Abstract
The paper presents a technique for micropropagation of endangered in Europe and extinct in Poland Pulsatilla vulgaris for ex situ conservation of the genetic resources. Genotype-dependent induction of somatic embryogenesis and rooting was revealed in series of two experiments (I and II) performed under the same experimental conditions. Shoot tips of seedlings were the best explants in both experiments and Murashige and Skoog (MS) medium supplemented with 0.25 or 0.5 mg L−1 BAP was suitable for induction of somatic embryos (SE) and adventitious shoots. Mass SE was obtained in experiment I after explants transfer on ½ MS (2% sucrose) + 0.45 mg L−1 B1 and extending culture to 2–3 months without passages. Rooting of adventitious shoots was a critical point. Out of seven rooting media used in experiment I, only two, ½ MS hormone free (2% sucrose) + 0.45 mg L−1 B1 or MS + 5 mg L−1 NAA + 3.76 mg L−1 B2 resulted in altogether 36.4% rooted shoots. In experiment II, somatic embryogenesis, rooting and acclimatization of adventitious shoots failed. Regenerated plantlets and seedlings converted from SE from experiment I were acclimatized to ex vitro conditions. Both genome size, determined by flow cytometry, and genetic diversity analyzed by ISSR markers, confirmed the compatibility of regenerants from experiment I with P. vulgaris initial seedlings and commercial cultivar. Regenerants obtained in experiment II differed genetically from the regenerants of experiment I and cultivar. Propagated in vitro tissues/organs (SE, adventitious shoots) of P. vulgaris could be a source of material for cryopreservation, artificial seed production and/or for acclimatization of regenerated plantlets and could be used for restoration of the extinct populations.
Key Message
The micropropagation technique via organogenesis and somatic embryogenesis of endangered in Europe pasqueflower was developed as a tool for species recovery. The critical point is that somatic embryogenesis is genotype-dependent, which affects the repeatability of the experiments and also imposes applying molecular techniques to confirm the genetic fidelity of the regenerants with the initial material and other genotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The preservation of endangered and/or threatened plant species by biotechnological methods such as in vitro culture is one of the ex situ conservation pathways. 42% of threatened European species are in ex situ collections (Sharrock and Jones 2011), which prevents loss of plant biodiversity and allows to recover genetic diversity from ex situ preserved material. Although the final step, translocation of the focal species or population from botanical gardens to the nature, is highly challenging task, 60% of cases of ex situ conservation were successful or partially successful (Abeli et al. 2020, 2021).
Nine occurring in Europe Pulsatilla species are rare, endangered and/or endemic, occupy small populations on local, light habitats acting as refugia (Akeroyd 1993). Overall species decline through Europe has been principally driven by the competition with grasses in the absence of appropriate management regimes, reduced grazing pressure, enhanced nitrogen deposition with acid rains and illegal transfer to gardens (Walker and Pinches 2011). In Poland main threat is a secondary succession from grassland through light, semi-open pine forest to shady, dense pine or pine-oak forest, maybe also flower picking and collecting herbarium specimens (Zarzycki et al. 2014).
Pulsatilla vulgaris Mill. of Ranunculaceae (2n = 2x = 16 or 2n = 4x = 32, Sramkó et al. 2019) was widespread throughout calcareous grasslands and open pine forests in western and central Europe, but now its populations are strongly declined (DiLeo et al. 2017). In Poland the species is extinct, its natural occurrence has not been confirmed in formerly known locations since 1930s (Zarzycki et al. 2014). Ex situ living collections from several Polish historical sites are cultivated in botanical gardens (Galera et al. 1999).
The effective technique to preserve genetic resources of Pulsatilla species is ex situ conservation using in vitro culture for plant multiplication via organogenesis and/or somatic embryogenesis (SE) (Šedivá 2002, 2012; Rybczyński and Mikuła 2006; Danova et al. 2009; Naumovski et al. 2009; Sauliene and Brinkyte 2009; Lin et al. 2011; Priede and Kļaviņa 2011; Šedivá and Žlebčík 2012). Development of synthetic seeds through encapsulation system (enclosing totipotent plant parts e.g., somatic embryos) provides a proficient method for continuous distribution and short/medium-term conservation of the germplasms (Gantait et al. 2015). In twenty-first century the germplasm of increasing number of endangered species is propagated via SE and protected as synthetic seeds or cryopreserved (e.g., Moon et al. 2013; Kang et al. 2014).
We aimed to: (1) develop an efficient micropropagation technique of P. vulgaris, extinct in Poland, using as an initial material indigenous plants from ex situ collection of botanical garden; (2) confirm genetic fidelity of regenerants with initial material by molecular markers; (3) estimate genome size of the source material and regenerants by flow cytometry. For practical purpose, the protocol could be used for mass micropropagation of P. vulgaris for restoration of Polish extinct populations.
Materials and methods
Plant material
The following material was used: (1) Pulsatilla vulgaris plants (Fig. 1a) from the collection of the Botanical Garden of the Polish Academy of Science – Center for Biological Diversity Conservation in Powsin (Powsin BG) as a source material; (2) seeds collected from these plants in two seasons (2019 for the in vitro experiment I and 2020 for the experiment II); (3) P. vulgaris cultivar commercially obtained from Łobzów Garden in Kraków (Poland) as a reference for plants from Powsin BG (P. vulgaris in Poland does not grow in nature); (4) P. pratensis growing close to Skołczanka nature reserve (Kraków, Poland) collected with the permission of the Regional Directorate for Environmental Protection. It was necessary to include this species in the study because in the vicinity of P. vulgaris in Powsin BG, specimens of P. pratensis grew and cross-pollination resulting in possible hybrid origin of the collected seeds could not have been excluded.
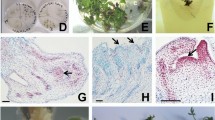
Organogenesis and somatic embryogenesis (SE), seedlings development and regenerated plants acclimatization of Pulsatilla vulgaris in experiment I. a Plant at blooming in the collection of Powsin Botanical Garden (Poland); b seeds on ½ MS medium with 2% sucrose for germination; c seedling used as explants for in vitro culture; d explants (shoot tips, cotyledons and hypocotyls with roots) on MS + 0.25 mg L−1 BAP inducing medium; e callus on shoot tips with developing SE (arrows), visible SE with cotyledons and embryonic root (asterisk) after callus transfer from MS + 0.25 L−1 BAP on hormone free ½ MS with 2% sucrose; f adventitious shoots developed on MS + 0.25 mg L−1 BAP inducing medium; g fragment of morphogenic callus with adventitious shoots (arrow) and somatic embryos (asterisk); h, i – mass of developing seedlings on ½ MS with 2% sucrose + 0.45 mg L−1 B1 inside the plastic container (h) and taken out of the container (i); j different stages of seedlings and juvenile plants converted from SE; k seedlings of converted SE in plastic containers filled with sterile perlite in ex situ conditions
Media and culture conditions
Media inducing somatic embryogenesis and organogenesis
Seeds harvested in 2019 and 2020 in Powsin BG were sterilized in 70% ethanol for 30 s and then in 50% commercial bleach Ace with sodium hypochlorite for 15 min, washed thrice with autoclaved distilled water. After 5 weeks of culture on ½ MS medium (Murashige and Skoog 1962) with 2% sucrose (w/v), solidified with 0.8% agar (w/v), medium pH 5.7–5.8, cotyledons, hypocotyls with roots and shoot tips were excised from the seedlings and used as explants. All explants were placed on inducing media solidified with 0.8% agar (w/v): MS + 0.25 or 0.5 mg L−1 BAP (according to the protocol of organogenesis in P. patens, Priede and Kļaviņa 2011) in Petri dishes. The first passage on the same fresh media was after 3 weeks from the culture establishment, next after 6 weeks.
In the experiment II (2020) for SE and organogenesis the same conditions and media were used based on the results of experiment I.
Rooting media
In the experiment I adventitious shoots or clusters of adventitious shoots (shoots’ separation from callus and their split into separate shoot was difficult and causing their damage) developing on morphogenic callus were transferred on 7 rooting media after next 4–6 weeks: (1) ½ MS (2% sucrose) + 0.45 mg L−1 B1; (2) MS + 5 mg L−1 NAA + 3.76 mg L−1 B2 (5 or 7 days in darkness according to Priede and Kļaviņa 2011); (3) ½ MS (2% sucrose) + 2 mg L−1 IBA + 0.45 mg L−1 B1; (4) MS (2% sucrose) + 2.5 mg L−1 NAA + 0.45 mg L−1 B1; (5) ½ MS + 2 mg L−1 IBA; (6) ½ MS + 0.5 mg L−1 IBA; 7) MS + 10 mg L−1 NAA. Half of shoots on media with NAA (nos. 4, 7) after 5 or 7 days was transferred on hormone free medium, the rest of explants were maintained on medium with NAA. All cultures were maintained in a growth chamber at 25 ± 3 °C under a 16 h photoperiod (cool-white fluorescent lamps, 60–90 μmol m−2 s−1).
In the experiment II for shoot rooting ½ MS (2% sucrose) + 0.45 mg L−1 B1 was used based on the results of experiment I.
Acclimatization to ex vitro conditions
Plants developed from somatic embryos in experiment I and rooted adventitious shoots obtained in the experiment I and II were transferred into sterile plastic containers filled with sterile perlite, perlite or sand mixed with garden soil (1:1) and maintained in a growth chamber for several days. Then the containers were opened in non-sterile conditions for ex vitro acclimatization.
Histological analysis for the confirmation of somatic embryogenesis and organogenesis
Paraffin method was used for histological analysis. Morphogenic/embryogenic callus produced on explants in the experiment I was fixed in the mixture of glacial acetic acid and 96% ethanol (1/3, v/v) dehydrated in an ethanol series, embedded in paraffin, and sectioned in 10–12 μm slices on a rotary microtome (Adamas Instrumenten BV, HM 340E). Sections were stained with Ehrlich’s hematoxylin (FLUKA, Switzerland) and alcian blue (FLUKA, Switzerland) and mounted in Entellan (Sigma-Aldrich, USA).
Genome size estimation
Leaves of seedlings of P. vulgaris obtained from seeds collected in 2019 in Powsin BG, adventitious shoots obtained in experiments I and II, leaves of plants of P. vulgaris cultivar, and of P. pratensis were used to establish genome size by flow cytometry (FCM), using buffer for nuclei isolation developed by Marie and Brown (1993); propidium iodide (50 μg/mL) was applied for DNA staining, and Secale cereale (2C = 16.19 pg; Doležel et al. 1992) served as an internal standard (for details of sample preparation and analysis see Żabicki et al. 2019). CV of the G0/G1 peak of Pulsatilla ranged between 2.51 and 4.58%. Nuclear DNA content was calculated using the linear relationship between the ratio of the 2C peak positions of Pulsatilla/internal standard on a histogram of fluorescence intensities.
Genetic diversity estimation by ISSR markers
DNA isolation
Leaves of three P. vulgaris plants from Powsin BG, six of cultivar, 12 seedlings of both experiments, 10 adventitious shoots obtained in experiment I and four in experiment II, three of P. pratensis collected for molecular analysis were dried in silica gel (silicon dioxide; F.H.U. “DOR-CHEM”, Poland). DNA was isolated using CTAB extraction method (Gavel and Jarret 1991). Quality of DNA was tested on 1% agarose gel.
ISSR markers
Six primers of ISSR markers were selected (Gupta et al. 1994; Stepansky et al. 1999) (Table 1). The analysis with ISSR markers was based on protocols developed by Żabicka et al. (2020); the same reagents and laboratory equipment were used. The PCR products were separated in 1% agarose gel with 1X TBE and SimplySafe (EURx Sp. z o.o. Gdansk, Poland) for about 90 min at 120 V. Band patterns of 38 samples were captured with a MultiDoc-It™ Imaging System with VisionWorks® LS Analysis Software (UVP, Upland, CA, USA). PCR reaction cycle were repeated on selected samples to test reproducibility of obtained results.
Split phylogenetic network (NeighborNet) was constructed in SplitsTree v. 4.6 (Huson and Bryant 2006) based on Dice coefficient to evaluate the relationships between examined individuals, bootstrap was calculated on 1000 replicates.
Statistics
To evaluate the significance of media composition influence on the culture results of the experiment II, ANOVA followed by Tukey HSD test post hoc (P = 0.05) was used. All calculations were performed in Statistica 13.3 software (TIBCO Software Inc., Palo Alto, California, USA).
Statistical analysis of genome size was performed using ANOVA followed by Tukey HSD test post hoc for different N (P = 0.05) in Statistica 13.3 software (TIBCO Software Inc., Palo Alto, California, USA).
Results
Somatic embryogenesis and organogenesis induction, regenerants acclimatization
In the experiment I, due to poor seed germination, only several seedlings were used as explants. Dissected shoot tips of seedlings were the best explants and MS + 0.25 mg L−1 BAP the most suitable inducing medium. Cotyledons, hypocotyls + roots poorly responded to in vitro conditions, becoming brownish and finally degenerated (Fig. 1b–d). Mass indirect (via callus) and direct somatic embryos (SE) development was obtained from shoot tips after callus or explants transfer from inducing medium (MS + 0.25 mg L−1 BAP) onto ½ MS (2% sucrose) + 0.45 mg L−1 B1 and extending the culture to 2–3 months without passage. In indirect pathway, callus induction and proliferation on shoot tips started after two weeks of culture (Fig. 1e). On one explant dozens, uncountable SE developed that converted into seedlings (Fig. 1e, g–j). Simultaneously with SE induction on the inducing medium, adventitious shoots were formed on the same explants (Fig. 1g). Both SE (Fig. 2a–c) and organogenesis (Fig. 2d–f) were confirmed on histological sections.
Somatic embryogenesis (SE) and adventitious shoot formation on histological sections of Pulsatilla vulgaris callus in experiment I. a Globular SE (arrowhead) and meristematic center (arrow), visible callus cells differing in shape and size; b Heart-stage SE; c Torpedo SE with embryonic root (arrow) and cotyledon (asterisk); d Multiple meristematic centers (arrows) in callus located along the cells differentiating into vascular tissue (arrowhead); e, f – Different stages of adventitious shoots development (arrows). Bar = 100 µm (a–c, e, f); 200 µm (d)
From the mass of seedlings converted from SE, 121 differing in size and organ development stage were separated, and 55 of the largest were moved to ex vitro conditions (Fig. 1k). The smallest seedlings were transferred to hormone-free MS supplemented with 2% sucrose to grow before being moved to ex vitro conditions, but their survivability was close to zero.
The rate of individual shoots and clusters of adventitious shoots rooting on ½ MS hormone free, 2% sucrose + 0.45 mg L−1 B1 (40 of 108) or MS + 5 mg L−1 NAA + 3.76 mg L−1 B2 (3 of 10) reached altogether 36.4% (43 of 118). Finally, 35 the most vital rooted shoots were transferred to non-sterile conditions for gradual acclimatization. On the remaining five media, none adventitious shoots had formed roots.
Based on the results of the experiment I, the experiment II was performed using more initial explants in several repetitions (Suppl. Table 1). Although for adventitious shoots and SE induction the same, most efficient media, as in the experiment I (MS + 0.25 or 0.5 mg L−1 BAP), the same type of explants (shoot tips, cotyledons, hypocotyls + roots) were applied under the same culture conditions as in experiment I, the results were different. Among explants used, shoot tips produced the highest frequency of adventitious shoots (97.2%), hypocotyls + roots and cotyledons responded with lower frequency (21.8% and 8.8%, respectively). There were no differences between media, the differences were evident between type of explants. The shoot tips responded with the highest frequency to the in vitro culture. Somatic embryos were not induced on any of the explant and medium (Suppl. Table 1). Out of 146 individual shoots and clusters of uncountable adventitious shoots transferred to rooting medium (½ MS hormone free, 2% sucrose + 0.45 mg L−1 B1), only five shoots (3.42%) formed roots. Their acclimatization to ex vitro conditions filed.
Genome size-based uniformity of regenerants and initial seedlings
Flow cytometry revealed a difference in genome size between P. vulgaris cultivar (2C = 25.94 pg) and P. pratensis (2C = 12.80 pg) that corresponds well with their ploidy level (2n = 4x = 32 and 2n = 2x = 16, respectively) (Table 2). Genome size of regenerants obtained in the experiment I (2C = 25.72 pg) did not differ statistically from 2C-value established for P. vulgaris seedlings obtained from seeds collected in Powsin BG and for the commercial cultivar but differed from the value established for regenerants obtained in the experiment II (2C = 25.14 pg). All adventitious shoots regenerated in the experiment II differed also in their genome size from the cultivar but not from initial seedlings from Powsin BG.
ISSR markers-based differences between the regenerants of experiments I and II
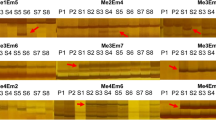
NeighborNet of 21 Pulsatilla vulgaris sensu lato (three plants from Powsin BG—P; 12 seedlings from seeds of experiments I and II—NP and 2NP, respectively; six plants of cultivar from Łobzów Garden—L), 14 regenerants (10 from the experiment I—REG; four from the experiment II—2Reg) and three P. pratensis (Pr) had shown significant divergence of P. pratensis specimens, with 94.2 branch support (Fig. 3). All regenerants from the experiment I (REG) clustered together with P. vulgaris specimens, including live collected plants (L, P) and seedlings (NP). Specimens regenerated during the experiment II (2Reg) formed a third significantly distinct group with 88.4 branch support (Fig. 3).
NeigborNet of Pulsatilla vulgaris sensu lato (Powsin BG—P; seedlings from seeds of I and II experiments—NP and 2NP, respectively; plants of cultivar from Łobzów Garden—L), regenerants (from the experiment I—REG and experiment II—2Reg) and P. pratensis (Pr) constructed on 93 ISSR loci using Dice distance. Branch support values are based on bootstrap analysis with 1000 replicates
Discussion
The main conclusion from the studies is that it was possible to achieve mass somatic embryogenesis in P. vulgaris but the developed protocol was not reproducible. Although the initial material for in vitro culture (seedlings) in experiment I and II originated from seeds produced by plants of the same clump of P. vulgaris growing in the Powsin BG, mass somatic embryogenesis was obtained exclusively in experiment I.
Is somatic embryogenesis in P. vulgaris genotype-dependent?
Somatic embryogenesis is a multi-step, molecular and biochemical process of embryo formation based on cell totipotency in which the vegetative cells acquire embryogenic competence under stress conditions (Pérez-Núñez et al. 2009; Thorpe 2012; Gulzar et al. 2020). This process is regulated by a set of genes of which only a few have been extensively studied, and chromatin remodeling by different mechanisms regulates these genes activity (reviewed by Gulzar et al. 2020).
As the stress induced by plant growth regulators, nutrient, oxygenic, desiccation and other signalling elements during in vitro culture lead to vegetative cells transformation into embryos via activation/deactivation of genes and transcriptional networks (Feher 2015), different species/genotype/explant reaction is easy to explain. Genotype-dependent somatic embryogenesis was described in several ornamental and economically important species (e.g., Chengalrayan et al. 1998; Mishra and Khurana 2003; Fiuk and Rybczyński 2008; Santos et al. 2018; Sánchez-Romero 2021). In olive several different protocols for somatic embryogenesis were developed with varied efficiency, depending on the genotype, limiting standardization and the applicability (Sánchez-Romero 2021).
The question arises whether different respond of P. vulgaris explants in experiment I and II could be the case of genotype-dependent? Could individuals representing genetic variation resulting from intraspecific crossing be considered as distinct genotypes? In living collection of Powsin BG P. vulgaris clumps (clones) grew close to P. pratensis, but the occurrence of other Pulsatilla species in garden collection is also possible. Hence the seeds could have originated from cross-pollination within P. vulgaris but their hybrid origin cannot be also excluded. Based on ISSR molecular markers, initial material (seedlings) used for in vitro culture in experiment I and II originating from seeds collected in two seasons, was genetically close to P. vulgaris living collection in Powsin BG and cultivar from Łobzów Garden and distinct from P. pratensis (Fig. 3). This eliminates hybrid origin of seeds and P. pratensis as one putative parent.
Pulsatilla vulgaris is predominately self-incompatible species, cross-pollination increases genetic diversity. In German populations high within-population variability (84.4%) and a weak, but significant, differentiation among populations were detected (Hensen et al. 2005). Genetic diversity of pasqueflower seems to be strongly affected by demographic changes i.e., decline, expansion and fragmentation (Walker and Pinches 2011). We could thus conclude that seedlings of P. vulgaris used in two experiments were different genotypes resulted from cross-pollination.
Genome size as an indication of somaclonal variation leading to genome multiplication/reduction under in vitro culture
Somaclonal variation is an important drawback of organogenesis/somatic embryogenesis, providing genetic variation in regenerated plantlets as a consequence of the change of chromosome number, chromosomal aberrations, genetic alterations, or epigenetic modification (Larkin and Scowcroft 1981). Regenerants converted from SE and adventitious shoots of P. vulgaris obtained in experiments I possessed the same genome size as initial seedlings and cultivar from Łobzów Garden. They possesed genome size of tetraploid cytotype of P. vulgaris (2C = 25.72 pg; 2n = 4x = 32), which was twice as large as in diploid P. pratensis (2C = 12.80 pg; 2n = 2x = 16). Genome size of adventitious shoots produced in experiment II (2C = 25.14 pg), although significantly differed from regenerated shoots of experiment I, possessed mean 2C DNA content similar to tetraploid P. vulgaris and not to diploid P. pratensis. Based on this characteristic they could be considered as regenerants of P. vulgaris and thus could be reintroduced to the nature.
ISSR molecular markers separated regenerants of experiment II from P. vulgaris
Combined results of genome size value and ISSR markers clearly indicated that regeneration via organogenesis and SE successfully achieved in experiment I is suitable procedure for obtaining plants for reintroduction. Surprisingly, regenerated shoots of experiment II although representing genome size similar to P. vulgaris, genetically formed a separate, strongly (88.4) supported branch (Fig. 3). Their origin is difficult to reconstruct—genetic variation is too high to explain it as genome alteration influenced by culture conditions (somaclonal variation). The genetic distance from P. vulgaris and P. pratensis is very long what eliminates these two species as putative parents of the seeds. It could be hypothesized that the initial seedlings for the experiment II (not used for ISSR analysis) originated from hybrid seeds of P. vulgaris with other tetraploid Pulsatilla species growing in Powsin BG. In Pulsatilla hybridization and introgression played a crucial role in the evolutionary history of the genus, and interspecific hybrids or species of hybrid origin occur in nature. Hybrids are fertile due to weak genetic isolation mechanisms (Szczecińska et al. 2017; Torzewski 2018; Li et al. 2019).
Ex situ conservation of P. vulgaris using seed collections is highly disputable
In natural populations and in ex situ collections seed set may be pollen-limited (Jonsson et al. 1991). P. vulgaris as an early flowering plant may face disadvantages of low pollinator abundances and unfavorable weather conditions influencing seed set. Pollinator exclusion, self-pollination and wind pollination only resulted in a very low percentage of seed set (Kehrberger and Holzschuh 2019). The limitation is also low seed germination frequency of P. vulgaris from Powsin BG on sterile media (MS, filter paper, lignin) which did not exceed 20% and was slightly higher (~ 30%) in non-sterile garden soil (present studies; data not shown). Field experiments have indicated that increasing the sizes of local populations of P. vulgaris by transplanting ex situ grown seedlings or just by sowing seeds was hardly ever successful in Switzerland (Pfeifer et al. 2002) but successful in three English populations (Gargiulo et al. 2019).
Rare and endangered with extinction species are effectively propagated via SE and could be protected as artificial seeds or cryopreserved (e.g., Rybczyński and Mikuła 2006; Moon et al. 2013; Kang et al. 2014; Streczynski et al. 2019). Among Pulsatilla species, only one species, specifically Pulsatilla koreana has been propagated via SE so far (Lin et al. 2011). In the recent paper we have revealed that SE is also possible for P. vulgaris however usage of seeds from living collections should be done with cautious.
Conclusion
-
(1)
Mass somatic embryogenesis, rooting of adventitious shoots and regenerants acclimatization of P. vulgaris was strongly genotype-dependent; the developed protocol was not reproducible, although the same culture conditions, explants, plant growth regulators, media were used. The critical points demonstrated here will be helpful in further studies on the micropropagation of P. vulgaris aiming to save germplasm of this endangered in Europe species.
-
(2)
For restoration of extinct P. vulgaris populations it is important to confirm genetic compatibility of regenerated plantlets with the initial material by molecular markers to avoid introducing new variability to natural populations.
-
(3)
Genome size alone cannot be used for identification of genome qualitative alteration; regenerants with similar genome size might be genetically highly different as was shown in P. vulgaris regenerants in experiment II.
-
(4)
The study highlights the necessity to be cautious when using material from botanical gardens. In the case of P. vulgaris it was not possible to obtain different material from natural, local genetic resources as this species is extinct in Poland.
References
Abeli T, Dalrymple S, Godefroid S, Mondoni A, Müller JV, Rossi G, Orsenigo S (2020) Ex situ collections and their potential for the restoration of extinct plants. Conserv Biol 34:303–313. https://doi.org/10.1111/cobi.13391
Abeli T, Albani Rocchetti G, Barina Z et al (2021) Seventeen ‘extinct’ plant species back to conservation attention in Europe. Nat Plants 7:282–286. https://doi.org/10.1038/s41477-021-00878-1
Akeroyd JR (1993) Pulsatilla Miller. Flora Eur 1:264–266
Bolkovskikh ZV, Grif VG, Zakhareva OI, Matveeva TC, (eds) (1969) Chromosome numbers of flowering plants. Brooker RW, Carlsson BÅ & Call, Nauka
Chengalrayan K, Mhaske V, Hazra S (1998) Genotypic control of peanut somatic embryogenesis. Plant Cell Rep 17:522–525. https://doi.org/10.1007/s002990050435
Danova K, Bertoli A, Pistelli L, Dimitrov D, Pistelli L (2009) In vitro culture of Balkan endemic and rare Pulsatilla species for conservational purposes and secondary metabolites production. Bot Serbica 33:157–162
DiLeo M, Rico Y, Boehmer H, Wagner H (2017) An ecological connectivity network maintains genetic diversity of a flagship wildflower, Pulsatilla vulgaris. Biol Conserv 212:12–21. https://doi.org/10.1016/j.biocon.2017.05.026
Doležel J, Sgorbati S, Lucretti S (1992) Comparison of three DNA fluorochromes for flow cytometric estimation of nuclear DNA content in plants. Physiol Plant 85:625–631. https://doi.org/10.1111/j.1399-3054.1992.tb04764.x
Feher A (2015) Somatic embryogenesis - stress-induced remodeling of plant cell fate. Biochim Biophys Acta 1849:385–402. https://doi.org/10.1016/j.bbagrm.2014.07.005
Fiuk A, Rybczyński JJ (2008) Genotype and plant growth regulator-dependent response of somatic embryogenesis from Gentiana spp. leaf explants. In Vitro Cell Dev Biol Plant 44(2):90–99. https://doi.org/10.1007/s11627-008-9124-3
Galera H, Puchalski J, Gawryś W (1999) Polskie kolekcje roślin chronionych i zagrożonych oraz endemitów i reliktów. Część I. Gatunki objęte ochroną prawną. Biuletyn Ogrodów Botanicznych, Muzeów i Zbiorów 8:41–83
Gantait S, Kundu S, Ali N, Sahu NC (2015) Synthetic seed production of medicinal plants: a review on influence of explants, encapsulation agent and matrix. Acta Physiol Plant 37:98. https://doi.org/10.1007/s11738-015-1847-2
Gargiulo R, Worswick G, Arnold C, Pike LJ, Cowan RS, Hardwick KA, Chapman T, Fay MF (2019) Conservation of the threatened species, Pulsatilla vulgaris Mill. (pasqueflower), is aided by reproductive system and polyploidy. J Hered 110:618–628. https://doi.org/10.1093/jhered/esz035
Gavel NJ, Jarret RL (1991) A modified CTAB DNA extraction procedure for Musa and Ipomoea. Plant Mol Biol Rep 9:262–266. https://doi.org/10.1007/BF02672076
Gulzar B, Mujib A, Malik MQ et al (2020) Genes, proteins and other networks regulating somatic embryogenesis in plants. J Genet Eng Biotechnol 18:31. https://doi.org/10.1186/s43141-020-00047-5
Gupta M, Chyi YS, Romero-Severson J, Owen JL (1994) Amplification of DNA markers from evolutionary diverse genomes using single primers of simple-sequence repeats. Theor Appl Genet 89:998–1006. https://doi.org/10.1007/BF00224530
Hensen I, Oberprieler C, Wesche K (2005) Genetic structure, population size, and seed production of Pulsatilla vulgaris Mill. (Ranunculaceae) in Central Germany. Flora 200:3–14. https://doi.org/10.1016/j.flora.2004.05.001
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Jonsson O, Rosquist G, Widén B (1991) Operation of dichogamy and herkogamy in five taxa of Pulsatilla. Ecography 14:260–271. https://doi.org/10.1111/j.1600-0587.1991.tb00660.x
Kang HD, Moon HK, Lee SK (2014) Micropropagation via somatic embryogenesis of rare and endangered species Acanthopanax seoulenses Nakai. For Sci Technol 10:190–196. https://doi.org/10.1080/21580103.2014.913536
Kehrberger S, Holzschuh A (2019) How does timing of flowering affect competition for pollinators, flower visitation and seed set in an early spring grassland plant? Sci Rep 9:15593. https://doi.org/10.1038/s41598-019-51916-0
Larkin PJ, Scowcroft WR (1981) Somaclonal variation—a novel source of variability from cell cultures for plant improvement. Theor Appl Genet 60:197–214. https://doi.org/10.1007/BF02342540
Li Q-j, Wang X, Wang J-r, Su N, Zhang L, Ma Y-p, Chang Z-y, Zhao L, Potter D (2019) Efficient identification of Pulsatilla (Ranunculaceae) using DNA barcodes and micro-morphological characters. Front Plant Sci 10:1196. https://doi.org/10.3389/fpls.2019.01196
Lin GZ, Zhao XM, Hong SK, Lian YI (2011) Somatic embryogenesis and shoot organogenesis in the medicinal plant Pulsatilla koreana Nakai. Plant Cell Tiss Organ Cult 106:93–103. https://doi.org/10.1007/s11240-010-9897-z
Marhold K, Mereda P, Mráz P, Hodálowá I, Kolník M, Kučera J, Lihowá J, Mártonefi P, Mrázowá V, Perný M (eds) (2005) Karyological database of ferns and flowering plants of Slovakia. http://147.213.100.121/webapp/
Marie D, Brown SC (1993) A cytometric exercise in plant DNA histograms, with 2C values for 70 species. Biol Cell 78:41–51. https://doi.org/10.1016/0248-4900(93)90113-S
Mishra A, Khurana P (2003) Genotype dependent somatic embryogenesis and regeneration from leaf base cultures of Sorghum bicolor. J Plant Biochem Biotechnol 12:53–56. https://doi.org/10.1007/BF03263160
Moon HK, Kim YW, Hong YP, Park SY (2013) Improvement of somatic embryogenesis and plantlet conversion in Oplopanax elatus, an endangered medicinal woody plant. Springerplus 2:428. https://doi.org/10.1186/2193-1801-2-428
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Naumovski D, Radič S, Pevalek-Kozlina B (2009) In vitro micropropagation of Pulsatilla pratensis (L.) Miller ssp. nigricans (Störck) Zämelis. Propag Ornam Plants 9:16–20
Pérez-Núñez MT, Souza R, Sáenz L, Chan JL, Zúñiga-Aguilar JJ, Oropeza C (2009) Detection of a SERK-like gene in coconut and analysis of its expression during the formation of embryogenic callus and somatic embryos. Plant Cell Rep 28:11–19. https://doi.org/10.1007/s00299-008-0616-8
Pfeifer E, Holdereggeret R, Matthies D, Rutishauser R (2002) Investigation on the population biology of a flagship species of dry meadows: Pulsatilla vulgaris Mill. in north-eastern Switzerland. Bot Helv 112:153–171
Priede G, Kļaviņa D (2011) In vitro cultivation and root initiation of the endangered plant Pulsatilla patens. Environ Exp Biol 9:71–74
Rybczyński JJ, Mikuła A (2006) Engagement of biotechnology in the protection of threatened plant species in Poland. Biodiv Res Conserv 3–4:361–368
Sánchez-Romero C (2021) Somatic embryogenesis in olive. Plants 10:433. https://doi.org/10.3390/plants10030433
Santos IR, Maximiano MR, Almeida RF, da Cunha RNV, Lopes R, Scherwinski-Pereira JE et al (2018) Genotype-dependent changes of gene expression during somatic embryogenesis in oil palm hybrids (Elaeis oleifera x E. guineensis). PLoS ONE 13(12):e0209445. https://doi.org/10.1371/journal.pone.0209445
Sauliene I, Brinkyte E (2009) The impact of phytohormones on pasgueflower (Pulsatilla) regeneration in vitro. Acta Biol Univ Daugavp 9:249–254
Šedivá J (2002) In vitro seed propagation of some Pulsatilla species (Pulsatilla L.). Acta Pruhoniciana 73:48–51
Šedivá J (2012) In vitro root formation in Pulsatilla vernalis (L.) Mill. Propag Ornam Plants 12:96–101
Šedivá J, Žlebčík J (2012) Summary of findings from a propagation and ex situ conservation of Pulsatilla vernalis, P. pratensis ssp. bohemica, P. patens and P. grandis. Acta Pruhoniciana 100:155–160
Sharrock S, Jones M (2011) Saving Europe’s threatened flora: progress towards GSPC Target 8 in Europe. Biodivers Conserv 20:325–333. https://doi.org/10.1007/s10531-010-9912-z
Sramkó G, Laczkób L, Volkovac PA, Bateman RM, Mlinarec J (2019) Evolutionary history of the Pasque-flowers (Pulsatilla, Ranunculaceae): molecular phylogenetics, systematics and rDNA evolution. Mol Phylogenet Evol 135:45–61. https://doi.org/10.1016/j.ympev.2019.02.015
Stepansky A, Kovalski I, Perl-Treves R (1999) Intraspecific classification of melons (Cucumis melo L.) in view of their phenotypic and molecular variation. Plant Syst Evol 217:313–332
Streczynski R, Clark H, Whelehan LM, Ang ST, Hardstaff LK, Funnekotter B, Bunn OCA, Sommerville KD, Mancera L (2019) Current issues in plant cryopreservation and importance for ex situ conservation of threatened Australian native species. Aust J Bot 67(1):1–15. https://doi.org/10.1071/BT18147
Szczecińska M, Łazarski G, Bilska K (2017) The complete plastid genome and nuclear genome markers provide molecular evidence for the hybrid origin of Pulsatilla × hackelii Pohl. Turk J Bot 41:329–337. https://doi.org/10.3906/bot-1610-2
Thorpe T (2012) History of plant tissue culture. Methods Mol Biol 877:9–27. https://doi.org/10.1385/1-59259-959-1:009
Torzewski K (2018) Occurrence of Pulsatilla hybrids in the Polish flora. Steciana 22:123–127. https://doi.org/10.12657/steciana.022.014
Walker KJ, Pinches CE (2011) Reduced grazing and the decline of Pulsatilla vulgaris Mill. (Ranunculaceae) in England, UK. Biol Conserv 144:3098–3105. https://doi.org/10.1016/j.biocon.2011.10.006
Żabicka J, Migdałek G, Słomka A, Sliwinska E, Mackiewicz L, Keczyński A, Kuta E (2020) Interspecific hybridization and introgression influence biodiversity—based on genetic diversity of Central European Viola epipsila-V. palustris complex. Diversity 12:1–23. https://doi.org/10.3390/d12090321
Żabicki P, Sliwinska E, Mitka J, Sutkowska A, Tuleja M, Migdałek G, Żabicka J, Słomka A, Kwiatkowska M, Kuta E (2019) Does somaclonal variation play advantageous role in conservation practice of endangered species?: comprehensive genetic studies of in vitro propagated plantlets of Viola stagnina Kit. (Violaceae). Plant Cell Tiss Organ Cult 136:339–352. https://doi.org/10.1007/s11240-018-1519-1
Zarzycki K, Kaźmierczakowa R, Mirek Z (eds) (2014) Polska Czerwona Księga Roślin. Paprotniki i rośliny kwiatowe. 3rd ed. Instytut Ochrony Przyrody PAN: Kraków, Poland, 895pp.
Acknowledgements
The Authors thank Mrs. Teresa Stokłosa from Jagiellonian University in Krakow for technical help in histological analyses and MSc Stefan Gawroński for assistance in P. pratensis collection. The photo of Pulsatilla vulgaris comes from Adam Kapler private archive.
Funding
The studies were performed in the frame of BioGalmany project no. POIS.02.04.00-00-0043/17-00.
Author information
Authors and Affiliations
Contributions
JŻ and PŻ: planned and carried out the in vitro experiments, data analysis and interpretation of the data. AS: conception and design, performance of experiments, critical reading of the manuscript. MJ-K and TN: material collection, result discussion. ES: genome size analysis and interpretation of the data. AK: seeds and leaves collection from Powsin Botanical Garden, discussion on Pulsatilla vulgaris status. GM: analysis and interpretation of molecular data. EK: Supervision and interpretation of the results, drafting the article. All authors read the manuscript, critically revised and approved final version of the paper.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Additional information
Communicated by Amita Bhattacharya.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Żabicka, J., Żabicki, P., Słomka, A. et al. Genotype-dependent mass somatic embryogenesis: a chance to recover extinct populations of Pulsatilla vulgaris Mill.. Plant Cell Tiss Organ Cult 146, 345–355 (2021). https://doi.org/10.1007/s11240-021-02074-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-021-02074-7