Abstract
Metallo-nucleic acids have been investigated for their applications in the field of nanodevices and genetic expansion. The cytosine-Ag+-cytosine mismatch base pair interactions and their stability in nucleic acids have attracted the attention of chemists. We report a systematic study of canonical, mismatch, Ag+ mediated system with CC, AT, and GC base pairs computationally. The stability of such mismatch base pairs is dependent on the pH range ~ 5 to 9 and the duplexes beyond this range are unstable. The DFT calculations performed with the model DNA duplexes comprising of such mismatch pairs reveal the stability trend while varying the pH conditions. The stability of canonical Watson–Crick ATGC base pairs was compared with the CCAT and CCGC mismatch base pairs and the calculated results at B3LYP-D3/6-31G(d) level of theory in the aqueous phase suggest that the base stacking and hydrogen bonding are well maintained in the former case, however, the larger perturbations in the geometry are observed with the mispair and relatively unstable. The calculated binding energy B3LYP-D3/6-31G(d) level of theory of ATGC is energetically more stable (~ 20 kcal/mol) than the mismatch base pairs. The Ag+ mediated mismatch base pairs i.e., C_CAT and C_CGC examined at the same level of theory suggest that the CC mismatch base pairs complexed with proper alignment to the Ag+ ion and the AT and GC bases maintained the hydrogen bonding interactions. The relative binding energies have been calculated with 6–311 + G(d,p) basis set using B3LYP-D3 and wb97xd DFT functionals in the study. The mismatched base pair duplex systems i.e., C_CAT and C_CGC are structurally similar to the canonical Watson–Crick base pairs and energetically stable by ~ 40 and ~ 50 kcal/mol compared to the canonical ATGC base pairs. The experimental report on the thermal transition profile in 5’-(A)10C(A)10–3’ and 5’ (T)10C(T)10–3’ duplexes showed remarkable stability and corroborate the calculated results (Ono et al. in Chem Commun 4825−4827, 2008). The stability of Ag+ mediated mismatch bases at the higher pH 9 was also examined and the nucleobases such as guanine and thymine would be deprotonated under this condition. The calculated results suggest that the CCA_T and CCG_C duplexes are largely distorted with the complexation of Ag+ with the AT and GC base pairs and would in turn denature the duplex. The AIM analysis performed at B3LYP-D3/6-31G(d) level of theory for all the studied Ag+ mediated complexes reveals that the Ag+ interaction with the corresponding nucleobases was electrostatic in nature. The role of pH in governing the stability of C–Ag+-C complex formation in mismatch base nucleic acids is crucial for their application of genetic expansion and nucleic acid-based nanodevices.
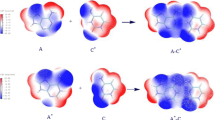
Graphical abstract









Similar content being viewed by others
References
Ono A, Cao S, Togashi H et al (2008) Specific interactions between silver(i) ions and cytosine-cytosine pairs in DNA duplexes. Chem Commun 4825–4827. https://doi.org/10.1039/b808686a
Ono A, Togashi H (2004) Highly selective oligonucleotide-based sensor for mercury(II) in aqueous solutions. Angew Chemie - Int Ed 43:4300–4302. https://doi.org/10.1002/anie.200454172
Torigoe H, Ono A, Kozasa T (2010) HgII ion specifically binds with T: T mismatched base pair in duplex DNA. Chem - A Eur J 16:13218–13225. https://doi.org/10.1002/chem.201001171
Ono A, Torigoe H, Tanaka Y, Okamoto I (2011) Binding of metal ions by pyrimidine base pairs in DNA duplexes. Chem Soc Rev 40:5855–5866. https://doi.org/10.1039/c1cs15149e
Krishnan Y, Simmel FC (2011) Nucleic acid based molecular devices. Angew Chemie - Int Ed 50:3124–3156. https://doi.org/10.1002/anie.200907223
Takezawa Y, Shionoya M (2012) Metal-mediated DNA base pairing: alternatives to hydrogen-bonded Watson-Crick base pairs. Acc Chem Res 45:2066–2076. https://doi.org/10.1021/ar200313h
Torigoe H, Okamoto I, Dairaku T et al (2012) Thermodynamic and structural properties of the specific binding between Ag+ ion and C: C mismatched base pair in duplex DNA to form C-Ag-C metal-mediated base pair. Biochimie 94:2431–2440. https://doi.org/10.1016/j.biochi.2012.06.024
Torigoe H, Miyakawa Y, Ono A, Kozasa T (2011) Thermodynamic properties of the specific binding between Ag+ ions and C: C mismatched base pairs in duplex DNA. Nucleosides, Nucleotides Nucleic Acids 30:149–167. https://doi.org/10.1080/15257770.2011.553210
Urata H, Yamaguchi E, Nakamura Y, Wada S (2011) Pyrimidine-pyrimidine base pairs stabilized by silver(i) ions. Chem Commun 47:941–943. https://doi.org/10.1039/c0cc04091f
Funai T, Miyazaki Y, Aotani M et al (2012) Ag I ion mediated formation of a C-A mispair by DNA polymerases. Angew Chemie - Int Ed 51:6464–6466. https://doi.org/10.1002/anie.201109191
Kondo J, Sugawara T, Saneyoshi H, Ono A (2017) Crystal structure of a DNA duplex containing four Ag(i) ions in consecutive dinuclear Ag(i)-mediated base pairs: 4-thiothymine-2Ag(i)-4-thiothymine. Chem Commun 53:11747–11750. https://doi.org/10.1039/c7cc06153f
Mei H, Yang H, Röhl I, Seela F (2014) Silver arrays inside DNA duplexes constructed from silver(I)-mediated pyrrolo-dc-pyrrolo-dc base pairs. ChemPlusChem 79:914–918. https://doi.org/10.1002/cplu.201402060
Yang H, Mei H, Seela F (2015) Pyrrolo-dC metal-mediated base pairs in the reverse Watson-Crick double helix: enhanced stability of parallel DNA and impact of 6-pyridinyl residues on fluorescence and silver-ion binding. Chem - A Eur J 21:10207–10219. https://doi.org/10.1002/chem.201500582
Kondo J, Tada Y, Dairaku T et al (2015) High-resolution crystal structure of a silver(I)-RNA hybrid duplex containing Watson-Crick-like C-silver(I)-C metallo-base pairs. Angew Chemie - Int Ed 54:13323–13326. https://doi.org/10.1002/anie.201507894
Dairaku T, Furuita K, Sato H et al (2016) Structure determination of an AgI-mediated cytosine–cytosine base pair within DNA duplex in solution with1H/15N/109Ag NMR spectroscopy. Chem - A Eur J 22:13028–13031. https://doi.org/10.1002/chem.201603048
Chen X, Karpenko A, Lopez-Acevedo O (2017) Silver-mediated double helix: structural parameters for a robust DNA building block. ACS Omega 2:7343–7348. https://doi.org/10.1021/acsomega.7b01089
Fortino M, Marino T, Russo N (2015) Theoretical study of silver-ion-mediated base pairs: the case of C-Ag-C and C-Ag-A systems. J Phys Chem A 119:5153–5157. https://doi.org/10.1021/jp5096739
Swasey SM, Leal LE, Lopez-Acevedo O et al (2015) Silver (I) as DNA glue: Ag+-mediated guanine pairing revealed by removing Watson-Crick constraints. Sci Rep 5:10163. https://doi.org/10.1038/srep10163
Becke AD (1993) Density-functional thermochemistry. III. The role of exact exchange. J Chem Phys 98:5648–5652. https://doi.org/10.1063/1.464913
Binkley JS, Pople JA, Hehre WJ (1980) Self-consistent molecular orbital methods. 21. Small split-valence basis sets for first-row elements. J Am Chem Soc 102:939–947. https://doi.org/10.1021/ja00523a008
Kawahara SI, Uchimaru T (2000) Basis set effect on hydrogen bond stabilization energy estimation of the Watson-Crick type nucleic acid base pairs using medium-size basis sets: Single point MP2 evaluations at the HF optimized structures. Phys Chem Chem Phys 2:2869–2872. https://doi.org/10.1039/b001507p
Grimme S, Antony J, Ehrlich S, Krieg H (2010) A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J Chem Phys 132:154104. https://doi.org/10.1063/1.3382344
Chai JD, Head-Gordon M (2008) Long-range corrected hybrid density functionals with damped atom-atom dispersion corrections. Phys Chem Chem Phys 10:6615–6620. https://doi.org/10.1039/b810189b
Marek PH, Szatylowicz H, Krygowski TM (2019) Stacking of nucleic acid bases: optimization of the computational approach—the case of adenine dimers. Struct Chem 30:351–359. https://doi.org/10.1007/s11224-018-1253-7
Murray JS, Politzer P (2011) The electrostatic potential: an overview. Wiley Interdiscip Rev Comput Mol Sci 1:153–163. https://doi.org/10.1002/wcms.19
Bader RFW (1991) A quantum theory of molecular structure and its applications. Chem Rev 91:893–928. https://doi.org/10.1021/cr00005a013
Lu T, Chen F (2012) Multiwfn: A multifunctional wavefunction analyzer. J Comput Chem 33:580–592. https://doi.org/10.1002/jcc.22885
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Had M (2009) Gaussian 09 (revision B1). Gaussian Inc, Wallingford
Drew HR, Wing RM, Takano T et al (1981) Structure of a B-DNA dodecamer: conformation and dynamics. Proc Natl Acad Sci U S A 78:2179–2183. https://doi.org/10.1073/pnas.78.4.2179
Lee JS, Woodsworth ML, Latimer LJP, Morgan AR (1984) Poly(pyrimidine) poly(purine) synthetic DNAs containing 5-methylcytosine form stable triplexes at neutral pH. Nucleic Acids Res 12:6603–6614. https://doi.org/10.1093/nar/12.16.6603
Verdolino V, Cammi R, Munk BH, Schlegel HB (2008) Calculation of pKa values of nucleobases and the guanine oxidation products guanidinohydantoin and spiroiminodihydantoin using density functional theory and a polarizable continuum model. J Phys Chem B 112:16860–16873. https://doi.org/10.1021/jp8068877
Wang Y, Luan BQ, Yang Z et al (2014) Single molecule investigation of Ag+ interactions with single cytosine-, methylcytosine- and hydroxymethylcytosine-cytosine mismatches in a nanopore. Sci Rep 4:5883. https://doi.org/10.1038/srep05883
Zacharias M (2020) Base-pairing and base-stacking contributions to double-stranded DNA formation. J Phys Chem B 124:10345–10352. https://doi.org/10.1021/acs.jpcb.0c07670
Acknowledgements
S.B is thankful to the Academy of Scientific and Innovative Research (AcSIR), Ghaziabad 201002, Uttar Pradesh, India, for his Ph.D. registration. S.B acknowledges CSIR, New Delhi, India for CSIR-SRF fellowship. We thank the reviewers for their valuable comments and suggestions that have helped us to improve the paper. CSIR-CSMCRI 145/2021 registration number.
Funding
S.B. and B.G were financially supported by the Department of biotechnology (DBT), New Delhi (Grant no. BT/PR12730/BID/7/523/2015), and Department of Science & Technology (DST).
Author information
Authors and Affiliations
Contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhai, S., Ganguly, B. Role of pH in the stability of cytosine-cytosine mismatch and canonical AT and GC base pairs mediated with silver ion: a DFT study. Struct Chem 33, 35–47 (2022). https://doi.org/10.1007/s11224-021-01814-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-021-01814-x