Abstract
Learning the structure of Bayesian networks from data is known to be a computationally challenging, NP-hard problem. The literature has long investigated how to perform structure learning from data containing large numbers of variables, following a general interest in high-dimensional applications (“small n, large p”) in systems biology and genetics. More recently, data sets with large numbers of observations (the so-called “big data”) have become increasingly common; and these data sets are not necessarily high-dimensional, sometimes having only a few tens of variables depending on the application. We revisit the computational complexity of Bayesian network structure learning in this setting, showing that the common choice of measuring it with the number of estimated local distributions leads to unrealistic time complexity estimates for the most common class of score-based algorithms, greedy search. We then derive more accurate expressions under common distributional assumptions. These expressions suggest that the speed of Bayesian network learning can be improved by taking advantage of the availability of closed-form estimators for local distributions with few parents. Furthermore, we find that using predictive instead of in-sample goodness-of-fit scores improves speed; and we confirm that it improves the accuracy of network reconstruction as well, as previously observed by Chickering and Heckerman (Stat Comput 10: 55–62, 2000). We demonstrate these results on large real-world environmental and epidemiological data; and on reference data sets available from public repositories.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Bayesian networks (BNs; Pearl 1988) are a class of graphical models defined over a set of random variables \(\mathbf {X}= \{X_1, \ldots , X_N\}\), each describing some quantity of interest, that are associated with the nodes of a directed acyclic graph (DAG) \(\mathcal {G}\). (They are often referred to interchangeably.) Arcs in \(\mathcal {G}\) express direct dependence relationships between the variables in \(\mathbf {X}\), with graphical separation in \(\mathcal {G}\) implying conditional independence in probability. As a result, \(\mathcal {G}\) induces the factorisation
in which the joint probability distribution of \(\mathbf {X}\) (with parameters \(\varTheta \)) decomposes in one local distribution for each \(X_i\) (with parameters \(\varTheta _{X_i}\), \(\bigcup _{\mathbf {X}} \varTheta _{X_i}= \varTheta \)) conditional on its parents \(\varPi _{X_i}\).
While in principle there are many possible choices for the distribution of \(\mathbf {X}\), the literature has focused mostly on three cases. Discrete BNs (Heckerman et al. 1995) assume that both \(\mathbf {X}\) and the \(X_i\) are multinomial random variables. Local distributions take the form
their parameters are the conditional probabilities of \(X_i\) given each configuration of the values of its parents, usually represented as a conditional probability table for each \(X_i\). Gaussian BNs (GBNs; Geiger and Heckerman 1994) model \(\mathbf {X}\) with a multivariate normal random variable and assume that the \(X_i\) are univariate normals linked by linear dependencies. The parameters of the local distributions can be equivalently written (Weatherburn 1961) as the partial Pearson correlations \(\rho _{X_i, X_j {\text {|}}\varPi _{X_i}\setminus X_j}\) between \(X_i\) and each parent \(X_j\) given the other parents; or as the coefficients \(\varvec{\beta }_{X_i}\) of the linear regression model
so that \(X_i {\text {|}}\varPi _{X_i}\sim N(\mu _{X_i} + \varPi _{X_i}\varvec{\beta }_{X_i}, \sigma ^2_{X_i})\). Finally, conditional linear Gaussian BNs (CLGBNs; Lauritzen and Wermuth 1989) combine discrete and continuous random variables in a mixture model:
-
discrete \(X_i\) are only allowed to have discrete parents (denoted \(\varDelta _{X_i}\)), are assumed to follow a multinomial distribution parameterised with conditional probability tables;
-
continuous \(X_i\) are allowed to have both discrete and continuous parents (denoted \(\varGamma _{X_i}\), \(\varDelta _{X_i}\cup \varGamma _{X_i}= \varPi _{X_i}\)), and their local distributions are
$$\begin{aligned} X_i {\text {|}}\varPi _{X_i}\sim N \left( \mu _{X_i, \delta _{X_i}} + \varGamma _{X_i}\varvec{\beta }_{X_i, \delta _{X_i}}, \sigma ^2_{X_i, \delta _{X_i}}\right) \end{aligned}$$which can be written as a mixture of linear regressions
$$\begin{aligned} X_i= & {} \mu _{X_i, \delta _{X_i}} + \varGamma _{X_i}\varvec{\beta }_{X_i, \delta _{X_i}} + \varepsilon _{X_i, \delta _{X_i}}, \\&\varepsilon _{X_i, \delta _{X_i}} \sim N \left( 0, \sigma ^2_{X_i, \delta _{X_i}}\right) , \end{aligned}$$against the continuous parents with one component for each configuration \(\delta _{X_i}\in \mathrm {Val}(\varDelta _{X_i})\) of the discrete parents. If \(X_i\) has no discrete parents, the mixture reverts to a single linear regression.
Other distributional assumptions, such as mixtures of truncated exponentials (Moral et al. 2001) or copulas (Elidan 2010), have been proposed in the literature but have seen less widespread adoption due to the lack of exact conditional inference and simple closed-form estimators.
The task of learning a BN from a data set \(\mathcal {D}\) containing n observations is performed in two steps:
Structure learning consists in finding the DAG \(\mathcal {G}\) that encodes the dependence structure of the data, thus maximising \({\text {P}}(\mathcal {G}{\text {|}}\mathcal {D})\) or some alternative goodness-of-fit measure; parameter learning consists in estimating the parameters \(\varTheta \) given the \(\mathcal {G}\) obtained from structure learning. If we assume parameters in different local distributions are independent (Heckerman et al. 1995), we can perform parameter learning independently for each node following (1) because
Furthermore, if \(\mathcal {G}\) is sufficiently sparse each node will have a small number of parents; and \(X_i {\text {|}}\varPi _{X_i}\) will have a low-dimensional parameter space, making parameter learning computationally efficient.
On the other hand, structure learning is well known to be both NP-hard (Chickering and Heckerman 1994) and NP-complete (Chickering 1996), even under unrealistically favourable conditions such as the availability of an independence and inference oracle (Chickering et al. 2004).Footnote 1 This is despite the fact that if we take
again following (1) we can decompose the marginal likelihood \({\text {P}}(\mathcal {D}{\text {|}}\mathcal {G})\) into one component for each local distribution
and despite the fact that each component can be written in closed form for discrete BNs (Heckerman et al. 1995), GBNs (Geiger and Heckerman 1994) and CLGBNs (Bøttcher 2001). The same is true if we replace \({\text {P}}(\mathcal {D}{\text {|}}\mathcal {G})\) with frequentist goodness-of-fit scores such as BIC (Schwarz 1978), which is commonly used in structure learning because of its simple expression:
Compared to marginal likelihoods, BIC has the advantage that it does not depend on any hyperparameter, while converging to \(\log {\text {P}}(\mathcal {D}{\text {|}}\mathcal {G})\) as \(n \rightarrow \infty \).
These score functions, which we will denote with \(\mathrm {Score}(\mathcal {G}, \mathcal {D})\) in the following, have two important properties:
-
they decompose into one component for each local distribution following (1), say
$$\begin{aligned} \mathrm {Score}(\mathcal {G}, \mathcal {D}) = \sum _{i = 1}^{N}\mathrm {Score}(X_i, \varPi _{X_i}, \mathcal {D}), \end{aligned}$$thus allowing local computations (decomposability);
-
they assign the same score value to DAGs that encode the same probability distributions and can therefore be grouped in an equivalence classes (score equivalence; Chickering 1995).Footnote 2
Structure learning via score maximisation is performed using general-purpose optimisation techniques, typically heuristics, adapted to take advantage of these properties to increase the speed of structure learning. The most common are greedy search strategies that employ local moves designed to affect only few local distributions, to that new candidate DAGs can be scored without recomputing the full \({\text {P}}(\mathcal {D}{\text {|}}\mathcal {G})\). This can be done either in the space of the DAGs with hill climbing and tabu search (Russell and Norvig 2009), or in the space of the equivalence classes with Greedy Equivalent Search (GES; Chickering 2002). Other options that have been explored in the literature are genetic algorithms (Larranaga et al. 1996) and ant colony optimisation (Campos et al. 2002). Exact maximisation of \({\text {P}}(\mathcal {D}{\text {|}}\mathcal {G})\) and BIC has also become feasible for small data sets in recent years thanks to increasingly efficient pruning of the space of the DAGs and tight bounds on the scores (Cussens 2012; Suzuki 2017; Scanagatta et al. 2015).
In addition, we note that it is also possible to perform structure learning using conditional independence tests to learn conditional independence constraints from \(\mathcal {D}\), and thus identify which arcs should be included in \(\mathcal {G}\). The resulting algorithms are called constraint-based algorithms, as opposed to the score-based algorithms we introduced above; for an overview and a comparison of these two approaches see Scutari and Denis (2014). Chickering et al. (2004) proved that constraint-based algorithms are also NP-hard for unrestricted DAGs; and they are in fact equivalent to score-based algorithms given a fixed topological ordering when independence constraints are tested with statistical tests related to cross-entropy (Cowell 2001). For these reasons, in this paper we will focus only on score-based algorithms while recognising that a similar investigation of constraint-based algorithms represents a promising direction for future research.
The contributions of this paper are:
-
1.
to provide general expressions for the (time) computational complexity of the most common class of score-based structure learning algorithms, greedy search, as a function of the number of variables N, of the sample size n, and of the number of parameters \(|\varTheta |\);
-
2.
to use these expressions to identify two simple yet effective optimisations to speed up structure learning in “big data” settings in which \(n \gg N\).
Both are increasingly important when using BNs in modern machine learning applications, as data sets with large numbers of observations (the so-called “big data”) are becoming as common as classic high-dimensional data (“small n, large p”, or “small n, large N” using the notation introduced above). The vast majority of complexity and scalability results (Kalisch and Bühlmann 2007; Scanagatta et al. 2015) and computational optimisations (Scutari 2017) in the literature are derived in the latter setting and implicitly assume \(n \ll N\); they are not relevant in the former setting in which \(n \gg N\). Our contributions also complement related work on advanced data structures for machine learning applications, which include ADtrees (Moore and Lee 1998), frequent sets (Goldenberg and Moore 2004) and more recently bitmap representations combined with radix sort (Karan et al. 2018). Such literature develops a framework for caching sufficient statistics, but concentrates on discrete variables, whereas we work in a more general setting in which data can include both discrete and continuous variables.
The material is organised as follows. In Sect. 2, we will present in detail how greedy search can be efficiently implemented thanks to the factorisation in (1), and we will derive its computational complexity as a function N; this result has been mentioned in many places in the literature, but to the best of our knowledge its derivation has not been described in depth. In Sect. 3, we will then argue that the resulting expression does not reflect the actual computational complexity of structure learning, particularly in a “big data” setting where \(n \gg N\); and we will re-derive it in terms of n and \(|\varTheta |\) for the three classes of BNs described above. In Sect. 4, we will use this new expression to identify two optimisations that can markedly improve the overall speed of learning GBNs and CLGBNs by leveraging the availability of closed-form estimates for the parameters of the local distributions and out-of-sample goodness-of-fit scores. Finally, in Sect. 5 we will demonstrate the improvements in speed produced by the proposed optimisations on simulated and real-world data, as well as their effects on the accuracy of learned structures.
2 Computational complexity of greedy search

A state-of-the-art implementation of greedy search in the context of BN structure learning is shown in Algorithm 1. It consists of an initialisation phase (steps 1 and 2) followed by a hill climbing search (step 3), which is then optionally refined with tabu search (step 4) and random restarts (step 5). Minor variations of this algorithm have been used in large parts of the literature on BN structure learning with score-based methods (some notable examples are Heckerman et al. 1995; Tsamardinos et al. 2006; Friedman 1997).
Hill climbing uses local moves (arc additions, deletions and reversals) to explore the neighbourhood of the current candidate DAG \(\mathcal {G}_{ max }\) in the space of all possible DAGs in order to find the DAG \(\mathcal {G}\) (if any) that increases the score \(\mathrm {Score}(\mathcal {G}, \mathcal {D})\) the most over \(\mathcal {G}_{ max }\). That is, in each iteration hill climbing tries to delete and reverse each arc in the current optimal DAG \(\mathcal {G}_{ max }\); and to add each possible arc that is not already present in \(\mathcal {G}_{ max }\). For all the resulting DAGs \(\mathcal {G}^*\) that are acyclic, hill climbing then computes \(S_{\mathcal {G}^*} = \mathrm {Score}(\mathcal {G}^*, \mathcal {D})\); cyclic graphs are discarded. The \(\mathcal {G}^*\) with the highest \(S_{\mathcal {G}^*}\) becomes the new candidate DAG \(\mathcal {G}\). If that DAG has a score \(S_{\mathcal {G}} > S_{ max }\) then \(\mathcal {G}\) becomes the new \(\mathcal {G}_{ max }\), \(S_{ max }\) will be set to \(S_{\mathcal {G}}\), and hill climbing will move to the next iteration.
This greedy search eventually leads to a DAG \(\mathcal {G}_{ max }\) that has no neighbour with a higher score. Since hill climbing is an optimisation heuristic, there is no theoretical guarantee that \(\mathcal {G}_{ max }\) is a global maximum. In fact, the space of the DAGs grows super-exponentially in N (Harary and Palmer 1973); hence, multiple local maxima are likely present even if the sample size n is large. The problem may be compounded by the existence of score-equivalent DAGs, which by definition have the same \(S_{\mathcal {G}}\) for all the \(\mathcal {G}\) falling in the same equivalence class. However, Gillispie and Perlman (2002) have shown that while the number of equivalence classes is of the same order of magnitude as the space of the DAGs, most contain few DAGs and as many as \(27.4\%\) contain just a single DAG. This suggests that the impact of score equivalence on hill climbing may be limited. Furthermore, greedy search can be easily modified into GES to work directly in the space of equivalence classes by using different set of local moves, side-stepping this possible issue entirely.
In order to escape from local maxima, greedy search first tries to move away from \(\mathcal {G}_{ max }\) by allowing up to \(t_0\) additional local moves. These moves necessarily produce DAGs \(\mathcal {G}^*\) with \(S_{\mathcal {G}*} \leqslant S_{ max }\); hence, the new candidate DAGs are chosen to have the highest \(S_\mathcal {G}\) even if \(S_\mathcal {G}< S_{ max }\). Furthermore, DAGs that have been accepted as candidates in the last \(t_1\) iterations are kept in a list (the tabu list) and are not considered again in order to guide the search towards unexplored regions of the space of the DAGs. This approach is called tabu search (step 4) and was originally proposed by Glover and Laguna (1998). If a new DAG with a score larger than \(\mathcal {G}_{ max }\) is found in the process, that DAG is taken as the new \(\mathcal {G}_{ max }\) and greedy search returns to step 3, reverting to hill climbing.
If, on the other hand, no such DAG is found then greedy search tries again to escape the local maximum \(\mathcal {G}_{ max }\) for \(r_0\) times with random non-local moves, that is, by moving to a distant location in the space of the DAGs and starting the greedy search again; hence, the name random restart (step 5). The non-local moves are typically determined by applying a batch of \(r_1\) randomly chosen local moves that substantially alter \(\mathcal {G}_{ max }\). If the DAG that was perturbed was indeed the global maximum, the assumption is that this second search will also identify it as the optimal DAG, in which case the algorithm terminates.
We will first study the (time) computational complexity of greedy search under the assumptions that are commonly used in the literature (see, for instance, Tsamardinos et al. 2006; Spirtes et al. 2001) for this purpose:
-
1.
We treat the estimation of each local distribution as an atomic O(1) operation; that is, the (time) complexity of structure learning is measured by the number of estimated local distributions.
-
2.
Model comparisons are assumed to always add, delete and reverse arcs correctly with respect to the underlying true model which happens asymptotically for \(n \rightarrow \infty \) since marginal likelihoods and BIC are globally and locally consistent (Chickering 2002).
-
2.
The true DAG \(\mathcal {G}_{\mathrm{REF}}\) is sparse and contains O(cN) arcs, where c is typically assumed to be between 1 and 5.
In steps 1 and 2, greedy search computes all the N local distributions for \(\mathcal {G}_0\). In step 3, each iteration tries all possible arc additions, deletions and reversals. Since there are \({N \atopwithdelims ()2}\) possible arcs in a DAG with N nodes, this requires \(O(N^2)\) model comparisons. If we assume \(\mathcal {G}_0\) is the empty DAG (that is, a DAG with no arcs), hill climbing will gradually add all the arcs in \(\mathcal {G}_{\mathrm{REF}}\), one in each iteration. Assuming \(\mathcal {G}_{\mathrm{REF}}\) is sparse, and assuming that arcs are removed or reversed a negligible number of times, the overall computational complexity of hill climbing is then \(O(cN^3)\) model comparisons. Step 4 performs \(t_0\) more iterations and is therefore \(O(t_0 N^2)\). Therefore, the combined time complexity of steps 3 and 4 is \(O(cN^3 + t_0 N^2)\). Each of the random restarts involves changing \(r_1\) arcs, and thus we can expect that it will take \(r_1\) iterations of hill climbing to go back to the same maximum, followed by tabu search; and that happens for \(r_0\) times. Overall, this adds \(O(r_0 (r_1N^2 + t_0 N^2))\) to the time complexity, resulting in an overall complexity g(N) of
The leading term is \(O(cN^3)\) for some small constant c, making greedy search cubic in complexity.
Fortunately, the factorisation in (1) makes it possible to recompute only one or two local distributions for each model comparison:
-
Adding or removing an arc only alters one parent set; for instance, adding \(X_j \rightarrow X_i\) means that \(\varPi _{X_i}= \varPi _{X_i}\cup X_j\), and therefore \({\text {P}}(X_i {\text {|}}\varPi _{X_i})\) should be updated to \({\text {P}}(X_i {\text {|}}\varPi _{X_i}\cup X_j)\). All the other local distributions \({\text {P}}(X_k {\text {|}}\varPi _{X_j}), X_k \ne X_i\) are unchanged.
-
Reversing an arc \(X_j \rightarrow X_i\) to \(X_i \rightarrow X_j\) means that \(\varPi _{X_i}= \varPi _{X_i}\setminus X_j\) and \(\varPi _{X_j} = \varPi _{X_j} \cup X_i\), and so both \({\text {P}}(X_i {\text {|}}\varPi _{X_i})\) and \({\text {P}}(X_j {\text {|}}\varPi _{X_j})\) should be updated.
Hence, it is possible to dramatically reduce the computational complexity of greedy search by keeping a cache of the score values of the N local distributions for the current \(\mathcal {G}_{ max }\)
and of the \(N^2 - N\) score differences
where \(\varPi _{X_i}^{ max }\) and \(\varPi _{X_i}^{\mathcal {G}^*}\) are the parents of \(X_i\) in \(\mathcal {G}_{ max }\) and in the \(\mathcal {G}^*\) obtained by removing (if present) or adding (if not) \(X_j \rightarrow X_i\) to \(\mathcal {G}_{ max }\). Only N (for arc additions and deletions) or 2N (for arc reversals) elements of \(\varDelta \) need to be actually computed in each iteration; those corresponding to the variable(s) whose parent sets were changed by the local move produced the current \(\mathcal {G}_{ max }\) in the previous iteration. After that, all possible arc additions, deletions and reversals can be evaluated without any further computational cost by adding or subtracting the appropriate \(\varDelta _{ij}\) from the \(B_i\). Arc reversals can be handled as a combination of arc removals and additions (e.g. reversing \(X_i \rightarrow X_j\) is equivalent to removing \(X_i \rightarrow X_j\) and adding \(X_j \rightarrow X_i\)). As a result, the overall computational complexity of greedy search reduces from \(O(cN^3)\) to \(O(cN^2)\). Finally, we briefly note that score equivalence may allow further computational saving because many local moves will produce new \(\mathcal {G}^*\) that are in the same equivalence class as \(\mathcal {G}_{ max }\); and for those moves necessarily \(\varDelta _{ij} = 0\) (for arc reversals) or \(\varDelta _{ij} = \varDelta _{ji}\) (for adding or removing \(X_i \rightarrow X_j\) and \(X_j \rightarrow X_i\)).
3 Revisiting computational complexity
In practice, the computational complexity of estimating a local distribution \({\text {P}}(X_i {\text {|}}\varPi _{X_i})\) from data depends on three of factors:
-
the characteristics of the data themselves (the sample size n, the number of possible values for categorical variables);
-
the number of parents of \(X_i\) in the DAG, that is, \(|\varPi _{X_i}|\);
-
the distributional assumptions on \({\text {P}}(X_i {\text {|}}\varPi _{X_i})\), which determine the number of parameters \(|\varTheta _{X_i}|\).
3.1 Computational complexity for local distributions
If n is large, or if \(|\varTheta _{X_i}|\) is markedly different for different \(X_i\), different local distributions will take different times to learn, violating the O(1) assumption from the previous section. In other words, if we denote the computational complexity of learning the local distribution of \(X_i\) as \(O(f_{\varPi _{X_i}}(X_i))\), we find below that \(O(f_{\varPi _{X_i}}(X_i)) \ne O(1)\).
3.1.1 Nodes in discrete BNs
In the case of discrete BNs, the conditional probabilities \(\pi _{ik {\text {|}}j}\) associated with each \(X_i {\text {|}}\varPi _{X_i}\) are computed from the corresponding counts \(n_{ijk}\) tallied from \(\{X_i, \varPi _{X_i}\}\); hence, estimating them takes \(O(n(1 + |\varPi _{X_i}|))\) time. Computing the marginals counts for each configuration of \(\varPi _{X_i}\) then takes \(O(|\varTheta _{X_i}|)\) time; assuming that each discrete variable takes at most l values, then \(|\varTheta _{X_i}| \leqslant l^{1 + |\varPi _{X_i}|}\) leading to
3.1.2 Nodes in GBNs
In the case of GBNs, the regressions coefficients for \(X_i {\text {|}}\varPi _{X_i}\) are usually computed by applying a QR decomposition to the augmented data matrix \([1 \, \varPi _{X_i}]\):
which can be solved efficiently by backward substitution since \(\mathbf {R}\) is upper-triangular. This approach is the de facto standard approach for fitting linear regression models because it is numerically stable even in the presence of correlated \(\varPi _{X_i}\) (see Seber 2008, for details). Afterwards, we can compute the fitted values \({\hat{x}}_i = \varPi _{X_i}\hat{\varvec{\beta }}_{X_i}\) and the residuals \(X_i - {\hat{x}}_i\) to estimate \({\hat{\sigma }}^2_{X_i} \propto (X_i - {\hat{x}}_i)^T (X_i - {\hat{x}}_i)\). The overall computational complexity is
with leading term \(O((n + 1)(1 + |\varPi _{X_i}|)^2)\).
3.1.3 Nodes in CLGBNs
As for CLGBNs, the local distributions of discrete nodes are estimated in the same way as they would be in a discrete BN. For Gaussian nodes, a regression of \(X_i\) against the continuous parents \(\varGamma _{X_i}\) is fitted from the \(n_{\delta _{X_i}}\) observations corresponding to each configuration of the discrete parents \(\varDelta _{X_i}\). Hence, the overall computational complexity is
with leading term \(O\left( (n + l^{|\varDelta _{X_i}|})(1 + |\varGamma _{X_i}|)^2\right) \). If \(X_i\) has no discrete parents, then (5) simplifies to (4) since \(|\mathrm {Val}(\varDelta _{X_i})| = 1\) and \(n_{\delta _{X_i}} = n\).
3.2 Computational complexity for the whole BN
Let’s now assume without loss of generality that the dependence structure of \(\mathbf {X}\) can be represented by a DAG \(\mathcal {G}\) with in-degree sequence \(d_{X_1} \leqslant d_{X_2} \leqslant \ldots \leqslant d_{X_N}\). For a sparse graph containing cN arcs, this means \(\sum _{i = 1}^{N}d_{X_i}= cN\). Then if we make the common choice of starting greedy search from the empty DAG, we can rewrite (2) as
because:
-
parents are added sequentially to each of the N nodes;
-
if a node \(X_i\) has \(d_{X_i}\) parents then greedy search will perform \(d_{X_i}+ 1\) passes over the candidate parents;
-
for each pass, \(N - 1\) local distributions will need to be relearned as described in Sect. 2.
The candidate parents in the (\(d_{X_i}+ 1\))th pass are evaluated but not included in \(\mathcal {G}\), since no further parents are accepted for a node after its parent set \(\varPi _{X_i}\) is complete. If we drop the assumption from Sect. 2 that each term in the expression above is O(1), and we substitute it with the computational complexity expressions we derived above in this section, then we can write
where \(O(f_{jk}(X_i)) = O(f_{\varPi _{X_i}^{(j - 1)} \cup X_k}(X_i))\), the computational complexity of learning the local distribution of \(X_i\) conditional of \(j - 1\) parents \(\varPi _{X_i}^{(j)}\) currently in \(\mathcal {G}\) and a new candidate parent \(X_k\).
3.2.1 Discrete BNs
For discrete BNs, \(f_{jk}(X_i)\) takes the form shown in (3) and
The second term is an arithmetic progression,
and the third term is a geometric progression
leading to
Hence, we can see that \(O(g(N, \mathbf {d}))\) increases linearly in the sample size. If \(\mathcal {G}\) is uniformly sparse, all \(d_{X_i}\) are bounded by a constant b (\(d_{X_i}\leqslant b\), \(c \leqslant b\)) and
so the computational complexity is quadratic in N. Note that this is a stronger sparsity assumption than \(\sum _{i = 1}^{N}d_{X_i}= cN\), because it bounds individual \(d_{X_i}\) instead of their sum; and it is commonly used to make challenging learning problems feasible (e.g. Cooper and Herskovits 1992; Friedman and Koller 2003). If, on the other hand, G is dense and \(d_{X_i}= O(N)\), then \(c = O(N)\)
and \(O(g(N, \mathbf {d}))\) is more than exponential in N. In between these two extremes, the distribution of the \(d_{X_i}\) determines the actual computational complexity of greedy search for a specific types of structures. For instance, if \(\mathcal {G}\) is a scale-free DAG (Bollobás et al. 2003) the in-degree of most nodes will be small and we can expect a computational complexity closer to quadratic than exponential if the probability of large in-degrees decays quickly enough compared to N.
3.2.2 GBNs
If we consider the leading term of (4), we obtain the following expression:
Noting the arithmetic progression
we can write
which is again linear in n but cubic in the \(d_{X_i}\). We note, however, that even for dense networks (\(d_{X_i}= O(N)\)) computational complexity remains polynomial
which was not the case for discrete BNs. If, on the other hand \(d_{X_i}\leqslant b\),
which is quadratic in N.
3.2.3 CLGBNs
Deriving the computational complexity for CLGBNs is more complicated because of the heterogeneous nature of the nodes. If we consider the leading term of (5) for a BN with \(M < N\) Gaussian nodes and \(N - M\) multinomial nodes, we have
The first term can be computed using (7) since discrete nodes can only have discrete parents, and thus cluster in a subgraph of \(N - M\) nodes whose in-degrees are completely determined by other discrete nodes; and the same considerations we made in Sect. 3.2.1 apply.
As for the second term, we will first assume that all \(D_i\) discrete parents of each node are added first, before any of the \(G_i\) continuous parents (\(d_{X_i}= D_i + G_i\)). Hence, we write
We further separate discrete and continuous nodes in the summations over the possible \(N - 1\) candidates for inclusion or removal from the current parent set, so that substituting (5) we obtain
Finally, combining all terms we obtain the following expression:
While it is not possible to concisely describe the behaviour resulting from this expression given the number of data-dependent parameters (\(D_i\), \(G_i\), M), we can observe that:
-
\(O(g(N, \mathbf {d}))\) is always linear in the sample size;
-
unless the number of discrete parents is bounded for both discrete and continuous nodes, \(O(g(N, \mathbf {d}))\) is again more than exponential;
-
if the proportion of discrete nodes is small, we can assume that \(M \approx N\) and \(O(g(N, \mathbf {d}))\) is always polynomial.
4 Greedy search and big data
In Sect. 3, we have shown that the computational complexity of greedy search scales linearly in n, so greedy search is efficient in the sample size and it is suitable for learning BNs from big data. However, we have also shown that different distributional assumptions on \(\mathbf {X}\) and on the \(d_{X_i}\) lead to different complexity estimates for various types of BNs. We will now build on these results to suggest two possible improvements to speed up greedy search.
4.1 Speeding up low-order regressions in GBNs and CLGBNs
Firstly, we suggest that estimating local distributions with few parents can be made more efficient; if we assume that \(\mathcal {G}\) is sparse, those make up the majority of the local distributions learned by greedy search and their estimation can potentially dominate the overall computational cost of Algorithm 1. As we can see from the summations in (6), the overall number of learned local distributions with j parents is
that is, it is inversely proportional to the number of nodes for which \(d_{X_i}\) is less than \(j- 1\) in the DAG we are learning. If that subset of nodes represents large fraction of the total, as is the case for scale-free networks and for networks in which all \(d_{X_i}\leqslant b\), (8) suggests that a correspondingly large fraction of the local distributions we will estimate in Algorithm 1 will have a small number j of parents. Furthermore, we find that in our experience BNs will typically have a weakly connected DAG (that is, with no isolated nodes); and in this case local distributions with \(j = 0, 1\) will need to be learned for all nodes, and those with \(j = 2\) for all non-root nodes.
In the case of GBNs, local distributions for \(j = 0, 1, 2\) parents can be estimated in closed form using simple expressions as follows:
-
\(j = 0\) corresponds to trivial linear regressions of the type
$$\begin{aligned} X_i = \mu _{X_i} + \varepsilon _{X_i}. \end{aligned}$$in which the only parameters are the mean and the variance of \(X_i\).
-
\(j = 1\) corresponds to simple linear regressions of the type
$$\begin{aligned} X_i = \mu _{X_i} + X_j \beta _{X_j} + \varepsilon _{X_i}, \end{aligned}$$for which there are the well-known (e.g. Draper and Smith 1998) closed-form estimates
$$\begin{aligned} {\hat{\mu }}_{X_i}&= {\bar{x}}_i - {\hat{\beta }}_{X_j}{\bar{x}}_j, \\ {\hat{\beta }}_{X_j}&= \frac{{\text {COV}}(X_i, X_j)}{{\text {VAR}}(X_i)}, \\ {\hat{\sigma }}^2_{X_i}&= \frac{1}{n - 2}(X_i - {\hat{x}}_i)^T (X_i - {\hat{x}}_i); \end{aligned}$$where \({\text {VAR}}(\cdot )\) and \({\text {COV}}(\cdot , \cdot )\) are empirical variances and covariances.
-
for \(j = 2\), we can estimate the parameters of
$$\begin{aligned} X_i = \mu _{X_i} + X_j \beta _{X_j} + X_k \beta _{X_k} + \varepsilon _{X_i} \end{aligned}$$using their links to partial correlations:
$$\begin{aligned} \rho _{X_i X_j {\text {|}}X_k}&= \frac{\rho _{X_i X_j} - \rho _{X_i X_k} \rho _{X_j X_k}}{\sqrt{1 - \rho _{X_i X_k}^2}\sqrt{1 - \rho _{X_j X_k}^2}} \\&= \beta _{j} \frac{\sqrt{1 - \rho _{X_j X_k}^2}}{\sqrt{1 - \rho _{X_i X_k}^2}}; \\ \rho _{X_i X_k {\text {|}}X_j}&= \beta _{k} \frac{\sqrt{1 - \rho _{X_j X_k}^2}}{\sqrt{1 - \rho _{X_i X_j}^2}}; \end{aligned}$$for further details we refer the reader to Weatherburn (1961). Simplifying these expressions leads to
$$\begin{aligned} {\hat{\beta }}_{X_j}&= \frac{1}{d} \big [{\text {VAR}}(X_k){\text {COV}}(X_i, X_j) \\&\quad - {\text {COV}}(X_j, X_k){\text {COV}}(X_i, X_k)\big ], \\ {\hat{\beta }}_{X_k}&= \frac{1}{d} \big [{\text {VAR}}(X_j){\text {COV}}(X_i, X_k) \\&\quad - {\text {COV}}(X_j, X_k){\text {COV}}(X_i, X_j)\big ]; \end{aligned}$$with denominator
$$\begin{aligned} d = {\text {VAR}}(X_j){\text {VAR}}(X_k) - {\text {COV}}(X_j, X_k). \end{aligned}$$Then, the intercept and the standard error estimates can be computed as
$$\begin{aligned} {\hat{\mu }}_{X_i}&= {\bar{x}}_i - {\hat{\beta }}_{X_j}{\bar{x}}_j - {\hat{\beta }}_{X_k}{\bar{x}}_k, \\ {\hat{\sigma }}^2_{X_i}&= \frac{1}{n - 3}(X_i - {\hat{x}}_i)^T (X_i - {\hat{x}}_i). \end{aligned}$$
All these expressions are based on the variances and the covariances of \((X_i, \varPi _{X_i})\), and therefore can be computed in
This is faster than (4) for the same number of parents, albeit in the same class of computational complexity
and it suggests that learning low-order local distributions in this way can be markedly faster, thus driving down the overall computational complexity of greedy search without any change in its behaviour. We also find that issues with singularities and numeric stability, which are one of the reasons to use the QR decomposition to estimate the regression coefficients, are easy to diagnose using the variances and the covariances of \((X_i, \varPi _{X_i})\); and they can be resolved without increasing computational complexity again.
As for CLGBNs, similar improvements in speed are possible for continuous nodes. Firstly, if a continuous \(X_i\) has no discrete parents (\(\varDelta _{X_i}= \varnothing \)) then the computational complexity of learning its local distribution using QR is again given by (4) as we noted in Sect. 3.1.3; and we are in the same setting we just described for GBNs. Secondly, if \(X_i\) has discrete parents (\(D_{X_i} > 0\)) and j continuous parents (\(G_{X_i} = j\)), the closed-form expressions above can be computed for all the configurations of the discrete parents in
time, which is faster than the estimator from (5):
Interestingly we note that (10) does not depend on \(D_{X_i}\), unlike (5); the computational complexity of learning local distributions with \(G_{X_i} \leqslant 2\) does not become exponential even if the number of discrete parents is not bounded.
4.2 Predicting is faster than learning
BNs are often implicitly formulated in a prequential setting (Dawid 1984), in which a data set \(\mathcal {D}\) is considered as a snapshot of a continuous stream of observations and BNs are learned from that sample with a focus on predicting future observations. Chickering and Heckerman (2000) called this the “engineering criterion” and set
as the score function to select the optimal \(\mathcal {G}_{ max }\), effectively maximising the negative cross-entropy between the “correct” posterior distribution of \(\mathbf {X}^{(n + 1)}\) and that determined by the BN with DAG \(\mathcal {G}\). They showed that this score is consistent and that even for finite sample sizes it produces BNs which are at least as good as the BNs learned using the scores in Sect. 1, which focus on fitting \(\mathcal {D}\) well. Allen and Greiner (2000) and later Peña et al. (2005) confirmed this fact by embedding k-fold cross-validation into greedy search, and obtaining both better accuracy both in prediction and network reconstruction. In both papers, the use of cross-validation was motivated by the need to make the best use of relatively small samples, for which the computational complexity was not a crucial issue.
However, in a big data setting it is both faster and accurate to estimate (11) directly by splitting the data into a training and test set and computing
that is, we learn the local distributions on \(\mathcal {D}^{train}\) and we estimate the probability of \(\mathcal {D}^{test}\). As is the case for many other models (e.g., deep neural networks; Goodfellow et al. 2016), we note that prediction is computationally much cheaper than learning because it does not involve solving an optimisation problem. In the case of BNs, computing (12) is:
-
\(O(N|\mathcal {D}^{test}|)\) for discrete BNs, because we just have to perform an O(1) look-up to collect the relevant conditional probability for each node and observation;
-
\(O(cN|\mathcal {D}^{test}|)\) for GBNs and CLGBNs, because for each node and observation we need to compute \(\varPi _{X_i}^{(n+1)}{\hat{\beta }}_{X_i}\) and \({\hat{\beta }}_{X_i}\) is a vector of length \(d_{X_i}\).
In contrast, using the same number of observations for learning in GBNs and CLGBNs involves a QR decomposition to estimate the regression coefficients of each node in both (4) and (5); and that takes longer than linear time in N.
Conditional Linear Gaussian BN from Vitolo et al. (2018). Yellow nodes are multinomial, blue nodes are Gaussian, and green nodes are conditional linear Gaussian
Hence by learning local distributions only on \(\mathcal {D}^{train}\) we improve the speed of structure learning because the per-observation cost of prediction is lower than that of learning; and \(\mathcal {D}^{train}\) will still be large enough to obtain good estimates of their parameters \(\varTheta _{X_i}\). Clearly, the magnitude of the speed-up will be determined by the proportion of \(\mathcal {D}\) used as \(\mathcal {D}^{test}\). Further improvements are possible by using the closed-form results from Sect. 4.1 to reduce the complexity of learning local distributions on \(\mathcal {D}^{train}\), combining the effect of all the optimisations proposed in this section.
5 Benchmarking and simulations
We demonstrate the improvements in the speed of structure learning and we discussed in Sects. 4.1 and 4.2 using the MEHRA data set from Vitolo et al. (2018), which studied 50 million observations to explore the interplay between environmental factors, exposure levels to outdoor air pollutants, and health outcomes in the English regions of the UK between 1981 and 2014. The CLGBN learned in that paper is shown in Fig. 1: It comprises 24 variables describing the concentrations of various air pollutants (O3, PM2.5, PM10, SO2, NO2, CO) measured in 162 monitoring stations, their geographical characteristics (latitude, longitude, latitude, region and zone type), weather (wind speed and direction, temperature, rainfall, solar radiation, boundary layer height), demography and mortality rates.
The original analysis was performed with the bnlearn R package (Scutari 2010), and it was complicated by the fact that many of the variables describing the pollutants had significant amounts of missing data due to the lack of coverage in particular regions and years. Therefore, Vitolo et al. (2018) learned the BN using the Structural EM algorithm (Friedman 1997), which is an application of the expectation-maximisation algorithm (EM; Dempster et al. 1977) to BN structure learning that uses hill climbing to implement the M step.
For the purpose of this paper, and to better illustrate the performance improvements arising from the optimisations from Sect. 4, we will generate large samples from the CLGBN learned by Vitolo et al. (2018) to be able to control sample size and to work with plain hill climbing on complete data. In particular:
-
1.
we consider sample sizes of 1, 2, 5, 10, 20 and 50 millions;
-
2.
for each sample size, we generate 5 data sets from the CLGBN;
-
3.
for each sample, we learn back the structure of the BN using hill climbing using various optimisations:
-
QR: estimating all Gaussian and conditional linear Gaussian local distributions using the QR decomposition, and BIC as the score function;
-
1P: using the closed-form estimates for the local distributions that involve 0 or 1 parents, and BIC as the score function;
-
2P: using the closed-form estimates for the local distributions that involve 0, 1 or 2 parents, and BIC as the score functions;
-
PRED: using the closed-form estimates for the local distributions that involve 0, 1 or 2 parents for learning the local distributions on \(75\%\) of the data and estimating (12) on the remaining \(25\%\).
-
For each sample and optimisation, we run hill climbing 5 times and we average the resulting running times to reduce the variability of each estimate. Furthermore, we measure the accuracy of network reconstruction using the Structural Hamming Distance (SHD; 2006), which measures the number of arcs that differ between the CPDAG representations of the equivalence classes of two network structures. In our case, those we learn from the simulated data and the original network structure from Vitolo et al. (2018). All computations are performed with the bnlearn package in R 3.3.3 on a machine with two Intel Xeon CPU E5-2690 processors (16 cores) and 384GB of RAM.
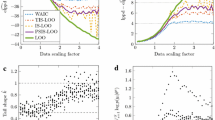
The running times for 1P, 2P and PRED, normalised using those for QR as a baseline, are shown in Fig. 2. As expected, running times decrease with the level of optimisation: 1P (pink) is \(\approx 20\%\) faster than QR, 2P (green) is \(\approx 25\%\) faster and PRED (red) is \(\approx 60\%\) faster, with minor variations at different sample sizes. PRED exhibits a larger variability because of the randomness introduced by the subsampling of \(\mathcal {D}^{test}\) and provides smaller speed-ups for the smallest considered sample size (1 million). Furthermore, we confirm the results from Chickering and Heckerman (2000) on network reconstruction accuracy. In Table 1, we report the sums of the SHDs between the network structures learned by BIC and that from Vitolo et al. (2018), and the corresponding sum for the networks learned using PRED, for the considered sample sizes. Overall, we find that BIC results in 13 errors over the 30 learned DAGs, compared to 4 for (12). The difference is quite marked for samples of size 1 million (11 errors versus 2 errors). On the other hand, neither score results in any error for samples with more than 10 million observations, thus confirming the consistency of PRED. Finally we confirm that the observed running times increase linearly in the sample size as we show in Sect. 3.
In order to verify that these speed increases extend beyond the MEHRA data set, we considered five other data sets from the UCI Machine Learning Repository (Dheeru and Karra Taniskidou 2017) and from the repository of the Data Exposition Session of the Joint Statistical Meetings (JSM). These particular data sets have been chosen because of their large sample sizes and because they have similar characteristics to MEHRA (continuous variables, a few discrete variables, 20-40 nodes overall; see Table 2 for details). However, since their underlying “true DAGs” are unknown, we cannot comment on the accuracy of the DAGs we learn from them. For the same reason, we limit the density of the learned DAGs by restricting each node to have at most 5 parents; this produces DAGs with 2.5N to 3.5N arcs depending on the data set. The times for 1P, 2P and PRED, again normalised by those for QR, are shown in Fig. 3. Overall, we confirm that PRED is \(\approx 60\%\) faster on average than QR. Compared to MEHRA, 1P and 2P are to some extent slower with average speed-ups of only \(\approx 15\%\) and \(\approx 22\%\), respectively. However, it is apparent by comparing Figs. 2 and 3 that the reductions in running times are consistent over all the data sets considered in this paper and hold for a wide range of sample sizes and combinations of discrete and continuous variables.
Running times for the data sets in Table 2, normalised using the baseline implementation based on the QR decomposition (blue), for 1P (pink), 2P (green) and PRED (red). Bars represent 95% confidence intervals. Average running times are reported for QR
6 Conclusions
Learning the structure of BNs from large data sets is a computationally challenging problem. After deriving the computational complexity of the greedy search algorithm in closed form for discrete, Gaussian and conditional linear Gaussian BNs, we studied the implications of the resulting expressions in a “big data” setting where the sample size is very large, and much larger than the number of nodes in the BN. We found that, contrary to classic characterisations, computational complexity strongly depends on the class of BN being learned in addition to the sparsity of the underlying DAG. Starting from this result, we suggested two for greedy search with the aim to speed up the most common algorithm used for BN structure learning. Using a large environmental data set and five data sets from the UCI Machine Learning Repository and the JSM Data Exposition, we show that it is possible to reduce the running time greedy search by \(\approx 60\%\).
Notes
Interestingly, some relaxations of BN structure learning are not NP-hard; see for example Claassen et al. (2013) on learning the structure of causal networks.
All DAGs in the same equivalence class have the same underlying undirected graph and v-structures (patterns of arcs like \(X_i \rightarrow X_j \leftarrow X_k\), with no arcs between \(X_i\) and \(X_k\)).
References
Allen, T.V., Greiner, R.: Model Selection Criteria for Learning Belief Nets: An Empirical Comparison. In: Proceedings of the 17th International Conference on Machine Learning (ICML), pp. 1047–1054 (2000)
Baldi, P., Sadowski, P., Whiteson, D.: Searching for Exotic Particles in High-energy Physics with Deep Learning. Nature Communications 5 (4308), (2014). https://doi.org/10.1038/ncomms5308
Baldi, P., Cranmer, K., Faucett, T., Sadowski, P., Whiteson, D.: Parameterized neural networks for high-energy physics. The Euro. Phys. J. C 76, 235 (2016)
Bollobás, B., Borgs, C., Chayes, J., Riordan, O.: Directed scale-free graphs. In: Proceedings of the 14th Annual ACM-SIAM Symposium on Discrete Algorithms, pp. 132–139 (2003)
Bøttcher, S. G.: Learning Bayesian Networks with Mixed Variables. In: Proceedings of the 8th International Workshop in Artificial Intelligence and Statistics (2001)
Campos, L.M.D., Fernández-Luna, J.M., Gámez, J.A., Puerta, J.M.: Ant Colony optimization for learning bayesian networks. Int. J. Approx. Reason. 31(3), 291–311 (2002)
Chickering, D.M.: A Transformational Characterization of Equivalent Bayesian Network Structures. In: Proceedings of the 11th Conference on Uncertainty in Artificial Intelligence, pp. 87–98 (1995)
Chickering, D.M.: Learning Bayesian networks is NP-complete. In: Fisher, D., Lenz, H. (eds.) Learning from Data: Artificial Intelligence and Statistics V, pp. 121–130. Springer, Berlin (1996)
Chickering, D.M.: Optimal structure identification with greedy search. J. Mach. Learn. Res. 3, 507–554 (2002)
Chickering, D.M., Heckerman, D.: Learning Bayesian networks is NP-hard. Tech. Rep. MSR-TR-94-17, Microsoft Corporation (1994)
Chickering, D.M., Heckerman, D.: A comparison of scientific and engineering criteria for Bayesian model selection. Stat. Comput. 10, 55–62 (2000)
Chickering, D.M., Heckerman, D., Meek, C.: Large-sample learning of Bayesian networks is NP-hard. J. Mach. Learn. Res. 5, 1287–1330 (2004)
Claassen, T., Mooij, J.M., Heskes, T.: Learning Sparse Causal Models is not NP-hard. In: Proceedings of the 29th Conference on Uncertainty in Artificial Intelligence, pp. 172–181 (2013)
Cooper, G., Herskovits, E.: A Bayesian method for the induction of probabilistic networks from data. Mach. Learn. 9, 309–347 (1992)
Cowell, R.: Conditions Under Which Conditional Independence and Scoring Methods Lead to Identical Selection of Bayesian Network Models. In: Proceedings of the 17th Conference on Uncertainty in Artificial Intelligence, pp. 91–97 (2001)
Cussens, J.: Bayesian Network Learning with Cutting Planes. In: Proceedings of the 27th Conference on Uncertainty in Artificial Intelligence, pp. 153–160 (2012)
Dawid, A.P.: Present position and potential developments: some personal views: statistical theory: the prequential approach. J. R. Stat. Soc. Ser. A 147(2), 278–292 (1984)
Dempster, A.P., Laird, N.M., Rubin, D.B.: Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. Ser. B 39(1), 1–38 (1977)
Dheeru D, Karra Taniskidou E (2017) UCI Machine Learning Repository. http://archive.ics.uci.edu/ml
Draper, N.R., Smith, H.: Applied Regression Analysis, 3rd edn. Wiley, London (1998)
Elidan, G.: Copula bayesian networks. In: Lafferty JD, Williams CKI, Shawe-Taylor J, Zemel RS, Culotta A (eds) Advances in Neural Information Processing Systems 23, pp. 559–567 (2010)
Fonollosa, J., Sheik, S., Huerta, R., Marco, S.: Reservoir computing compensates slow response of chemosensor arrays exposed to fast varying gas concentrations in continuous monitoring. Sens. Actuators B: Chem. 215, 618–629 (2015)
Friedman, N.: Learning Belief Networks in the Presence of Missing Values and Hidden Variables. In: Proceedings of the 14th International Conference on Machine Learning (ICML), pp. 125–133 (1997)
Friedman, N., Koller, D.: Being Bayesian about network structure: a Bayesian approach to structure discovery in Bayesian networks. Mach. Learn. 50, 95–125 (2003)
Geiger, D., Heckerman, D.: Learning Gaussian Networks. In: Proceedings of the 10th Conference on Uncertainty in Artificial Intelligence, pp. 235–243 (1994)
Gillispie, S., Perlman, M.: The size distribution for Markov equivalence classes of acyclic digraph models. Artif. Intell. 14, 137–155 (2002)
Glover, F., Laguna, M.: Tabu search. Springer, Berlin (1998)
Goldenberg, A., Moore, A.: Tractable Learning of Large Bayes Net Structures from Sparse Data. In: Proceedings of the 21st International Conference on Machine Learning (ICML), pp. 44–52 (2004)
Goodfellow, I., Bengio, Y., Courville, A.: Deep Learning. MIT Press, Cambridge (2016)
Harary, F., Palmer, E.M.: Graphical Enumeration. Academic Press, Edinburgh (1973)
Heckerman, D., Geiger, D., Chickering, D.M.: Learning Bayesian Networks: The Combination of Knowledge and Statistical Data. Machine Learning 20(3):197–243, Available as Technical Report MSR-TR-94-09 (1995)
JSM, the Data Exposition Session (2009) Airline on-time performance. http://stat-computing.org/dataexpo/200/
Kalisch, M., Bühlmann, P.: Estimating high-dimensional directed acyclic graphs with the PC-Algorithm. J. Mach. Learn. Res. 8, 613–636 (2007)
Karan, S., Eichhorn, M., Hurlburt, B., Iraci, G., Zola, J.: Fast Counting in Machine Learning Applications. In: Proceedings of the 34th Conference on Uncertainty in Artificial Intelligence, pp. 540–549 (2018)
Larranaga, P., Poza, M., Yurramendi, Y., Murga, R.H., Kuijpers, C.M.H.: Structure learning of Bayesian networks by genetic algorithms: a performance analysis of control parameters. IEEE Trans. Pattern Anal. Mach. Intell. 18(9), 912–926 (1996)
Lauritzen, S.L., Wermuth, N.: Graphical models for associations between variables, some of which are qualitative and some quantitative. The Ann. Stat. 17(1), 31–57 (1989)
Moore, A., Lee, M.S.: Cached sufficient statistics for efficient machine learning with large datasets. J. Artif. Intell. Res. 8, 67–91 (1998)
Moral, S., Rumi, R., Salmerón, A.: Mixtures of Truncated Exponentials in Hybrid Bayesian Networks. In: Symbolic and Quantitative Approaches to Reasoning with Uncertainty (ECSQARU), Springer, Lecture Notes in Computer Science, vol 2143, pp. 156–167 (2001)
Pearl, J.: Probabilistic Reasoning in Intelligent Systems: Networks of Plausible Inference. Morgan Kaufmann (1988)
Peña, J.M., Björkegren, J., Tegnèr, J.: Learning dynamic Bayesian network models via cross-validation. Patter Recognit. Lett. 26, 2295–2308 (2005)
Russell, S.J., Norvig, P.: Artificial Intelligence: A Modern Approach, 3rd edn. Prentice Hall, Englewood Cliffs (2009)
Scanagatta, M., de Campos, C.P., Corani, G., Zaffalon, M.: Learning Bayesian networks with thousands of variables. Adv. Neural Inf. Process. Syst. 28, 1864–1872 (2015)
Schwarz, G.: Estimating the dimension of a model. The Ann. Stat. 6(2), 461–464 (1978)
Scutari, M.: Learning Bayesian networks with the bnlearn R Package. J. Stat. Softw. 35(3), 1–22 (2010)
Scutari, M.: Bayesian network constraint-based structure learning algorithms: parallel and optimised implementations in the bnlearn R Package. J. Stat. Softw. 77(2), 1–20 (2017)
Scutari, M., Denis, J.B.: Bayesian Networks with Examples in R. Chapman & Hall, London (2014)
Seber, G.A.F.: A Matrix Handbook for Stasticians. Wiley, New York (2008)
Spirtes, P., Glymour, C., Scheines, R.: Causation, Prediction, and Search, 2nd edn. MIT Press, Cambridge (2001)
Suzuki, J.: An efficient Bayesian network structure learning strategy. N. Gener. Comput. 35(1), 105–124 (2017)
Tsamardinos, I., Brown, L.E., Aliferis, C.F.: The Max-Min Hill-Climbing Bayesian network structure learning algorithm. Mach. Learn. 65(1), 31–78 (2006)
Vitolo, C., Scutari, M., Ghalaieny, M., Tucker, A., Russell, A.: Modelling Air Pollution, Climate and Health Data Using Bayesian Networks: a Case Study of the English Regions. Earth and Space Science 5, submitted (2018)
Weatherburn, C.E.: A First Course in Mathematical Statistics. Cambridge University Press, Cambridge (1961)
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
OpenAccess This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Scutari, M., Vitolo, C. & Tucker, A. Learning Bayesian networks from big data with greedy search: computational complexity and efficient implementation. Stat Comput 29, 1095–1108 (2019). https://doi.org/10.1007/s11222-019-09857-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11222-019-09857-1