Abstract
Genetically modified (GM) crops undergo large scale multi-location field trials to characterize agronomics, composition, and the concentration of newly expressed protein(s) [herein referred to as transgenic protein(s)]. The concentration of transgenic proteins in different plant tissues and across the developmental stages of the plant is considered in the safety assessment of GM crops. Reference or housekeeping proteins are expected to maintain a relatively stable expression pattern in healthy plants given their role in cellular functions. Understanding the effects of genotype, growth stage and location on the concentration of endogenous housekeeping proteins may provide insight into the contribution these factors could have on transgenic protein concentrations in GM crops. The concentrations of three endogenous proteins (actin, elongation factor 1-alpha, and glyceraldehyde 3-phosphate dehydrogenase) were measured in several different maize hybrids grown across multiple field locations over 2 years. Leaf samples were collected from healthy plants at three developmental stages across the growing seasons, and protein concentrations were quantified by indirect enzyme-linked immunosorbent assay (ELISA) for each protein. In general, the concentrations of these three endogenous proteins were relatively consistent across hybrid backgrounds, when compared within one growth stage and location (2–26%CV), whereas the concentrations of proteins in the same hybrid and growth stage across different locations were more variable (12–64%CV). In general, the protein concentrations in 2013 and 2014 show similar trends in variability. Some degree of variability in protein concentrations should be expected for both transgenic and endogenous plant-expressed proteins. In the case of GM crops, the potential variation in protein concentrations due to location effects is captured in the current model of multi-location field testing.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prior to commercialization, genetically modified (GM) crops undergo a comprehensive safety assessment to determine food, feed, and environmental safety. Large scale multi-location field trials are conducted to evaluate agronomic performance, compositional equivalence, and trait efficacy, including the quantification of newly expressed protein(s) [herein referred to as transgenic protein(s)]. A range of geographic locations and representative environments are selected for field studies to capture the potential variability in agronomic performance and phenotypic characteristics that may be due to the environment and other external factors (EFSA 2015). Agronomic performance of a hybrid is known to be affected by environmental factors like temperature, rainfall, soil type, and soil nutrients. Similarly, protein concentrations may also be influenced by external factors at each field location in addition to tissue type and growth stage (Dong and Li 2007; Trtikova et al. 2015; Wan et al. 2005). There are several examples in the literature demonstrating the potential variability in transgenic proteins. For example, the concentration of crystalline (cry) proteins from Bacillus thuringiensis (Bt) in GM cotton were previously observed to be influenced by the environment or field location, growth stage and tissue type (Greenplate 1999; Pettigrew and Adamczyk 2006; Siebert et al. 2009). Likewise, GM maize grown in field trials in India (Siebert et al. 2009) and in Germany (Zhang et al. 2016) both reported variable Bt protein concentrations, depending on the tissue type, growth stage, and field location. And, in a separate study, Bt protein concentrations in maize were shown to be variable under optimal and stressed environments (Trtikova et al. 2015).
While there is a general consensus that the concentration of transgenic proteins will have some inherent variability over time and across locations, the degree to which these external factors affect transgenic protein expression remains largely unknown and also depends on the characteristics of each protein and the promoter used to drive expression. Hence, to support safety assessments, transgenic proteins are typically quantified from multiple tissue types (e.g., leaf, stalk, root, pollen) and growth stages (e.g., early vegetative, reproductive and senescent stages) of the plant, and across multiple representative environments to capture potential variability.
Endogenous genes that encode for housekeeping proteins involved in basic cellular processes are typically assumed to have stable expression within a plant and are not thought to be influenced to a large extent by external factors in healthy plants. Hence, housekeeping genes are commonly utilized as internal controls for normalization in gene expression analyses (Czechowski et al. 2005), and housekeeping proteins can be used for normalization in protein analytical methods (Li et al. 2011). For example, Glyceraldehyde 3-Phosphate Dehydrogenase (GAPDH), an important component of the glycolytic pathway, is commonly used as an internal reference gene (Thellin et al. 1999) and can also be used as a reference protein (Li et al. 2011). Elongation factor 1-alpha (EF1-alpha) helps regulate protein synthesis through its role in the elongation of amino acid chains and has been identified as a stable reference protein for rice protein analysis (Li et al. 2011). Similarly, β-actin, which is essential for the structure of the cytoskeleton, is commonly used as an internal control in gene expression studies (Thellin et al. 1999). However, several studies have demonstrated that these, as well as other endogenous genes and proteins, can be influenced by a variety of factors (Czechowski et al. 2005; Li et al. 2011; Thellin et al. 1999).
The objective of this study was to characterize the effects of genotype, growth stage and location on the concentration of three endogenous housekeeping proteins [actin, EF1-alpha, and GAPDH] by indirect ELISA. Several traditionally bred, commercially available maize hybrids grown at multiple field trial locations in India over 2 years were used in this study. Experimental design and field study parameters comparable with those used to support India field trial requirements for GM crops prior to regulatory approval [Biosafety Research Level trials, which are governed by the Review Committee on Genetic Manipulation (RCGM) and the Genetic Engineering Approval Committee (GEAC), respectively; (IGMORIS 2008)] were used to assess the concentration of these endogenous housekeeping proteins in these traditional hybrids. Effort was made to follow similar field study planting, maintenance, and tissue sampling protocols as those used for GM crop field trials. Understanding the effects of genotype, growth stage and location on the variability of these endogenous housekeeping proteins may give some insight into the degree of variability that may be expected for transgenic proteins in GM crops. Although the variability observed in the concentration of the three endogenous proteins is not expected to be fully reflective of the variability that could be observed for all other proteins (including transgenic or other endogenous proteins), these housekeeping proteins are used in this study as an example to better understand the influence of genotype, growth stage and location on protein concentration in general.
Materials and methods
Field trials
Field trials were conducted in India, during the rainy season in 2013 and 2014. In 2013, six maize hybrids were grown in six locations (Aurangabad, Bengaluru, Davangere, Hyderabad, Jalandhar, and Udaipur; Online Resource 1 and 2). In 2014, six different maize hybrids were grown in four of the same locations of previous year along with two additional locations, Patna and Shigaon (Online Resource 1 and 2). Hybrids were sown in a randomized complete block design (RCBD) containing three blocks, with each block containing six row plots of twenty plants per each row. The maize hybrids were selected based on diverse genetic backgrounds and variable maturity. Field trial locations were selected to be representative of the environment in India where commercial maize hybrids are grown and based on their suitability for the relative maturity of the hybrids. All of the common agronomic practices (land preparation, inter-cultivation, irrigation, fertilizer application, herbicide and insecticide applications, etc.) were applied uniformly across locations.
Sample collection and sample preparation
Five healthy maize plants from each plot were randomly selected for sampling at three growth stages, early vegetative (VS), grand growth (GS) and reproductive (RS) stages representing the V3–V4, V10–V12, and R1–R2 stages of crop growth, respectively (Ritchie et al. 2005). From each plant, the second or third leaf from the top of the plant was excised, approximately 6–8 cm of the middle portion of the leaf (excluding the mid-rib) was separated, and the samples were pooled in a tube and stored immediately on dry ice. Samples were shipped to the laboratory on dry ice, lyophilized, finely ground, and stored at −80 °C until further analysis.
Confirmation of antibody specificity
To confirm the specificity of the commercially available antibodies to their respective proteins, total soluble protein was extracted from representative leaf tissues (GS stage from hybrids grown in 2014) and was subjected to western blot analysis. Homogenized leaf tissue (10 mg) was weighed into 1.2 ml polypropylene tubes and 600 µl of carbonate buffer (pH 9.6) was added. Sterilized steel beads were added to the tube and agitated for 1 min at 1200 strokes using a genogrinder (Spex, USA). The mix was centrifuged at 4000 rpm at 4 °C for 10 min, and 280 µl of supernatant was collected and mixed with 80 µl of 5× sample buffer (250 mM Tris HCl, pH 6.8, 10% SDS, 30% (v/v) glycerol, 10 mM DTT, 0.05% (w/v) Bromophenol blue) and 40 µl of 10X NuPage sample reducing agent (Invitrogen, USA). The samples were incubated at 95 °C for 5 min, and 30 µl of each sample was loaded on to a 12% SDS-PAGE gel (Invitrogen, USA) after a brief centrifugation at room temperature.
Gel electrophoresis and western blot analysis were performed to characterize actin, GAPDH and EF1-alpha proteins. Following gel electrophoresis (35 mA for 1.5 h), the separated proteins were blotted onto a nitrocellulose membrane with an Invitrogen iBlot system (Invitrogen, USA). The membrane was blocked with 5% skimmed milk, followed by incubation with an appropriate primary antibody diluted in 1% skimmed milk for two hours at room temperature. After washing three times with phosphate buffer saline containing 0.05% tween 20 (PBST), the membranes were incubated for an hour with an appropriate secondary antibody diluted in 1% skimmed milk. The details of primary and the secondary antibodies dilutions for detecting each protein are provided in Online Resource 3. After washing three times with PBST, the membrane was washed for 10 min in PBS, and was subsequently developed using a SuperSignal West Pico Chemiluminiscent substrate (Thermo Scientific, USA). Images were captured in Image Quant LAS 4000 system (GE Healthcare, Japan).
Protein quantification by ELISA
Preparation of standard curves
A relative quantification method was developed to quantify the expression of the three endogenous proteins. To prepare a standard curve for actin and GAPDH proteins, lyophilized leaf tissue from maize samples at RS stage were pooled to create a reference sample. To prepare a standard curve for EF-1 alpha, rice culm tissue at active tillering stage (25–35 days after transplantation) was used. The reference sample was weighed (10 mg) into 1.2 ml tubes and extracted with 600 µl of carbonate buffer (pH 9.6). Following centrifugation, supernatant was collected into new tubes and serially diluted, and an arbitrary number (units/ml) was assigned to the top dilution. Proportionate numbers (units/ml) were assigned to the remaining dilutions deriving a relative standard curve. For actin, the standard curve dilutions ranged from 1:200, 1:400, 1:500, 1:600, and 1:800, which were represented by 5, 2.5, 2.0, 1.67, and 1.25 units/ml, respectively. For GAPDH, the standard curve dilutions ranged from 1:100, 1:200, 1:400, 1:800, 1:1600 and 1:2000, which were represented by 10, 5, 2.5, 1.25, 0.625, and 0.5 units/ml, respectively. For EF-1 alpha, the standard curve dilutions ranged from 1:800, 1:1600, 1:2000, 1:2400 and 1:3200, 1:4000 and 1:5000, which were represented by 0.125, 0.0625, 0.05, 0.042, 0.03125, 0.025, and 0.02 units/ml, respectively.
ELISA
On 96-well plates, three replicates of the appropriate standard curve, and duplicates for each test sample were loaded (100 µl per well), and the plates were incubated at room temperature for 1 h followed by an overnight incubation at 4 °C. The coating solutions were aspirated and the plates were blocked with 100 µl of 1% skimmed milk, 0.5% skimmed milk, and 300 µl of sodium caseinate solution (0.2% sodium caseinate and 5% sucrose) for actin, EF1-alpha, and GAPDH proteins, respectively. Following blocking, the plates were incubated with 100 µl of primary antibody diluted in PBST (Online Resource 3) at 24 °C for 1–2 h. The plates were washed three times with PBST and incubated with secondary antibody diluted in PBST (Online Resource 3) at 24 °C for 1–2 h. After washing three times with PBST, plates were incubated with 100 µl per well of TMB for 30 min, and the reaction was stopped with 1 N HCl. The resultant color intensity was read with Spectramax M5 Spectrophotometer (Molecular Devices, USA) at wavelength settings of 450 nm minus 650 nm. Softmax Gxp software was used to generate the quadratic curve or 4-parameter logistics curve for the standard curves for each protein, and the resultant optical density (OD) values from samples were converted to arbitrary units/ml by interpolating to the generated standard curve. All the standard curves have shown R2 value > 0.98.
Statistical analysis
For a given endogenous protein in a given growth stage, data was analyzed using the following linear mixed model: yijk = µ + hi + ℓj + bk(j) + (hℓ)ij + εijk; {hi ~ iid N(0, σ 2 Hybrid ), ℓj ~ iid N(0, σ 2 Location ), bk(j) ~ iid N(0, σ 2 Block ), (hℓ)ij ~ iid N(0, σ 2 Hybrid× Location ), and εijk ~ iid N(0, σ 2 Error )}. Where µ denotes the overall mean (fixed effect), hi denotes the effect of the ith hybrid (random effect), ℓj denotes the effect of the jth location (random effect), bk(j) denotes the effect of the kth block within the jth location (random effect), (hℓ)ij denotes the interaction between the ith hybrid and the jth location (random effect), and εijk denotes the random error associated with the sample collected from the plot assigned the ith hybrid in the kth block of the jth location (random effect or residual). Notation ~ iid N(0, σ 2 a ) indicates random variables that are independent and identically distributed (iid) as normal with zero mean and variance σ 2 a . Subscript a represents the corresponding source of variation. Statistical analyses were conducted using SAS software, Version 9.3 (SAS Institute Inc., Cary, NC, USA). SAS PROC MIXED was utilized to fit the model and to generate variance component estimates; therefore, this linear mixed model analysis is also called variance component analysis. σ2Hybrid is the variance component for hybrids, which is also understood as genotypic variance. σ2Location is the variance component for location, which is also understood as environmental variance. σ 2 Hybrid ×Location is the variance component for hybrid and location interactions, which is also understood as G × E (genotype by environment) variance. σ 2 Block and σ 2 Error are variance components for block effects and plot effects within location, which are understood as local environmental variance. These variance components add up to the total variance for a given endogenous protein in a given growth stage. To evaluate and compare relative contribution of each variance component, estimated variance values were normalized to 100%.
Results
Confirmation of antibody specificity
Western blot analysis was used to confirm the specificity of the antibodies to their respective proteins. A single prominent band was observed for the actin, GAPDH, and EF1-alpha proteins with the expected molecular weight of approximately 41.3, 37, and 49.1 kDa, respectively (Online Resource 4). Based on these results, these commercial antibodies were determined to be specific for the respective endogenous proteins.
Relative expression of actin, EF1-alpha and GAPDH proteins across locations and hybrids
The effect of hybrid background on the concentration of each endogenous protein was determined comparing relative concentration of the protein (units/ml) in all the hybrids grown at a single location. The coefficient of variation (%CV—Hybrid) represents the variability in the concentration of endogenous proteins that is related to hybrid background (i.e., genotypic variance). To determine the relative effect of location on the concentration of endogenous proteins, the expression of a protein (units/ml) in each hybrid background was compared across the locations. The coefficient of variation (%CV—Location) represents the variability in the concentration of endogenous proteins that is related to location (i.e., environmental variance). The results from 2013 to 2014 growing seasons and each growth stage were kept separate and were not pooled for the analyses.
The relative effect of hybrid background on the concentration of actin (units/ml), represented as %CV—Hybrid ranged between 5–22% and 5–24% across all growth stages, in the years of 2013 and 2014, respectively (Table 1). The relative effect of location on the concentration of actin (units/ml), represented as %CV—Location ranged between 22–36% and 31–51% across all growth stages, in the years of 2013 and 2014, respectively (Table 1). The general trend observed for the concentration of actin were consistent for the GAPDH and the EF1-alpha proteins, with hybrid background contributing less than location to the total variability in relative protein concentration. The %CV—Hybrid for GAPDH ranged from 4–25% to 2–26% in 2013 and 2014, respectively, whereas the %CV—Location ranged from 20–42% in 2013 to 15–49% in 2014 (Table 2). Similarly, the %CV—Hybrid for EF1-alpha ranged from 4–21% in 2013 to 5–25% in 2014, whereas the %CV—Location ranged from 13–31% in 2013 to 12–64% in 2014 (Table 3). In general, the results for all three endogenous proteins in the 2013 and 2014 growing seasons provide evidence that both hybrid background and location contribute to the observed variation in their concentration; however, in both years, location contributed more to the total variability observed in this study.
Relative concentration of actin, EF1-alpha and GAPDH proteins across growth stages
Variability in the concentration of the three endogenous proteins at each growth stage was analyzed across all the locations and all the hybrids (Table 4). The effect of growth stage on the relative concentration of the GAPDH was highest, with %CV ranging from 35% in 2013 to 55% in 2014. The %CV for actin ranged from 19% in 2013 to 12% in 2014, and the %CV for EF1-alpha ranged from 15% in 2013 to 11% in 2014.
Variance component analysis of endogenous protein concentrations
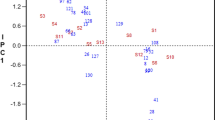
Variance component analysis of the relative concentration of each protein was used to characterize the amount of variation contributed by location, hybrid, location × hybrid, block (replication) and residual across all growth stages (Fig. 1). The results from 2013 to 2014 growing seasons and each growth stage were assessed separately. In 2013, the variation in the concentration of actin protein that was contributed by location was estimated to be 47, 75, and 74% for the VS, GS, and RS growth stages, respectively. Similarly, for the EF1-alpha and GAPDH proteins, location contributed 47, 62, and 40%, and 51, 65, and 40% of the variation, respectively for these three growth stages. The % variation due to hybrid background was very low relative to the other factors considered, ranging from 0 to 8% for all the three proteins. The amount of variation contributed by location x hybrid interaction was also low, with the exception of the EF1-alpha and GAPDH proteins at the RS growth stage, where it contributed 20 and 25% of the total variability, respectively. Block-to-block variability contributed 0–17% variation across the three growth stages and across the three proteins.
Variance component analysis of the contribution of location, hybrid, location × hybrid, block (replication) and residual to the variation in relative Actin, GAPDH and EF1-alpha protein concentrations across growth stages [vegetative (VS), grand growth (GS) and reproductive stage (RS), which correspond to the V3–V4, V10–V12, and R1–R2 stages of crop growth, respectively]. Panels a and b represent the analysis of 2013 and 2014 years, respectively
In 2014, the amount of variation in the concentration of the actin protein that was contributed by location was estimated to be 79, 85, and 74% for the VS, GS, and RS growth stages, respectively. Similarly, for the EF1-alpha and GAPDH proteins, location contributed 40, 74, and 63%, and 31, 84 and 25% of the variation, respectively for the same growth stages. The % variation due to hybrid background in 2014 was very low as it was in 2013, ranging from 1 to 7% for all three proteins, with a few exceptions. For example, hybrid background contributed 17% of the total variability at the at VS stage for EF1-alpha and 19% at RS stage for GAPDH proteins. The amount of variation contributed by the location x hybrid interaction and replication were both low, with %CV ranging from 1–10% to 0–14%, respectively.
Discussion
For the safety assessment of a GM crop, the concentration of a transgenic protein is measured across representative growing environments, within relevant plant tissues, and throughout the growing season. Agronomic performance of a crop may be influenced, to varying degrees, by biotic and abiotic factors in a growing season. Similarly, transgenic proteins may also be influenced by temporal and spatial differences, in addition to these factors (Dong and Li 2007). Variability in protein expression is not unique to transgenic proteins in GM crops, and variability across the growing season, across locations, or within different plant tissues is expected for all plant-expressed proteins.
Endogenous housekeeping proteins were used as an example in this study to demonstrate the degree to which plant-expressed proteins can vary over time and across different environments. Housekeeping proteins typically maintain relatively stable concentrations in healthy plants, due to their role in cellular functions, however they also may be influenced to some degree by external environmental factors without affecting the health and safety of the plant. Understanding the factors that contribute to the variability in concentration of these housekeeping proteins may provide insight to explain the variability that is expected for transgenic proteins in GM crops. Actin, EF1-alpha and GAPDH were selected as example proteins for assessment because they are well studied (Brunner et al. 2004; Cruz et al. 2009; Gimeno et al. 2014; Gutha et al. 2010; Li et al. 2011; Paolacci et al. 2009), and indirect quantitative ELISA methods could be developed with commercially available antibodies.
In this study, the concentrations of all three endogenous proteins were influenced by growth stage, hybrid background, and location, to some extent. The concentrations of these three endogenous proteins were relatively consistent across hybrid backgrounds, when compared within one growth stage and location (2–26%CV), whereas concentrations of the proteins in the same hybrid and growth stage across different locations were more variable (12–64%CV). In general, the effects of genotype on the variability of these stably expressed housekeeping proteins was small, relative to total protein variability, and the observed trends were relatively consistent in both 2013 and 2014. The inherent variability of endogenous plant-expressed proteins is not expected to affect the health or safety of the plant, and degree of variability observed for these endogenous proteins is important to consider with respect to the variability that can be expected for other plant-expressed proteins, including transgenic proteins.
Like endogenous proteins, the growth stage of the plant is expected to influence transgenic protein concentrations, and there are many examples where transgenic proteins have been variable across the growing season in different crops (Kranthi et al. 2005; Nguyen and Jehle 2007; Zhang et al. 2011). The degree to which growth stage affects protein concentration will be different depending on the characteristics of each protein, the promotor that drives the protein expression, and the crop (Dong and Li 2007; Trtikova et al. 2015; Wan et al. 2005). Additionally, as reviewed by Dong and Li (2007), soil, microclimate, pest pressure, and crop management practices may all affect the concentration of transgenic proteins. There are also many examples of studies that have identified location to be a factor influencing transgenic protein concentrations in GM crops (Adamczyk et al. 2001; Kamath et al. 2010; Nguyen and Jehle 2007; Zhang et al. 2016). Because some degree of protein variability is expected, transgenic protein concentrations in GM plants are extensively characterized across growth stages, relevant tissue types, and multiple growing environments. Locations for regulatory field trials of GM crops are selected to be representative of a wide range of growing conditions, as they may differ in many meteorological as well as agronomic conditions (Pettigrew and Adamczyk 2006).
The concentrations of transgenic proteins in GM plants are measured across representative growing environments, within relevant plant tissues, and throughout the growing season. Protein concentrations in relevant plant tissues are used to estimate food, feed and environmental exposure, as part of the safety assessment of GM crops. Hazard studies are typically conducted at very high doses, relative to the estimated exposure. Any potential site to site, hybrid to hybrid, and seasonal variability in transgenic protein concentrations is very small, relative to the high doses tested as part of the safety assessment. Therefore, while some degree of variability in protein concentrations should be expected for both transgenic and endogenous plant-expressed proteins, this variability is captured in the current model of multi-location field testing, and will not affect the food, feed and environmental safety assessment of a GM crop.
References
Adamczyk J, Hardee D, Adams L, Sumerford D (2001) Correlating differences in larval survival and development of bollworm (Lepidoptera: Noctuidae) and fall armyworm (Lepidoptera: Noctuidae) to differential expression of Cry1A (c) δ-endotoxin in various plant parts among commercial cultivars of transgenic Bacillus thuringiensis cotton. J Econ Entomol 94:284–290. https://doi.org/10.1603/0022-0493-94.1.284
Brunner AM, Yakovlev IA, Strauss SH (2004) Validating internal controls for quantitative plant gene expression studies. BMC Plant Biol 4:1. https://doi.org/10.1186/1471-2229-4-14
Cruz F et al (2009) Evaluation of coffee reference genes for relative expression studies by quantitative real-time RT-PCR. Mol Breed 23:607–616. https://doi.org/10.1007/s11032-009-9259-x
Czechowski T, Stitt M, Altmann T, Udvardi MK, Scheible W-R (2005) Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol 139:5–17. https://doi.org/10.1104/pp.105.063743
Dong H, Li W (2007) Variability of endotoxin expression in Bt transgenic cotton. J Agron Crop Sci 193:21–29. https://doi.org/10.1111/j.1439-037X.2006.00240.x
EFSA (2015) Guidance on the agronomic and phenotypic characterisation of genetically modified plants. EFSA J 13:4128
Gimeno J, Eattock N, Van Deynze A, Blumwald E (2014) Selection and validation of reference genes for gene expression analysis in switchgrass (Panicum virgatum) using quantitative real-time RT-PCR. PLoS ONE 9:e91474. https://doi.org/10.1371/journal.pone.0091474
Greenplate JT (1999) Quantification of Bacillus thuringiensis insect control protein Cry1Ac over time in Bollgard Cotton fruit and terminals. J Econ Entomol 92:1377–1383. https://doi.org/10.1093/jee/92.6.1377
Gutha LR, Casassa LF, Harbertson JF, Naidu RA (2010) Modulation of flavonoid biosynthetic pathway genes and anthocyanins due to virus infection in grapevine (Vitis vinifera L.) leaves. BMC Plant Biol 10:187. https://doi.org/10.1186/1471-2229-10-187
IGMORIS (2008) Guidelines for the conduct of confined field trials of regulated, GE plants. Indian GMO Research Information System, New Delhi
Kamath S, Anuradha S, Vidya H, Mohan K, Dudin Y (2010) Quantification of Bacillus thuringiensis Cry1Ab protein in tissues of YieldGard® (MON810) corn hybrids tested at multiple field locations in India. Crop Prot 29:921–926. https://doi.org/10.1016/j.cropro.2010.04.018
Kranthi KR et al (2005) Temporal and intra-plant variability of Cry1Ac expression in Bt-cotton and its influence on the survival of the cotton bollworm, Helicoverpa armigera (Hübner) (Noctuidae:Lepidoptera). Curr Sci 89:291–298
Li X et al (2011) Identification and validation of rice reference proteins for western blotting. J Exp Bot 62:4763–4772. https://doi.org/10.1093/jxb/err084
Nguyen H, Jehle J (2007) Quantitative analysis of the seasonal and tissue-specific expression of Cry1Ab in transgenic maize Mon810. J Plant Dis Prot 114:82–87. https://doi.org/10.1007/BF03356208
Paolacci AR, Tanzarella OA, Porceddu E, Ciaffi M (2009) Identification and validation of reference genes for quantitative RT-PCR normalization in wheat. BMC Mol Biol 10:27. https://doi.org/10.1186/1471-2199-10-11
Pettigrew W, Adamczyk J (2006) Nitrogen fertility and planting date effects on lint yield and Cry1Ac (Bt) endotoxin production. Agron J 98:691–697. https://doi.org/10.2134/agronj2005.0327
Ritchie S, Hanway J, Benson G (2005) How a corn plant develops. Iowa State University of Science and Technology Cooperative Extension Service, Ames, IA, USA; Special Report No. 48
Siebert MW et al (2009) Quantification of Cry1Ac and Cry1F Bacillus thuringiensis insecticidal proteins in selected transgenic cotton plant tissue types. J Econ Entomol 102:1301–1308. https://doi.org/10.1603/029.102.0357
Thellin O et al (1999) Housekeeping genes as internal standards: use and limits. J Biotech 75:291–295. https://doi.org/10.1016/S0168-1656(99)00163-7
Trtikova M, Wikmark OG, Zemp N, Widmer A, Hilbeck A (2015) Transgene expression and Bt protein content in transgenic Bt maize (MON810) under optimal and stressful environmental conditions. PLoS ONE 10:e0123011. https://doi.org/10.1371/journal.pone.0123011
Wan P, Zhang Y, Wu K, Huang M (2005) Seasonal expression profiles of insecticidal protein and control efficacy against Helicoverpa armigera for Bt cotton in the Yangtze River valley of China. J Econ Entomol 98:195–201. https://doi.org/10.1603/0022-0493-98.1.195
Zhang Y, Li Y, Zhang Y, Chen Y, Wu K, Peng Y, Guo Y (2011) Seasonal expression of Bt proteins in transgenic rice lines and the resistance against Asiatic rice borer Chilo suppressalis (Walker). Environ Entomol 40:1323–1330. https://doi.org/10.1603/en11035
Zhang Y, Zhang J, Lan J, Wang J, Liu J, Yang M (2016) Temporal and spatial changes in Bt toxin expression in Bt-transgenic poplar and insect resistance in field tests. J For Res 27:1249–1256. https://doi.org/10.1007/s11676-016-0254-x
Acknowledgements
We thank John Duesing and Mary Locke for their help in conceiving and conduct of the entire study. We acknowledge Ramya Rajagopal and Srikanth Vemula for their help in conducting the field experiments. We also profusely thank Ashok Nandyala, Raghu Bheemanathi, and Rajani Talluri for their technical help in conducting ELISA experiments. We thank many others for critical review of the manuscript.
Author information
Authors and Affiliations
Contributions
SB and LRG conceived and coordinated the study. LRG, DP, AV, and SB designed the lab experiments, and DP and AV performed them. JS provided suggestions for standardization of ELISA methods. ST, CE and AV performed the western blot experiments. VK and HR designed and coordinated the field experiment. BH performed statistical analysis. LRG, ST, and JA wrote the manuscript. All the authors reviewed and accepted the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gutha, L.R., Purushottam, D., Veeramachaneni, A. et al. Expression of endogenous proteins in maize hybrids in a multi-location field trial in India. Transgenic Res 27, 331–342 (2018). https://doi.org/10.1007/s11248-018-0077-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-018-0077-y