Abstract
Aims
Incorporation of flower strips is an agricultural measure to increase aboveground biodiversity and ecosystem services. Although soil communities are key components of terrestrial biodiversity and drive important ecosystem services, their abundance, diversity, and composition in flower strips remain largely unexplored. Here, we shed light on earthworms and soil microorganisms in flower strips.
Methods
We sowed a grassy field margin vegetation as well as two annual and two perennial flower strip mixtures in fully randomized plots of 9 × 28 m in three different types of soil in Germany. Two years following sowing, we determined earthworm communities using chemical extraction and investigated the soil microbiome using real-time PCR (archaea, bacteria, fungi, and soil-N-cycling genes) and amplicon sequencing (bacteria and fungi).
Results
Different plant mixtures (i.e. field margin, annual and perennial flower strips) harbored distinct earthworm and soil microbial communities. Earthworm density and biomass declined or remained unaffected in annual flower strips but increased in perennial flower strips as compared to the field margins. Arbuscular mycorrhizal fungi showed greater diversity and relative abundance in non-tilled (i.e. field margin and perennial flower strips) than in tilled plant mixtures (i.e. annual flower strips).
Conclusions
We attribute changes in earthworm and microbial communities mainly to the effect of tillage and plant diversity. Overall, we suggest that perennial flower strips serve as refugia. Future studies should compare soil biota in perennial flower strips to those in adjacent fields and investigate whether the promotion of soil communities extends into adjacent fields (‘spillover’).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The global loss of biodiversity has far-reaching negative impacts on ecosystem functions (Tilman et al. 2014) and consequently humanity (Cardinale et al. 2012). Agricultural intensification significantly contributes to the loss of biodiversity in agroecosystems (e.g. Kleijn et al. 2009). In the European Union, one of the financially supported measures to mitigate the loss of aboveground biodiversity and associated ecosystem services in agroecosystems is the integration of semi-natural habitats such as uncultivated herbaceous or woody strips within or along arable fields (EU Regulation No 1307/2013). Among semi-natural habitats, incorporation of flower strips along field edges is a common practice. For example, in Germany, flower strips accounted for approximately 1% of the total arable land in 2018 (Schütz et al. 2022). Flower strips are known to increase, maintain or restore aboveground biodiversity and its related ecosystem functions in agroecosystems. For example, flower strips provide habitat and food resources for pollinators and therefore promote their abundance and diversity (e.g. Geppert et al. 2020). The magnitude of the effects of flower strips on pollination services and crop yield in adjacent croplands is variable and depends on the age of the flower strip and its plant diversity (i.e. perennial and old flower strips with high plant diversity promote pollination services most effectively) (Albrecht et al. 2020). Furthermore, flower strips can increase the abundance of natural enemies of pests and promote pest control services (e.g. Tschumi et al. 2015). A recent data synthesis revealed that flower strips enhance pest control services in adjacent croplands by 16% on average (Albrecht et al. 2020).
Although soil communities are a key component of terrestrial biodiversity and their diversity and composition determine ecosystem multifunctionality (Wagg et al. 2014), soil biota in flower strips remain largely unexplored. However, several studies investigated the effect of other types of semi-natural habitats on soil biota. For example, compared to adjacent arable fields, grassy field margins have been shown to harbor greater abundance as well as diversity of earthworms (e.g. Smith et al. 2008; Crittenden et al. 2015) and certain groups of soil-dwelling insects (e.g. Smith et al. 2008). Furthermore, a study conducted by Sechi et al. (2017) revealed differences in trait-dependent (i.e. eco-physiological, behavioral, and morphological traits) responses of belowground microfauna (i.e. nematodes) and mesofauna (i.e. collembolans, mites, and enchytraeids) to semi-natural field margins as compared to adjacent arable fields. Furthermore, Sechi et al. (2017) showed that semi-natural field margins favor fungal and bacterial biomass. The findings of D’Acunto et al. (2016) further suggest that compared to a conventionally managed soybean field, adjacent herbaceous field margins harbor functionally distinct microbial communities. Besides the effect of plant composition of semi-natural habitats on soil biota, management strategy (i.e. tillage) of semi-natural habitats has been shown to alter the abundance and diversity of soil macrofauna (Smith et al. 2008). Considering the known effects of semi-natural habitats and their management on soil biota, it is reasonable to assume that flower strips alter the belowground biota as well.
In their role as ecosystem engineers, earthworms contribute to several beneficial soil functions (e.g. water infiltration (e.g. Ehlers 1975; Ernst et al. 2009; Capowiez et al. 2015), suppression of phytopathogens (e.g. Wolfarth et al. 2011; Euteneuer et al. 2019; Plaas et al. 2019), and cycling of nutrients (e.g. Reichle 1977; Blouin et al. 2013; Medina-Sauza et al. 2019)) and enhance soil fertility (e.g. Tomati and Galli 1995; Bhadauria and Saxena 2010; Ahmed and Al-Mutairi 2022). Overall, earthworms are suitable biological indicators for sustainable soil management in agriculture (Paoletti 1999). More than two decades ago, Kohli et al. (1999) conducted one of the first studies on earthworms in flower strips. The authors showed that conversion of a tilled maize field into a non-tilled wild flower strip increased the abundance of earthworms already after one year and reached a plateau after two years of absence of tillage (Kohli et al. 1999). Besides tillage, the impacts of plant diversity and biomass on earthworm communities have frequently been studied in grasslands. While some studies revealed a positive impact of plant diversity and biomass on earthworm density and biomass (Zaller and Arnone 1999; Spehn et al. 2000; Eisenhauer et al. 2013), other studies were not able to confirm this (Wardle et al. 1999; Hedlund et al. 2003). These discrepancies among studies may be related to, inter alia, interactions with other soil biota (Milcu et al. 2006) and plant community composition (Gastine et al. 2003; Milcu et al. 2006, 2008; Eisenhauer et al. 2009). Yet, comprehensive experimental data on the effects of flower strips on earthworm communities and their functions are lacking.
Soil microbial communities regulate fundamental biogeochemical cycles (e.g. Rousk and Bengtson 2014) and drive plant productivity (e.g. van der Heijden et al. 2008) and therefore provide existential functions for agriculture. Although flower strips are widely applied and cover large areas, to our knowledge, data on microbial communities in flower strips are missing in the scientific literature. With respect to diversification of agroecosystems, the question of whether flower strips promote beneficial soil microorganisms arises. For example, Burrows and Pfleger (2002) found positive relationships between plant diversity and spore number and volume of arbuscular mycorrhizal fungi (AMF). Thus, it is reasonable to assume that higher plant diversity in flowers strips benefits AMF. Furthermore, as reduced tillage promotes AMF (e.g. Bowles et al. 2017), it can be expected that non-tilled perennial flower strips benefit AMF more than tilled annual flower strips. AMF form symbiotic associations with the majority of terrestrial plants and, inter alia, enhance nutrient acquisition by associated plants (Clark and Zeto 2000). Whether flower strips promote AMF remains yet to be tested.
In this work, we shed light on representatives of soil biota under flower strips. For the first time, we investigated soil archaea, bacteria, fungi, and earthworms under a grassy field margin vegetation versus four different types of flower strip mixtures (two annual and two perennial flower strip mixtures comprising 11 to 13 and 30 to 51 plant species, respectively). The five plant mixtures were sown in fully randomized plots of 9 × 28 m and soil biota were studied two years following sowing. Our experimental design was replicated on three study sites with three different soil types to evaluate whether belowground responses to flower strip mixtures are soil type-specific. We hypothesized that i) flower strip mixtures increase the abundance and alter the composition of soil microbial communities as well as increase the population size of earthworms compared to grassy field margin vegetation. We further expected that ii) perennial flower strip mixtures promote soil biota more effectively than annual flower strip mixtures due to the absence of soil management (annual flower strip mixtures were re-established) and larger plant richness.
Materials & methods
Study sites and study design
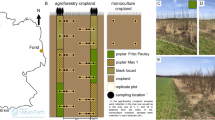
Our study was conducted at three study sites (near Lippetal on a Gleyic Podzol, at the experimental research station of the South Westphalia University of Applied Sciences near Merklingsen on a Gleyic Luvisol, and near Ense on a Stagnic Cambisol (IUSS Working Group WRB 2015); Fig. 1; see Table SI 1 for site description and general soil properties)) in the federal state of North Rhine-Westphalia, Germany. We refer to the study sites by their soil group (i.e. Podzol, Luvisol, and Cambisol soil).
In 2020, five different plant mixtures were sown at each site at a seeding rate of 10 kg ha−1. One of the mixtures was a field margin vegetation comprising four grasses commonly found in field margins at our study region (referred to as ‘field margin’). This mixture was sown in autumn 2020. In spring 2020, four different flower strip mixtures were sown. The flower strip mixtures comprised two annual flower strip mixtures (comprising 11 and 13 plant species, referred to as ‘annual flower strip 1’ and ‘annual flower strip 2’, respectively) and two perennial flower strip mixtures (comprising 30 and 51 plant species, referred to as ‘perennial flower strip 1’ and ‘perennial flower strip 2’, respectively) (Fig. 1). The floral composition of the five different plant mixtures at sowing (2020) and during our year of sampling (2022) are given in Table SI 2 and File SI 1, respectively. At each site, each plant mixture was sown in three replicate plots of 9 × 28 m in a completely randomized design (3 study sites × 5 plant mixtures × 3 replicate plots = 45 replicate plots across sites) (Fig. 1). Prior to the experiment, the sites were conventionally managed croplands (Podzol and Cambisol soil) or fallow (Luvisoil soil). Prior to sowing, soils were tilled twice using a grubber and rotary harrow due to weed pressure.
At each site, the annual flower strips were re-established (flower strips were mulched and the soil was tilled twice (grubber and rotary harrow) prior to resowing) in April 2021 and 2022. The field margin and perennial flower strips were topped at 15 cm height in March 2022 and not further managed, except in the Luvisol soil where all flower strips had to be re-established in spring 2021 due to high weed pressure. None of the replicate plots received fertilizer or plant protection products during the experiment.
Determination of general soil properties
Soil samples for the analysis of general soil properties were collected from July 15 to 16 2022. Soil bulk density was determined at 0 – 5 cm soil depth with 250 cm3 stainless steel cylinders using the soil core method (Blake and Hartge 1986). Soil pH, soil organic C (SOC), total N, double lactate-extractable P (PDL) and K (KDL), calcium chloride-extractable Mg (MgCaCl2), effective cation exchange capacity (CECeff), and soil texture were determined at 0 – 30 cm soil depth. Samples at 0 – 30 cm soil depth were collected using a stainless-steel auger (⌀ 3.5 cm). Three soil subsamples were collected and thoroughly homogenized in a sterile polyethylene bag to obtain one composite soil sample for 0 – 30 cm soil depth for each replicate plot. Composite soil samples were air-dried and sieved to < 2 mm. Soil pH was determined in demineralized H2O at a ratio of 1:2.5 (soil:water (w/v)). Prior to the determination of SOC, carbonates were removed from the samples using acid fumigation as per Harris et al. (2001). SOC and total N were determined using a CNS elemental analyzer (Vario EL Cube, Elementar, Germany). PDL and KDL were determined as per (VDLUFA 1991a) and MgCaCl2 as per (VDLUFA 1991b). Soil texture and CECeff were determined as per DIN 19683–2 (1997) and DIN ISO 11260 (1997), respectively.
Earthworm extraction and species identification
Earthworm communities were sampled from October 16 to 18 2022 using Allyl isothiocyanate (AITC) expulsion as described previously (Vaupel et al. 2023). Briefly, within each replicate plot, earthworms were expelled from two subplots in order to account for spatial heterogeneity. Squared aluminum frames (50 × 50 cm) were embedded approx. 5 cm into the soil and 5 L of a 0.01% (v/v in tap water) AITC solution were poured into the frames. Emerging earthworms were collected from the soil surface for 30 min, washed with tap water, and stored in tap water. In total, 2,250 earthworms were collected within 72 h. Within 12 h post sampling, earthworms were weighed (including gut content), species were determined based on morphology as per (Krück 2018), and all collected individuals were released. Earthworm counts and biomass from the two subplots were added up. Although the remaining gut content of earthworms may add bias to our earthworm biomass data, the large number of individuals restricted us from allowing earthworms to empty their gut prior to weighing. Seven different earthworm species were found across the three study sites: Allolobophora chlorotica, Aporrectodea caliginosa, Aporrectodea longa, Aporrectodea rosea, Aporrectodea trapezoides (also referred to as a subspecies of Aporrectodea caliginosa), Lumbricus rubellus, and Lumbricus terrestris. Earthworm species were classified into three ecological groups: epigeic (Lumbricus rubellus), endogeic (Allolobophora chlorotica, Aporrectodea caliginosa, Aporrectodea rosea, and Aporrectodea trapezoides), and anecic earthworms (Aporrectodea longa and Lumbricus terrestris), which were introduced by Bouché (1972). Earthworm data (i.e. count of individuals as well as total biomass per square meter) has been deposited at the BonaRes Repository (https://doi.org/10.20387/bonares-gx1f-bh69).
Soil DNA extraction
Soil samples for the analysis of soil microorganisms at 0 – 30 cm soil depth were collected on the same day as those for general soil properties (July 15 to 16 2022). An aliquot of approximately 50 g fresh soil from the composite samples of 0 – 30 cm soil depth (see Determination of general soil properties) was stored at -20 °C in the field. Upon arrival at the laboratory, frozen soil samples were stored at -20 °C until freeze-drying. Frozen soil samples were freeze-dried for 72 h and thoroughly homogenized using a vortexer as described previously (Beule et al. 2019). DNA was extracted from 50 mg finely ground soil using a cetyltrimethylammonium bromide (CTAB)-based protocol as per (Beule et al. 2021). Quantity and quality of the DNA extracts were assessed on 1.7% (w/v) agarose gels stained with SYBR Green I solution (Thermo Fisher Scientific GmbH, Dreieich, Germany).
Quantification of soil microbial groups using real-time PCR
Prior to real-time PCR, DNA extracts were diluted 1:50 (v/v) in double distilled H2O (ddH2O) to overcome PCR inhibition (Guerra et al. 2020). Soil bacteria and fungi were quantified as described previously (Beule et al. 2020). Soil archaea were quantified using the primer pair 340F / 100R (Gantner et al. 2011) using the identical master mix composition as for fungi (Beule et al. 2020). The thermocycling conditions of archaea were as follows: initial denaturation at 95 °C for 120 s followed by 40 cycles of 95 °C for 20 s, 60 °C for 30 s, and 68 °C for 30 s, and final elongation at 68 °C for 5 min. Genes involved in soil nitrogen (N)-cycling (nitrification: ammonia-oxidizing archaea (AOA) and bacteria (AOB) amoA genes; denitrification: nirK, nirS, and nosZ clade I and II genes) were quantified to estimate the population size of N-cycling microorganisms as per (Beule et al. 2019). All reactions were carried out in 4 µL reaction volumes in a Peqstar 96Q thermocycler (PEQLAB, Erlangen, Germany). Melting curves were generated as described previously (Beule et al. 2019).
Amplicon sequencing of the soil microbiome
Soil bacteria and fungi were amplified using the primer pair 341F (5′-CCTACGGGNGGCWGCAG-3′) / 785R (5′-GACTACHVGGGTATCTAAKCC-3′ (Klindworth et al. 2013) and ITS1-F_KYO2 (5’-TAGAGGAAGTAAAAGTCGTAA-3’) (Toju et al. 2012) / ITS86R (5’-TTCAAAGATTCGATGATTCA-3’) (Vancov and Keen 2009), respectively. Prior to PCR, DNA extracts were diluted 1:50 (v/v) in ddH2O to overcome PCR inhibition (Guerra et al. 2020). Amplification was carried out in 25 µL reaction volume in an Eppendorf Mastercycler EP Gradient S thermocycler (Eppendorf, Hamburg, Germany). Bacteria and fungi were each amplified within one PCR run using the same mastermix for all samples. The reaction volume contained 18.75 µL mastermix and 6.25 µL template DNA or ddH2O for a negative control. The mastermix comprised ddH2O, buffer (10 mM Tris–HCl, 50 mM KCl, 2.0 mM MgCl2, pH 8.3 at 25 °C), 100 µM of each deoxynucleoside triphosphate (New England Biolabs, Beverly, Massachusetts, USA), 0.5 µM of each primer, 1 mg mL−1 bovine serum albumin, and 0.03 u µL−1 Hot Start Taq DNA Polymerase (New England Biolabs, Beverly, Massachusetts, USA). Each primer was a mixture of primer with (50%) and without (50%) Illumina TruSeq 5’-end adapters (5’-GACGTGTGCTCTTCCGATCT-3’ for the forward primer and 5’-ACACGACGCTCTTCCGATCT-3’ for the reverse primer). Bacteria and fungi were amplified using a touch-up PCR protocol (Beule and Karlovsky 2021) with initial denaturation at 95 °C for 2 min, 3 touch-up cycles (95 °C for 20 s, 50 °C for 30 s, and 68 °C for 60 s), 22 or 25 cycles (95 °C for 20 s, 58 °C for 30 s, and 68 °C for 60 s) for bacteria and fungi, respectively, and final elongation at 68 °C for 10 min. Amplification success was verified on 1.7% (w/v) agarose gel stained with SYBR Green I solution (Thermo Fisher Scientific GmbH, Dreieich, Germany) and libraries were shipped to LGC Genomics (Berlin, Germany). A second amplification with standard i7- and i5- sequencing adapters was performed at the facilities of LGC Genomics. Libraries were multiplexed and sequenced on an Illumina MiSeq (V3 chemistry, 2 × 300 bp) (Illumina, Inc., San Diego, CA, USA). Amplicon sequencing data have been deposited at NCBI’s Short Read Archive (BioProject PRJNA905898 for bacteria and PRJNA905904 for fungi).
Bioinformatic processing of amplicon sequencing data
Paired-end sequencing data of bacteria and fungi were demultiplexed using Illumina’s bcl2fast version 2.20 (Illumina, San Diego, CA, USA). One-sided and conflicting barcodes as well as barcodes containing more than two mismatches were removed. Sequencing adapter and primer sequences were clipped and reads with < 100 bp were discarded. Afterwards, sequencing reads were processed in QIIME 2 version 2022.2 (Bolyen et al. 2019). Quality scores were manually inspected using the ‘q2-demux’ plugin. Sequence reads were quality filtered (allowing two expected errors), merged, and cleaned from chimeric sequences and singletons using DADA2 (Callahan et al. 2016). Obtained amplicon sequencing variants (ASVs) of bacteria and fungi were taxonomically classified against the SILVA ribosomal RNA gene database version 138 (Quast et al. 2013) and UNITE database version 8.3 QIIME developer release (Abarenkov et al. 2021), respectively. Classification was achieved utilizing a scikit-learn Naive Bayes machine-learning classifier (‘q2-fit-classifier-naive-bayes’ and ‘q2- classify-sklearn’ plugin) in the ‘balanced’ configuration ([7,7]; 0.7 for bacteria and [6,6]; 0.96 for fungi as suggested by (Bokulich et al. 2018)). Following classification, non-bacterial and non-fungal sequence reads were discarded from the bacterial and fungal data sets. Scaling with ranked subsampling (SRS) (Beule and Karlovsky 2020) using the ‘SRS’ R package version 0.2.3 (Heidrich et al. 2021) was used to normalize the bacterial and fungal ASV table to 19,219 and 18,318 sequence counts per sample, respectively. The normalized data sets contained 44,009 bacterial and 3,648 fungal ASVs.
Statistical analysis
To test the effect of plant mixtures (i.e. field margin and different flower strips) on soil properties (soil pH, bulk density, SOC, total N, PDL, KDL, and MgCaCl2) per site (i.e. soil type), we used one-way analysis of variance (ANOVA) followed by Tukey honestly significant difference (HSD) test. To test the effect of plant mixtures on the abundance of soil communities per site, we calculated the relative change of earthworm density and biomass as well as the abundance of archaea, bacteria, fungi, and N-cycling genes in response to the flower strips as follows:
where a is the observed response and b is the average response of field margin per site. Differences in the relative change of earthworm density and biomass as well as absolute abundance of archaea, bacteria, fungi, and N-cycling genes per site were determined using one-way ANOVA followed by Tukey HSD test.
We determined alpha diversity indices covering entropy (Shannon index (H’)), species richness (Chao1 index) and evenness (Pielou’s evenness (J ‘)) of bacterial and fungal communities using the ‘vegan’ R-package (version 2.5–7) (Oksanen et al. 2019). We then tested the effect of plant mixtures on alpha diversity indices per site using one-way ANOVA followed by Tukey HSD test. Differences in relative abundance of microbial taxa obtained from amplicon sequencing among treatments per site were determined from log(x + 1)-transformed data and tested using one-way ANOVA followed by Tukey HSD test. Prior to running one-way ANOVA tests, all data were manually inspected for homoscedasticity and normal distribution of the residuals and tested using Levene’s and Shapiro–Wilk test, respectively. Relationships among different parameters were tested using Spearman rank correlations.
Compositional differences of bacterial and fungal amplicon sequencing data were calculated using pairwise Bray–Curtis dissimilarities and visualized using non-metric multidimensional scaling (NMDS) as implemented in the ‘vegan’ R-package. To test the effects of site and plant mixture on the bacterial and fungal community composition, we performed permutational multivariate analysis of variance (PERMANOVA) on Bray–Curtis dissimilarities using 999 permutations [adonis2(dissimilarity matrix ~ soil type + plant mixture + soil type:plant mixture, nperm = 999)]. Additionally, we tested the effect of plant mixture on the bacterial and fungal community composition per site by running subsets of the datasets per site [adonis2(dissimilarity matrix ~ plant mixture, nperm = 999)]. Complementary to each PERMANOVA model, we assessed the dispersion of samples in each group using multivariate homogeneity of group dispersions.
All statistical analyses were performed in R (version 4.1.2) (R Core Team 2017). For all statistical tests, statistical significance was considered at p < 0.05.
Results
General soil properties
Within each soil type, soil properties remained unaffected by the recent introduction of flower strips. Flower strips did not affect soil pH, bulk density, SOC, total N, PDL, KDL, and MgCaCl2.
Earthworm communities
Earthworm density and biomass were strongly correlated with each other (r = 0.95; p < 0.0001) and increased from the Podzol to the Luvisol to the Cambisol soil (Fig. 2 A, Figure SI 1). The classification of the species into ecological groups (i.e. anecic, endogeic, and epigeic) revealed that earthworm community composition was site-specific. In the Podzol soil, anecic earthworms were absent and epigeic earthworms accounted for a large share of the community. In contrast, epigeic earthworms were not present in the Luvisol soil. The Cambisol soil harbored all three ecological groups (Fig. 2 B).
Earthworm communities. Population densities of ecological groups of earthworms (A) and their relative abundance within the earthworm communities (B). Bars represent individual replicate plots (n = 3). Relative change of earthworm density (C) and biomass (D) in response to flower strips. Non-transparent dots and triangles represent means and vertical bars range from the minimum to the maximum value. Transparent dots and triangles represent individual data points (i.e. replicate plots). Images are courtesy of the Integration and Application Network (ian.umces.edu/media-library)
Perennial flower strips strongly promoted earthworm population density and biomass across soils (Fig. 2A, C, D). In contrast, annual flower strips showed consistently lower density and biomass than the field margin in the Podzol and Luvisol soil (Fig. 2A, C, D). In these two soils, earthworms were almost absent under the annual flower strips (Fig. 2A). In the Podzol soil, the perennial flower strip 2 increased earthworm density and biomass by a factor of 3.7 to 17.5 compared to the field margin and the annual flower strips (p ≤ 0.031; t-ratio = -4.7 to -3.9), which was mainly driven by the increased occurrence of epigeic earthworms in the perennial flower strip 2. Earthworm density in the flower strips in the Luvisol soil did not differ statistically significant from the field margin. However, earthworm densities were 79 to 99 times larger in perennial than in annual flower strips (p ≤ 0.036; t-ratio = -4.4 to -3.5). In the same soil, earthworm biomass was 15.4 to 23.3 times larger in perennial flower strips and 9.2 to 12.8 times larger in the field margin (p ≤ 0.025; t-ratio = -8.0 to 4.2) as compared to annual flower strips. The Cambisol soil was the only soil in which annual flower strips showed earthworm densities and biomass similar to those in the field margin. In this soil, perennial flower strips increased earthworm density by 171 to 247% as compared to the annual flower strips and field margin (p ≤ 0.018; t-ratio = -5.8 to -3.9).
Soil microbiome
Gene copy numbers of archaea, bacteria, fungi, and functional groups involved in soil N-cycling were not affected by flower strips (Figure SI 2, Figure SI 3). Across soils, soil bacterial communities were dominated by the phyla of Actinobacteriota (29.4 ± 6.1%), Proteobacteria (16.4 ± 1.8%), and Acidobacteriota (12.5 ± 1.8%). The dominating bacterial classes were Actinobacteria (20.1 ± 6.6%), Alphaproteobacteria (11.3 ± 1.1%), and Planctomycetes (7.9 ± 2.3%) (Fig. 3A). The fungal community was dominated by Ascomycota (65.4 ± 14.0%), Mortierellomycota (12.3 ± 9.5%), and Basidiomycota (12.0 ± 8.2%) on phylum level and Sordariomycetes (41.5 ± 16.0%), Dothideomycetes (17.2 ± 11. 6%), and Mortierellomycetes (12.2 ± 8.1%) on class level (Fig. 3E). Alpha diversity indices (Shannon index (H’), Chao1 index, and Pielou’s evenness (J’)) were not affected by flower strips (Fig. 3B, C, D, F, G, H), except fungal Shannon diversity in the Luvisol soil which was higher in the perennial flower strips and the field margin compared to the annual flower strip 2 (p = 0.036; t-ratio = -3.9 to 4.3) (Fig. 3F).
Community composition and alpha diversity of soil bacteria and fungi. Mean relative abundance of bacterial (A) and fungal classes (E) per plant mixture and soil type. Alpha diversity indices of bacterial (B, C, D) and fungal communities (F, G, H). Non-transparent dots and triangles represent means and vertical bars range from the minimum to the maximum value (n = 3). Transparent dots and triangles represent individual data points (i.e. replicate plots). Images are courtesy of the Integration and Application Network (ian.umces.edu/media-library)
Soil type (i.e. Podzol, Luvisol, and Cambisol soil) and plant mixture (i.e. field margin and different flower strips) affected community composition of both bacteria and fungi (Table 1, Figure SI 4). For both communities, the effect of soil type on community composition was stronger than the effect of plant mixture (Table 1). Plant mixture effects per site were visualized using NMDS (Fig. 4). In the Luvisol and Cambisol soil the field margin, the annual flower strips, and the perennial flower strips each formed a distinct cluster in the NMDS for both bacteria and fungi (Fig. 4B, C, E, F). In the Podzol soil, two clusters emerged comprising the non-tilled plant mixtures (i.e. the field margin and the perennial flower strips) and the tilled plant mixtures (i.e. the annual flower strips) for bacterial communities (Fig. 4A).
Non-metric multidimensional scaling (NMDS) of Bray–Curtis dissimilarities of soil bacterial and fungal communities. NMDS plots of bacterial (A, B, C) and fungal communities (D, E, F) within each soil type. Dots and triangles represent individual data points (i.e. replicate plots) (n = 3) which are connected with the centroid of their respective plant mixture
The relative abundance of several bacterial phyla was affected by the plant mixtures (Fig. 5; see Table SI 3 for p-values) and reflected the clustering in the NMDS. For example, relative abundance of Desulfobacterota in the Cambisol soil were greater in the field margin than in the flower strips (p ≤ 0.0001; t-ratio = 7.6 to 8.5). In the same soil, the field margin increased the relative abundance of Methylomirabilota and Latescibacterota as compared to the annual flower strips (p ≤ 0.015; t-ratio = 4.0 to 5.0). In contrast, compared to the field margin, annual flower strips promoted the relative abundance of Actinobacteria, Bdellovibrionota, and Proteobacteria in the Cambisol soil (p ≤ 0.037; t-ratio = -6.7 to -3.6). In the Luvisol soil, Latescibacterota showed greater relative abundance in the field margin than in the flower strips (p ≤ 0.0011; t-ratio = 5.9 to 10.3). Likewise, in the Luvisol soil, relative abundances of Abditibacteriota and Gemmatimonadota were greater in the annual flower strips than in the field margin (p ≤ 0.043; t-ratio = -4.0 to -3.4). In all soil types, relative abundance of Bacteroidota were greater in the annual flower strips than in the field margin (p ≤ 0.021; t-ratio = -6.9 to -3.9).
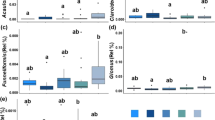
Within the fungal community, the abundance and diversity of affiliates of the monophyletic phylum Glomeromycota (containing all AMF), were altered by the plant mixtures (Fig. 6). Patterns of relative abundance of AMF were not consistent across soil types, except that the relative abundance of AMF was greater in the field margin mixture as compared to the annual flower strip mixtures (p ≤ 0.007; t-ratio = 1.9 to 2.9). Furthermore, in the Cambisol soil, relative abundance of AMF in the perennial flower strips was lower than in the field margin (p ≤ 0.0003; t-ratio = 2.3 to 2.4).
Arbuscular mycorrhizal fungi (AMF – Glomeromycota) in flower strips. Relative abundance of AMF orders in three different soil types (A). Bars represent individual replicate plots (n = 3). Shannon (H’) (B) and Chao1 index (C) of AMF. Non-transparent dots and triangles represent means and vertical bars range from the minimum to the maximum value (n = 3). Transparent dots and triangles represent individual data points (i.e. replicate plots). Images are courtesy of the Integration and Application Network (ian.umces.edu/media-library)
Across sites and plant mixtures, 249 ASVs were assigned to AMF, covering three orders, namely Archaeosporales, Glomerales, and Paraglomerales (Fig. 6A). Relative abundance of Archaeosporales in the Luvisol and the Cambisol soil was greater in the field margin as compared to the annual and perennial flower strips (p ≤ 0.016; t-ratio = 0.4 to 0.7). Furthermore, in the Luvisol soil, relative abundance of Glomerales was greater in the field margin and perennial flower strip 1 than in the annual flower strips (p ≤ 0.043; t-ratio = 1.3 to 1.7). Relative abundance of Glomerales in the Cambisol soil was greater in the field margin compared to the annual and perennial flower strips (p ≤ 0.0008; t-ratio = 2.0 to 2.6). In the Podzol soil, plant mixtures only affected the community share of Paraglomerales which was greater in the field margin compared to the annual flower strips (p ≤ 0.036; t-ratio = 2.1). In the Cambisol soil, relative abundance of Paraglomerales was lower in the annual and perennial flower strips as compared to the field margin (p ≤ 0.0007; t-ratio = 1.4 to 1.5). In contrast, community share of Paraglomerales did not differ among plant mixtures in the Luvisol soil.
Alpha diversity (Shannon index (H’) and Chao1 index) of AMF differed significantly among the plant mixtures (Fig. 6B, C). In each soil type, alpha diversity of AMF was greater in the field margin as compared to the annual flower strips (p ≤ 0.0053; t-ratio = 3.2 to 4.2 and p ≤ 0.0066; t-ratio = 23.7 to 53.5 for Shannon index and Chao1, respectively). Furthermore, alpha diversity of AMF did not differ between field margin and the perennial flower strips in the Podzol and Luvisol soil. In the Cambisol soil, however, Chao1 index was greater in the field margin than in the perennial flower strips (p ≤ 0.0001; t-ratio = 39.2 to 40.0). According to Shannon index, alpha diversity of AMF was greater in the perennial flower strips as compared to the annual flower strip 1 in all soil types (p ≤ 0.012; t-ratio = 2.4 to 3.6).
Discussion
Earthworm communities
We collected earthworms using AITC extraction without hand-sorting (Vaupel et al. 2023). While using exclusively AITC extraction works well for adult anecic earthworms, endogeic species and juveniles may be recovered with reduced efficacy without additional hand-sorting (Čoja et al. 2008; Pelosi et al. 2009). Chemical extraction is a non-destructive sampling technique that is often preferred over hand-sorting when additional data needs to be collected at the sampling location (e.g. Lees et al. 2016; Tóth et al. 2020).
In the present study, croplands or fallow were sown with a mixture of four grasses commonly found in field margins, annual flower strip, and perennial flower strip mixtures. In the Podzol and Luvisol soil, annual flower strips showed the lowest earthworm density and biomass (Fig. 2A, Figure SI 1), which we attribute to their annual re-establishment that included tillage (grubber and rotary harrow). Although all plots in the Luvisol soil had to be re-established one and a half years prior to earthworm sampling due to high weed pressure, differences among tillage regimes were already apparent following just one tillage operation (Fig. 2A, Figure SI 1). Such rapid recovery of earthworm populations following tillage agrees with previous studies showing population recoveries within few months post tillage (Marinissen 1992; Boström 1995). Tillage is well-known to affect density, biomass, and community composition of earthworms (Chan 2001; Ernst and Emmerling 2009). While density of anecic species generally decreases under tillage due to physical damage and the removal of plant litter from the soil surface (e.g. Ernst and Emmerling 2009), responses of endogeic species to tillage are rather inconsistent. While some studies showed that the density of endogeic species is either unaffected (e.g. Pelosi et al. 2014; Torppa and Taylor 2022) or greater in ploughed soils as compared to soils under reduced or no tillage (e.g.Ernst and Emmerling 2009; Capowiez et al. 2009), due to the incorporation of plant residues that serve as a food resource, other studies found a negative impact of tillage on endogeic earthworm density (e.g. Edwards and Lofty 1982; Simonsen et al. 2010). In view of these inconsistent results, Briones and Schmidt (2017) recently conducted a global meta-analysis on the effects of tillage on earthworm abundance and biomass. Their results revealed that the population densities of all three ecological groups benefit from reduced tillage and that epigeic and anecic species benefit more than endogeic (Briones and Schmidt 2017). Their results agree with our findings of a decline in all three ecological groups of earthworms (epigeic, endogeic, and anecic) under the tilled annual flower strips as compared to the non-tilled field margin and perennial flower strips (Fig. 2A).
Although differences in tillage regimes can explain the low earthworm densities in the annual flower strips, they do not explain the increased population densities in the non-tilled perennial flower strips as compared to the non-tilled field margin (Fig. 2A, C, D). The impacts of plant richness and biomass on earthworm communities have frequently been studied in grasslands. While some studies revealed a positive impact of plant richness and biomass on earthworm density and biomass (Zaller and Arnone 1999; Spehn et al. 2000; Eisenhauer et al. 2013), other studies were not able to confirm this (Wardle et al. 1999; Hedlund et al. 2003). These discrepancies among studies may be related to, inter alia, interactions with other soil biota (Milcu et al. 2006) and plant community composition (Gastine et al. 2003; Milcu et al. 2006, 2008; Eisenhauer et al. 2009). In our study, perennial flower strip mixtures showed higher plant species richness of sown plants than field margin mixtures (File SI 1), possibly contributing to the higher earthworm density and biomass observed in all three soil types (Fig. 2A, C, D, Figure SI 1). Although plant biomass was not determined in our study, previous studies showed that plant biomass production (and consequently plant litter production) generally increases with plant richness (e.g. Cardinale et al. 2007). Thus, we suggest that compared to the field margin, earthworm communities in the perennial flower strips benefited from higher quantities of above- and belowground plant litter (i.e. food resources). We further suggest that perennial flower strips not just increase the quantity of food input but also alter its quality which may be even more important for soil decomposer communities (e.g. Milcu et al. 2006; Eisenhauer and Reich 2012).
The spatial design of our study may have also affected the community dynamics of earthworms. At each study site, earthworm communities are likely to respond to a change in management through differential recruitment from the local species pool. However, although our plots were fully randomized, our study design did not restrict influxes of earthworms from spatially close soil environments. In this context, habitat connectivity represents an important driver of earthworm migration (Palm et al. 2013).
Soil microbiome
Plant mixture (i.e. field margin and different flower strips) was identified as a strong driver of bacterial and fungal community composition (Table 1, Fig. 4). Dissimilarities in community composition of bacteria and fungi between the annual flower strips and the other plant mixtures in each soil type (Fig. 4) may be related to tillage during the re-establishment of the annual flower strips. There is compelling evidence of not only changes in microbial population size (Mathew et al. 2012) but also in community composition of bacteria and fungi in response to tillage intensity (e.g. Degrune et al. 2016; Smith et al. 2016; Yin et al. 2017; Frøslev et al. 2022). For example, a global meta-analysis revealed that conservation tillage benefits bacterial and fungal biomass in soil (Chen et al. 2020). Another meta-analysis from the same year showed that the absence of tillage increased the relative abundance of Acidobacteria but decreased the relative abundance of Actionobacteria (Li et al. 2020). These results agree with our findings on these two phyla (Fig. 5, Table SI 3). Furthermore, a recent study was able to show that tillage also changes the vertical distribution of bacterial and fungal communities in soil (Sun et al. 2018). In light of the strong impact of tillage on soil structure (Pagliai et al. 2004) and the subsequent consequences for soil as a biological habitat (Young and Ritz 2000), it is conclusive that tillage can affect the composition of soil microbiomes.
Besides differences in soil management, differences in plant species composition as well as richness of the plant mixtures (field margin < annual flower strips < perennial flower strips; see File SI 1) likely contributed to the observed changes in community composition. Considering the plant richness, this assumption is supported by the differences in community composition between the non-tilled field margin and the non-tilled perennial flower strips. Although not determined in our study, differences in plant biomass among treatments may also have affected community composition. There are numerous interactions between plants and soil microorganisms that shape the soil microbiome. For example, plant root exudates shape the soil microbiome (especially in the rhizosphere) by recruiting plant-beneficial microorganisms (Vives-Peris et al. 2020). The quantity and quality of root exudates depend on abiotic and biotic stressors but also plant species and age (Badri and Vivanco 2009).Thus, it is reasonable to assume that microbial community composition was driven by the variation in the root exudation due to differences in plant species composition of the different plant mixtures. Indeed, a recent microcosm experiment proposed root exudates as an important link between plant richness and the soil microbiome (Steinauer et al. 2016). Furthermore, differences in plant species composition are expected to result in differences in the quantity and quality of above- (leaves, stalks) and belowground (roots) plant litter among plant mixtures which have been identified as a driver of soil microbial communities (e.g. Allison et al. 2005) and could thus have contributed to the observed community shifts.
The soil bacterial community composition was strongly affected by the plant mixture at phylum level (Fig. 5). There are several studies showing that microbial community composition drives ecosystem functions (Wagg et al. 2014), whereas the concept of functional redundancy within microbial systems (Louca et al. 2018) challenges this relationship. Currently, there is no consensus on the relationship between microbial composition and microbiome functionally as some recent studies from distinct environments demonstrated (Fierer et al. 2013; Galand et al. 2018). Another issue connected to this is that linking microbial identities to functional potentials of the microbiome remains challenging (Fierer 2017). Although there are several tools to predict functional potential profiles from the taxonomical profiles of microbiome data sets (Djemiel et al. 2022), we decided to not use these tools because microbiome data generated from short-read amplicons may not be suitable to accurately predict microbiome functions (Heidrich and Beule 2022). Instead, we suggest that future studies should measure actual microbial processes in flower strips and link these with microbiome data in order to test whether flower strips alter the functionality of the soil microbiome. The plant mixtures (Figure SI 3) did not alter functional genes involved in soil-N cycling. Abundances of soil-N-cycling genes have shown pronounced temporal dynamics, which are likely to be determined by plant growth stages (Regan et al. 2017). Since our study comprised of a single sampling point in time, we recommend that future studies should quantify N-cycling microorganisms repeatedly across the vegetation period to capture temporal dynamics of these communities. Finally, we argue that complementing microbiome data obtained from amplicon sequencing with absolute quantification of functional groups of microorganisms is a step towards understanding microbial functions in environmental systems.
In contrast to the differences in beta diversity (i.e. compositional dissimilarities among plant mixtures) discussed above, overall alpha diversity of bacteria and fungi remained mostly unaffected by the plant mixtures (Fig. 3). These results agree with the findings of Prober et al. (2015) who found that plant richness in grasslands is a predictor of beta but not alpha diversity. Alpha diversity of AMF, however, was affected by the plant mixtures (Fig. 6B, C). In addition to the diversity of AMF, plant mixtures also affected the relative abundance of AMF (Fig. 6A). The greater community share and diversity of AMF in the non-tilled (field margin and perennial flower strips) than in the tilled (annual flower strips) plant mixtures (Fig. 6) agrees with previous studies that showed that reduced tillage favors AMF (e.g.Säle et al. 2015; Bowles et al. 2017). Recently, Holden et al. (2019) compared AMF communities in field margins to those in arable land and found that field margins alter AMF community composition and increase AMF diversity as compared to arable land. Few years earlier, Verbruggen et al. (2012) proposed that AMF colonization could take place via different nearby landscape elements such as field margins. Although neighboring croplands were not investigated in this study, we hypothesize that perennial flower strips serve as a reservoir for AMF and enhance AMF colonization of neighboring crops.
Conclusion
Grassy field margins, annual and perennial flower strips harbor distinct earthworm and soil microbial communities. Compared to field margins, earthworm density and biomass declined or remained unaffected in annual flower strips but increased in perennial flower strips. Soil type was the strongest predictor of bacterial and fungal community composition. However, plant mixture (i.e. field margin, annual and perennial flower strips) affected microbiome assembly within each soil type. Although overall alpha diversity of bacteria and fungi remained mostly unaffected by the plant mixtures, AMF showed greater diversity and community share in non-tilled (i.e. field margin and perennial flower strips) as compared to tilled plant mixtures (i.e. annual flower strips). We attribute the observed changes in soil biota mainly to differences in tillage and plant richness. Overall, our data suggests that perennial flower strips serve as refugia for soil biota in agricultural landscapes. Thus, future studies should compare the population size, diversity, and functionality of soil biota in flower strips to those in adjacent agricultural fields in order to assess the belowground benefits of flower strips. Furthermore, we suggest to investigate whether beneficial effects on belowground biota are restricted to the perennial flower strips or extend into adjacent agricultural fields (‘spillover’) as they do for certain aboveground biota. We hope that our work provides a starting point for research on the biodiversity and function of belowground communities in flower strips.
Data Availability
We uploaded our data to different repositories (NCBI and BonaRes Data Centre) and indicated this in the manuscript.
References
Abarenkov K, Zirk A, Piirmann T et al (2021) UNITE QIIME release for eukaryotes. Version 10.05.2021. UNITE Community. https://doi.org/10.15156/BIO/1264819
Ahmed N, Al-Mutairi KA (2022) Earthworms Effect on Microbial Population and Soil Fertility as Well as Their Interaction with Agriculture Practices. Sustainability 14:7803. https://doi.org/10.3390/su14137803
Albrecht M, Kleijn D, Williams NM et al (2020) The effectiveness of flower strips and hedgerows on pest control, pollination services and crop yield: a quantitative synthesis. Ecol Lett 23:1488–1498. https://doi.org/10.1111/ele.13576
Allison VJ, Miller RM, Jastrow JD et al (2005) Changes in Soil Microbial Community Structure in a Tallgrass Prairie Chronosequence. Soil Sci Soc Am J 69:1412–1421. https://doi.org/10.2136/sssaj2004.0252
Badri DV, Vivanco JM (2009) Regulation and function of root exudates. Plant Cell Environ 32:666–681. https://doi.org/10.1111/j.1365-3040.2008.01926.x
Beule L, Arndt M, Karlovsky P (2021) Relative Abundances of Species or Sequence Variants Can Be Misleading: Soil Fungal Communities as an Example. Microorganisms 9:589. https://doi.org/10.3390/microorganisms9030589
Beule L, Corre MD, Schmidt M et al (2019) Conversion of monoculture cropland and open grassland to agroforestry alters the abundance of soil bacteria, fungi and soil-N-cycling genes. PLOS ONE 14:e0218779. https://doi.org/10.1371/journal.pone.0218779
Beule L, Karlovsky P (2021) Tree rows in temperate agroforestry croplands alter the composition of soil bacterial communities. PLOS ONE 16:e0246919. https://doi.org/10.1371/journal.pone.0246919
Beule L, Karlovsky P (2020) Improved normalization of species count data in ecology by scaling with ranked subsampling (SRS): application to microbial communities. PeerJ 8:e9593. https://doi.org/10.7717/peerj.9593
Beule L, Lehtsaar E, Corre MD et al (2020) Poplar Rows in Temperate Agroforestry Croplands Promote Bacteria, Fungi, and Denitrification Genes in Soils. Front Microbiol 10:3108. https://doi.org/10.3389/fmicb.2019.03108
Bhadauria T, Saxena KG (2010) Role of Earthworms in Soil Fertility Maintenance through the Production of Biogenic Structures. Appl Environ Soil Sci 2010:e816073. https://doi.org/10.1155/2010/816073
Blake GR, Hartge KH (1986) Bulk Density. In: Klute A (ed) Methods of Soil Analysis: Part 1—Physical and Mineralogical Methods sss a book series. Wiley, Hoboken, NJ, USA, pp. 363–375. https://doi.org/10.2136/sssabookser5.1.2ed.c13
Blouin M, Hodson ME, Delgado EA et al (2013) A review of earthworm impact on soil function and ecosystem services. Eur J Soil Sci 64:161–182. https://doi.org/10.1111/ejss.12025
Bokulich NA, Kaehler BD, Rideout JR et al (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6:90. https://doi.org/10.1186/s40168-018-0470-z
Bolyen E, Rideout JR, Dillon MR et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Boström U (1995) Earthworm populations (Lumbricidae) in ploughed and undisturbed leys. Soil Tillage Res 35:125–133. https://doi.org/10.1016/0167-1987(95)00489-0
Bouché M (1972) Lombriciens de France. Ecologie et Systématique INRA, Paris
Bowles TM, Jackson LE, Loeher M, Cavagnaro TR (2017) Ecological intensification and arbuscular mycorrhizas: a meta-analysis of tillage and cover crop effects. J Appl Ecol 54:1785–1793. https://doi.org/10.1111/1365-2664.12815
Briones MJI, Schmidt O (2017) Conventional tillage decreases the abundance and biomass of earthworms and alters their community structure in a global meta-analysis. Glob Change Biol 23:4396–4419. https://doi.org/10.1111/gcb.13744
Burrows RL, Pfleger FL (2002) Arbuscular mycorrhizal fungi respond to increasing plant diversity. Can J Bot 80:120–130. https://doi.org/10.1139/b01-138
Callahan BJ, McMurdie PJ, Rosen MJ et al (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Capowiez Y, Bottinelli N, Sammartino S et al (2015) Morphological and functional characterisation of the burrow systems of six earthworm species (Lumbricidae). Biol Fertil Soils 51:869–877. https://doi.org/10.1007/s00374-015-1036-x
Capowiez Y, Cadoux S, Bouchant P et al (2009) The effect of tillage type and cropping system on earthworm communities, macroporosity and water infiltration. Soil Tillage Res 105:209–216. https://doi.org/10.1016/j.still.2009.09.002
Cardinale BJ, Duffy JE, Gonzalez A et al (2012) Biodiversity loss and its impact on humanity. Nature 486:59–67. https://doi.org/10.1038/nature11148
Cardinale BJ, Wright JP, Cadotte MW et al (2007) Impacts of plant diversity on biomass production increase through time because of species complementarity. Proc Natl Acad Sci 104:18123–18128. https://doi.org/10.1073/pnas.0709069104
Chan KY (2001) An overview of some tillage impacts on earthworm population abundance and diversity — implications for functioning in soils. Soil Tillage Res 57:179–191. https://doi.org/10.1016/S0167-1987(00)00173-2
Chen H, Dai Z, Veach AM et al (2020) Global meta-analyses show that conservation tillage practices promote soil fungal and bacterial biomass. Agric Ecosyst Environ 293:106841. https://doi.org/10.1016/j.agee.2020.106841
Clark RB, Zeto SK (2000) Mineral acquisition by arbuscular mycorrhizal plants. J Plant Nutr 23:867–902. https://doi.org/10.1080/01904160009382068
Čoja T, Zehetner K, Bruckner A et al (2008) Efficacy and side effects of five sampling methods for soil earthworms (Annelida, Lumbricidae). Ecotoxicol Environ Saf 71:552–565. https://doi.org/10.1016/j.ecoenv.2007.08.002
Crittenden SJ, Huerta E, de Goede RGM, Pulleman MM (2015) Earthworm assemblages as affected by field margin strips and tillage intensity: An on-farm approach. Eur J Soil Biol 66:49–56. https://doi.org/10.1016/j.ejsobi.2014.11.007
D’Acunto L, Semmartin M, Ghersa CM (2016) Uncultivated margins are source of soil microbial diversity in an agricultural landscape. Agr Ecosyst Environ 220:1–7. https://doi.org/10.1016/j.agee.2015.12.032
Degrune F, Theodorakopoulos N, Dufrêne M et al (2016) No favorable effect of reduced tillage on microbial community diversity in a silty loam soil (Belgium). Agr Ecosyst Environ 224:12–21. https://doi.org/10.1016/j.agee.2016.03.017
DIN 19683–2 (1997) Methods of soil analysis for water management for agricultural purposes - Physical laboratory tests - Determination of grain size distribution after pretreatment using sodium pyrophosphate. Beuth Verlag GmbH, Berlin
DIN Iso 11260 (1997) Soil quality - Determination of effective cation exchange capacity and base saturation level using barium chloride solution. Beuth Verlag GmbH, Berlin
Djemiel C, Maron P-A, Terrat S et al (2022) Inferring microbiota functions from taxonomic genes: a review. Gigascience 11:giab090. https://doi.org/10.1093/gigascience/giab090
Edwards CA, Lofty JR (1982) The Effect of Direct Drilling and Minimal Cultivation on Earthworm Populations. J Appl Ecol 19:723–734. https://doi.org/10.2307/2403277
Ehlers W (1975) Observations on Earthworm Channels and Infiltration on Tilled and Untilled Loess Soil. Soil Sci 119:242–249
Eisenhauer N, Dobies T, Cesarz S et al (2013) Plant diversity effects on soil food webs are stronger than those of elevated CO2 and N deposition in a long-term grassland experiment. Proc Natl Acad Sci 110:6889–6894. https://doi.org/10.1073/pnas.1217382110
Eisenhauer N, König S, Sabais ACW et al (2009) Impacts of earthworms and arbuscular mycorrhizal fungi (Glomus intraradices) on plant performance are not interrelated. Soil Biol Biochem 41:561–567. https://doi.org/10.1016/j.soilbio.2008.12.017
Eisenhauer N, Reich PB (2012) Above- and below-ground plant inputs both fuel soil food webs. Soil Biol Biochem 45:156–160. https://doi.org/10.1016/j.soilbio.2011.10.019
Ernst G, Emmerling C (2009) Impact of five different tillage systems on soil organic carbon content and the density, biomass, and community composition of earthworms after a ten year period. Eur J Soil Biol 45:247–251. https://doi.org/10.1016/j.ejsobi.2009.02.002
Ernst G, Felten D, Vohland M, Emmerling C (2009) Impact of ecologically different earthworm species on soil water characteristics. Eur J Soil Biol 45:207–213. https://doi.org/10.1016/j.ejsobi.2009.01.001
EU Regulation No 1307/2013 of the European Parliament and of the Council of 17 December 2013 establishing rules for direct payments to farmers under support schemes within the framework of the common agricultural policy and repealing Council Regulation (EC) No 637/2008 and Council Regulation (EC) No 73/2009
Euteneuer P, Wagentristl H, Steinkellner S et al (2019) Earthworms affect decomposition of soil-borne plant pathogen Sclerotinia sclerotiorum in a cover crop field experiment. Appl Soil Ecol 138:88–93. https://doi.org/10.1016/j.apsoil.2019.02.020
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590. https://doi.org/10.1038/nrmicro.2017.87
Fierer N, Ladau J, Clemente JC et al (2013) Reconstructing the Microbial Diversity and Function of Pre-Agricultural Tallgrass Prairie Soils in the United States. Science 342:621–624. https://doi.org/10.1126/science.1243768
Frøslev TG, Nielsen IB, Santos SS et al (2022) The biodiversity effect of reduced tillage on soil microbiota. Ambio 51:1022–1033. https://doi.org/10.1007/s13280-021-01611-0
Galand PE, Pereira O, Hochart C et al (2018) A strong link between marine microbial community composition and function challenges the idea of functional redundancy. ISME J 12:2470–2478. https://doi.org/10.1038/s41396-018-0158-1
Gantner S, Andersson AF, Alonso-Sáez L, Bertilsson S (2011) Novel primers for 16S rRNA-based archaeal community analyses in environmental samples. J Microbiol Methods 84:12–18. https://doi.org/10.1016/j.mimet.2010.10.001
Gastine A, Scherer-Lorenzen M, Leadley PW (2003) No consistent effects of plant diversity on root biomass, soil biota and soil abiotic conditions in temperate grassland communities. Appl Soil Ecol 24:101–111. https://doi.org/10.1016/S0929-1393(02)00137-3
Geppert C, Hass A, Földesi R et al (2020) Agri-environment schemes enhance pollinator richness and abundance but bumblebee reproduction depends on field size. J Appl Ecol 57:1818–1828. https://doi.org/10.1111/1365-2664.13682
Guerra V, Beule L, Lehtsaar E et al (2020) Improved Protocol for DNA Extraction from Subsoils Using Phosphate Lysis Buffer. Microorganisms 8:532. https://doi.org/10.3390/microorganisms8040532
Harris D, Horwáth WR, van Kessel C (2001) Acid fumigation of soils to remove carbonates prior to total organic carbon or CARBON-13 isotopic analysis. Soil Sci Soc Am J 65:1853. https://doi.org/10.2136/sssaj2001.1853
Hedlund K, Santa Regina I, Van der Putten WH et al (2003) Plant species diversity, plant biomass and responses of the soil community on abandoned land across Europe: idiosyncracy or above-belowground time lags. Oikos 103:45–58. https://doi.org/10.1034/j.1600-0706.2003.12511.x
Heidrich V, Beule L (2022) Are short-read amplicons suitable for the prediction of microbiome functional potential? A critical perspective. iMeta 1:e38. https://doi.org/10.1002/imt2.38
Heidrich V, Karlovsky P, Beule L (2021) ‘SRS’ R Package and ‘q2-srs’ QIIME 2 Plugin: Normalization of Microbiome Data Using Scaling with Ranked Subsampling (SRS). Appl Sci 11:11473. https://doi.org/10.3390/app112311473
Holden J, Grayson RP, Berdeni D et al (2019) The role of hedgerows in soil functioning within agricultural landscapes. Agr Ecosyst Environ 273:1–12. https://doi.org/10.1016/j.agee.2018.11.027
IUSS Working Group WRB (2015) World Reference Base for Soil Resources 2014, update 2015 International soil classification system for naming soils and creating legends for soil maps. World Soil Resources Reports No. 106. FAO, Rome
Kleijn D, Kohler F, Báldi A et al (2009) On the relationship between farmland biodiversity and land-use intensity in Europe. Proc R Soc B: Biol Sci 276:903–909. https://doi.org/10.1098/rspb.2008.1509
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1. https://doi.org/10.1093/nar/gks808
Kohli L, Daniel O, Schönholzer F et al (1999) Miscanthus sinensis and wild flowers as food resources of Lumbricus terrestris L. Appl Soil Ecol 11:189–197. https://doi.org/10.1016/S0929-1393(98)00137-1
Krück SS (2018) Bildatlas zur Regenwurmbestimmung: mit einem Kompendium der Regenwurmfauna des Norddeutschen Tieflands. Rangsdorf, Natur+Text
Lees KJ, McKenzie AJ, Newell Price JP et al (2016) The effects of soil compaction mitigation on below-ground fauna: How earthworms respond to mechanical loosening and power harrow cultivation. Agr Ecosyst Environ 232:273–282. https://doi.org/10.1016/j.agee.2016.07.026
Li Y, Song D, Liang S et al (2020) Effect of no-tillage on soil bacterial and fungal community diversity: A meta-analysis. Soil Tillage Res 204:104721. https://doi.org/10.1016/j.still.2020.104721
Louca S, Polz MF, Mazel F et al (2018) Function and functional redundancy in microbial systems. Nat Ecol Evol 2:936–943. https://doi.org/10.1038/s41559-018-0519-1
Marinissen JCY (1992) Population dynamics of earthworms in a silt loam soil under conventional and “integrated” arable farming during two years with different weather patterns. Soil Biol Biochem 24:1647–1654. https://doi.org/10.1016/0038-0717(92)90164-S
Mathew RP, Feng Y, Githinji L et al (2012) Impact of No-Tillage and Conventional Tillage Systems on Soil Microbial Communities. Appl Environ Soil Sci 2012:548620. https://doi.org/10.1155/2012/548620
Medina-Sauza RM, Álvarez-Jiménez M, Delhal A et al (2019) Earthworms Building Up Soil Microbiota, a Review. Front Environ Sci 7:81. https://doi.org/10.3389/fenvs.2019.00081
Milcu A, Partsch S, Langel R, Scheu S (2006) The response of decomposers (earthworms, springtails and microorganisms) to variations in species and functional group diversity of plants. Oikos 112:513–524. https://doi.org/10.1111/j.0030-1299.2006.14292.x
Milcu A, Partsch S, Scherber C et al (2008) Earthworms and Legumes Control Litter Decomposition in a Plant Diversity Gradient. Ecology 89:1872–1882. https://doi.org/10.1890/07-1377.1
Pagliai M, Vignozzi N, Pellegrini S (2004) Soil structure and the effect of management practices. Soil Tillage Res 79:131–143. https://doi.org/10.1016/j.still.2004.07.002
Palm J, van Schaik NLMB, Schröder B (2013) Modelling distribution patterns of anecic, epigeic and endogeic earthworms at catchment-scale in agro-ecosystems. Pedobiologia 56:23–31. https://doi.org/10.1016/j.pedobi.2012.08.007
Paoletti MG (1999) The role of earthworms for assessment of sustainability and as bioindicators. Agr Ecosyst Environ 74:137–155. https://doi.org/10.1016/S0167-8809(99)00034-1
Pelosi C, Bertrand M, Capowiez Y et al (2009) Earthworm collection from agricultural fields: Comparisons of selected expellants in presence/absence of hand-sorting. Eur J Soil Biol 45:176–183. https://doi.org/10.1016/j.ejsobi.2008.09.013
Pelosi C, Pey B, Hedde M et al (2014) Reducing tillage in cultivated fields increases earthworm functional diversity. Appl Soil Ecol 83:79–87. https://doi.org/10.1016/j.apsoil.2013.10.005
Plaas E, Meyer-Wolfarth F, Banse M et al (2019) Towards valuation of biodiversity in agricultural soils: A case for earthworms. Ecol Econ 159:291–300. https://doi.org/10.1016/j.ecolecon.2019.02.003
Prober SM, Leff JW, Bates ST et al (2015) Plant diversity predicts beta but not alpha diversity of soil microbes across grasslands worldwide. Ecol Lett 18:85–95. https://doi.org/10.1111/ele.12381
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Regan K, Stempfhuber B, Schloter M et al (2017) Spatial and temporal dynamics of nitrogen fixing, nitrifying and denitrifying microbes in an unfertilized grassland soil. Soil Biol Biochem 109:214–226. https://doi.org/10.1016/j.soilbio.2016.11.011
Reichle DE (1977) The Role of Soil Invertebrates in Nutrient Cycling. Ecol Bull 25:145–156
Rousk J, Bengtson P (2014) Microbial regulation of global biogeochemical cycles. Front Microbiol 5:103. https://doi.org/10.3389/fmicb.2014.00103
Säle V, Aguilera P, Laczko E et al (2015) Impact of conservation tillage and organic farming on the diversity of arbuscular mycorrhizal fungi. Soil Biol Biochem 84:38–52. https://doi.org/10.1016/j.soilbio.2015.02.005
Schütz L, Wenzel B, Rottstock T et al (2022) How to promote multifunctionality of vegetated strips in arable farming: A qualitative approach for Germany. Ecosphere 13:e4229. https://doi.org/10.1002/ecs2.4229
Sechi V, De Goede RGM, Rutgers M et al (2017) A community trait-based approach to ecosystem functioning in soil. Agr Ecosyst Environ 239:265–273. https://doi.org/10.1016/j.agee.2017.01.036
Simonsen J, Posner J, Rosemeyer M, Baldock J (2010) Endogeic and anecic earthworm abundance in six Midwestern cropping systems. Appl Soil Ecol 44:147–155. https://doi.org/10.1016/j.apsoil.2009.11.005
Smith CR, Blair PL, Boyd C et al (2016) Microbial community responses to soil tillage and crop rotation in a corn/soybean agroecosystem. Ecol Evol 6:8075–8084. https://doi.org/10.1002/ece3.2553
Smith J, Potts SG, Woodcock BA, Eggleton P (2008) Can arable field margins be managed to enhance their biodiversity, conservation and functional value for soil macrofauna? J Appl Ecol 45:269–278. https://doi.org/10.1111/j.1365-2664.2007.01433.x
Spehn EM, Joshi J, Schmid B et al (2000) Plant diversity effects on soil heterotrophic activity in experimental grassland ecosystems. Plant Soil 224:217–230. https://doi.org/10.1023/A:1004891807664
Steinauer K, Chatzinotas A, Eisenhauer N (2016) Root exudate cocktails: the link between plant diversity and soil microorganisms? Ecol Evol 6:7387–7396. https://doi.org/10.1002/ece3.2454
Sun R, Li W, Dong W et al (2018) Tillage Changes Vertical Distribution of Soil Bacterial and Fungal Communities. Front Microbiol 9:699. https://doi.org/10.3389/fmicb.2018.00699
Tilman D, Isbell F, Cowles JM (2014) Biodiversity and Ecosystem Functioning. Annu Rev Ecol Evol Syst 45:471–493. https://doi.org/10.1146/annurev-ecolsys-120213-091917
Toju H, Tanabe AS, Yamamoto S, Sato H (2012) High-Coverage ITS Primers for the DNA-Based Identification of Ascomycetes and Basidiomycetes in Environmental Samples. PLOS ONE 7:e40863. https://doi.org/10.1371/journal.pone.0040863
Tomati U, Galli E (1995) Earthworms, soil fertility and plant productivity. Acta Zool Fennica 196:11–14
Torppa KA, Taylor AR (2022) Alternative combinations of tillage practices and crop rotations can foster earthworm density and bioturbation. Appl Soil Ecol 175:104460. https://doi.org/10.1016/j.apsoil.2022.104460
Tóth Z, Szlavecz K, Epp Schmidt DJ et al (2020) Earthworm assemblages in urban habitats across biogeographical regions. Appl Soil Ecol 151:103530. https://doi.org/10.1016/j.apsoil.2020.103530
Tschumi M, Albrecht M, Entling MH, Jacot K (2015) High effectiveness of tailored flower strips in reducing pests and crop plant damage. Proc R Soc B: Biol Sci 282:20151369. https://doi.org/10.1098/rspb.2015.1369
van der Heijden MGA, Bardgett RD, van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310. https://doi.org/10.1111/j.1461-0248.2007.01139.x
Vancov T, Keen B (2009) Amplification of soil fungal community DNA using the ITS86F and ITS4 primers. FEMS Microbiol Lett 296:91–96. https://doi.org/10.1111/j.1574-6968.2009.01621.x
Vaupel A, Bednar Z, Herwig N et al (2023) Tree-distance and tree-species effects on soil biota in a temperate agroforestry system. Plant Soil 2023:1–18. https://doi.org/10.1007/s11104-023-05932-9
VDLUFA (1991a) Determination of phosphorus and potassium in the double lactate (DL) extract. In: VDLUFA method book I, A 6.2.1.2. VDLUFA-Verlag (in German), Darmstadt, Germany
VDLUFA (1991b) Determination of plant-available magnesium in the calcium chloride extract. In: VDLUFA method book I, A 6.2.4.1. VDLUFA-Verlag (in German), Darmstadt, Germany
Verbruggen E, Van Der HEIJDENMGA, Weedon JT et al (2012) Community assembly, species richness and nestedness of arbuscular mycorrhizal fungi in agricultural soils. Mol Ecol 21:2341–2353. https://doi.org/10.1111/j.1365-294X.2012.05534.x
Vives-Peris V, de Ollas C, Gómez-Cadenas A, Pérez-Clemente RM (2020) Root exudates: from plant to rhizosphere and beyond. Plant Cell Rep 39:3–17. https://doi.org/10.1007/s00299-019-02447-5
Wagg C, Bender SF, Widmer F, van der Heijden MGA (2014) Soil biodiversity and soil community composition determine ecosystem multifunctionality. PNAS 111:5266–5270. https://doi.org/10.1073/pnas.1320054111
Wardle DA, Bonner KI, Barker GM et al (1999) Plant Removals in Perennial Grassland: Vegetation Dynamics, Decomposers, Soil Biodiversity, and Ecosystem Properties. Ecol Monogr 69:535–568. https://doi.org/10.1890/0012-9615(1999)069[0535:PRIPGV]2.0.CO;2
Wolfarth F, Schrader S, Oldenburg E et al (2011) Earthworms promote the reduction of Fusarium biomass and deoxynivalenol content in wheat straw under field conditions. Soil Biol Biochem 43:1858–1865. https://doi.org/10.1016/j.soilbio.2011.05.002
Yin C, Mueth N, Hulbert S et al (2017) Bacterial Communities on Wheat Grown Under Long-Term Conventional Tillage and No-Till in the Pacific Northwest of the United States. Phytobiomes Journal 1:83–90. https://doi.org/10.1094/PBIOMES-09-16-0008-R
Young IM, Ritz K (2000) Tillage, habitat space and function of soil microbes. Soil Tillage Res 53:201–213. https://doi.org/10.1016/S0167-1987(99)00106-3
Zaller JG, Arnone JA (1999) Earthworm responses to plant species’ loss and elevated CO2 in calcareous grassland. Plant Soil 208:1–8. https://doi.org/10.1023/A:1004424720523
Acknowledgements
The authors would like to thank Josef Beule for participating in soil sampling.
Funding
Open Access funding enabled and organized by Projekt DEAL. AV was supported by the joint project MonViA—the German Farmland Biodiversity Monitoring that has been funded by the Federal Ministry of Food and Agriculture (BMEL). The funder had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Open access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
ZB, AV, SB, VHK, and LB contributed to the conception and design of the study. ZB, AV, and LB performed the field work. ZB, AV, and LB performed the laboratory work. ZB, AV, and LB performed the statistical analysis. ZB, AV, SB, and LB wrote the first draft of the manuscript. NH, BH, and VHK contributed resources and critically revised the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Additional information
Responsible Editor: Rémi Cardinael.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11104_2023_6166_MOESM2_ESM.pdf
Supplementary file2 Earthworm biomass. Biomass (g m−2) of ecological groups of earthworms. Bars represent individual replicate plots (n = 3). (PDF 27 KB)
11104_2023_6166_MOESM3_ESM.pdf
Supplementary file3 Relative change of (A) soil archaea, (B) bacteria, and (C) fungi in response to flower strips. Non-transparent dots and triangles represent means and vertical bars range from the minimum to the maximum value (n = 3). Transparent dots and triangles represent individual data points (i.e. replicate plots). Archaea, bacteria, and fungi were quantified by using real-time PCR (see Quantification of soil microbial groups using real-time PCR for details). See Statistical analysis for details regarding the calculation of the relative change. Images are courtesy of the Integration and Application Network (ian.umces.edu/media-library). (PDF 4426 KB)
11104_2023_6166_MOESM4_ESM.pdf
Supplementary file4 Relative change of ammonia-oxidizing archaea (AOA) amoA (A), nirS (B), nosZ clade I (C), and nosZ clade II genes (D) in response to flower strips. Non-transparent dots and triangles represent means and vertical bars range from the minimum to the maximum value (n = 3). Transparent dots and triangles represent individual data points (i.e. replicate plots). AOA amoA, nirS, and nosZ clade I and II genes were quantified by using real-time PCR (see Quantification of soil microbial groups using real-time PCR for details). See Statistical analysis for details regarding the calculation of the relative change. Images are courtesy of the Integration and Application Network (ian.umces.edu/media-library). (PDF 4432 KB)
11104_2023_6166_MOESM5_ESM.pdf
Supplementary file5 Non-metric multidimensional scaling (NMDS) of Bray-Curtis dissimilarities of soil bacterial (A) and fungal (B) communities. Dots and triangles represent individual data points (i.e. replicate plots) (n = 3) which are connected with the centroid of their respective plant mixture. (PDF 49 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bednar, Z., Vaupel, A., Blümel, S. et al. Earthworm and soil microbial communities in flower strip mixtures. Plant Soil 492, 209–227 (2023). https://doi.org/10.1007/s11104-023-06166-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-06166-5