Abstract
Crystal structures of mouse thymidylate synthase (mTS) in complexes with (1) sulfate anion, (2) 2′-deoxyuridine 5′-monophosphate (dUMP) and (3) 5-fluoro-dUMP (FdUMP) and N 5,10-methylenetetrahydrofolate (meTHF) have been determined and deposited in Protein Data Bank under the accession codes 3IHI, 4E5O and 5FCT, respectively. The structures show a strong overall similarity to the corresponding structures of rat and human thymidylate synthases (rTS and hTS, respectively). Unlike with hTS, whose unliganded and liganded forms assume different conformations (“inactive” and “active,” respectively) in the loop 181–197, in each of the three mTS structures, the loop 175–191, homologous to hTS loop 181–197, populates the active conformer, with catalytic Cys 189 buried in the active site and directed toward C(6) of the pyrimidine ring of dUMP/FdUMP, pointing to protein’s inability to adopt the inactive conformation. The binary structures of either dUMP- or sulfate-bound mTS, showing the enzyme with open active site and extended C-terminus, differ from the structure of the mTS–5-FdUMP–meTHF ternary complex, with the active site closed and C-terminus folded inward, thus covering the active site cleft. Another difference pertains to the conformation of the Arg44 side chain in the active site-flanking loop 41–47, forming strong hydrogen bonds with the dUMP/FdUMP phosphate moiety in each of the two liganded mTS structures, but turning away from the active site entrance and loosing the possibility of H-bonding with sulfate in the sulfate-bound mTS structure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Thymidylate synthase (TS; EC 2.1.1.45), a target in chemotherapy, catalyzes the conversion of deoxyuridine monophosphate (dUMP) and R-N 5,10 -methylenetetrahydrofolate (meTHF) to deoxythymidine monophosphate and dihydrofolate, via reductive methylation, in which meTHF serves as both methylene group donor and reducing agent [1, 2]. As the sole de novo source of thymidylate synthesis in eukaryotic cells, for over 50 years, the enzyme has been a target in chemotherapy [3], with active forms of drugs being dUMP or N 5,10-meTHF analogues that inhibit the enzyme [4, 5]. Between dUMP analogues active in chemotherapy, the most prominent is 5-fluoro-dUMP (FdUMP), a strong thymidylate synthase inhibitor being the active form of drugs used in chemotherapy, such as 5-fluorouracil, 5-fluoro-2′-deoxyuridine and 5-fluorocytosine [2].
The first step of the reaction catalyzed by TS is the nucleophilic addition of the active site cysteine (residue 189 in mouse TS) to the pyrimidine C(6) of the nucleotide. Of essential role at this stage of catalysis are the proper binding position and orientation of dUMP. The first is secured by anchoring the phosphate of dUMP via several conservative hydrogen contacts from both TS subunits, while the latter by the way of hydrogen bonding between the conserved asparagine (Asn 220 in mTS) and the O4 and N3–H moieties of the pyrimidine ring. Properly bound dUMP provides the binding surface for the cofactor, resulting in the formation of the ternary complex TS–dUMP–meTHF [2]. Following activation of dUMP by the nucleophilic attack at the pyrimidine C(6), the negative charge of cysteine residue is delocalized, probably toward the C(4)=O of dUMP, and the corresponding enolate anion is formed (Fig. 1) with its C(5) position believed to become strongly nucleophilic. The latter facilitates a consecutive attack of the methylene residue, in the iminium ion form, resulting from opening of the imidazolidine ring of the cofactor. This leads to formation of the covalent intermediate, ternary complex of the enzyme with substrate and cofactor. Next, the C(5) hydrogen dissociates, as proton, from dUMP pyrimidine ring, with concomitant β-elimination of tetrahydrofolate, still remaining bound in the active center. The latter enables hydride transfer from the pteridine C(6), resulting in reduction of the pyrimidine C(5) methylene group to methyl one. The reaction is completed by regeneration of the pyrimidine 5,6 double bond, with concomitant elimination to form dTMP and enzyme [1, 2].
Structure determination of human TS complexes revealed a unique feature. In contrast to structures from other species, human TS loop 181–197 was observed to populate two conformers, one of them, apparently inactive, stabilized by a pair of hydrogen bonds from Arg163 to the carbonyls of Ala191 and Leu192. As mouse TS protein contains lysine residue at the position corresponding to arginine residue in human TS (Lys157 in mTS vs. Arg163 in hTS), it has been hypothesized unable to populate the inactive conformer [6]. In order to test that hypothesis, learning the crystal structure of the mouse enzyme was of interest.
An initial account of a part of the present results has been published in a special issue of Pteridines [7], covering materials of the 14th International Symposium on Pteridines and Folates (June 7–12, 2009, Jeju, Korea).
Materials and methods
Reagents
K/Na tartrate, PEG 3350, PEG 4000, Li2SO4, dUMP were purchased from Sigma, dithiothreitol (DTT) was from Carl Roth GmbH.
Recombinant mouse thymidylate synthase
This was overexpressed and purified as described previously [8, 9]. The enzyme was judged to be near homogeneous by denaturing polyacrylamide slab gel electrophoresis using a 40, 20 and 10 μg sample. Enzyme activity was measured spectrophotometrically [10], but at the last purification step, the assay measuring tritium release from [5-3H]dUMP was used as previously described [11]. The specific activity of purified enzyme was found to be 1.75 μmol min−1 mg −1 for conversion of dUMP to dTMP at 37 °C.
Separation of non-phosphorylated fraction of purified recombinant thymidylate synthase preparation
The enzyme preparation was separated into phosphorylated and non-phosphorylated fractions as previously described [12]. Only the non-phosphorylated fraction was used for crystallization.
Crystallization and data collection
Purified protein was dialyzed against 5 mM Tris HCl, pH 7.5 buffer containing 5 mM DTT and then concentrated using Amicon Centricon centrifugal filter. Crystals were grown by the vapor diffusion method in the hanging drops at 4 °C under following conditions. For TS, equal volumes of protein solution at 15 mg/ml concentration and well solution were mixed and allowed to equilibrate with 0.5 ml of well solution, containing 0.1 M Tris HCl, pH 8.5, 0.2 M Li2SO4 and 25 % (w/v) PEG 4000. For TS–dUMP complex, a drop of protein solution at 25 mg/ml concentration, containing 5.9 mM dUMP, was mixed as before 1:1 with well solution and allowed to equilibrate with 0.5 ml of well solution containing 0.2 M K Na Tartrate and 20 % PEG 3350. For TS–dUMP–meTHF complex, equal volumes of mTS solution (25 mg/ml), containing 5 mM FdUMP and 10 mM meTHF and well solution (0.1 M Tris HCl, pH 8.5, 0.1 M MgCl2 and 19.5 % (w/v) PEG 4000) were mixed and allowed to equilibrate with 0.5 ml of well solution. Crystallization conditions are summarized in Table 1.
The crystals (Figs. 3, 4) were transferred for a few seconds to cryoprotectant solution containing mother liquor and 25 % butanodiol and 30 % glicerol for mTS–dUMP and mTS, respectively, and flash-cooled in N2 vapors.
X-ray diffraction data were collected from three single flash-frozen crystals either at the Max-Lab Lund University Synchrotron (mTS and mTS–dUMP), using X-ray wavelengths of 1.038 and 0.908 Å, or at the BESSY Synchrotron (mTS–FdUMP–meTHF), using X-ray wavelength of 0.918 Å.
Data processing, structure determination and refinement
Data were processed with Denzo and Scalepack [13]. Both structures were determined by molecular replacement carried out with the CCP4 package [14], using the rat TS ternary complex without ligands as the search model. The crystal structures of mTS, mTS–dUMP and mTS–FdUMP–meTHF were determined at resolutions of 1.94, 1.70 and 1.55 Å, respectively. The correctness of the structures was evaluated using Sfcheck and Procheck from the CCP4 suite. Some X-ray data and model refinement parameters are summarized in Table 2.
Results and discussion
The structures, consisting of either one dimer (mTS and mTS–FdUMP–meTHF) or three dimers (mTS–dUMP) per an asymmetric part of the unit cell, showed an overall similarity to the corresponding structures of human and rat TSs (hTS and rTS, respectively). The Cα RMSD values between the liganded and unliganded mTS structures presented here, as well as between these structures and the crystal structures of other ternary complexes with dUMP and Tomudex, including those with hTS [15, 16], rTS [17] and mTS [18], are shown in Table 3. The superimposition of the mTS, mTS–dUMP and mTS–FdUMP–meTHF structures is shown in Fig. 2. The similarity of the corresponding TS structures is hardly unexpected in view of highly conserved primary structures of different TS proteins, as reflected by mTS amino acid sequence being 90 % identical with those of hTS [19] and rTS [20], and about 40–55 % identical with prokaryotic and phage TS sequences [19]. With respect to the latter, a recent comparison of amino acid sequences between TS of the parasitic nematode Trichinella spiralis and 33 other TS proteins, both eukaryotic and prokaryotic, showed 43–68 % identities and 56–79 % similarities [21].
The mTS–dUMP structure showed the enzyme in the active, open-state conformation. The dUMP pyrimidine ring C(6) and the catalytic Cys189 were separated by distances of 3.19, 3.22, 3.19, 3.17, 3.28 and 2.94 Å (subunits A, B, C, D, E and F, respectively), precluding covalent bondings. The substrate molecule was bound in a manner similar to that observed in the corresponding complexes of other mammalian and bacterial TSs, with the ligand anchored in the active site by several H-bonds to its phosphate moiety from four arginines from both subunits of the enzyme dimer (Arg44 and Arg209 from the first subunit and Arg169′ and Arg170′ from the other subunit) and single serine (Ser210). The orientation of dUMP was secured by H-bonds between the conserved Asn 220 and the O4 and N(3)-H moieties of the pyrimidine ring.
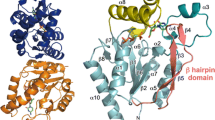
In the mTS–FdUMP–meTHF structure, the enzyme was found in the active, closed state conformation, with the catalytic Cys189 located in the active site and extended toward FdUMP. The distances between mTS Cys189 γS and FdUMP pyrimidine C(6) in the two subunits (A and B) were equal, each 1.82 Å, pointing to covalent bonding. However, the distances between the FdUMP pyrimidine C(5) and meTHF C(11) atoms were 2.28 Å (subunit A) and 1.55 Å (subunit B), indicating covalent bond between the two ligands to have been formed only in the subunit B. FdUMP was bound slightly shifted (~0.9 Å), compared to the position of dUMP in the mTS–dUMP structure, but preserved all the non-covalent interactions of the latter molecule. The conformation of the meTHF molecule was stabilized by favorable contacts with Ile102 and Asn106 from a long loop–helix–loop segment (100–127), flanking the entrance to the active site cleft, Ala305 and Met306 from C-terminus, Leu215 from a short loop 213–216 and Phe219 from central α-helix 217–234 (Fig. 3). It should be added that inhibition of thymidylate synthase by FdUMP involves time-dependent formation of a ternary covalently bound complex of the enzyme with FdUMP and mTHF, in a reaction similar to that involving dUMP (in fact FdUMP mimicks a substrate better than dUMP [22, 23]). However, at this step, the reaction stops, as the C(5) fluorine fails to dissociate (due to the strength of the C–F bond), as it happens with C(5) hydrogen in dUMP. What results is a slowly reversible enzyme inactivation. Studies on the mechanism of thymidylate synthase inhibition by FdUMP were a rich source of information about the catalytic mechanism [1, 2].
Non-bonded contacts stabilizing the conformation of the meTHF molecule in the subunit B of the mTS–FdUMP–meTHF complex structure. Two alternative conformations of the meTHF molecule are shown as ball and sticks, and other residues/moieties as sticks. Atom coloring is by element: gray for carbons, red for oxygens, blue for nitrogens, gold for sulfur and green for fluorine. Hydrogen atoms are omitted for clarity. The alternative conformers of meTHF and Lys 302 are colored cyan. The methylene bridge between FdUMP and meTHF is colored purple. The hydrophobic contacts and hydrogen bonds between meTHF and protein residues are colored shades of violet and green, respectively (Color figure online)
The active site in the mTS structure held a single sulfate anion, bound at nearly the same location as the phosphate moieties of dUMP and FdUMP in the mTS–dUMP and mTS–FdUMP–meTHF structures, respectively, and stabilized by H-bonds with the pair of arginine residues (Arg170′ and Arg209), belonging to the quartet coordinating the nucleotide phosphate in each of the liganded structures. The presence at the active site of the sulfate anion, an isosteric analogue of phosphate anion, is not surprising, as 0.1 M Li2SO4 was present in the crystallization medium.
The structures of sulfate-bound mTS, and of the dUMP–mTS and the mTS–FdUMP–meTHF complexes, differ in the open/closed state equilibrium, revealing either an open active site and extended C-terminus (mTS–dUMP and, to a lower degree due to a shorter C-terminus, mTS) or closed active site, with C-terminus folded inward and covering the active site entrance (mTS–FdUMP–meTHF) (Fig. 4). The latter is the most striking difference between the overall similar structures, as reflected in modest Cα RMSD values for dimers, as well as individual subunits (Table 3). Another, smaller difference pertains to the conformation of the loop 41–47, located at one of the regions flanking the active site cleft. In the liganded systems, this loop penetrates the active site, with the side chain of Arg44 extending toward, and forming H-bonds with the phosphate moiety of dUMP/FdUMP. In the absence of ligands (and despite the presence of sulfate, placed similarly as the phosphate moieties of dUMP/FdUMP), Arg44 turns away from the active site cleft and “uncovers” the active site entrance (Fig. 5).
The closed-conformation structure of the mTS–FdUMP–meTHF complex strongly resembles a mixed, yet basically closed conformation observed in the AB dimer of the previously reported mTS–dUMP-Tomudex (4EB4) crystal structure (cf. RMSD in Table 3). However, although still moderate, the RMS differences with respect to the more open CD dimer, and in particular to subunit C of the 4EB4 structure, are distinctly more pronounced. Those differences are related to the open/closed active site equilibrium and include the positions of C-terminus (RMSD of 2.8–5.5 Å) and two segments on the opposite flanks of the active site entrance, loop 41–47 (RMSD of 2.4–3.3 Å) and loop-helix-loop segment 100–127 (RMSD of 2.0–3.5 Å) (Fig. 6).
Superimposition of subunit A from the mTS–FdUMP–meTHF structure (red) and subunit C from the mTS–dUMP-Tomudex structure (yellow). Most significant differences can be observed in the conformations of C-terminus as well as two AS-flanking regions: 41–47 and 100–127. Proteins are shown as solid ribbons and ligands as sticks (Color figure online)
In human thymidylate synthase, loop 181–197 can populate two major conformations, active and inactive, related to each other by a 180° rotation [24, 25]. While catalytic Cys 195 in the former conformation is present in the active site, directed toward dUMP pyrimidine ring C(6) and forming with it a thiol adduct, in the latter it is located more than 10 Å away and directed toward the enzyme dimer interface (Fig. 2). The equilibrium between both conformations has been shown to be shifted toward the inactive conformation in the presence of phosphate or sulfate ions, with the active conformation promoted in the presence of dUMP [25].
In the mTS–dUMP and mTS–FdUMP–meTHF structures, loop 175–191, equivalent to human loop 181–197, populates the active conformation, with catalytic Cys189 either located at a close, but non-covalent distance from dUMP, or covalently bound to FdUMP. Interestingly, in the mTS structure, loop 175–191 populates the same (active) conformation, despite the absence of dUMP/FdUMP in the active site, containing only bound sulfate anion. This is in contrast to the sulfate-bound human TS structure (1HW4; Phan et al., 2001b), where corresponding loop 181–197 occupies the inactive conformation (Fig. 7).
Superimposition of the sulfate-bound structures of mTS (3IHI; green) and hTS (1HW4; red). Shown are fragments containing loops 175–191 from mTS (lime), with catalytic Cys189 inside of the active site (active conformation), and 181–197 from hTS (pink), with catalytic Cys195 outside of the active site (inactive conformation). Cys195 (hTS) and Cys189 (mTS), as well as the sulfate anions from both structures, are shown as sticks, and the rest of structures as solid ribbon (Color figure online)
In the human TS, the inactive conformer is stabilized by a pair of hydrogen bonds from Arg163 to the carbonyls of Ala191 and Leu192 (Fig. 3). The presence of lysine, instead of arginine, at the position 157 in mTS (equivalent to hTS 163) disallows the occurrence of the corresponding hydrogen bonding (with Ala185 and Leu186) due to shorter side chain and lower adaptability to multiple H-bonding of the former, compared to the latter residue, as concluded from the comparison of the mTS and hTS crystal structures (Fig. 8). This observation supports the above mentioned hypothesis by Lovelace et al. [6].
Superimposition of the sulfate-bound structure of mTS (3IHI; green) and the structure of hTS in a complex with non-peptidic inhibitor (4E28; red). Cme denotes a modified cysteine residue, SS-(2-hydroxyethyl)thiocysteine. Inactive conformation of human loop 181–197 is stabilized by hydrogen bonds between Ala191 and Leu192, and Arg163′ from the other subunit. Mouse loop 175–191 is in the active conformation, related to the inactive conformer by a 180° rotation. Lys157′ from the other subunit of mTS is unable to form hydrogen bonds equivalent to those in the human enzyme, i.e., with Ala185 and Leu186 (Color figure online)
In view of the foregoing conclusion of obvious interest is potential functional consequence of this mechanism, as well as possibility to find structural features enabling the inactive conformation stabilization (arginine residue in a position homologous to hTS 163) in other TS structures. Considering strong influences of posttranslational modifications on kinetics and inhibition of TS-catalyzed reaction [9, 26, 27] and resulting wide variability of properties of the enzyme preparations of the same specific origin [2, 9, 27], a simple comparison of mTS and hTS properties would not answer the former question. However, Gibson et al. [28] obtained a mutant R163K of the human enzyme and found its catalytic activity to be higher than that of the parental hTS. Furthermore, the native mutant protein crystallized in the active conformation. Thus, hTS Arg163 appears to result in certain conformational plasticity, associated with somewhat different catalytic properties. Based on analysis of so far determined mammalian TS protein sequences, indicating arginine to be present at position homologous to hTS Arg163 in all primates, except of prosimians, Luo et al. [29] suggested this enzyme form to evolve from a common ancestral mammalian form (with homologous Lys) following the divergence of prosimians and simians. In accord, recently published mammalian TS sequences show Lys at position homologous to hTS Arg163 (NCBI Accessions EHB12689, ELK31636, EPY81457 and EGW14385), with one exception of marmoset (NCBI Accession JAB35844), a representative of New World Primates, showing corresponding arginine. The letter confirms that this mechanism of local conformational plasticity may be unique for primate TSs, at least between mammalian enzyme forms. It should be mentioned, though, that it may be not unique between other TSs, as at least the enzyme sequences corresponding to both bacterium Bacillus subtilis b and fungus Cryptococcus neoformans show arginine residue in a position homologous to hTS 163 [1].
References
Carreras CW, Santi DV (1995) The catalytic mechanism and structure of thymidylate synthase. Annu Rev Biochem 64:721–762
Rode W, Leś A (1996) Molecular mechanism of thymidylate synthase-catalyzed reaction and interaction of the enzyme with 2- and/or 4-substituted analogues of dUMP and 5-fluoro-dUMP. Acta Biochim Pol 43:133–142
Berger FG, Berger SH (2006) Thymidylate synthase as a chemotherapeutic drug target: where are we after fifty years? Cancer Biol Therapy 5:1238–1241
Costi MP, Ferrari S, Venturelli A, Calò S, Tondi D, Barlocco D (2005) Thymidylate synthase structure, function and implication in drug discovery. Curr Med Chem 12:2241–2258
Jarmuła A (2010) Antifolate inhibitors of thymidylate synthase as anticancer drugs. Mini-Rev Med Chem 10:1211–1222
Lovelace LL, Gibson LM, Lebioda L (2007) Cooperative inhibition of human thymidylate synthase by mixtures of active site binding and allosteric inhibitors. Biochemistry 46:2823–2830
Dowierciał A, Jarmuła A, Rypniewski W, Sokołowska M, Frączyk T, Cieśla J, Rode W (2009) Crystal structures of substrate- and sulfate-bound mouse thymidylate synthase. Pteridines 20(Special Issue):163–167
Cieśla J, Golos B, Walajtys-Rode E, Jagielska E, Plucienniczak A, Rode W (2002) The effect of Arg209 to Lys mutation in mouse thymidylate synthase. Acta Biochim Pol 49:651–658
Ludwiczak J, Maj P, Wilk P, Frączyk T, Ruman T, Kierdaszuk B, Jarmuła A, Rode W (2016) Phosphorylation of thymidylate synthase affects slow-binding inhibition by 5-fluoro-dUMP and N4-hydroxy-dCMP. Mol BioSyst 12:1333–1341
Wahba A, Friedkin M (1962) The enzymatic synthesis of thymidylate. J Biol Chem 237:3794–3801
Rode W, Kulikowski T, Kedzierska B, Jastreboff M, Shugar D (1984) Inhibition of mammalian tumour thymidylate synthetase by 5-alkylated 2′-deoxyuridine 5′-phosphates. Biochem Pharmacol 33:2699–2705
Frączyk T, Ruman T, Wilk P, Palmowski P, Rogowska-Wrzesinska A, Cieśla J, Zieliński Z, Nizioł J, Jarmuła A, Maj P, Gołos B, Wińska P, Ostafil S, Wałajtys-Rode E, Shugar D, Rode W (2015) Properties of phosphorylated thymidylate synthase. Biochim Biophys Acta Proteins Proteom 1854:1922–1934
Otwinowski Z, Minor W (1997) Processing of X-ray diffraction data collected in oscillation mode. In: Carter CW, Jr., Sweet RM (eds) Methods in enzymology: macromolecular crystallography part A, vol 276. Academic Press, San Diego, pp 307–326
The CCP4 suite: programs for protein crystallography (1994) Collaborative computational project, number 4. Acta Crystallogr D Biol Crystallogr 50:760–763
Phan J, Koli S, Minor W, Dunlap RB, Berger SH, Lebioda L (2001) Human thymidylate synthase is in the closed conformation when complexed with dUMP and Raltitrexed, an antifolate drug. Biochemistry 40:1897–1902
Almog R, Waddling CA, Maley F, Maley G, Van Roey P (2001) Crystal structure of a deletion mutant of human thymidylate synthase Δ(7–29) and its ternary complex with Tomudex and dUMP. Prot Sci 10:988–996
Sotelo-Mundo RR, Ciesla J, Dzik JM, Rode W, Maley F, Maley GF, Hardy LW, Montfort WR (1998) Crystal structures of rat thymidylate synthase inhibited by Tomudex, a potent anticancer drug. Biochemistry 38:1087–1094
Dowierciał A,Wilk P, Rypniewski W, Rode W, Jarmuła A (2014) Crystal structure of mouse thymidylate synthase in tertiary complex with dUMP and raltitrexed reveals N-terminus architecture and two different active site conformations. BioMed Res Int, Article ID 945803, pp 1–7
Perryman SM, Rossana C, Deng TL, Vanin EF, Johnson LF (1986) Sequence of a cDNA for mouse thymidylate synthase reveals striking similarity with the prokaryotic enzyme. Mol Biol Evol 3:313–321
Cieśla J, Weiner KX, Weiner RS, Reston JT, Maley GF, Maley F (1995) Isolation and expression of rat thymidylate synthase cDNA: phylogenetic comparison with human and mouse thymidylate synthases. Biochim Biophys Acta 1261:233–242
Dąbrowska M, Jagielska E, Cieśla J, Płucienniczak A, Kwiatowski J, Wranicz M, Boireau P, Rode W (2004) Trichinella spiralis thymidylate synthase: cDNA cloning and sequencing, and developmental pattern of mRNA expression. Parasitology 128:209–221
Jarmuła A, Anulewicz R, Leś A, Cyrański MK, Adamowicz L, Bretner M, Felczak K, Kulikowski T, Krygowski TM, Rode W (1998) Crystal structures of 5-fluoro-dUrd and its 2 and/or 4-thio analogues: models of substituted dUMP pyrimidine ring interacting with thymidylate synthase. Biochim Biophys Acta 1382:277–286
Jarmuła A, Cyrański MK, Leś A, Krygowski TM, Rode W (1998) Interaction of thymidylate synthase with 5-fluoro-substituted dUMP analogues in view of the pyrimidine ring structure. Pol J Chem 72:1958–1962
Luo B, Repalli J, Yousef A-M, Johnson SR, Lebioda L, Berger SH (2011) Human thymidylate synthase with loop 181–197 stabilized in an inactive conformation: ligand interactions, phosphorylation, and inhibition profiles. Protein Sci 20:87–94
Phan J, Steadman DJ, Koli S, Ding WC, Minor W, Dunlap RB, Berger SH, Lebioda L (2001) Structure of human thymidylate synthase suggests advantages of chemotherapy with noncompetitive inhibitors. J Biol Chem 276:14170–14177
Cieśla J, Frączyk T, Zieliński Z, Sikora J, Rode W (2006) Altered mouse leukemia L1210 thymidylate synthase, associated with cell resistance to 5-fluoro-dUrd, is not mutated but rather reflects posttranslational modification. Acta Biochim Pol 53:189–198
Frączyk T, Ruman T, Wilk P, Palmowski P, Rogowska-Wrzesinska A, Cieśla J, Zieliński Z, Nizioł J, Jarmuła A, Maj P, Gołos B, Wińska P, Ostafil S, Wałajtys-Rode E, Shugar D, Rode W (2015) Properties of phosphorylated thymidylate synthase. Biochim Biophys Acta: Proteins Proteom 1854:1922–1934
Gibson LM, Lovelace LL, Lebioda L (2008) The R163 K mutant of human thymidylate synthase is stabilized in an active conformation: structural asymmetry and reactivity of cysteine 195. Biochemistry 47:4636–4643
Luo B, Johnson SR, Lebioda L, Berger SH (2011) Evolution of metamorphism in thymidylate synthases within the primate lineages. J Mol Evol 72:306–314
Acknowledgments
Supported by the Ministry of Science and Higher Education (Grant No. N301 3948 33) and National Science Centre (Grant No. 2011/01/B/NZ6/01781).
Author information
Authors and Affiliations
Corresponding author
Additional information
Anna Dowierciał and Adam Jarmuła have contributed equally to the present work.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Dowierciał, A., Jarmuła, A., Wilk, P. et al. Mouse thymidylate synthase does not show the inactive conformation, observed for the human enzyme. Struct Chem 28, 667–674 (2017). https://doi.org/10.1007/s11224-016-0840-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-016-0840-8