Abstract
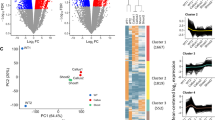
Plant hormones not only play important roles in regulating plant growth and development, but they also promote cell dedifferentiation and redifferentiation, which play an important role in tissue culture. In the present study, an efficient method of promoting callus plant regeneration for the tissue culture of Brassica juncea (L.) was proposed. We discovered that the callus could be efficiently induced into redifferentiation by 6.8 × 10− 3 µmol/l thidiazuron (TDZ), a cytokinin analogue. To analyze the molecular mechanism of tissue culture, we sequenced the transcriptome of the callus under different culture conditions at two developmental stages. A total of 5362 differentially expressed genes were identified in the TDZ treatments; among them, the expression of 4676 genes was upregulated and that of 686 was downregulated. In the cytokinin signal transduction pathway, seven genes in the Arabidopsis (Arabidopsis thaliana) response regulator (ARR) family were upregulated. In addition, cytokinin affected gene expression in other plant hormone signaling pathways, such as the auxin, ethylene, and brassinosteroid pathways. Most of these genes, such as auxin/indole-3-acetic acid (Aux/IAA) genes and ethylene response factors (ERF-1, ERF-2, and ERF-6), generally showed upregulated expression, which also participated in the regulation of callus plant regeneration. This study provides novel insight into the molecular mechanism of callus plant regeneration in Brassica juncea, which could also provide reference for other plant tissue cultures.
Key message
In Brassica juncea, 6.8 × 10− 3 µmol/l TDZ resulted in the highest regeneration rate of callus, and the genes involved in cytokinin and auxin signal transduction pathways participated in the redifferentiation progress.






Similar content being viewed by others
Data availability
The raw data used in this study had been submitted to the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra/) under accession PRJNA562990.
Abbreviations
- 2,4-D:
-
2,4-Dichlorophenoxyacetic acid
- NAA:
-
Naphthalene acetic acid
- TDZ:
-
Thidiazuron
- MS:
-
Murashige and Skoog
- CGM:
-
Concentration gradient medium
- H0D15:
-
Callus grown on the CGM containing no hormone for 15 days
- HaD15:
-
Callus grown on the CGM containing only 5.4 × 10−4 µmol/l NAA for 15 days
- HbD15:
-
Callus grown on the CGM containing only 6.8 × 10−3 µmol/l TDZ for 15 days
- H0D21:
-
Callus grown on the CGM containing no hormone for 21 days
- HaD21:
-
Callus grown on the CGM containing only 5.4 × 10−4 µmol/l NAA for 21 days
- HbD21:
-
Callus grown on the CGM containing only 6.8 × 10−3 µmol/l TDZ for 21 days
- T0N0:
-
Callus grown on the CGM containing no hormone
- T0N0.1:
-
Callus grown on the CGM containing only 5.4 × 10−4 µmol/l NAA
- T1.5N0:
-
Callus grown on the CGM containing only 6.8 × 10−3 µmol/l TDZ
References
Barrero JM, Millar AA, Griffiths J, Czechowski T, Scheible WR, Udvardi M, Reid JB, Ross JJ, Jacobsen JV, Gubler F (2010) Gene expression profiling identifies two regulatory genes controlling dormancy and ABA sensitivity in Arabidopsis seeds. Plant J 61:611–622. https://doi.org/10.1111/j.1365-313X.2009.04088.x
Benjamins R, Scheres B (2008) Auxin: the looping star in plant development. Annu Rev Plant Biol 59:443–465. https://doi.org/10.1146/annurev.arplant.58.032806.103805
Bennett MJ, Marchant A, Green HG, May ST, Ward SP, Millner PA, Walker AR, Schulz B, Feldmann KA (1996) Arabidopsis AUX1 gene: a permease-like regulator of root gravitropism. Science 273:948–950. https://doi.org/10.1126/science.273.5277.948
Bohn-Courseau I (2010) Auxin: a major regulator of organogenesis. C R Biol 333:290–296. https://doi.org/10.1016/j.crvi.2010.01.004
Buechel S, Leibfried A, To JP, Zhao Z, Andersen SU, Kieber JJ, Lohmann JU (2010) Role of A-type ARABIDOPSIS RESPONSE REGULATORS in meristem maintenance and regeneration. Eur J Cell Biol 89:279–284. https://doi.org/10.1016/j.ejcb.2009.11.016
Chen C, Xia R, Chen H, He YT, Xia R (2018) a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. bioRxiv.https://doi.org/10.1101/289660
Choi G, Yi H, Lee J, Kwon YK, Soh MS, Shin B, Luka Z, Hahn TR, Song PS (1999) Phytochrome signalling is mediated through nucleoside diphosphate kinase 2. Nature 401:610. https://doi.org/10.1038/44176
Chone RMS, Rocha DI, Monte-Bello CC, Rocha DI, Monte-Bello CC, Pinheiro HP, Dornelas MC, Haddad CRB, Almeida JAS (2018) Brassinosteroid increases the cytokinin efficiency to induce direct somatic embryogenesis in leaf explants of Coffea arabica L. (Rubiaceae). Plant Cell Tissue Organ Cult 135:63–71. https://doi.org/10.1007/s11240-018-1443-4
Cifuentes-Esquivel N, Bou-Torrent J, Galstyan A, Gallemí M, Sessa G, Martret MS, Roig-Villanova I, Ruberti I, Martínez-García JF (2013) The bHLH proteins BEE and BIM positively modulate the shade avoidance syndrome in Arabidopsis seedlings. Plant J 75:989–1002. https://doi.org/10.1111/tpj.12264
Davies PJ (2010) The plant hormones: their nature, occurrence, and functions. In: Plant Horm. Springer, pp 1–15. https://doi.org/10.1007/978-1-4020-2686-7_1
Dewitte W, Riou-Khamlichi C, Scofield S, Healy JS, Jacqmard A, Kilby NJ, Murray JA (2003) Altered cell cycle distribution, hyperplasia, and inhibited differentiation in Arabidopsis caused by the D-type cyclin CYCD3. Plant Cell 15:79–92. https://doi.org/10.1105/tpc.004838
Gordon SP, Heisler MG, Reddy GV, Ohno C, Das P, Meyerowitz EM (2007) Pattern formation during de novo assembly of the Arabidopsis shoot meristem. Development 134:3539–3548. http://dev.biologists.org/content/134/19/3539
Grefen C, Harter K (2004) Plant two-component systems: principles, functions, complexity and cross talk. Planta 219:733–742. https://doi.org/10.1007/s00425-004-1316-4
Hu Y, Bao F, Li J (2000) Promotive effect of brassinosteroids on cell division involves a distinct CycD3-induction pathway in Arabidopsis. Plant J 24:693–701. https://doi.org/10.1046/j.1365-313x.2000.00915.x
Huang SJ, Chang CL, Wang PH, Tsai MC, Hsu PH, Chang IF (2013) A type III ACC synthase, ACS7, is involved in root gravitropism in Arabidopsis thaliana. J Exp Bot 64:4343–4360. https://doi.org/10.1093/jxb/ert241
Hwang I, Sheen J (2001) Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413:383. https://doi.org/10.1038/35096500
Hwang I, Chen HC, Sheen J (2002) Two-component signal transduction pathways in Arabidopsis. Plant Physiol 129:500–515. https://doi.org/10.1104/pp.005504
Hwang I, Sheen J, Müller B (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63:353–380. https://doi.org/10.1146/annurev-arplant-042811-105503
Ikeuchi M, Ogawa Y, Iwase A, Sugimoto K (2016) Plant regeneration: cellular origins and molecular mechanisms. Development 143(9):1442–1451. https://doi.org/10.1242/dev.134668
Iwama A, Yamashino T, Tanaka Y, Sakakibara H, Kakimoto T, Sato S, Kato T, Tabata S, Nagatani A, Mizuno T (2007) AHK5 histidine kinase regulates root elongation through an ETR1-dependent abscisic acid and ethylene signaling pathway in Arabidopsis thaliana. Plant Cell Physiol 48:375–380. https://doi.org/10.1093/pcp/pcl065
Ji XH, Zhang R, Wang N, Yang L, Chen XS (2015) Transcriptome profiling reveals auxin suppressed anthocyanin biosynthesis in red-fleshed apple callus (Malus sieversii f. niedzwetzkyana). Plant Cell Tiss Organ Cult 123:389–404. https://doi.org/10.1007/s11240-015-0843-y
Kende H (1993) Ethylene biosynthesis. Annu Rev Plant Biol 44:283–307. https://doi.org/10.1146/annurev.pp.44.060193.001435
Khehra G, Mathias R (1992) The interaction of genotype, explant and media on the regeneration of shoots from complex explants of Brassica napus L. J Exp Bot 43:1413–1418. https://doi.org/10.1093/jxb/43.11.1413
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357. https://doi.org/10.1038/nmeth.3317
Kim J, Kang J, Park MY, Song M, Kim YC, Kim SY (2019) Arabidopsis zinc finger homeodomain protein ZHD5 promotes shoot regeneration and confers other cytokinin-related phenotypes when overexpressed. Plant Cell Tiss Organ Cult 137:181–185. https://doi.org/10.1007/s11240-018-01546-7
Laskowski M, Biller S, Stanley K, Kajstura T, Prusty R (2006) Expression profiling of auxin-treated Arabidopsis roots: toward a molecular analysis of lateral root emergence. Plant Cell Physiol 47:788–792. https://doi.org/10.1093/pcp/pcj043
Leyser HO, Pickett FB, Dharmasiri S, Estelle M (1996) Mutations in the AXR3 gene of Arabidopsis result in altered auxin response including ectopic expression from the SAUR-AC1 promoter. Plant J 10:403–413. https://doi.org/10.1046/j.1365-313x.1996.10030403.x
Li PT, Chen TT, Lu QW, Ge Q, Gong WK, Liu AY, Gong JW, Shang HH, Deng XY, Li JW, Li SQ, Xiao XH, Liu RX, Zhang Q, Duan L, Zou XY, Zhang Z, Jiang X, Zhang Y, Peng RH, Shi YZ, Yuan YL (2019) Transcriptomic and biochemical analysis of upland cotton (Gossypium hirsutum) and a chromosome segment substitution line from G. hirsutum × G. barbadense in response to Verticillium dahliae infection. BMC Plant Biol 19:19–43. https://doi.org/10.1186/s12870-018-1619-4
Liao Y, Smyth GK, Shi W (2013) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930. https://doi.org/10.1093/bioinformatics/btt656
Lin YY, Wang Y, Iqbal A, Shi P, Li J, Yang YD, Lei XT (2017) Optimization of culture medium and temperature for the in vitro germination of oil palm pollen. Sci Hortic 220:134–138. https://doi.org/10.1016/j.scienta.2017.03.040
Lincoln JE, Campbell A, Oetiker J, Rottmann W, Oeller P, Shen N, Theologis A (1993) LE-ACS4, a fruit ripening and wound-induced 1-aminocyclopropane-1-carboxylate synthase gene of tomato (Lycopersicon esculentum). Expression in Escherichia coli, structural characterization, expression characteristics, and phylogenetic analysis. J Biol Chem 268:19422–19430
Liu H, Zhang H, Dong YX, Hao YJ, Zhang XS (2018) DNA METHYLTRANSFERASE1-mediated shoot regeneration is regulated by cytokinin‐induced cell cycle in Arabidopsis. New Phytol 217:219–232. https://doi.org/10.1111/nph.14814
Liu L, Liu F, Chu J, Yi X, Fan W, Tang T, Chen G, Zhao QHXX (2019) A transcriptome analysis reveals a role for the indole GLS-linked auxin biosynthesis in secondary dormancy in rapeseed (Brassica napus L.). BMC Plant Biol 19(1):264–282. https://doi.org/10.1186/s12870-019-1866-z
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2–∆∆CT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Mantiri FR, Kurdyukov S, Lohar DP, Sharopova N, Saeed NA, Wang XD, VandenBosch KA, Rose RJ (2008) The transcription factor MtSERF1 of the ERF subfamily identified by transcriptional profiling is required for somatic embryogenesis induced by auxin plus cytokinin in Medicago truncatula. Plant Physiol 146:1622–1636. https://doi.org/10.1104/pp.107.110379
Mason MG, Mathews DE, Argyros DA, Maxwell BB, Kieber JJ, Alonso JM, Schaller GE (2005) Multiple type-B response regulators mediate cytokinin signal transduction in Arabidopsis. Plant Cell 17:3007–3018. https://doi.org/10.1105/tpc.105.035451
Matsui K, Tanaka H, Ohme-Takagi M (2004) Suppression of the biosynthesis of proanthocyanidin in Arabidopsis by a chimeric PAP1 repressor. Plant Biotechnol J 2:487–493. https://doi.org/10.1111/j.1467-7652.2004.00094.x
Merchante C, Alonso JM, Stepanova AN (2013) Ethylene signaling: simple ligand, complex regulation. Curr Opin Plant Biol 16:554–560. https://doi.org/10.1016/j.pbi.2013.08.001
Mira-Rodado V, Veerabagu M, Witthöft J, Teply J, Harter K, Desikan R (2012) Identification of two-component system elements downstream of AHK5 in the stomatal closure response of Arabidopsis thaliana. Plant Signaling Behav 7:1467–1476. https://doi.org/10.4161/psb.21898
Mitchell J, Mandava N, Worley J, Plimmer J, Smith M (1970) Brassins—a new family of plant hormones from rape pollen. Nature 225:1065. https://doi.org/10.1038/2251065a0
Moreira S, Bishopp A, Carvalho H, Campilho A (2013) AHP6 inhibits cytokinin signaling to regulate the orientation of pericycle cell division during lateral root initiation. PLoS ONE 8:e56370. https://doi.org/10.1371/journal.pone.0056370
Nakazawa M, Yabe N, Ichikawa T, Yamamoto YY, Yoshizumi T, Hasunuma K, Matsui M (2001) DFL1, an auxin-responsive GH3 gene homologue, negatively regulates shoot cell elongation and lateral root formation, and positively regulates the light response of hypocotyl length. Plant J 25:213–221. https://doi.org/10.1111/j.1365-313X.2001.00957.x
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. The Plant Cell 7:173–182. https://doi.org/10.1105/tpc.7.2.173
Pernisová M, Klíma P, Horák J, Válková M, Malbeck J, Souček P, Reichman P, Hoyerová K, Dubová J, Friml J, žímalová EZ, Hejátko J (2009) Cytokinins modulate auxin-induced organogenesis in plants via regulation of the auxin efflux. Proc Natl Acad Sci 106:3609–3614. https://doi.org/10.1073/pnas.0811539106
Pimentel H, Bray NL, Puente S, Melsted P, Pachter L (2017) Differential analysis of RNA-seq incorporating quantification uncertainty. Nat Methods 14(7):687–690. https://doi.org/10.1038/nmeth.4324
Radoeva T, Weijers D (2014) A roadmap to embryo identity in plants. Trends Plant Sci 19(11):709–716. https://doi.org/10.1016/j.tplants.2014.06.009
Rogg LE, Lasswell J, Bartel B (2001) A gain-of-function mutation in IAA28 suppresses lateral root development. Plant Cell 13:465–480. https://doi.org/10.1105/tpc.13.3.465
Rupps A, Raschke J, Rümmler M, Linke B, Zoglauer K (2016) Identification of putative homologs of Larix decidua to Babyboom (BBM), leafy cotyledon1 (LEC1), Wuschel-related Homeobox2 (WOX2) and somatic embryogenesis receptor-like kinase (SERK) during somatic embryogenesis. Planta 243(2):473–488. https://doi.org/10.1007/s00425-015-2409-y
Sakai H, Aoyama T, Oka A (2000) Arabidopsis ARR1 and ARR2 response regulators operate as transcriptional activators. Plant J 24:703–711. https://doi.org/10.1111/j.1365-313X.2000.00909.x
Schaller GE, Kieber JJ, Shiu SH (2008) Two-component signaling elements and histidyl-aspartyl phosphorelays. The Arabidopsis Book/American Society of Plant Biologists 6. https://doi.org/10.1199/tab.0112
Solano R, Stepanova A, Chao Q, Ecker JR (1998) Nuclear events in ethylene signaling: a transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev 12:3703–3714. https://doi.org/10.1101/gad.12.23.3703
Su YH, Liu YB, Zhang XS (2011) Auxin–cytokinin interaction regulates meristem development. Mol Plant 4:616–625. https://doi.org/10.1093/mp/ssr007
Sugiyama M (2015) Historical review of research on plant cell dedifferentiation. J Plant Res 128(3):349–359. https://doi.org/10.1007/s10265-015-0706-y
Sweere U, Eichenberg K, Lohrmann J, Mira-Rodado V, Bäurle I, Kudla J, Nagy F, Schäfer E, Harter K (2001) Interaction of the response regulator ARR4 with phytochrome B in modulating red light signaling. Science 294:1108–1111. https://doi.org/10.1126/science.1065022
Toonen MA, Hendriks T, Schmidt ED, Verhoeven HA, van Kammen A, de Vries SC (1994) Description of somatic-embryo-forming single cells in carrot suspension cultures employing video cell tracking. Planta 194:565–572. https://doi.org/10.1007/BF00714471
Uehara T, Okushima Y, Mimura T, Tasaka M, Fukaki H (2008) Domain II mutations in CRANE/IAA18 suppress lateral root formation and affect shoot development in Arabidopsis thaliana. Plant Cell Physiol 49:1025–1038. https://doi.org/10.1093/pcp/pcn079
Urao T, Yamaguchi-Shinozaki K, Shinozaki K (2000) Two-component systems in plant signal transduction. Trends Plant Sci 5:67–74. https://doi.org/10.1016/S1360-1385(99)01542-3
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10:57. https://doi.org/10.1038/nrg2484
Wang W, Bai MY, Wang ZY (2014) The brassinosteroid signaling network—a paradigm of signal integration. Curr Opin Plant Biol 21:147–153. https://doi.org/10.1016/j.pbi.2014.07.012
Xu P, Cao SQ, Hu KN, Wang XH, Huang W, Wang G, Lv ZW, Liu ZS, Wen J, Yi B, Ma CZ, Tu JX, Fu TD, Shen JX (2017) Trilocular phenotype in Brassica juncea L. resulted from interruption of CLAVATA1 gene homologue (BjMc1) transcription. Sci Rep 7:3498. https://doi.org/10.1038/s41598-017-03755-0
Yang JH, Liu DY, Wang XW, Ji CM, Cheng F, Liu BN, Hu ZY, Chen S, Pental D, Ju YH, Yao P, Li XM, Xie K, Zhang JH, Wang JL, Liu F, Ma WW, Shopan J, Zheng HK, Mackenzie SA, Zhang MF (2016) The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection. Nat Genet 48:1225. https://doi.org/10.1038/ng.3657
Yang Y, Zhu K, Li H, Han S, Meng Q, Khan SU, Fan CC, Xie KB, Zhou YM (2018) Precise editing of CLAVATA genes in Brassica napus L. regulates multilocular silique development. Plant Biotechnol J 16(7):1322–1335. https://doi.org/10.1111/pbi.12872
Ye J, Zhang Y, Cui HH, Liu JW, Wu YQ, Cheng Y, Xu HX, Huang XX, Li ST, Zhou A, Zhang XQ, Bolund L, Chen Q, Wang J, Yang H, Fang L, Shi CM (2018) WEGO 2.0: a web tool for analyzing and plotting GO annotations, 2018 update. Nucleic Acids Res 46(W1):W71–W75. https://doi.org/10.1093/nar/gky400
Zhai LL, Xu L, Wang Y, Zhu XW, Feng HY, Li C, Liu L, Muleke ME, Liu LW (2016) Transcriptional identification and characterization of differentially expressed genes associated with embryogenesis in radish (Raphanus sativus L.). Sci Rep 6:21652. https://doi.org/10.1038/srep21652
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (NSFC, Grant No. 31571698); the National Key Research and Development Program of China (Grant No. 2016YFD0101300) and the Program for Modern Agricultural Industrial Technology System of China (Grant No. CARS-12).
Author information
Authors and Affiliations
Contributions
JS and PX conceived and designed the experiments. HL performed the experiments and wrote the manuscript. HL, KH and QX analyzed the data. JW, BY, CM, JT, TF supervised this study. PX and JS edited and revised the manuscript.
Corresponding author
Additional information
Communicated by Alison M.R. Ferrie.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lu, H., Xu, P., Hu, K. et al. Transcriptome profiling reveals cytokinin promoted callus regeneration in Brassica juncea. Plant Cell Tiss Organ Cult 141, 191–206 (2020). https://doi.org/10.1007/s11240-020-01779-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-020-01779-5