Abstract
When double-strand breaks (DSBs) in DNA remain unrepaired, catastrophic loss of genes occurs, leading to translocations, mutations and carcinogenesis. If a sister chromatid is not available at the DNA DSB, non-homologous end joining (NHEJ) is used to join broken ends. The NHEJ pathway comprises synapsis, end processing and ligation. Here, we ask how DSBs in DNA are repaired efficiently. We suggest that colocation of proteins is achieved over time by the following components: stages, where the main actors are assembled, scaffolds that are erected quickly around broken parts to give access, and strings that tether proteins together. In NHEJ, a stage is provided by the Ku heterodimer interacting with DSBs and several other proteins including DNA-PKcs, APLF, BRCA1 and PAXX. A further stage, DNA-PKcs, links the kinase with DNA, Ku, PARP1, BRCA1 and Artemis. A temporary scaffold facilitates repair and is constructed from XRCC4/XLF filaments that bridge Ku bound at DSB ends. LigIV bound to XRCC4 C-termini likely terminates the scaffold, bringing LigIV close to the DNA broken ends. A string, provided by the Artemis C-terminal region, is intrinsically disordered but includes short linear “epitopes” that recognise DNA-PKcs, LigIV and PTIP, so keeping these components nearby. We show that these stages, scaffolds and strings facilitate colocation and efficient DSB repair. Understanding these processes provides insight into the biology of DNA repair and possible therapeutic intervention in cancer and other diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA double-strand breaks (DSBs) are the most severe form of DNA damage in eukaryotic cells and are generated by ionising radiation, reactive oxygen species and DNA replication across nicks [1]. DSBs can lead to cell death, and erroneous repair may result in carcinogenesis through chromosomal translocation or modification. Non-homologous end joining (NHEJ) and homologous recombination (HR) are the two major pathways of DSB repair in human cells. HR functions mainly in the late S/G2 phases of the cell cycle due to the requirement for a sister chromatid. In contrast, NHEJ repairs DSBs directly without a DNA template and can function throughout G1/early S phases [1].
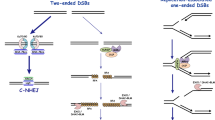
The NHEJ process comprises DNA synapsis, end processing and ligation [2]. During synapsis, Ku70/Ku80 heterodimers assemble around and maintain proximity of broken DNA ends [3]. DNA-dependent protein kinase catalytic subunit (DNA-PKcs), a PI3-kinase-related kinase, is recruited through interaction with the Ku80 C-terminus to form DNA-PK [4, 5]. Two DNA-PK complexes are thought to hold DNA ends close together [6]. DNA-PKcs phosphorylates itself and various other proteins, including NHEJ components. End processing involves nucleases such as Artemis, which exhibits endonuclease activity after activation by DNA-PKcs phosphorylation [7]. The final ligation step is mediated by DNA ligase IV (LigIV) in a stable complex with dimeric XRCC4 [8]. XLF/Cernunnos also interacts with XRCC4 and enhances LigIV DNA ligation, promoting NHEJ [9, 10].
Structural and functional studies of individual components in NHEJ previously reported include DNA-PKcs [11], LigIV [12], XLF [13] and XRCC4 [14]. We have investigated the following binary complexes: LigIV/XRCC4 [14], XRCC4/XLF [15], LigIV/Artemis [16] and DNA-PKcs/Ku80 C-terminus [11; Sibanda BL, Chirgadze DY, Ascher D, Blundell TL, In preparation]. Last year we discovered PAXX [17], an additional component involved in NHEJ (Fig. 1).
These studies and others (e.g. [18, 19]) still leave unanswered questions, including how NHEJ components are coordinated across space and time throughout synapsis, end processing and ligation. We observed three different mechanisms that may ensure appropriate spatial colocation of components: stages that are preformed stable structures where the main actors gather and engage, scaffolds that are constructed quickly and disassembled easily to facilitate cell responses, and strings that tether components together. For NHEJ, these include:
-
1.
A stage comprised of Ku heterodimer, interacting with DNA broken ends and binding a variety of components including DNA-PKcs, aprataxin- and PNKP-like factor (APLF), XLF, BRCA1 and PAXX.
-
2.
A further stage, DNA-PKcs, which interacts with Ku heterodimer, DNA, PARP1, BRCA1, Artemis and other components, often depending on post-translational modification and binding of other factors.
-
3.
A scaffold assembled from protein filaments of alternating XRCC4 and XLF dimers to bridge Ku heterodimers. This bridging is regulated by LigIV, which the tails of XRCC4 bind tightly, therefore bringing LigIV close to DNA broken ends.
-
4.
An intrinsically disordered string comprising the 300-residue C-terminal region of Artemis. Foldable regions of the Artemis C-terminus interact with DNA-PKcs, LigIV and PTIP, which may keep these components close to the DNA repair process in order to be available to function when needed.
Apart from ensuring correct colocation at the appropriate time, we have contended that the complexity of such assemblies is selectively advantageous [20–22]. Binary interactions in regulatory or signalling systems would occur opportunistically in the crowded environment of the cell, giving rise to noise in the system. Conversely, cooperative formation of multiprotein systems is less likely to form opportunistically, especially if they have many components and ordered-assembly mechanisms.
Here, we show that such stages, scaffolds and strings have complementary roles, often binding to the same partners but operating in different ways over space and time. Understanding spatial and temporal organisation of NHEJ may provide insight into whether such mechanisms are universal for mediating colocation in other complex regulatory systems.
Assembling and coordinating the actors in NHEJ
Spatial organisation of molecular assemblies in NHEJ has been studied using nanoelectrospray mass spectrometry, small-angle X-ray scattering (SAXS), X-ray crystallography, NMR and cryo-electron microscopy (cryo-EM). Cumulative insight from these techniques allows us to understand further how NHEJ proceeds efficiently over time.
A stage: Ku heterodimer forms a platform with DNA
The Ku heterodimer assembles around broken DNA ends, forming a stage in which the core ring structure is responsible for interacting with APLF, WRN, XLF, PAXX and BRCA1, and the C-terminal domain of Ku80 interacts with DNA-PKcs [4, 5, 23]. We are using X-ray crystal-structure analysis, cryo-EM and biophysical approaches to define the interactions between components and describe the subsequent structural changes. We comment here on two of the proteins that assemble on this stage and whose interactions have been topics of recent discussion.
PAXX, identified recently in our lab as PAralog of XRCC4 and XLF by using structure-based homology searches, interacts with Ku to promote DNA-DSB repair [17, 24, 25] (Fig. 2). Using an in vitro pull-down assay and biochemical analysis, we showed that the PAXX C-terminus (residues 177–204) interacts with the core ring of Ku in a DNA-dependent manner. We suggested this interaction might be regulated by phosphorylation of the PAXX C-terminus by DNA-PK. In collaboration with the laboratory of Dr. Steve Jackson [17], we demonstrated that PAXX is recruited to DNA-damage sites in cells, and PAXX depletion leads to DSB-repair defects and hypersensitivity to ionising radiation. Using RNA interference and CRISPR-Cas9 to generate PAXX-/- human cells, we showed that PAXX functions with XRCC4 and XLF to mediate DSB repair and cell survival in response to DSB-inducing agents. PAXX promotes Ku-dependent DNA ligation in vitro and assembly of core non-homologous end-joining (NHEJ) components on damaged chromatin in cells.
Two groups have reported an interaction between BRCA1 and Ku [26, 27]. BRCA1 functions as a tumour suppresser gene, and patients carrying the germline or somatic mutations have high risk of developing breast and ovarian cancer [28]. The expression and phosphorylation of BRCA1 is cell cycle-dependent with the highest level at S and M phases [29]. BRCA1 recruitment is important for influencing Ku70/80 retention at the DNA-damage sites and promoting the chromosomal G1 phase of NHEJ. The exact region in BRCA1 responsible for this interaction with Ku was reported as amino acids 1–200 in one report [27] but more recently as 262–552 in the other [26]. Our ongoing objective is to define the region of BRCA1 responsible for the interaction with Ku.
A second stage: DNA-PKcs
In 2010, we described the crystal structure of the ~4000 amino-acid chain of DNA-PKcs complexed with the C-terminus of Ku80 at low resolution, in which the two molecules in the asymmetric unit have slightly different relative positions of the head domain and the large circular structure of HEAT repeats that comprise the stage [11]. More recently, we extended the study to a higher resolution, allowing the chain to be traced. We are using both X-ray crystallography and cryo-EM to define how other molecules, such as Ku and DNA, assemble on this stage. Interestingly, in the S phase, BRCT domains of BRCA1 (see “The strings” section) are reported to interact with DNA-PKcs (near S2056) in a phosphorylation-independent manner to block DNA-PKcs autophosphorylation, reducing NHEJ and increasing HR [30].
A scaffold: XRCC4/XLF filaments
XRCC4 and XLF are both homodimers with similar architectures, including the N-terminal head domains and C-terminal tail regions [13, 14]. Our group [12, 15] and others [31–35] have shown by mutagenesis study, size chromatography, SAXS, native mass spectrometry and X-ray crystallography that XRCC4 and XLF dimers interact through their head domains, leading to the formation of XRCC4/XLF filaments, elongation of which is regulated by LigIV [12]. The alternating XRCC4/XLF helical polymers are flexible, and multiple copies of filament can assemble into cylinders with a diameter that may accommodate and colocate other components of NHEJ (Fig. 3). As a result, XRCC4/XLF filaments may function as a scaffold to allow the alignment of the two broken DNA ends and the local assembly of NHEJ components to enhance LigIV function.
XRCC4 and XLF. In our crystal structure, alternating XRCC4/XLF helical polymers can assemble into cylinders with diameters that may accommodate and colocate other components of NHEJ (PDB ID: 3W03). a XLF and XRCC4 dimers interact through their head domains to form a helical XRCC4/XLF filament. b The filaments assemble as a cylinder in the crystal structure. The view perpendicular to the filament axis is shown with a single filament. c The central space created by the filament could potentially accommodate nucleosome, Ku and DNA-PKcs
The strings
Artemis C-terminal region
Artemis, which belongs to the metallo-β-lactamase superfamily, is the major nuclease involved in the end-processing step of NHEJ. It has been estimated that Artemis repairs 10–15 % of DSBs generated by IR [36]. The Artemis gene DCLRE1C was first discovered mutated in patients who had radiosensitive-severe combined immunodeficiency (RS-SCID) and Athabascan SCID (SCIDA) [7]. It was later found that Artemis is the essential endonuclease that opens the hairpin intermediate structure upon stimulation by DNA-PKcs during V(D)J recombination [37]. The nuclease also has intrinsic 5′ exonuclease activity [38] and weak endonuclease activity on ssDNA in vitro, which is likely stimulated when forming a complex with DNA-PKcs [39].
The Artemis structure can be divided into two regions, each accounting for approximately half of its size (Fig. 4). These regions correspond to the N-terminal nuclease, adopting β-metallo-lactamase and β-CASP domains [40] homologous to RNaseJ, Apollo and SNM1A, and a C-terminal region, which has no enzyme activity but is nevertheless indispensable for normal Artemis function [41]. IUPRED (Prediction of Intrinsically Unstructured Proteins) [42] predicts the C-terminal region of Artemis to be mostly intrinsically disordered, and ANCHOR (Prediction of Protein Binding Regions in Disordered Proteins) [43] analysis predicts several short regions within the intrinsically disordered C-terminal region to bind globular or structured protein partners (Fig. 5).
IUPred and ANCHOR analysis of Artemis with sequence alignment. The C-terminal region of Artemis is predicted to be extensively intrinsically disordered according to IUPred. ANCHOR analysis shows that within the intrinsically disordered region, several segments are likely to interact with other proteins. Sequence alignment also shows that many of these segments are conserved
We previously showed that Artemis residues 485–495 undergo concerted folding and binding with the first two helices of LigIV to form a three-helix bundle (Fig. 6). Residue W489 of Artemis forms a hydrogen bond with D18 of LigIV, stabilised by interactions between F492/F493 of Artemis and F49/F42 of LigIV. Moreover, P487 of Artemis resides in the hydrophobic pocket formed by L53, A52 and F49 of LigIV [16].
Interaction between LigIV and Artemis. The crystal structure of the catalytic domains of LigIV (orange) and the Artemis fragment (amino-acid residues 485–495; silver) is shown on the left. The interaction of the first two helices of LigIV with Artemis is shown in close view on the right (Color figure online)
Although Artemis was predicted to interact with DNA-PKcs via residues 399–404 [44], recent pull-down results from our group show that residues 399–404 are not sufficient. While peptides containing residues 399–408 and 399–426 of Artemis could not pull down DNA-PKcs, residues 385–413 of Artemis could do so, and even showed a stronger interaction with DNA-PKcs compared to full-length Artemis, indicating that residues that are N-terminal to 399–404 are needed for the Artemis/DNA-PKcs interaction. Furthermore, residues 641–660 of Artemis may bind to the second BRCT domain of an adaptor protein, PTIP, via phosphorylation [43].
Thus, it is likely that the Artemis C-terminus binds to several key components, including DNA-PKcs, LigIV and PTIP [45], facilitating colocation of these components with broken DNA ends. The list may be longer since other studies reveal that the Artemis C-terminal region interacts with other proteins. For example, it was shown that Artemis binds to the Mre11/Rad50/Nbs1 complex (MRN complex) in an ATM-dependent manner under ionising radiation [46]. Although the precise binding region is not defined, it is likely that the Artemis C-terminus is involved given that hyperphosphorylation of Artemis, especially S645 of the SQ/TQ site on the C-terminal region, is dependent on ATM. Additionally, Artemis was co-immunoprecipitated with Nbs1, and Nbs1/Mre11 were co-immunoprecipitated with Artemis reciprocally [46].
APLF
APLF (Aprataxin- and-PNK-Like Factor) similarly includes multiple binding sites throughout the protein, which facilitate its function both as a stabiliser for the NHEJ holo-complex at DNA-damage sites and as a stimulator of DNA-end ligation [23, 47]. The N-terminal FHA domain of APLF interacts with XRCC4 in a phosphorylation-dependent manner [48]. The amino acid sequence of a large region of APLF in between the structured N- and C-terminal regions is predicted to be intrinsically disordered and is poorly conserved in evolution (Fig. 7). However, a short sequence (residues 182–192) in this region is conserved in evolution from the sponge (Amphimedon queenslandica) to human. This conserved sequence is important for the interaction with Ku [23, 49–51], likely through concerted folding and binding. Consequently, this binding site within the intrinsically disordered region of APLF facilitates colocation of the Ku heterodimer with regions of APLF that have globular structures.
BRCA1
BRCA1 contains 1863 residues with the two most conserved domains in BRCA1 located at the N- and C-termini. The N-terminal RING domain, which forms a heterodimer with another RING domain from BARD1 [52], functions as an E3 ubiquitin ligase [53, 54]. The N-terminal region (1–304) was also recently found to bind the protein OLA1 during centrosome regulation [55]. The C-terminal region of BRCA1 contains tandem BRCT domains, which appear to bind DNA-PKcs in a non-phosphorylation-dependent manner, as discussed above, and also bind phosphopeptides containing the pSXXF motif [56–58]. We recently determined the structure of the complex of BRCA1 with the C-terminal phosphopeptide of Abraxas (Fig. 8) [59]. Abraxas is part of BRCA1-A complex facilitating the recruitment of BRCA1 to the DNA-damaged site. In collaboration with Dr. Bin Wang (MD Anderson Cancer Center, Houston, TX), we demonstrated that ionising-radiation-induced and ATM-dependent phosphorylation of a serine residue of Abraxas next to the pSPxF motif induces dimerisation of the BRCT/Abraxas complex. Mutation of the ionising-radiation-induced phosphorylation site of Abraxas leads to a deficiency in BRCA1 accumulation and cellular sensitivity to ionising radiation. These findings suggest a novel role of the BRCT/Abraxas complex in the DNA-damage response [46].
Dimerisation of two BRCT/Abraxas complexes. The surface representation of two BRCT/Abraxas complexes is shown in pink and blue, respectively (PDB code: 4Y18). The motif-interaction site, which contains phosphorylated S406 (pS406), is indicated by the circle formed from a dashed line. Phosphorylation at S404 is indicated as pS404 (Color figure online)
Protein interactions within the N-terminal and C-terminal domains of BRCA1 facilitate its role in the DNA-damage response and DNA repair, while the long and less conserved flexible region between these domains provides additional sites for more dynamic and regulated protein interactions (Fig. 9). This disordered region contains the nuclear localisation signal (NLS; amino-acid residues 500–508) [60], PALB-interaction site (amino-acid residues 1397–1424) [61, 62] and phosphorylation sites that are targeted by ATM, Chk2, Cdk2 and DNA-PK at different phases of the cell cycle [63–67]. Although the function of these phosphorylation events is not understood completely, key phosphorylation events include the following: S1423 is phosphorylated by ATM for check-point G2/M function [66], and S988 is phosphorylated by Chk2 [65] for release from Chk2 and HR activity [65, 68]. Thus, the flexible inter-domain region of BRCA1 functions as a string to tether proteins for use as needed.
Discussion
In order for NHEJ to function accurately, spatial and temporal organisation of NHEJ proteins should be as efficient as possible. This is achieved by colocation of components through macromolecular stages, scaffolds and strings. Despite efforts to understand the spatial assembly of NHEJ complexes by observing their structures, the exact coordination of NHEJ proteins at a DSB site remains unclear. This is due to the dynamic and complex nature of NHEJ proteins, which should gather in a DSB-dependent manner and also disassemble quickly after DNA damage is repaired. Therefore, many NHEJ protein–protein interactions are weak or require the presence of DNA. Moreover, the proteins may assemble differently depending on the type of DNA damage. To observe such a flexible and dynamic biological system, we need to design DNA structures that we would like to study, “freeze” intermediate states of the NHEJ process, for instance, using mutations or crosslinking, and sort heterogeneous samples according to structural state. These are extremely challenging tasks for X-ray crystallography. However, recent advances in cryo-EM offer an attractive approach to overcome these difficulties.
Nevertheless, the spatial organisation of NHEJ complexes is better understood than the temporal organisation of NHEJ. Varying models for NHEJ recruitment have been proposed, and our view is that NHEJ is not a pre-programmed process, though it does require the sequential recruitment of stages, scaffolds and strings. The scaffold proteins may have multiple weak interactions with the stage proteins, therefore permitting stabilisation of the scaffolds only when the complete stage is assembled. Alternatively, the interactions may be regulated through allostery involving conformational change; DNA-PKcs conformational changes appear to be activated by post-translational modification, including auto-phosphorylation. To study the temporal organisation of NHEJ proteins, it will be necessary to exploit single-molecule approaches. Indeed, the synapsis by core NHEJ proteins [69] and a dynamic nature of the filament formation of XRCC4 and XLF, which are hard to be visualised by current structural techniques, have been investigated by those approaches [70, 71]. In combination with structural studies of NHEJ complexes, single-molecule approaches may reveal how NHEJ proteins are recruited to DSB sites. However, one of the pitfalls of such in vitro approaches is the assumption that all NHEJ proteins are known. The discovery of PAXX in the past 2 years underlines the possibility that there are components of NHEJ that remain unknown.
Apart from advancing our knowledge of this important DNA DSB-repair system, understanding the interactions of NHEJ components across space and time will indicate novel druggable targets for therapeutic intervention in cancer and a number of rare genetic diseases. Linear foldable regions within intrinsically disordered polypeptides bind to ligandable sites with well-defined pockets [72, 73]. In HR, we have exploited this feature by using fragment-based design to target Rad51-BRCA2 and develop nanomolar drug candidates [73]. We continue to explore the ligandability of Artemis-LigIV and other interactions mediated by the intrinsically disordered C-terminus of Artemis (Esswein SE, Ochi T and Blundell TL, unpublished).
Our research not only sheds light on the spatial and temporal organisation of NHEJ, but also invites speculation as to whether similar stages, scaffolds and strings operate in other complex regulatory systems. The existence of many globular proteins or complexes that bind to multiple components (stages), systems that assemble to form rods and other cell structural elements (scaffolds), and areas of concerted folding and binding within intrinsically disordered regions that facilitate colocation of other globular proteins (strings) suggest that our model may be generic. If so, defining these properties will further explain how other multicomponent complexes are coordinated across space and time in cells and also guide future efforts to regulate these pathways. Such regulation may be possible through modulation of component colocation, particularly by targeting the sites where intrinsically disordered regions interact with globular proteins.
References
O’Driscoll M, Jeggo PA (2006) The role of double-strand break repair—insights from human genetics. Nat Rev Genet 7(1):45–54
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211. doi:10.1146/annurev.biochem.052308.093131
Walker JR, Corpina RA, Goldberg J (2001) Structure of the Ku heterodimer bound to DNA and its implications for double-strand break repair. Nature 412(6847):607–614. doi:10.1038/35088000
Gell D, Jackson SP (1999) Mapping of protein–protein interactions within the DNA-dependent protein kinase complex. Nucleic Acids Res 27(17):3494–3502
Singleton BK, Torres-Arzayus MI, Rottinghaus ST, Taccioli GE, Jeggo PA (1999) The C terminus of Ku80 activates the DNA-dependent protein kinase catalytic subunit. Mol Cell Biol 19(5):3267–3277
Spagnolo L, Rivera-Calzada A, Pearl LH, Llorca O (2006) Three-dimensional structure of the human DNA-PKcs/Ku70/Ku80 complex assembled on DNA and its implications for DNA DSB repair. Mol Cell 22(4):511–519. doi:10.1016/j.molcel.2006.04.013
Moshous D, Callebaut I, de Chasseval R, Corneo B, Cavazzana-Calvo M, Le Deist F, Tezcan I, Sanal O, Bertrand Y, Philippe N, Fischer A, de Villartay JP (2001) Artemis, a novel DNA double-strand break repair/V(D)J recombination protein, is mutated in human severe combined immune deficiency. Cell 105(2):177–186
Grawunder U, Wilm M, Wu X, Kulesza P, Wilson TE, Mann M, Lieber MR (1997) Activity of DNA ligase IV stimulated by complex formation with XRCC4 protein in mammalian cells. Nature 388(6641):492–495. doi:10.1038/41358
Buck D, Moshous D, de Chasseval R, Ma Y, le Deist F, Cavazzana-Calvo M, Fischer A, Casanova JL, Lieber MR, de Villartay JP (2006) Severe combined immunodeficiency and microcephaly in siblings with hypomorphic mutations in DNA ligase IV. Eur J Immunol 36(1):224–235. doi:10.1002/eji.200535401
Ahnesorg P, Smith P, Jackson SP (2006) XLF interacts with the XRCC4-DNA ligase IV complex to promote DNA nonhomologous end-joining. Cell 124(2):301–313. doi:10.1016/j.cell.2005.12.031
Sibanda BL, Chirgadze DY, Blundell TL (2010) Crystal structure of DNA-PKcs reveals a large open-ring cradle comprised of HEAT repeats. Nature 463(7277):118–121. doi:10.1038/nature08648
Ochi T, Wu Q, Chirgadze DY, Grossmann JG, Bolanos-Garcia VM, Blundell TL (2012) Structural insights into the role of domain flexibility in human DNA ligase IV. Structure 20(7):1212–1222. doi:10.1016/j.str.2012.04.012
Li Y, Chirgadze DY, Bolanos-Garcia VM, Sibanda BL, Davies OR, Ahnesorg P, Jackson SP, Blundell TL (2008) Crystal structure of human XLF/Cernunnos reveals unexpected differences from XRCC4 with implications for NHEJ. EMBO J 27(1):290–300. doi:10.1038/sj.emboj.7601942
Sibanda BL, Critchlow SE, Begun J, Pei XY, Jackson SP, Blundell TL, Pellegrini L (2001) Crystal structure of an Xrcc4-DNA ligase IV complex. Nat Struct Biol 8(12):1015–1019. doi:10.1038/nsb725
Wu Q, Ochi T, Matak-Vinkovic D, Robinson CV, Chirgadze DY, Blundell TL (2011) Non-homologous end-joining partners in a helical dance: structural studies of XLF-XRCC4 interactions. Biochem Soc Trans 39(5):1387–1392. doi:10.1042/BST0391387 (suppl 1382 p following 1392)
Ochi T, Gu X, Blundell TL (2013) Structure of the catalytic region of DNA ligase IV in complex with an Artemis fragment sheds light on double-strand break repair. Structure 21(4):672–679. doi:10.1016/j.str.2013.02.014
Ochi T, Blackford AN, Coates J, Jhujh S, Mehmood S, Tamura N, Travers J, Wu Q, Draviam VM, Robinson CV, Blundell TL, Jackson SP (2015) DNA repair. PAXX, a paralog of XRCC4 and XLF, interacts with Ku to promote DNA double-strand break repair. Science 347(6218):185–188. doi:10.1126/science.1261971
Radhakrishnan SK, Jette N, Lees-Miller SP (2014) Non-homologous end joining: emerging themes and unanswered questions. DNA Repair (Amst) 17:2–8. doi:10.1016/j.dnarep.2014.01.009
Rambo RP, Tainer JA (2013) Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496(7446):477–481. doi:10.1038/nature12070
Blundell TL, Burke DF, Chirgadze D, Dhanaraj V, Hyvonen M, Innis CA, Parisini E, Pellegrini L, Sayed M, Sibanda BL (2000) Protein–protein interactions in receptor activation and intracellular signalling. Biol Chem 381(9–10):955–959. doi:10.1515/BC.2000.117
Bolanos-Garcia VM, Wu Q, Ochi T, Chirgadze DY, Sibanda BL, Blundell TL (2012) Spatial and temporal organization of multi-protein assemblies: achieving sensitive control in information-rich cell-regulatory systems. Philos Trans A Math Phys Eng Sci 370(1969):3023–3039. doi:10.1098/rsta.2011.0268
Blaszczyk M, Harmer NJ, Chirgadze DY, Ascher DB, Blundell TL (2015) Achieving high signal-to-noise in cell regulatory systems: spatial organization of multiprotein transmembrane assemblies of FGFR and MET receptors. Prog Biophys Mol Biol 118(3):103–111. doi:10.1016/j.pbiomolbio.2015.04.007
Grundy GJ, Rulten SL, Zeng Z, Arribas-Bosacoma R, Iles N, Manley K, Oliver A, Caldecott KW (2013) APLF promotes the assembly and activity of non-homologous end joining protein complexes. EMBO J 32(1):112–125. doi:10.1038/emboj.2012.304
Xing M, Yang M, Huo W, Feng F, Wei L, Jiang W, Ning S, Yan Z, Li W, Wang Q, Hou M, Dong C, Guo R, Gao G, Ji J, Zha S, Lan L, Liang H, Xu D (2015) Interactome analysis identifies a new paralogue of XRCC4 in non-homologous end joining DNA repair pathway. Nat Commun 6:6233. doi:10.1038/ncomms7233
Craxton A, Somers J, Munnur D, Jukes-Jones R, Cain K, Malewicz M (2015) XLS (c9orf142) is a new component of mammalian DNA double-stranded break repair. Cell Death Differ 22(6):890–897. doi:10.1038/cdd.2015.22
Jiang G, Plo I, Wang T, Rahman M, Cho JH, Yang E, Lopez BS, Xia F (2013) BRCA1-Ku80 protein interaction enhances end-joining fidelity of chromosomal double-strand breaks in the G1 phase of the cell cycle. J Biol Chem 288(13):8966–8976. doi:10.1074/jbc.M112.412650
Wei L, Lan L, Hong Z, Yasui A, Ishioka C, Chiba N (2008) Rapid recruitment of BRCA1 to DNA double-strand breaks is dependent on its association with Ku80. Mol Cell Biol 28(24):7380–7393. doi:10.1128/MCB.01075-08
Futreal PA, Liu Q, Shattuck-Eidens D, Cochran C, Harshman K, Tavtigian S, Bennett LM, Haugen-Strano A, Swensen J, Miki Y et al (1994) BRCA1 mutations in primary breast and ovarian carcinomas. Science 266(5182):120–122
Chen Y, Farmer AA, Chen CF, Jones DC, Chen PL, Lee WH (1996) BRCA1 is a 220-kDa nuclear phosphoprotein that is expressed and phosphorylated in a cell cycle-dependent manner. Cancer Res 56(14):3168–3172
Davis AJ, Chi L, So S, Lee KJ, Mori E, Fattah K, Yang J, Chen DJ (2014) BRCA1 modulates the autophosphorylation status of DNA-PKcs in S phase of the cell cycle. Nucleic Acids Res 42(18):11487–11501. doi:10.1093/nar/gku824
Ropars V, Drevet P, Legrand P, Baconnais S, Amram J, Faure G, Marquez JA, Pietrement O, Guerois R, Callebaut I, Le Cam E, Revy P, de Villartay JP, Charbonnier JB (2011) Structural characterization of filaments formed by human Xrcc4-Cernunnos/XLF complex involved in nonhomologous DNA end-joining. Proc Natl Acad Sci USA 108(31):12663–12668. doi:10.1073/pnas.1100758108
Hammel M, Rey M, Yu Y, Mani RS, Classen S, Liu M, Pique ME, Fang S, Mahaney BL, Weinfeld M, Schriemer DC, Lees-Miller SP, Tainer JA (2011) XRCC4 protein interactions with XRCC4-like factor (XLF) create an extended grooved scaffold for DNA ligation and double strand break repair. J Biol Chem 286(37):32638–32650. doi:10.1074/jbc.M111.272641
Andres SN, Vergnes A, Ristic D, Wyman C, Modesti M, Junop M (2012) A human XRCC4-XLF complex bridges DNA. Nucleic Acids Res 40(4):1868–1878. doi:10.1093/nar/gks022
Andres SN, Modesti M, Tsai CJ, Chu G, Junop MS (2007) Crystal structure of human XLF: a twist in nonhomologous DNA end-joining. Mol Cell 28(6):1093–1101. doi:10.1016/j.molcel.2007.10.024
Malivert L, Ropars V, Nunez M, Drevet P, Miron S, Faure G, Guerois R, Mornon JP, Revy P, Charbonnier JB, Callebaut I, de Villartay JP (2010) Delineation of the Xrcc4-interacting region in the globular head domain of cernunnos/XLF. J Biol Chem 285(34):26475–26483. doi:10.1074/jbc.M110.138156
Riballo E, Kuhne M, Rief N, Doherty A, Smith GC, Recio MJ, Reis C, Dahm K, Fricke A, Krempler A, Parker AR, Jackson SP, Gennery A, Jeggo PA, Lobrich M (2004) A pathway of double-strand break rejoining dependent upon ATM, Artemis, and proteins locating to gamma-H2AX foci. Mol Cell 16(5):715–724. doi:10.1016/j.molcel.2004.10.029
Ma Y, Pannicke U, Schwarz K, Lieber MR (2002) Hairpin opening and overhang processing by an Artemis/DNA-dependent protein kinase complex in nonhomologous end joining and V(D)J recombination. Cell 108(6):781–794
Li S, Chang HH, Niewolik D, Hedrick MP, Pinkerton AB, Hassig CA, Schwarz K, Lieber MR (2014) Evidence that the DNA endonuclease ARTEMIS also has intrinsic 5′-exonuclease activity. J Biol Chem 289(11):7825–7834. doi:10.1074/jbc.M113.544874
Gu J, Li S, Zhang X, Wang LC, Niewolik D, Schwarz K, Legerski RJ, Zandi E, Lieber MR (2010) DNA-PKcs regulates a single-stranded DNA endonuclease activity of Artemis. DNA Repair (Amst) 9(4):429–437. doi:10.1016/j.dnarep.2010.01.001
Callebaut I, Moshous D, Mornon JP, de Villartay JP (2002) Metallo-beta-lactamase fold within nucleic acids processing enzymes: the beta-CASP family. Nucleic Acids Res 30(16):3592–3601
Huang Y, Giblin W, Kubec M, Westfield G, St Charles J, Chadde L, Kraftson S, Sekiguchi J (2009) Impact of a hypomorphic Artemis disease allele on lymphocyte development, DNA end processing, and genome stability. J Exp Med 206(4):893–908. doi:10.1084/jem.20082396
Dosztanyi Z, Csizmok V, Tompa P, Simon I (2005) The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J Mol Biol 347(4):827–839. doi:10.1016/j.jmb.2005.01.071
Meszaros B, Simon I, Dosztanyi Z (2009) Prediction of protein binding regions in disordered proteins. PLoS Comput Biol 5(5):e1000376. doi:10.1371/journal.pcbi.1000376
Soubeyrand S, Pope L, De Chasseval R, Gosselin D, Dong F, de Villartay JP, Hache RJ (2006) Artemis phosphorylated by DNA-dependent protein kinase associates preferentially with discrete regions of chromatin. J Mol Biol 358(5):1200–1211. doi:10.1016/j.jmb.2006.02.061
Wang J, Aroumougame A, Lobrich M, Li Y, Chen D, Chen J, Gong Z (2014) PTIP associates with Artemis to dictate DNA repair pathway choice. Genes Dev 28(24):2693–2698. doi:10.1101/gad.252478.114
Chen L, Morio T, Minegishi Y, Nakada S, Nagasawa M, Komatsu K, Chessa L, Villa A, Lecis D, Delia D, Mizutani S (2005) Ataxia-telangiectasia-mutated dependent phosphorylation of Artemis in response to DNA damage. Cancer Sci 96(2):134–141. doi:10.1111/j.1349-7006.2005.00019.x
Rulten SL, Cortes-Ledesma F, Guo L, Iles NJ, Caldecott KW (2008) APLF (C2orf13) is a novel component of poly(ADP-ribose) signaling in mammalian cells. Mol Cell Biol 28(14):4620–4628. doi:10.1128/MCB.02243-07
Cherry AL, Nott TJ, Kelly G, Rulten SL, Caldecott KW, Smerdon SJ (2015) Versatility in phospho-dependent molecular recognition of the XRCC1 and XRCC4 DNA-damage scaffolds by aprataxin-family FHA domains. DNA Repair (Amst) 35:116–125. doi:10.1016/j.dnarep.2015.10.002
Iles N, Rulten S, El-Khamisy SF, Caldecott KW (2007) APLF (C2orf13) is a novel human protein involved in the cellular response to chromosomal DNA strand breaks. Mol Cell Biol 27(10):3793–3803. doi:10.1128/MCB.02269-06
Kanno S, Kuzuoka H, Sasao S, Hong Z, Lan L, Nakajima S, Yasui A (2007) A novel human AP endonuclease with conserved zinc-finger-like motifs involved in DNA strand break responses. EMBO J 26(8):2094–2103. doi:10.1038/sj.emboj.7601663
Macrae CJ, McCulloch RD, Ylanko J, Durocher D, Koch CA (2008) APLF (C2orf13) facilitates nonhomologous end-joining and undergoes ATM-dependent hyperphosphorylation following ionizing radiation. DNA Repair (Amst) 7(2):292–302. doi:10.1016/j.dnarep.2007.10.008
Wu LC, Wang ZW, Tsan JT, Spillman MA, Phung A, Xu XL, Yang MC, Hwang LY, Bowcock AM, Baer R (1996) Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat Genet 14(4):430–440. doi:10.1038/ng1296-430
Hashizume R, Fukuda M, Maeda I, Nishikawa H, Oyake D, Yabuki Y, Ogata H, Ohta T (2001) The RING heterodimer BRCA1–BARD1 is a ubiquitin ligase inactivated by a breast cancer-derived mutation. J Biol Chem 276(18):14537–14540. doi:10.1074/jbc.C000881200
Wu-Baer F, Lagrazon K, Yuan W, Baer R (2003) The BRCA1/BARD1 heterodimer assembles polyubiquitin chains through an unconventional linkage involving lysine residue K6 of ubiquitin. J Biol Chem 278(37):34743–34746. doi:10.1074/jbc.C300249200
Matsuzawa A, Kanno S, Nakayama M, Mochiduki H, Wei L, Shimaoka T, Furukawa Y, Kato K, Shibata S, Yasui A, Ishioka C, Chiba N (2014) The BRCA1/BARD1-interacting protein OLA1 functions in centrosome regulation. Mol Cell 53(1):101–114. doi:10.1016/j.molcel.2013.10.028
Yu X, Chini CC, He M, Mer G, Chen J (2003) The BRCT domain is a phospho-protein binding domain. Science 302(5645):639–642. doi:10.1126/science.1088753
Manke IA, Lowery DM, Nguyen A, Yaffe MB (2003) BRCT repeats as phosphopeptide-binding modules involved in protein targeting. Science 302(5645):636–639. doi:10.1126/science.1088877
Rodriguez M, Yu X, Chen J, Songyang Z (2003) Phosphopeptide binding specificities of BRCA1 COOH-terminal (BRCT) domains. J Biol Chem 278(52):52914–52918. doi:10.1074/jbc.C300407200
Wu Q, Paul A, Su D, Mehmood S, Foo TK, Ochi T, Bunting EL, Xia B, Robinson CV, Wang B, Blundell TL (2016) Structure of BRCA1-BRCT/abraxas complex reveals phosphorylation-dependent BRCT dimerization at DNA damage sites. Mol Cell 61(3):434–448. doi:10.1016/j.molcel.2015.12.017
Thakur S, Zhang HB, Peng Y, Le H, Carroll B, Ward T, Yao J, Farid LM, Couch FJ, Wilson RB, Weber BL (1997) Localization of BRCA1 and a splice variant identifies the nuclear localization signal. Mol Cell Biol 17(1):444–452
Sy SM, Huen MS, Chen J (2009) PALB2 is an integral component of the BRCA complex required for homologous recombination repair. Proc Natl Acad Sci USA 106(17):7155–7160. doi:10.1073/pnas.0811159106
Zhang F, Ma J, Wu J, Ye L, Cai H, Xia B, Yu X (2009) PALB2 links BRCA1 and BRCA2 in the DNA-damage response. Curr Biol 19(6):524–529. doi:10.1016/j.cub.2009.02.018
Cortez D, Wang Y, Qin J, Elledge SJ (1999) Requirement of ATM-dependent phosphorylation of brca1 in the DNA damage response to double-strand breaks. Science 286(5442):1162–1166
Greenberg RA, Sobhian B, Pathania S, Cantor SB, Nakatani Y, Livingston DM (2006) Multifactorial contributions to an acute DNA damage response by BRCA1/BARD1-containing complexes. Genes Dev 20(1):34–46. doi:10.1101/gad.1381306
Lee JS, Collins KM, Brown AL, Lee CH, Chung JH (2000) hCds1-mediated phosphorylation of BRCA1 regulates the DNA damage response. Nature 404(6774):201–204. doi:10.1038/35004614
Xu B, Kim S, Kastan MB (2001) Involvement of Brca1 in S-phase and G(2)-phase checkpoints after ionizing irradiation. Mol Cell Biol 21(10):3445–3450. doi:10.1128/MCB.21.10.3445-3450.2001
Ruffner H, Jiang W, Craig AG, Hunter T, Verma IM (1999) BRCA1 is phosphorylated at serine 1497 in vivo at a cyclin-dependent kinase 2 phosphorylation site. Mol Cell Biol 19(7):4843–4854
Zhang J, Willers H, Feng Z, Ghosh JC, Kim S, Weaver DT, Chung JH, Powell SN, Xia F (2004) Chk2 phosphorylation of BRCA1 regulates DNA double-strand break repair. Mol Cell Biol 24(2):708–718
Graham TG, Walter JC, Loparo JJ (2016) Two-stage synapsis of DNA ends during non-homologous end joining. Mol Cell 61(6):850–858. doi:10.1016/j.molcel.2016.02.010
Reid DA, Keegan S, Leo-Macias A, Watanabe G, Strande NT, Chang HH, Oksuz BA, Fenyo D, Lieber MR, Ramsden DA, Rothenberg E (2015) Organization and dynamics of the nonhomologous end-joining machinery during DNA double-strand break repair. Proc Natl Acad Sci USA 112(20):E2575–E2584. doi:10.1073/pnas.1420115112
Brouwer I, Sitters G, Candelli A, Heerema SJ, Heller I, de Melo AJ, Zhang H, Normanno D, Modesti M, Peterman EJ, Wuite GJ (2016) Sliding sleeves of XRCC4-XLF bridge DNA and connect fragments of broken DNA. Nature 535(7613):566–569. doi:10.1038/nature18643
Jubb H, Blundell TL, Ascher DB (2015) Flexibility and small pockets at protein–protein interfaces: new insights into druggability. Prog Biophys Mol Biol 119(1):2–9. doi:10.1016/j.pbiomolbio.2015.01.009
Scott DE, Coyne AG, Venkitaraman A, Blundell TL, Abell C, Hyvonen M (2015) Small-molecule inhibitors that target protein–protein interactions in the RAD51 family of recombinases. ChemMedChem 10(2):296–303. doi:10.1002/cmdc.201402428
Eustermann S, Brockmann C, Mehrotra PV, Yang JC, Loakes D, West SC, Ahel I, Neuhaus D (2010) Solution structures of the two PBZ domains from human APLF and their interaction with poly(ADP-ribose). Nat Struct Mol Biol 17(2):241–243. doi:10.1038/nsmb.1747
Dosztanyi Z, Meszaros B, Simon I (2009) ANCHOR: web server for predicting protein binding regions in disordered proteins. Bioinformatics 25(20):2745–2746. doi:10.1093/bioinformatics/btp518
Brzovic PS, Rajagopal P, Hoyt DW, King MC, Klevit RE (2001) Structure of a BRCA1–BARD1 heterodimeric RING–RING complex. Nat Struct Biol 8(10):833–837. doi:10.1038/nsb1001-833
Williams RS, Green R, Glover JN (2001) Crystal structure of the BRCT repeat region from the breast cancer-associated protein BRCA1. Nat Struct Biol 8(10):838–842. doi:10.1038/nsb1001-838
Acknowledgments
T.O, Q.W, B.L.S, D.C and D.A were supported by a Wellcome Trust Programme Grant (application No. 093167/Z/10/Z. D.B.A received a C. J. Martin Research Fellowship from the National Health and Medical Research Council of Australia (APP1072476), and S.R.E was supported by a Gates Cambridge Scholarship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liang, S., Esswein, S.R., Ochi, T. et al. Achieving selectivity in space and time with DNA double-strand-break response and repair: molecular stages and scaffolds come with strings attached. Struct Chem 28, 161–171 (2017). https://doi.org/10.1007/s11224-016-0841-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-016-0841-7