further differentiation. The Native Field is 100% RCG (Fig. 2a, inset; Anderson 2019) and, based on oral testimonies, contains native types of RCG (Anderson 2019). Our PCoA, DAPC and STRUCTURE analyses placed the Commercial Field, ‘Venture’ and ‘Palaton’ in one cluster while the second cluster contains all other Minnesota collections including the Native Field (Fig. 4a, b). The Native Field and the Commercial Field are in close proximity to each other (~2.86 km, Fig. 2a inset). However, our analysis showed that RCG samples from the Native Field are genetically closer to those of the Roseau River ~2.06 km away, as well as all other Minnesota Rivers, rather than the Commercial Field (~2.86km in distance; Fig. 2a, inset). This indicates no or minimal gene flow among both locations despite their relative sympatry. Similar findings have been reported in another grass, Phragmites australis (Saltonstall 2003), with minimal gene flow occurring among genetic lineages. This is surprising, since reed canarygrass is a wind-pollinated outcrossing species (with a tight gametophytic self incompatibility system; Casler et al. 2009) and it would be easy for gene exchange to take place among genotypes in the Commercial Field and the Native Field. While it is unknown the distance RCG pollen travels, the pollen is a known wind-borne allergen (Schumacher et al. 1968) although in production yield trials, the parents of ‘Vantage’ were closely planted (91 cm on center) to maximize seed production (Rincker et al. 1977). Seed migration can be aided by cultivation (Roseau, MN is a major site for RCG seed production in cultivated fields) or waterways that, when complemented with outcrossing was theorized to increase genetic variability (Lavergne and Molofsky 2007). This seems less likely with these populations, given the genetic similarity of the Native Field with all Minnesota Rivers. More likely this increased genetic variation holds for intercontinental germplasm (among N. American and European types; Casler et al. 2009). Cross-incompatibility could occur among the Native Field and Commercial Field genotypes, although it is unlikely to be the case for every possible hybridization event. It is possible that, due to the high density of RCG stands, inter-populational hybrid seeds may not germinate due to the lack of a niche or they are outcompeted by extant RCG plants, effectively preventing replacement by new genotypes. Likewise, Gifford et al. (2002) showed that most RCG propagates primarily as clones, rather than by seed. Jakubowski et al. (2011) indicated that breeding efforts did not produce invasive types of RCG while Jakubowski et al. (2010) reported that landscape modifications were a primary predictor of RCG invasions.
Analysis of molecular variance (AMOVA) showed that the majority of the total genetic variance was found within rather than among populations (Fig. 7), with a similar level of genetic variation found within Minnesota collections and the Minnesota Rivers and Czech Republic Rivers, 99% and 98%, respectively (Fig. 5). Similarly, high levels of genetic variation within populations were obtained in past RCG research with 84% (Jakubowski et al. 2011) ranging from 80% to 90%, depending on comparison scale (Nelson et al. 2014). Our SNP results, in conjunction with previous studies, confirm that RCG grass is genetically diverse within populations. However, Fst values (ranging from Fst = 0.003 to Fst = 0.026; Fig. 7) indicate that there is no significant genetic differentiation among all populations or groups (Fig. 5). The highest differentiation was observed between Cultivars and the Commercial Field collections and all other MN collections, in particular the wild Minnesota Rivers collection.
Management implications
Since all examined riparian, wetland, and native field accessions throughout the State of Minnesota are most likely native and not European, the implications of native riparian and wetland stands of RCG are cause for a reexamination in approach regarding its control. While it still remains, undeniably, an invasive wetland species, its native rather than exotic status challenges assumptions for its absolute control (Anderson et al. 2021), leaving land managers in potentially precarious decision-making and risk assessments. While numerous native N. American plants have been determined to be invasive, e.g. Typha, the aggressiveness of RCG’s spread throughout N. America and the Midwest (including Minnesota; Galatowitsch et al. 1999), in particular, has exacted a systematic program of elimination by land managers. While numerous factors have contributed to its extensive spread throughout N. America, including transport as hay during the Dust Bowl, construction of highway corridors (Constitutional Routes and interstate highways), elevated nitrogen levels in water tables and wetlands due to commercial agriculture (Galatowitsch et al. 1999; Reinhardt and Galatowitsch, 2004), as well as its planting for revegetation or as an ornamental/forage/biofuel crop, discussion of which populations present ecological risk and warrant control is needed. A future study of growth rate differences among and within Minnesota riparian populations may provide additional insight. It is possible that extensive landscape modifications contributed to RCG expansion (Jakubowski et al. 2010).
The implications of RCG as a native invasive will require differential shifts in land managers’ perspectives and approaches for control (Anderson et al. 2021), provided inexpensive and quick determination of native vs. exotic status is possible in the field. Particular differences may exist for Tribal Land Managers versus State or Provincial Departments of Natural Resources and private agencies, depending on whether the native stands are preserved or if all RCG is exterminated (as is the case at present). Additionally, regulatory challenges have yet to be legislated for control of a native invasive species such as RCG (Anderson et al. 2021). These opportunities to change attitudes and implement judicial control measures will serve as a template for other invasive species which are native to a region. A risk of these findings is that legislatively mandated funding for continued control of RCG may be rescinded although, arguably, that might be perilous in specific ecosystems. Clearly, in Minnesota at least, the findings of Lavergne and Molofsky (2007) for eastern N. American RCG populations do not hold wherein European populations recombined with native N. American types, resulting in highly invasive, clonal populations. Whether our findings in Minnesota hold true for elsewhere in the Midwest and Pacific Northwest areas of N. America await determination.
Summary
Minnesota wild rivers populations of RCG cluster together with two early herbarium specimens and other extant RCG collections with exception of cultivated RCG types. These results suggest that present wild MN rivers RCG population is most likely native to MN, with no distinct grouping for each MN river. In addition, gene flow from cultivated types of RCG is minimal, since those types were contained within its cultivation area. Comparison of MN and Czech Republic wild rivers RCG collection indicated the existence of two separate clusters, separating those RCG collections at the continental level. Herbarium specimens proved to be useful as a benchmark of native species status, however gDNA degradation can have a negative impact on specimen grouping, potentially creating false clustering of samples. Future work that will investigate genetic composition of RCG samples along MN highways as potential corridors of RCG spread. In addition, large, continental scale sampling of RCG herbarium specimens can reveal the level of genetic diversity and differentiation of RCG across continents.
References
Abbey M, Anderson NO, Yue C, Short G, Schermann M, Phelps N, Venturelli P, Vickers Z (2019) An analysis of strawberry (Fragaria ×ananassa) productivity in northern latitudinal aquaponic growing conditions. J Am Pomol Soc 73:22–37
Abu Zaitoun SY, Jamous RM, Shtaya MJ, Mallah OB, Eid IS, Ali-Shtayeh MS (2018) Characterizing Palestinian snake melon (Cucumis melo var flexuosus) germplasm diversity and structure using SNP and DArTseq markers. BMC Plant Biol 18:246. https://doi.org/10.1186/s12870-018-1475-2
Alderson J, Sharp WC, United States. Department of Agriculture (1994) Grass varieties in the United States. Washington, D.C.: U.S. Dept. of Agriculture
Alway FJ (1931). Early trials and use of reed-canary grass as a forage plant. [WWW Document]. https://www.cabdirect.org/cabdirect/abstract/19310700063. Accessed 10 July 2019
Ambros Z, Štykar J (1999) Geobiocenologie I. Mendelova zemědělská a lesnická univerzita v Brně, Brno, Czech Republic
Anderson NO (2019) Workshop: throwing out the bathwater but keeping the baby: lessons learned from purple loosestrife and reed canarygrass. Hort Technology 1:1–10. https://doi.org/https://doi.org/10.21273/HORTTECH04307-19
Anderson NO, Kávová T, Bastlová D, Čurn V, Kubátová B, Edwards KR, Januš V, Květ J (2016) Phenotypic and genotypic variation in Czech forage, ornamental and wild populations of reed canarygrass. Crop Sci. 56:2421–2435. https://doi.org/10.2135/cropsci2015.11.0705
Anderson NO, Smith AG, Noyszewski AK, Ito E, Dalbotten D, Pellerin H (2021) Management and Control Issues for Native, Invasive Species (Reed Canarygrass): evaluating Philosophical. HortTechnology (In Press), Management and Legislative Issues
Anonymous (2012). Grass Seed Production in NW Minnesota. Seed Today.
Apfelbaum S, Sams C (1987) Ecology and Control of Reed Canary Grass (Phalaris arundinacea L.). Nat Areas J 7
Barilli E, Cobos MJ, Carrillo E, Kilian A, Carling J, Rubiales D (2018) A high-density integrated DArTseq SNP-based genetic map of Pisum fulvum and identification of QTLs controlling rust resistance. Front Plant Sci. https://doi.org/10.3389/fpls.2018.00167
Besnard G, Gaudeul M, Lavergne S, Muller S, Rouhan G, Sukhorukov AP, Vanderpoorten A, Jabbour F (2018) Herbarium-based science in the twenty-first century. Bot Lett 165:323–327. https://doi.org/10.1080/23818107.2018.1482783
Bougeard S, Dray S (2018) Supervised multiblock analysis in R with the ade4 Package. J Stat Softw 86: 1-17. https://doi.org/https://doi.org/10.18637/jss.v086.i01
Burrell AS, Disotell TR, Bergey CM (2015) The use of museum specimens with high-throughput DNA sequencers. J Hum Evol. https://doi.org/10.1016/j.jhevol.2014.10.015
Carlson IT, Oram RN, Surprenant J, (1996). Reed canarygrass and other Phalaris species. In: L.E. Moser, D.R. Buxton, and M.D. Casler (Eds.). Cool-Season Forage Grasses., Agronomy Monograph. American Soc. of Agronomy, Madison, WI, pp. 569–604.
Carlson IT, Oram RN, Surprenant J (2015). Reed Canarygrass and Other Phalaris Species In: Cool-Season Forage Grasses. John Wiley & Sons, Ltd, pp. 569–604. https://doi.org/https://doi.org/10.2134/agronmonogr34.c18
Casler MD (2010) Genetics, breeding, and ecology of reed canarygrass 7
Casler MD, Phillips MM, Krohn AL (2009). DNA polymorphisms reveal geographic races of reed canarygrass.
Chekol T, Vough LR, Chaney RL (2002) Plant-soil-contaminant specificity affects phytoremediation of organic contaminants. Int. J. Phytoremediation 4:17–26. https://doi.org/10.1080/15226510208500070
Christenson N (2012) Personal communication in a conversation with N.O. Anderson. Roseau, MN. Unpublished text
Crawford PHC, Hoagland BW, Sadler J (2009) Can herbarium records be used to map alien species invasion and native species expansion over the past 100 years? J Biogeogr 36:651–661
Densmore F (2012) How Indians Use Wild Plants for Food. Courier Corporation, Medicine & Crafts
Dore WG, McNeill J (1980) Grasses of Ontario, 1st Edition, edition. Hull, Que, Canadian Government Pub Centre, Ottawa
Earl D, vonHoldt BM (2011) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Ekblom R, Galindo J (2011) Applications of next generation sequencing in molecular ecology of non-model organisms. Heredity 107:1–15. https://doi.org/10.1038/hdy.2010.152
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578. https://doi.org/10.1111/j.1471-8286.2007.01758.x
Fitzpatrick MA, Ozer EA, Hauser AR (2016) Utility of whole-genome sequencing in characterizing acinetobacter epidemiology and analyzing hospital outbreaks. J Clin Microbiol 54:593–612. https://doi.org/10.1128/JCM.01818-15
Galatowitsch SM, Anderson NO, Ascher PD (1999) Invasiveness in wetland plants in temperate North America. Wetlands 19:733–755. https://doi.org/10.1007/BF03161781
Garot E, Joët T, Combes M-C, Lashermes P (2019) Genetic diversity and population divergences of an indigenous tree ( Coffea mauritiana ) in Reunion Island: role of climatic and geographical factors. Heredity 122:833–847. https://doi.org/10.1038/s41437-018-0168-9
Gifford AL, Ferdy J-B, Molofsky J (2002) Genetic composition and morphological variation among populations of the invasive grass, Phalaris arundinacea. Can J Bot 80:779–785. https://doi.org/10.1139/b02-063
Graper A, Noyszewski AK, Anderson NO, Smith AG (2021) Variability in ITS1 and ITS2 sequences of historic (herbaria) and extant (fresh) Phalaris species. BMC Plant Biol (In Press).
Gruber B, Unmack PJ, Berry OF, Georges A (2018) dartr: an r package to facilitate analysis of SNP data generated from reduced representation genome sequencing. Mol Ecol Resour 18:691–699. https://doi.org/10.1111/1755-0998.12745
Hroudová Z, Hrivnák R, Chytrý M (2009) Classification of inland Bolboschoenus-dominated vegetation in Central Europe. Phytocoenologia. https://doi.org/10.1127/0340-269X/2009/0039-0205
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332. https://doi.org/10.1111/j.1755-0998.2009.02591.x
Jakubowski A, Casler M, Jackson R (2010) Landscape context predicts reed canarygrass invasion: implications for management. Wetlands 30:685–692. https://doi.org/10.1007/s13157-010-0078-y
Jakubowski AR, Casler MD, Jackson RD (2011) Has selection for improved agronomic traits made reed canarygrass invasive? PloS One 6:e25757–e25757. https://doi.org/10.1371/journal.pone.0025757
Jakubowski AR, Casler MD, Jackson RD (2013) Genetic evidence suggests a widespread distribution of native North American populations of reed canarygrass. Biol Invasions 15:261–268. https://doi.org/10.1007/s10530-012-0300-3
Jakubowski A, Jackson R, Casler M (2014) The history of reed canarygrass in north america: persistence of natives among invading eurasian populations. Crop Sci 54:210. https://doi.org/10.2135/cropsci2013.05.0342
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinforma Oxf Engl 27:3070–3071. https://doi.org/10.1093/bioinformatics/btr521
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94. https://doi.org/10.1186/1471-2156-11-94
Kávová T (2013) Genetická variabilita v populacích chrastice rákosovité (Phalaris arundinacea L.). M.S. Thesis. University of South Bohemia in České Budějovice, Faculty of Agriculture, České Budějovice, Czech Republic
Kávová T, Kubátová B, Čurn V, Anderson N (2017) Genetic Variability of US and Czech Phalaris Arundinacea L. New Perspect. Forage Crops, Wild and Cultivated Populations. https://doi.org/10.5772/intechopen.69669
Kindscher K, Noguera E (2002) Cultural use of plants from the Baker Wetlands. Prepared for the Technical outreach Services for Native American Communities (TOSNAC) at Haskell Indian Nations University. Kansas Biological Survey, University of Kansas, Manhattan
Lavergne S, Molofsky J (2004) Reed Canary Grass (Phalaris arundinacea) as a Biological Model in the Study of Plant Invasions. Crit Rev Plant Sci 23:415–429. https://doi.org/10.1080/07352680490505934
Lavergne S, Molofsky J (2007) Increased genetic variation and evolutionary potential drive the success of an invasive grass. Proc Natl Acad Sci 104:3883. https://doi.org/10.1073/pnas.0607324104
LaVoie C, Dufresne C, Delisle F (2005) The spread of reed canarygrass (Phalaris arundinacea) in Québec: a spatio-temporal perspective. Écoscience 12:366–375
Lewandowski I, Scurlock JMO, Lindvall E, Christou M (2003) The development and current status of perennial rhizomatous grasses as energy crops in the US and Europe. Biomass Bioenergy 25:335–361. https://doi.org/10.1016/S0961-9534(03)00030-8
Magnusson R (2012) Personal communication in a conversation with N.O. Anderson. Roseau, MN. Unpublished text
Merigliano MF, Lesica P (1998) The native status of reed canarygrass (Phalaris arundinacea L.) in the inland northwest, USA. Nat Areas J 18:223–230
Minnesota Department of Natural Resources (1968). Bulletin No. 25 - An Inventory of Minnesota Lakes. Minnesota. Department of Natural Resources, St. Paul, MN. https://files.dnr.state.mn.us/publications/waters/ALL_BULL25.pdf
Nelson MF, Anderson NO (2016). Adaptive responses to water stress in phalaris arundinacea, an invasive wetland grass. Invasive Species Ecol Manag Strateg Conserv 1–20
Nelson MF, Anderson NO, Casler MD, Jakubowski AR (2014) Population genetic structure of N. American and European Phalaris arundinacea L. as inferred from inter-simple sequence repeat markers. Biol Invasions 16:353–363. https://doi.org/10.1007/s10530-013-0525-9
Northwest Regional Development Commission (2014) Northwestern Minnesota: Roseau County History. Land of the Dancing Sky.
Noyszewski AK, Anderson NO, Smith AG, Kilian A, Dalbotten D, Ito E, Timm A, Pellerin H (2019) Workshop: challenges of establishing native versus exotic status of herbarium specimens. HortTechnology 1:1–5
Olsen FJ, Chong SK (1991) Reclamation of acid coal refuse. Landsc Urban Plan 20:309–313. https://doi.org/10.1016/0169-2046(91)90004-6
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Perdereau A, Klaus M, Barth S, Hodkinson TR (2017) Plastid genome sequencing reveals biogeographical structure and extensive population genetic variation in wild populations of Phalaris arundinacea L. in north-western Europe. GCB Bioenergy 9:46–56. https://doi.org/10.1111/gcbb.12362
Piper CV (1914) Forage plants and their culture. The rural text-book series, The Macmillan company, New York
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945
Prud’homme L (1916) Bulletin of the historical society of saint-boniface - 1916 - Vol. V, Part 2 [WWW Document]. http://shsb.mb.ca/en/node/1328. Accessed 10 Aug 2019.
RStudio Team (2015) RStudio: An open source and enterprise-ready professional software for the R statistical computing environment. Minnesota Supercomputing Institute, Minneapolis, MN. https://www.msi.umn.edu/sw/rstudio
Reinhardt C, Galatowitsch SM (2004) Best management practices for the invasive Phalaris arundinacea L. (Reed Canary Grass) in wetland restorations. Final Report 2004-2036.
Rincker CM, Carlson IT, Garrison CS (1977) Effects of two diverse environments on seed production characteristics of the parent clones of ‘vantage’ reed canarygrass1. Crop Sci. 17:625–628. https://doi.org/10.2135/cropsci1977.0011183X001700040036x
Robbana C, Kehel Z, Ben Naceur M, Sansaloni C, Bassi F, Amri A (2019) Genome-wide genetic diversity and population structure of tunisian durum wheat landraces based on dartseq technology. Int J Mol Sci. https://doi.org/10.3390/ijms20061352
Saltonstall K (2003) Microsatellite variation within and among North American lineages of Phragmites australis. Mol Ecol 12(7):1689–702. https://doi.org/10.1046/j.1365-294x.2003.01849.x
Samecka-Cymerman A, Kempers AJ (2001) Concentrations of heavy metals and plant nutrients in water, sediments and aquatic macrophytes of anthropogenic lakes (former open cut brown coal mines) differing in stage of acidification. Sci Total Environ 281:87–98. https://doi.org/10.1016/s0048-9697(01)00838-5
Sánchez-Sevilla JF, Horvath A, Botella MA, Gaston A, Folta K, Kilian A, Denoyes B, Amaya I (2015) Diversity arrays technology (DArT) marker platforms for diversity analysis and linkage mapping in a complex crop, the octoploid cultivated strawberry (Fragaria × ananassa). PLOS One 10:e0144960. https://doi.org/10.1371/journal.pone.0144960
Sawyer S, Krause J, Guschanski K, Savolainen V, Pääbo S (2012) Temporal Patterns of Nucleotide Misincorporations and DNA Fragmentation in Ancient DNA. Plos One 7:e34131. https://doi.org/10.1371/journal.pone.0034131
Schaeffer C (2019) Personal communication via email with N.O. Anderson. University of Minnesota, Dept. of Agronomy & Plant Genetics, St. Paul, MN. Unpublished text
Schoth HA, Harry A (1929) Reed canary grass. [WWW Document]. Digit. Libr. https://digital.library.unt.edu/ark:/67531/metadc5895/m1/2/. Accessed 16 Oct 2019.
Schumaker MJ, Griffith RD, O’Rourke MK (1968) Recognition of pollen and other particulate aeroantigens by immunoblot microscopy. J Allergy Clin Immunol 82(4):608–616. https://doi.org/10.1016/0091-6749(88)90972-4
Sheaffer CC, Rosen CJ, Gupta SC (2008) Reed canarygrass forage yield and nutrient uptake on a year-round wastewater application site. J Agron Crop Sci 194:465–469. https://doi.org/10.1111/j.1439-037X.2008.00327.x
Stiller M, Green RE, Ronan M, Simons JF, Du L, He W, Egholm M, Rothberg JM, Keates SG, Keats SG, Ovodov ND, Antipina EE, Baryshnikov GF, Kuzmin YV, Vasilevski AA, Wuenschell GE, Termini J, Hofreiter M, Jaenicke-Després V, Pääbo S (2006) Patterns of nucleotide misincorporations during enzymatic amplification and direct large-scale sequencing of ancient DNA. Proc Natl Acad Sci USA 103:13578–13584. https://doi.org/10.1073/pnas.0605327103
Turner NJ, Bouchard R, Kennedy DID (1980) Ethnobotany of the Okanagan-Colville Indians of British Columbia and Washington. British Columbia Provincial Museum
Voler R, Smith SG (1965) Changes in the aquatic vascular flora of Lake East Okoboji in historic times. Proc Iowa Acad Sci 72:65–72
Voshell SM, Hilu KW (2014) Canary grasses (Phalaris, Poaceae): biogeography, molecular dating and the role of floret structure in dispersal. Mol Ecol 23:212–224
Voshell SM, Baldini RM, Hilu KW (2015) Infrageneric treatment of Phalaris (Canary grasses, Poaceae) based on molecular phylogenetics and floret structure. Australian Syst Bot 28:355–367
Wandeler P, Smith S, Morin PA, Pettifor RA, Funk SM (2003) Patterns of nuclear DNA degeneration over time–a case study in historic teeth samples. Mol Ecol 12:1087–1093. https://doi.org/10.1046/j.1365-294x.2003.01807.x
Wu S-H, Rejmánek M, Grotkopp E, DiTomaso JM (2005) Herbarium records, actual distribution, and critical attributes of invasive plants: genus Crotalaria in Taiwan. Taxon 54:133–138. https://doi.org/10.2307/25065311
Acknowledgements
This research was supported by grants from the: Minnesota Invasive Terrestrial Plants and Pests Center (MITPPC; 2017–2020); Czech-US American Science Information Center (AMVIS), Ministry of Education, Czech Republic, Program KONTAKT II, project LH11039; Grant Agency of the University of South Bohemia in Českě Budějovice (GAJU 063/2013/Z); project Postdoc USB (Reg. No. CZ.1.07/2.3.00/30.0006) realized through the European Union Education for Competitiveness Operational Programme; United States, Department of State, Bureau of Educational and Cultural Affairs, J. William Fulbright Scholar Program and an Undergraduate Research Opportunities Program grant (UROP, 2019). Collection of samples along Minnesota rivers were under the auspices of a 2012 State of Minnesota, Department of Natural Resources, Division of Ecological & Water Resources, Scientific and Natural Areas Program Collection Permit. We would also like to thank the Bell Museum (University of Minnesota) for allowing destructive sampling and providing historic RCG samples; special thanks to Drs. Michael D. Casler and Andrew Jakubowski for their consultations regarding published research using historic herbaria samples. Additional thanks are extended to Alli Graper, Kaylie Niedźwiecki, and Sarah Gardner for their help in laboratory work.
Author information
Authors and Affiliations
Contributions
Research was conceived by all authors and managed by NA, AS, DB, EI, AT, and HP. Manuscript was written and edited by AKN, NA. Data collection and analysis was performed by AKN. RCG tissue collection was performed by NA, AKN, BK, TK, VJ, VC, DB, and JK (Minnesota) and NA, BK, Tk, VJ, VC, KE, DB, JK (Czech Republic). DArTseq genotyping was performed by AK. All authors read and approved the final manuscript.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noyszewski, A.K., Anderson, N.O., Smith, A.G. et al. Riparian populations of minnesota reed canarygrass (Phalaris arundinacea) are most likely native, based on SNPs (DArTseqLD). Wetlands Ecol Manage 29, 467–494 (2021). https://doi.org/10.1007/s11273-021-09795-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11273-021-09795-8