Abstract
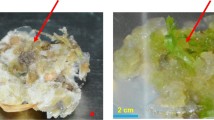
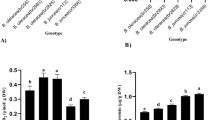
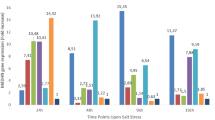
Soil salinity is one of the major abiotic stress factors that limits cotton (Gossypium hirsutum L.) production worldwide. Although cotton is categorised as a salt-tolerant plant, its tolerance level can vary depending on the cultivar. The present study aimed to evaluate the growth and biochemical responses of two cotton cultivars (Carmen and NM-503) to salt stress under tissue culture conditions by using an in vitro selection technique. Hypocotyl explants were cultured on callus formation medium (full-strength MS (Murashige and Skoog medium) macro- and micro-salts + B5 (Gamborg medium) vitamins + 0.1 mg/L 2,4-D (2,4-dichlorophenoxyacetic acid) + 0.5 mg/L kinetin) containing 0, 100, 200 or 400 mM NaCl for 4 weeks. Compared to Carmen cultivar, NM-503 cultivar showed an improved growth performance under non-stress control conditions; however, both cultivars showed similar growth and browning tendencies when exposed to salinity stress. Although both cultivars showed a significant decrease in biomass accumulation and relative growth rate following treatment with 200 and 400 mM NaCl, they were tolerant to NaCl stress up to 200 mM in terms of callus survival rate. Carmen cultivar showed the lowest photosynthetic pigment content after treatment with 400 mM NaCl; however, both cultivars showed no significant differences in the photosynthetic pigment content between NaCl concentrations. Furthermore, in contrast to Carmen cultivar, NM-503 cultivar did not accumulate proline in response to 200 and 400 mM NaCl treatment. Under salinity stress, NM-503 cultivar exhibited lower lipid peroxidation level than Carmen cultivar. Both cultivars showed no significant differences in superoxide dismutase activity in control and NaCl-treated groups, except for the 400 mM NaCl-treated group in Carmen cultivar. Apart from constitutive enzyme activity, both cultivars showed similar catalase activities at all concentrations of NaCl treatment. In both cultivars, peroxidase and ascorbate peroxidase activities increased in response to an increase in NaCl concentrations. While glutathione reductase activity gradually increased in NM-503 cultivar in response to NaCl treatment, it gradually decreased in Carmen cultivar. In conclusion, both cultivars showed similar growth response under salinity stress, and this growth response was significantly restricted by 200 and 400 mM NaCl treatment. Among all studied parameters, the most distinct response of the cultivars to salinity stress was reflected by the lipid peroxidation level, which indicates the differences in the mechanism of cellular antioxidant protection between the cultivars. Despite the absence of a salt exclusion mechanism used by intact plants, the high level of resilience shown by the calli of NM-503 cultivar against severe salt stress conditions may be attributed to its effective cellular antioxidant defence mechanism.
Key message
Salinity stress tolerance of two cotton cultivars (Carmen and NM-503) was evaluated by in vitro selection. Among the growth and biochemical parameters, the lipid peroxidation level was the most decisive parameter, and NM-503 cultivar was more salt tolerant than Carmen cultivar.




Similar content being viewed by others
Data Availability
All data generated or analysed during this study are included in this published article as tables and figures.
References
Ahmad S, Khan NI, Iqbal MZ, Hussain A, Hassan M (2002) Salt tolerance of cotton (Gossypium hirsutum L). Asian J Plant Sci 1(6):715–719. https://doi.org/10.3923/ajps.2002.715.719
Akhtar J, Saqib ZA, Sarfraz M, Saleem I, Haq MA (2010) Evaluating salt tolerant cotton genotypes at different levels of NaCl stress in solution and soil culture. Pak J Bot 42(4):2857–2866
Alhasnawi AN, Che Radziah CMZ, Kadhimi AA, Isahak A, Mohamad A, Yusoff WMW (2016) Enhancement of antioxidant enzyme activities in rice callus by ascorbic acid under salinity stress. Biol Plant 60(4):783–787. https://doi.org/10.1007/s10535-016-0603-9
Arnon DI (1949) Copper enzymes in isolated chloroplasts polyphenoloxidase in Beta vulgaris. Plant Physiol 24(1):1–15. https://doi.org/10.1104/pp.24.1.1
Ashraf M (2002) Salt tolerance of cotton: some new advances. Crit Rev Plant Sci 21(1):1–30. https://doi.org/10.1080/0735-260291044160
Ashraf M, Harris P (2004) Potential biochemical indicators of salinity tolerance in plants. Plant Sci 166(1):3–16. https://doi.org/10.1016/j.plantsci.2003.10.024
Aydin Y, Ipekci Z, Talas-Oğraş T, Zehir A, Bajrovic K, Gozukirmizi N (2004) High frequency somatic embryogenesis in cotton. Biol Plant 48(4):491–495. https://doi.org/10.1023/B:BIOP.0000047142.07987.e1
Baohong Z, Yun Z (1999) Effects of NaCI stress on cotton tissue culture and plant regeneration. Pak J Biol Sci 2:1085–1087. https://doi.org/10.3923/pjbs.1999.1085.1087
Basal H (2010) Response of cotton (Gossypium hirsutum L.) genotypes to salt stress. Pak J Bot 42(1):505–511
Basal H, Canavar O, Khan NU, Cerit CS (2011) Combining ability and heterotic studies through line × tester in local and exotic upland cotton genotypes. Pak J Bot 43(3):1699–1706
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. https://doi.org/10.1007/BF00018060
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44(1):276–287. https://doi.org/10.1016/0003-2697(71)90370-8
Bergmeyer HU, Gawehn K (1970) Methoden der enzymatischen Analyse, vol 432. Weinheim, Verlag Chemie
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Cakmak I, Horst WJ (1991) Effect of aluminium on lipid peroxidation, superoxide dismutase, catalase, and peroxidase activities in root tips of soybean (Glycine max). Physiol Plant 83(3):463–468. https://doi.org/10.1111/j.1399-3054.1991.tb00121.x
Carlberg I, Mannervik B (1985) Glutathione reductase. Meth Enzymol 113:484–490. https://doi.org/10.1016/S0076-6879(85)13062-4
Cha-um S, Somsueb S, Samphumphuang T, Kirdmanee C (2013) Salt tolerant screening in eucalypt genotypes (Eucalyptus spp.) using photosynthetic abilities, proline accumulation, and growth characteristics as effective indices. In Vitro Cell Dev Biol–Plant 49:611–619. https://doi.org/10.1007/s11627-013-9537-5
Chelli-Chaabouni A, Mosbah AB, Maalej M, Gargouri K, Gargouri-Bouzid R, Drira N (2010) In vitro salinity tolerance of two pistachio rootstocks: Pistacia vera L. and P. atlantica Desf. Environ Exp Bot 69(3):302–312. https://doi.org/10.1016/j.envexpbot.2010.05.010
El Yacoubi H, Ayolié K, Rochdi A (2010) In vitro cellular salt tolerance of Troyer citrange: changes in growth and solutes accumulation in callus tissue. Int J Agric Biol 12(2):187–193
El-Mahdy MT, Youssef M, Elazab DS (2022) In vitro screening for salinity tolerance in pomegranate (Punica granatum L.) by morphological and molecular characterization. Acta Physiol Plant 44:27. https://doi.org/10.1007/s11738-022-03361-2
Gamborg OL, Miller RA, Ojima K (1968) Nutrient requirements of suspension cultures of soybean root cells. Exp Cell Res 50(1):151–158. https://doi.org/10.1016/0014-4827(68)90403-5
Gandonou C, Abrini J, Idaomar M, Skali-Senhaji N (2005) Effects of NaCl on growth and ion and proline accumulation in sugarcane (Saccharum sp.) callus culture. Belg J Bot 138(2):173–180. https://doi.org/10.2307/20794582
Ghane SG, Lokhande VH, Nikam TD (2014) Growth, physiological, and biochemical responses in relation to salinity tolerance for in vitro selection in oil seed crop Guizotia abyssinica Cass. J Crop Sci Biotechnol 17:11–20. https://doi.org/10.1007/s12892-013-0084-8
Golkar P, Taghizadeh M (2018) In vitro evaluation of phenolic and osmolite compounds, ionic content, and antioxidant activity in safflower (Carthamus tinctorius L.) under salinity stress. Plant Cell Tiss Organ Cult 134:357–368. https://doi.org/10.1007/s11240-018-1427-4
González A, Steffen KL, Lynch JP (1998) Light and excess manganese: implications for oxidative stress in common bean. Plant Physiol 118(2):493–504. https://doi.org/10.1104/pp.118.2.493
Gossett DR, Millhollon EP, Lucas MC, Banks SW, Marney MM (1994a) The effects of NaCl on antioxidant enzyme activities in callus tissue of salt-tolerant and salt-sensitive cotton cultivars (Gossypium hirsutum L). Plant Cell Rep 13:498–503. https://doi.org/10.1007/BF00232944
Gossett DR, Millhollon EP, Lucas MC (1994b) Antioxidant response to NaCl stress in salt-tolerant and salt-sensitive cultivars of cotton. Crop Sci 34:706–714. https://doi.org/10.2135/cropsci1994.0011183X003400030020x
Guo H, Li S, Min W, Ye J, Hou Z (2019) Ionomic and transcriptomic analyses of two cotton cultivars (Gossypium hirsutum L.) provide insights into the ion balance mechanism of cotton under salt stress. PLoS ONE 14(12):e0226776. https://doi.org/10.1371/journal.pone.0226776
Hasanuzzaman M, Bhuyan MHMB, Anee TI, Parvin K, Nahar K, Mahmud JA, Fujita M (2019) Regulation of ascorbate-glutathione pathway in mitigating oxidative damage in plants under abiotic stress. Antioxidants 8(9):384. https://doi.org/10.3390/antiox8090384
Herzog V, Fahimi HD (1973) A new sensitive colorimetric assay for peroxidase using 3,3′-diaminobenzidine as hydrogen donor. Anal Biochem 55(2):554–562. https://doi.org/10.1016/0003-2697(73)90144-9
Hossain Z, Mandal AKA, Datta SK, Biswas AK (2007) Development of NaCl-tolerant line in Chrysanthemum morifolium Ramat. Through shoot organogenesis of selected callus line. J Biotechnol 129(4):658–667. https://doi.org/10.1016/j.jbiotec.2007.02.020
Juturu VN, Mekala GK, Kirti PB (2015) Current status of tissue culture and genetic transformation research in cotton (Gossypium spp). Plant Cell Tiss Organ Cult 120:813–839. https://doi.org/10.1007/s11240-014-0640-z
Kruglova NN, Seldimirova OA, Zinatullina AE (2018) In vitro callus as a model system for the study of plant stress-resistance to abiotic factors (on the example of cereals). Biol Bull Rev 8:518–526. https://doi.org/10.1134/S2079086418060063
Liang W, Ma X, Wan P, Liu L (2018) Plant salt-tolerance mechanism: a review. Biochem Biophys Res Commun 495(1):286–291. https://doi.org/10.1016/j.bbrc.2017.11.043
Lichtenthaler HK, Wellburn AR (1983) Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem Soc Trans 11(5):591–592. https://doi.org/10.1042/bst0110591
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444(2):139–158. https://doi.org/10.1016/j.abb.2005.10.018
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7(9):405–410. https://doi.org/10.1016/s1360-1385(02)02312-9
Mohanraj R (2016) In vitro regeneration of salt-tolerant plants. In: Anis M, Ahmad N (eds) Plant tissue culture: propagation, conservation and crop improvement. Springer, Singapore, pp 299–307. https://doi.org/10.1007/978-981-10-1917-3_13
Mohsin A, Tahmina N, Ayesha J, Yonghong Z, Jing L, Huangyang Z, Jie W, Chengbin X, Shenjie W, Alamin A (2022) Evaluation of Thellungiella halophila ST7 for improving salt tolerance in cotton. J Cotton Res 5:1. https://doi.org/10.1186/s42397-021-00108-1
Munawar W, Hameed A, Khan MKR (2021) Differential morphophysiological and biochemical responses of cotton genotypes under various salinity stress levels during early growth stage. Front Plant Sci 12:622309. https://doi.org/10.3389/fpls.2021.622309
Munns R, Tester M (2008) Mechanism of salinity tolerance. Annu Rev Plant Biol 59:651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15(3):473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Nascimento ECS, do Nascimento R, da Silva AAR, de Castro Bezerra CV, Batista MC, de Sá Almeida Veloso LL, de Araújo Pereira MC, Oliveira H (2019) Growth and photosynthetic pigments of cotton cultivars irrigated with saline water. Agric Sci 10(1): 81–91. https://doi.org/10.4236/as.2019.101007
Patade VY, Bhargava S, Suprasanna P (2012) Effects of NaCl and iso-osmotic PEG stress on growth, osmolytes accumulation and antioxidant defense in cultured sugarcane cells. Plant Cell Tissue Organ Cult 108:279–286. https://doi.org/10.1007/s11240-011-0041-5
Pérez-Clemente RM, Gómez-Cadenas A (2012) In vitro tissue culture, a tool for the study and breeding of plants subjected to abiotic stress conditions. In: Leva A, Rinaldi LMR (eds) Recent advances in plant in vitro culture. InTechOpen, Rijeka, Croatia, pp 91–108
Queirós F, Fidalgo F, Santos I, Salema R (2007) In vitro selection of salt tolerant cell lines in Solanum tuberosum L. Biol Plant 51:728–734. https://doi.org/10.1007/s10535-007-0149-y
Rai MK, Kalia RK, Singh R, Gangola MP, Dhawan AK (2011) Developing stress tolerant plants through in vitro selection – an overview of the recent progress. Environ Exp Bot 71(1):89–98. https://doi.org/10.1016/j.envexpbot.2010.10.021
Shibli RA, Al-Juboory K (2002) Comparative responses of “nabali” olive microshoot, callus, and suspension cell cultures to salinity and water deficit. J Plant Nutr 25(1):61–74. https://doi.org/10.1081/PLN-100108780
Singh D, Kaur S, Kumar A (2020) In vitro drought tolerance in selected elite clones of Eucalyptus tereticornis Sm. Acta Physiol Plant 42:17. https://doi.org/10.1007/s11738-019-3009-4
Trolinder NL, Goodin JR (1987) Somatic embryogenesis and plant regeneration in Gossypium hirstum L. Plant Cell Rep 6(3):231–234. https://doi.org/10.1007/BF00268487
Xiong Y, Liang H, Yan H, Guo B, Niu M, Chen S, Jian S, Ren H, Zhang X, Li Y, Zeng S, Wu K, Zheng F, da Silva JAT (2019) NaCl-induced stress: physiological responses of six halophyte species in in vitro and in vivo culture. Plant Cell Tissue Organ Cult 139:531–546. https://doi.org/10.1007/s11240-019-01697-1
Zhang Y, Zhang Y, Yu J, Zhang H, Wang L, Wang S, Guo S, Miao Y, Chen S, Liand Y, Dai S (2019) NaCl-responsive ROS scavenging and energy supply in alkali grass callus revealed from proteomic analysis. BMC Genomics 20:990–1006. https://doi.org/10.1186/s12864-019-6325-6
Zhang J, Zhang P, Huo X, Gao Y, Chen Y, Song Z, Wang F, Zhang J (2021) Comparative phenotypic and transcriptomic analysis reveals key responses of upland cotton to salinity stress during postgermination. Front Plant Sci 12. https://doi.org/10.3389/fpls.2021.639104
Zheng J, Zhang Z, Gong Z, Liang Y, Sang Z, Xu Y, Li X, Wang J (2022) Genome-wide association analysis of salt-tolerant traits in terrestrial cotton at seedling stage. Plants 11(1):97. https://doi.org/10.3390/plants11010097
Acknowledgements
We are grateful to Ege University Planning and Monitoring Coordination of Organizational Development and Directorate of Library and Documentation for their support in editing and proofreading service of this study.
Funding
This project was supported by the Ege University Scientific Research Projects Coordination Unit (project number: FYL-2018-20040).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Experiments were conducted by MS. Data collection and analysis were performed by MS, LYA and MB. The first draft of the manuscript was written by MB and critically edited by LYA and AG. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Informed consent
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Communicated by Christell van der Vyver.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sacu, M., Aktas, L.Y., Bayraktar, M. et al. Growth and antioxidant defence in hypocotyl-derived calli of two cotton cultivars with contrasting salt tolerance. Plant Cell Tiss Organ Cult 154, 297–309 (2023). https://doi.org/10.1007/s11240-023-02569-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-023-02569-5