Abstract
Scientific insights gained from the severe acute respiratory syndrome (SARS) and middle east respiratory syndrome (MERS) outbreaks have been assisting scientists and researchers in the quest of antiviral drug discovery process against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Coronaviruses and influenza viruses both rely on the host type 2 transmembrane serine protease, TMPRSS2, for entry and propagation. Recent studies report SARS-CoV-2 also uses TMPRSS2 to enter cells. In the current study, we employed structure-based virtual screening of 1,82,651 natural compounds downloaded from the zin database against the homology model of the TMPRSS2 protein, followed by a molecular dynamics-based simulation to identify potential TMPRSS2 hits. The virtual screening yielded 110 hits with docking scores ranging from −8.654 to −6.775 and glide energies ranging from −55.714 to −29.065 kcal/mol. The binding mode analysis revealed that the hit molecules made H-bond, Pi-Pi stacking and salt bridge contacts with the TMPRSS2 active site residues. MD simulations of the top two hits (ZINC000095912839 and ZINC000085597504) revealed to form a stable complex with TMPRSS2, with a minimal RMSD and RMSF fluctuation. Both the hit structures interacted strongly with the Asp180, Gln183, Gly184, Ser186, Gly207 and Gly209, as predicted by Glide XP docking, and formed a significant H-bond interaction with Ser181 in MD simulation. Among these two, ZINC000095912839 was having the most stable binding interaction with TMPRSS2 of the two molecules. The present study successfully identified TMPRSS2 ligands from a database of zinc natural molecules as potential leads for novel SARs-CoV-2 treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
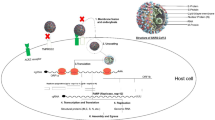
Scientific knowledge gained from the previous SARS and MERS outbreaks has hastened the quest for developing novel antiviral drugs against SARS-CoV-2. Non-structural viral proteins (3-chymotrypsin-like protease, papain-like protease, RNA-dependent RNA polymerase, and its helicase), viral structural proteins (S-glycoprotein) and host protein, transmembrane serine protease 2 (TMPRSS2), are the major antiviral targets identified for their druggability. Human coronaviruses can enter the cell via two pathways: the endosomal mediated entry (i.e. by cathepsins) and the cell-surface or an early endosomal pathway mediated by TMPRSS2 [1,2,3]. SARS-CoV-2 is said to use the latter path, where its spike glycoprotein (S) binds to host ACE2 and TMPRSS2 receptors to allow cell entry [4,5,6,7,8]. Uncoating allows genomic RNA to be used as mRNA to translate the replicase polyproteins. Polyprotein 1a (pp1a) and polyprotein 1ab (pp1ab) are produced by the translation of the replicase gene. Autoproteolytic cleavage of pp1a and pp1ab yields 11 non-structural proteins (nsp1–nsp11) and 15 non-structural proteins (nsp1–nsp10 and nsp12–nsp16), respectively. The nsp12 RNA polymerase is an RNA-dependent RNA polymerase (replicase, RdRp). The replicase employs genomic RNA as a template to generate negative-sense genomic RNAs (gRNAs), which are then used to prepare progeny positive-sense RNA genomes [7, 8]. Through discontinuous transcription of the genome, the replicase synthesises a nested set of sub-genomic RNAs (sgRNAs). Later, the sgRNAs are translated into structural and accessory proteins. The endoplasmic reticulum (ER) structural proteins S, M and E are transported to the ER–Golgi intermediate compartment (ERGIC) for virion assembly [9]. To form nucleocapsids, the N proteins bind to progeny genomic RNA. The smooth-walled vesicles transport the assembled virions from the ERGIC to the cell membrane, where the mature virus particles are released [9, 10].
TMPRSS2 is an S1A class of serine proteases like Factor Xa and trypsin that processes S-protein into two functional subunits, N-terminal receptor-binding domain (S1) and a C-terminal membrane fusion domain (S2) at the S1/S2 cleavage site. The S1 domain facilitates ACE2 recognition and initiates a conformational change in the S2 subunit, leading to the insertion of fusion peptides into the host cell membrane to facilitate membrane fusion and delivery of the viral nucleocapsid into the cytoplasm [1, 4, 10,11,12,13,14,15,16,17,18,19,20,21]. The S2 domain contains a fusion peptide (FP), a second proteolytic site (S2′), an internal fusion peptide (IFP) and two heptad-repeat domains (HR1 and HR2) before the transmembrane domain (TM) (Fig. 1). Further studies speculated that both FP and IFP involved in the viral entry process by cleaving S-protein at both S1/S2 and S2′ cleavage sites are essential [18,19,20].
Structure of TMPRSS2. S1 N-terminal receptor-binding domain; S2 C-terminal membrane fusion domain; SP signal peptide; NTD N-terminal domain (NTD), RBD receptor-binding domain; FP fusion peptide; IFP internal fusion peptide; HR1 heptad repeat 1; HR2 heptad repeat 2; TM transmembrane domain. The SP, S1↓S2 and S2′ cleavage sites are indicated by arrows
TMPRSS2 is expressed in the prostate, stomach, colon, salivary glands and gastrointestinal, urogenital and respiratory epithelia in humans [22]. Overexpressed TMPRSS2 was discovered to be controlled by androgen receptor signalling in prostate cancer. It initiates the metastatic cascades by activating the hepatocyte growth factor (HGF). Prostate cancer metastasis is reported to be inhibited by TMPRSS2 inhibitors [23,24,25]. Previous research has established that TMPRSS2 is an activating protease for respiratory influenza virus [26, 27]. In animal models of SARS-CoV and MERS-CoV infection, the role of host TMPRSS2 in spike protein activation was clearly demonstrated. The absence of TMPRSS2 (in TMPRSS2-knockout mice) significantly reduced airway infection and spread [13]. Furthermore, when Tmprss2(− / −) mice are infected with a re-assorted influenza A virus (IAV) H10 subtype hemagglutinin (HA), they exhibited no abnormal clinical signs, lung lesions, viral antigen, or body weight loss when compared to wild-type mice [28]. In another study, TMPRSS2 is an important HA-activating protease of IAV and IBV (influenza B virus) in primary human type II pneumocytes and human bronchial cells [12]. TMPRSS2-positive VeroE6 cells are highly susceptible to SARS-CoV-2 infection, indicating the role of TMPRSS2 in viral entry into the host cell [15]. SARS-CoV-2 receptors, ACE2 and TMPRSS2 have been found to be most abundant in bronchial transient secretory cells [16]. A recent study confirms that SARS-CoV-2 takes advantage of the host ACE2 for entry and the serine protease, TMPRSS2, for S-protein priming [10, 29,30,31]. Camostat, a TMPRSS2 inhibitor, exhibited inhibition of SARS-CoV-2 host cell entry [10, 32]. These findings strongly suggest that TMPRSS2 is a critical protein required for SARS-CoV-2 host cell entry and, thus, represents a treatment option. As the 3D crystal structure of TMPRSS2 is not available, we used the previously reported homology model of TMPRSS2, which was generated using TMPRSS15 (PDB ID. 4DGJ) [18].
Methodology
The structure-based virtual screening
The Schrödinger software suite’s virtual screening workflow 2018–3 version (Maestro 11.7, Schrödinger, LLC, New York, NY, 2020) was used to screen the Zinc database [33] against the active site of TMPRSS2.
Database and ligand preparation
In total, 1,82,651 molecules from the natural products category of Zinc database were downloaded in 2D SDF format. The Ligprep module of the software was used to prepare these two-dimensional structures. In brief, the molecules were desalted, converted from a 2D structure to a low energy 3D structure, tautomeric and ionised (between pH 6.8 and 7.2 using the Epik module), and all possible stereoisomeric states were generated. Using the OPLS 2005 force field, the energies of generated structures were minimised.
Homology model of TMPRSS2
The homology model of TMPRSS2 protein was obtained from the TMPRSS15 crystallographic structure (PDB ID. 4DGJ) (with 41% of similarity on their peptide sequence) [20]. The obtained template is an aligned sequences of all the available S1A proteases followed by the identification of TMPRSS15 as the most suitable for the current study. PDB id 4DGJ was selected to build the homology model using Prime module of Schrodinger. The generated model was further validated by performing the MD simulation of 100 ns.
Protein preparation and receptor grid generation
The Protein Preparation Wizard was used to generate the homology protein of TMPRSS2 from TMPRSS15 (PDB ID. 4DGJ). Bond order, missing atoms, tautomer/ionization states, water orientation and hydrogen bond networking were all examined in the protein. The OPLS 2005 force field was then applied for constrained energy minimization. The receptor grid was created using the previously prepared protein. The centroid of the workspace ligand (Benzamidine; A: BEN 245) was used to define the size and position of the receptor grid box, with a van der Waals scaling factor of 1.0 and a partial charge cut-off of 0.25.
Virtual screening
The Glide Virtual Screening Workflow was used to perform the virtual screening. As input structures, previously prepared ligands were used. Lipinski’s rule and reactive functional group criteria were used to predict and prefilter the ADME properties of these ligands prior to Glide Docking. In three stages, all ligands that passed through these prefilters were docked into the previously prepared receptor grid structure: The molecules were docked flexibly in Glide HTVS (High Throughput Virtual Screening) mode using the default settings in the first stage. In the second stage, 10% of the high-scoring hits from the previous step were docked flexibly in Glide SP (Standard Precision) mode. In the final stage, 10% of the good scoring hits from the previous step were docked flexibly in Glide XP (Extra Precision) mode.
MD simulation
Desmond software (Schrödinger software suite 2018–3 version) was used to run molecular dynamics (MD) simulations for 100 ns on the top two hits (ZINC000095912839 and ZINC000085597504). The system, which included an SPC solvent model, ligand and protein complex, was constructed in an orthorhombic boundary, considering the buffer distance of 10 Aͦ × 10 Aͦ × 10 Aͦ, and the charge was neutralised with counterions. The system was then minimised, and the simulation was performed for 100 ns using an NPT ensemble system at 300.0 K temperature and 1.0 bar pressure. The trajectory was recorded every 100.0 ps, and 1000 frames were captured to calculate the root mean square deviation (RMSD).
Results and discussion
The structure-based virtual screening
The HTVS docking of the prepared database ligands (1,82,651 molecules) to the active site of the homology model of TMPRSS2 resulted in 10,914 hits. The further docking of these hits in the SP mode yielded in 1091 hits. The final docking of these hits in the XP mode produced 110 hits with a docking score range of −8.654 to −6.775 and a glide energy range of −55.714 to −29.065 kcal/mol (Table 1; Fig. 2).
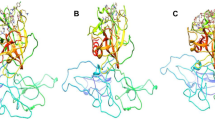
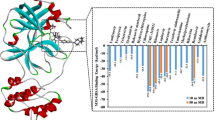
The analysis of the docking complexes of the top 10 hits reveals that the ligands form H-bond, Pi-Pi stacking and salt bridge-type interactions with the active site residues of TMPRSS2 (Table 1; Fig. 3). Both aliphatic and aromatic hydroxyl and amine groups present in these molecular structures formed H-bond interactions with active site residues Val25, His41, Lys87, Asp180, Gln183, Gly184, Ser186, Gly207, Gly209 and Val218 (Table 1; Fig. 4). The Pi-Pi stacking interaction was observed between the benzene moiety of ZINC000000526288 and the active site His41 residue. The salt bridge interaction was observed between the aliphatic amine group of ZINC000000077285 and Asp180 residues of the active site (Table 1; Fig. 3).
ADME properties
The in silico ADME analysis results of TMPRSS2 hits are given in Table 2. The molecules show properties within the permitted limits of Lipinski rule of 5 and Jorgensen’s rule of 3. The results, therefore, suggest that the hit molecules have acceptable ADME properties.
MD simulation
MD simulation of the selected hits was carried out to assess the physical movements of atoms and molecules of the ligand-receptor complex under physiological conditions to gain insights into the protein–ligand interactions. The MD simulation analysis of ligand-receptor complex of ZINC000095912839 and TMPRSS2 shows a stable ligand RMSD of 3.6 Å for the first 60 ns, which later changes to 4.4 Å. In contrast, the protein RMSD was steady at around 1.9 Å for the period of 80 ns and changes to 2.4 Å (Fig. 4). The protein RMSF graph shows that TMPRSS2 residues remain stable during the period of simulation (Fig. 4). Protein–ligand contact analysis shows that molecule ZINC000095912839 interacts with more than 20 active site residues of TMPRSS2. The interaction types observed include H-bonds, hydrophobic, ionic and water bridges. Among these, a prominent H-bond interaction lasting throughout the simulation period was observed with Asp180, Gly209 and Ser181 residues (Fig. 4).
The MD simulation results analysis of ZINC000085597504 and TMPRSS2 show a relatively fluctuating ligand RMSD which seems to be stable at around 8 Å for the first 28 ns, which later fluctuates till 78 ns and becomes stable at 28 Å for the rest of the simulation period. In contrast, the protein RMSD seems to be relatively stable at around 2.1 Å throughout the simulation period (Fig. 5). The protein RMSF graph shows that TMPRSS2 residues remain stable during the period of simulation (Fig. 5). Protein–ligand contact analysis shows that molecule ZINC000085597504 interacts with more than 50 active site residues of TMPRSS2. The interaction types observed include H-bonds, hydrophobic, ionic and water bridges. Among these, a prominent H-bond interaction was observed only with Asp180, Ser181, Ser183, Gly209 and Cys210 residues (Fig. 5).
Overall, the comparison of MD simulation results of the hits with those of Glide XP docking suggests a significant correction for ligand interaction with active site residues. The prominent residues commonly observed include Asp180, Gln 183, Gly184, Gly209, Ser186 and Gly209. Furthermore, MD simulation also identifies a notable H-bond interaction with Ser181 for the three hits, which was not recognized during Glide XP docking.
Conclusion
TMPRSS2 is a vital host target protein that has been recognised as an important antiviral drug target against SARS-CoV-2 infection. There are no solved crystal structures of TMPRSS2 in the protein databank at this time; however, a homology structure derived from TMPRSS15 has been useful in the discovery and development of lead molecules against this target. The potential hit molecules from the zinc natural molecule database were identified using structural-based virtual screening and molecular dynamics-based computational research in the current study. Furthermore, in vitro and in vivo studies with these molecules may shed more light on their potential benefits in the treatment of CoVID-19.
Availability of data and material
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
References
Shirato K, Kawase M, Matsuyama S (2018) Wild-type human coronaviruses prefer cell-surface TMPRSS2 to endosomal cathepsins for cell entry. Virology 517:9–15
Mollica V, Rizzo A, Massari F (2020) The pivotal role of TMPRSS2 in coronavirus disease 2019 and prostate cancer. Future Oncol 16(27):2029–2033
Nandi S, Kumar M, Saxena M, Saxena KA (2021) The antiviral and antimalarial drug REPURPOSING in quest of chemotherapeutics to combat COVID-19 utilizing structure-based molecular docking. Comb Chem High Throughput Screening 24(7):1055–1068
Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB (2020) Pharmacologic treatments for coronavirus disease 2019 (COVID-19): A Review. JAMA
Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 1282:1–23
Roberts DJ, Bebell LM, Edlow AG. SARS-CoV-2 ACE2 and TMPRSS2 (2021) receptor protein expression patterns throughout geSTATION. J Infect Dis
Krishnamurthy PT (2020) Coronavirus Disease 2019: virology and drug targets. Infect Disord Drug Targets
Fung TS, Liu DX (2014) Coronavirus infection, ER stress, apoptosis and innate immunity. Front Microbiol 5:296
Li F (2016) Structure, function, and evolution of coronavirus spike proteins. Annu Rev Virol 3(1):237–261
Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, Erichsen S et al (2019) SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 81(2):271–80 e8
Sakai K, Ami Y, Tahara M, Kubota T, Anraku M, Abe M et al (2014) The host protease TMPRSS2 plays a major role in in vivo replication of emerging H7N9 and seasonal influenza viruses. J Virol 88(10):5608–5616
Limburg H, Harbig A, Bestle D, Stein DA, Moulton HM, Jaeger J et al (2021) TMPRSS2 is the major activating protease of influenza A virus in primary human airway cells and influenza B virus in human type II pneumocytes. J Virol 93(21)
Iwata-Yoshikawa N, Okamura T, Shimizu Y, Hasegawa H, Takeda M, Nagata N (2019) TMPRSS2 contributes to virus spread and immunopathology in the airways of murine models after coronavirus infection. J Virol 93(6)
Abe M, Tahara M, Sakai K, Yamaguchi H, Kanou K, Shirato K et al (2013) TMPRSS2 is an activating protease for respiratory parainfluenza viruses. J Virol 87(21):11930–11935
Matsuyama S, Nao N, Shirato K, Kawase M, Saito S, Takayama I et al (2020) Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc Natl Acad Sci U S A 117(13):7001–7003
Lukassen S, Chua RL, Trefzer T, Kahn NC, Schneider MA, Muley T et al (2020) SARS-CoV-2 receptor ACE2 and TMPRSS2 are primarily expressed in bronchial transient secretory cells. EMBO J e105114
Stopsack KH, Mucci LA, Antonarakis ES, Nelson PS, Kantoff PW. TMPRSS2 and COVID-19 (2020) serendipity or opportunity for intervention? Cancer Discov
Huggins DJ (2020) Structural analysis of experimental drugs binding to the SARS-CoV-2 target TMPRSS2. J Mol Graph Model 100:107710
Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E (2020) The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res 176:104742
Jaimes JA, Andre NM, Chappie JS, Millet JK, Whittaker GR (2020) Phylogenetic analysis and structural modeling of SARS-CoV-2 spike protein reveals an evolutionary distinct and proteolytically sensitive activation loop. J Mol Biol 432(10):3309–3325
Fraser BJ, Beldar S, Seitova A, Hutchinson A, Mannar D, Li Y et al (2021) Structure, activity and inhibition of human TMPRSS2, a protease implicated in SARS-CoV-2 activation. bioRxiv 06.23.449282
Vaarala MH, Porvari K, Kyllonen A, Lukkarinen O, Vihko P (2001) The TMPRSS2 gene encoding transmembrane serine protease is overexpressed in a majority of prostate cancer patients: detection of mutated TMPRSS2 form in a case of aggressive disease. Int J Cancer 94(5):705–710
Tomlins SA, Laxman B, Varambally S, Cao X, Yu J, Helgeson BE et al (2008) Role of the TMPRSS2-ERG gene fusion in prostate cancer. Neoplasia 10(2):177–188
Stopsack KH, Mucci LA, Antonarakis ES, Nelson PS, Kantoff PW (2020) TMPRSS2 and COVID-19: serendipity or opportunity for intervention? Cancer Discov 10(6):779–782
Wang Z, Wang Y, Zhang J, Hu Q, Zhi F, Zhang S et al (2017) Significance of the TMPRSS2:ERG gene fusion in prostate cancer. Mol Med Rep 16(4):5450–5458
Laporte M, Naesens L (2017) Airway proteases: an emerging drug target for influenza and other respiratory virus infections. Curr Opin Virol 24:16–24
Hatesuer B, Bertram S, Mehnert N, Bahgat MM, Nelson PS, Pohlmann S et al (2013) Tmprss2 is essential for influenza H1N1 virus pathogenesis in mice. PLoS Pathog 9(12):e1003774
Lambertz RLO, Gerhauser I, Nehlmeier I, Leist SR, Kollmus H, Pohlmann S et al (2019) Tmprss2 knock-out mice are resistant to H10 influenza A virus pathogenesis. J Gen Virol 100(7):1073–1078
Mahmoud IS, Jarrar YB (2021) Targeting the intestinal TMPRSS2 protease to prevent SARS-CoV-2 entry into enterocytes-prospects and challenges. Mol Biol Rep
Lin H, Cherukupalli S, Feng D, Gao S, Kang D, Zhan P et al (2021) SARS-CoV-2 entry inhibitors targeting virus-ACE2 or virus-TMPRSS2 interactions. Curr Med Chem
Kim TY, Jeon S, Jang Y, Gotina L, Won J, Ju YH et al (2021) Platycodin D, a natural component of Platycodon grandiflorum, prevents both lysosome- and TMPRSS2-driven SARS-CoV-2 infection by hindering membrane fusion. Exp Mol
Kitagawa J, Arai H, Iida H, Mukai J, Furukawa K, Ohtsu S et al (2021) A phase I study of high dose camostat mesylate in healthy adults provides a rationale to repurpose the TMPRSS2 inhibitor for the treatment of COVID-19. Clin Transl Sci
Sterling T, Irwin JJ (2015) ZINC 15–Ligand Discovery for Everyone. J Chem Inf Model 55(11):2324–2337
Acknowledgements
The authors also thank Manipal – Schrödinger Centre for Molecular Simulations and Manipal College of Pharmaceutical Sciences, Manipal Academy of Higher Education, Manipal, India, for providing the facility to carry out this research work.
Funding
The authors received infrastructure support provided by DST-FIST, Govt. of India, New Delhi (Grant No. SR/FST/LSI-574/2013) to the Dept. of Pharmacology, JSS College of Pharmacy, Ooty.
Author information
Authors and Affiliations
Contributions
Conceptualization – K Sreedhara Ranganath Pai, Praveen T Krishnamurthy; investigation – Suman Manandhar, Ravi Kiran Ammu V V V, Kusuma Kumari Garikapati; supervision – K Sreedhara Ranganath Pai, Praveen T Krishnamurthy; writing – original draft – Suman Manandhar, Ravi Kiran Ammu V V V, Kusuma Kumari Garikapati; writing –
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Manandhar, S., Pai, K.R., Krishnamurthy, P.T. et al. Identification, virtual screening and molecular dynamic analysis of novel TMPRSS2 inhibitors from natural compound database as potential entry-blocking agents in SARS-CoV-2 therapy. Struct Chem 33, 1609–1617 (2022). https://doi.org/10.1007/s11224-022-01991-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-022-01991-3