Abstract
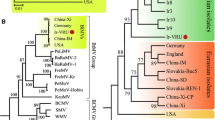
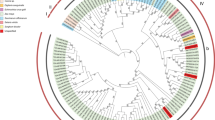
Two new strains of Abutilon mosaic virus (AbMV; Geminiviridae) from Germany (Stuttgart) and France (Paris) have been characterized by circomics, direct pyrosequencing of rolling circle amplification (RCA) products, as well as conventional cloning and Sanger sequencing. RCA combined with an analysis of restriction fragment length polymorphisms confirmed the completeness of the sequence determination and a close relationship of both isolates for DNA A with 99 % nucleotide sequence identity. Phylogenetic tree reconstruction supported their clustering with other AbMV strains in a clade with Middle American begomoviruses, whereas South American begomoviruses that infect Abutilon or Sida micrantha are less closely related. Comparing the coat protein (CP) genes of the AbMV cluster, with those of related Middle and South American begomoviruses revealed a remarkable overrepresentation for non-synonymous nucleotide exchanges for certain amino acid positions in the AbMV cluster. Projection of these positions to a structural model of the African cassava mosaic virus CP yielded a non-random distribution at the periphery and, most importantly, highlighted those amino acids that had been identified in whitefly-transmission experiments before. These results establish the basis for an analysis of the evolutionary liberty of certain amino acid positions of the CP, and their impact on the deciphering of insect transmission determinants is discussed.



Similar content being viewed by others
References
H. zur Jeske, H. de Hausen, E.-M. Villiers (eds.), Torque Teno virus: the still elusive human pathogens (Springer, Berlin, 2009), pp. 185–226
E. Regel, in Gartenflora, ed. by E. Regel (Ferdinand Enke, Stuttgart, 1868), p. 375
C. Wege, R.D. Gotthardt, T. Frischmuth, H. Jeske, Arch. Virol. 145, 2217–2225 (2000)
Jeske H., AAB Descr Plant Viruses No. 373, 2000
E. Regel, in Gartenflora, ed. by E. Regel (Ferdinand Enke, Stuttgart, 1875), pp. 116–117
E. Regel, in Gartenflora, ed. by E. Regel (Ferdinand Enke, Stuttgart, 1871), p. 310
H. Lindemuth, Verh. Bot. Ver. Prov. Brandenbg. 14, 32–37 (1872)
W. Hertzsch, Züchter 2, 195–199 (1930)
H. Lindemuth, Landwirthschaftliche Jahrb. H. 36, 807–861 (1907)
H. Lindemuth, Landwirthschaftliche Jahrb. H. 6, 887–939 (1878)
H. Lindemuth, Vegetative Bastarderzeugung durch Impfung (Hempel & Parey, Berlin, 1878)
J.Y. Keur, Phytopathology 24, 12–13 (1934)
J.Y. Keur, Bull. Torr. Bot. Club 61, 53–70 (1934)
W. Hertzsch, Z. Bot. 20, 65–80 (1928)
A.S. Costa, Annu. Rev. Phytopathol. 16, 429–449 (1976)
A.S. Costa, Phytopathol. Z. 24, 97–112 (1955)
E. Flores, K. Silberschmidt, Phytopathol. Z. 60, 181–195 (1967)
E. Flores, K. Silberschmidt, Phytopathology 53, 238 (1963)
K. Silberschmidt, E. Flores, L.R. Tommasi, Phytopathol. Z. 30, 387–414 (1957)
K. Silberschmidt, L.R. Tommasi, Anais. Acad. Brasil. Cienc. 27, 195–214 (1953)
A. Orlando, K. Silberschmidt, Arqu. Inst. Biol. Sao Paulo 17, 1–36 (1946)
A. Orlando, K. Silberschmidt, O Biol. XI, 139–140 (1945)
K. Silberschmidt, Arqu. Inst. Biol. Sao Paulo 16, 49–64 (1945)
H. Czosnek, M. Ghanim, S. Morin, G. Rubinstein, V. Fridman, M. Zeidan, Adv. Virus Res. 57, 291–322 (2001)
H. Jeske, D. Gotthardt, S. Kober, J. Virol. Methods 163, 301–308 (2010)
J. Jovel, W. Preiß, H. Jeske, Virus Res. 130, 63–70 (2007)
J. Jovel, G. Reski, D. Rothenstein, M. Ringel, T. Frischmuth, H. Jeske, Arch. Virol. 149, 829–841 (2004)
P.S. Wyant, S. Strohmeier, B. Schäfer, B. Krenz, I.P. Assuncao, G.S. Lima, H. Jeske, Virology 427, 151–157 (2012)
P.S. Wyant, D. Gotthardt, B. Schäfer, B. Krenz, H. Jeske, Arch. Virol. 156, 347–352 (2011)
T. Paprotka, V. Metzler, H. Jeske, Arch. Virol. 155, 813–816 (2010)
M. Höhnle, P. Höfer, I.D. Bedford, R.W. Briddon, P.G. Markham, T. Frischmuth, Virology 290, 164–171 (2001)
P. Höfer, I.D. Bedford, P.G. Markham, H. Jeske, T. Frischmuth, Virology 236, 288–295 (1997)
Z.C. Wu, J.S. Hu, J.E. Polston, D.E. Ullman, E. Hiebert, Phytopathology 86, 608–613 (1996)
S. Morin, M. Ghanim, I. Sobol, H. Czosnek, Virology 276, 404–416 (2000)
T. Frischmuth, G. Zimmat, H. Jeske, Virology 178, 461–468 (1990)
H. Jeske, D. Menzel, G. Werz, Phytopathol. Z. 89, 289–295 (1977)
A.A. Donnell, H.E. Ballard Jr, P.D. Cantino, Syst. Bot. 37, 712–722 (2012)
S. Unseld, M. Ringel, P. Höfer, M. Höhnle, H. Jeske, I.D. Bedford, P.G. Markham, T. Frischmuth, Arch. Virol. 145, 1449–1454 (2000)
S. Unseld, M. Ringel, A. Konrad, S. Lauster, T. Frischmuth, Virology 274, 179–188 (2000)
T. Frischmuth, M. Engel, S. Lauster, H. Jeske, J. Gen. Virol. 78, 2675–2682 (1997)
P. Höfer, M. Engel, H. Jeske, T. Frischmuth, J. Gen. Virol. 78, 1785–1790 (1997)
D. Pohl, C. Wege, J. Gen. Virol. 88, 1034–1040 (2007)
B. Krenz, C. Wege, H. Jeske, J. Virol. Methods 169, 129–137 (2010)
R.W. Briddon, B.L. Patil, B. Bagewadi, M.S. Nawaz-ul-Rehman, C.M. Fauquet, BMC Evol. Biol. 10, 97 (2010)
S.W. Qin, B.M. Ward, S.G. Lazarowitz, J. Virol. 72, 9247–9256 (1998)
D. Haible, S. Kober, H. Jeske, J. Virol. Methods 135, 9–16 (2006)
H. Jeske, S. Kober, B. Schäfer, S. Strohmeier, Virus Genes 49, 312–324 (2014)
T. Hall, GERF Bull. Biosci. 2, 60–61 (2011)
R.R. Stocsits, I.L. Hofacker, C. Fried, P.F. Stadler, BMC Bioinform. 6, 160 (2005)
B.G. Hall, Mol. Biol. Evol. 30, 1229–1235 (2013)
S.L. Kosakovsky Pond, S.D.W. Frost, S.V. Muse, Bioinformatics 21, 676–679 (2005)
B. Böttcher, S. Unseld, H. Ceulemans, R.B. Russell, H. Jeske, J. Virol. 78, 6758–6765 (2004)
S.L. Kosakovsky Pond, S.D.W. Frost, Mol. Biol. Evol. 22, 1208–1222 (2005)
E. Noris, A.M. Vaira, P. Caciagli, V. Masenga, B. Gronenborn, G.P. Accotto, J. Virol. 72, 10050–10057 (1998)
A. Kheyr-Pour, K. Bananej, G.A. Dafalla, P. Caciagli, E. Noris, A. Ahoonmanesh, H. Lecoq, B. Gronenborn, Phytopathology 90, 629–635 (2000)
S. Liu, I.D. Bedford, R.W. Briddon, P.G. Markham, J. Gen. Virol. 78, 1791–1794 (1997)
I. Grigoras, T. Timchenko, L. Katul, A. Grande-Perez, H.J. Vetten, B. Gronenborn, J. Virol. 83, 10778–10787 (2009)
I. Grigoras, T. Timchenko, A. Grande-Perez, L. Katul, H.J. Vetten, B. Gronenborn, J. Virol. 84, 9105–9117 (2010)
B.D. Harrison, M.M. Swanson, D. Fargette, Physiol. Mol. Plant Pathol. 60, 257–271 (2002)
J. Schubert, A. Habekuss, K. Kazmaier, H. Jeske, Virus Res. 127, 61–70 (2007)
Acknowledgments
The authors are grateful to Prof. C. Wege, Drs. P. Wyant and K. Hipp for critical comments on the manuscript and providing leaf samples, to B. Schäfer and S. Kober for excellent technical assistance, and to D. Gotthardt for taking care of the plants. The advices in botanical taxonomy of Profs. P. D. Cantino, Ohio, and J. G. Rohwer, Hamburg, were very helpful. This work was supported by a trilateral ERA-PG/BMBF Project (BMBF 0313986).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11262_2014_1125_MOESM1_ESM.pptx
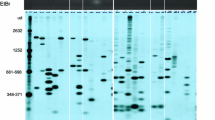
Supplementary material 1 Spreadsheet calculations to determine the fit between expected and observed fragments of the gels shown in Fig. 1 b, c, d, respectively. The logarithmic values of the molecular weights are plotted against migration distances transformed to Rf value probits as described previously [28].(PPTX 77 kb)
11262_2014_1125_MOESM2_ESM.xlsx
Supplementary material 2 Suppl. Table 1: Pairwise comparison of nucleotide sequence identities (SI) for AbMV DNA A and DNA B components after aligning using codaln algorithm. The values for AbMV D(S) and F(P) are highlighted.Suppl. Table 2: List of DNA A sequences compared to those of AbMV with accession numbers, long names and countries of collection as far as they can be retrieved from the data base entries.Suppl. Table 3: List of DNA B sequences compared to those of AbMV with accession numbers, long names and countries of collection as far as they can be retrieved from the data base entries.Suppl. Table 4: List of DNA A sequences of South American Abutilon-infecting and Sida micrantha mosaic viruses used for CP ORF comparisons, with accession numbers, long names and countries of collection as far as they can be retrieved from the data base entries.(XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Fischer, A., Strohmeier, S., Krenz, B. et al. Evolutionary liberties of the Abutilon mosaic virus cluster. Virus Genes 50, 63–70 (2015). https://doi.org/10.1007/s11262-014-1125-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-014-1125-1