Abstract
Purpose
The important role of viral communities in various environments is being continuously revealed. However, the current understanding of viruses in soil ecosystem remains poor, especially in plant continuous cropping soils.
Methods
Here, we provide detailed information about prokaryotic and viral diversity, the functional classes of viral ORFs and virus-host linkage in Coptis chinensis continuously cropped fields and uncultivated fields by analyzing physiochemical characteristics, 16S rRNA gene sequencing and viromic data.
Results
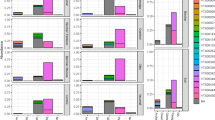
Among prokaryotes, some specific bacteria, especially plant pathogens of the order Burkholderiales, were enriched in soil due to the continuous cropping of C. chinensis. For viruses, we recovered 1,029 viral contigs from continuous cropping and uncultivated soil samples. Continuous cropping of C. chinensis significantly influenced the diversity and abundance of soil viral communities and viral protein functions. Furthermore, we linked 9.06% of viral contigs to putative hosts, spanning 7 bacterial phyla and 2 archaeal phyla. Unfortunately, the specific bacteria enriched in C. chinensis continuous cropping soils were not predicted to be prokaryotic hosts of the recovered viruses.
Conclusion
Overall, these findings reveal the effects of C. chinensis continuous cropping on the diversity and composition of the prokaryotic and viral communities, and indicated how virus-host interactions influence the soil microbes in continuous cropping systems.






Similar content being viewed by others
Abbreviations
- OTU:
-
Operational taxonomic unit
- ORF:
-
Open reading frame
- PCR:
-
Polymerase chain reaction
- DNA:
-
Deoxyribonucleic acid
- RNA:
-
Ribonucleic acid
References
Alami MM, Xue J, Ma Y, Zhu D, Abbas A, Gong Z, Wang X (2020a) Structure, function, diversity, and composition of fungal communities in rhizospheric soil of coptis chinensis Franch under a successive cropping system. Plants 9:244
Alami MM, Xue J, Ma Y, Zhu D, Gong Z, Shu S, Wang X (2020b) Structure, diversity, and composition of bacterial communities in rhizospheric soil of coptis chinensis franch under continuously cropped fields. Diversity 12:57
Bland C, Ramsey TL, Sabree F, Lowe M, Brown K, Kyrpides NC, Hugenholtz P (2007) CRISPR recognition tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinformatics 8:1–8
Calero-Cáceres W, Balcázar JL (2019) Antibiotic resistance genes in bacteriophages from diverse marine habitats. Sci Total Environ 654:452–455
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:1–9
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Chan PP, Lin BY, Mak AJ, Lowe TM (2021) tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res 49:9077–9096
Chen Y, Chen Y, Shi C, Huang Z, Zhang Y, Li S, Li Y, Ye J, Yu C, Li Z (2018) SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience 7:gix120
Chen P, Wang YZ, Liu QZ, Zhang YT, Li XY, Li HQ, Li WH (2020) Phase changes of continuous cropping obstacles in strawberry (Fragaria × ananassa Duch.) production. Appl Soil Ecol 155:103626
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642
Coutinho FH, Rosselli R, Rodriguez-Valera F (2019) Trends of microdiversity reveal depth-dependent evolutionary strategies of viruses in the Mediterranean. mSystems 4:e00554-19
Dubey A, Malla MA, Khan F, Chowdhary K, Yadav S, Kumar A, Sharma S, Khare PK, Khan ML (2019) Soil microbiome: a key player for conservation of soil health under changing climate. Biodivers Conserv 28:2405–2429
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol 7:e1002195
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
El-Tarabily KA, Sivasithamparam K (2006) Non-streptomycete actinomycetes as biocontrol agents of soil-borne fungal plant pathogens and as plant growth promoters. Soil Biol Biochem 38:1505–1520
Emerson JB (2019) Soil viruses: a new hope. mSystems 4:e00120-e119
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152
Gao Z, Han M, Hu Y, Li Z, Liu C, Wang X, Tian Q, Jiao W, Hu J, Liu L (2019) Effects of continuous cropping of sweet potato on the fungal community structure in rhizospheric soil. Front Microbiol 10:2269
Hendrix RW, Smith MC, Burns RN, Ford ME, Hatfull GF (1999) Evolutionary relationships among diverse bacteriophages and prophages: all the world’s a phage. Proc Natl Acad Sci USA 96:2192–2197
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, von Mering C, Bork P (2017) Fast genome-wide functional annotation through orthology assignment by eggNOG-mapper. Mol Biol Evol 34:2115–2122
Hyatt D, LoCascio PF, Hauser LJ, Uberbacher EC (2012) Gene and translation initiation site prediction in metagenomic sequences. Bioinformatics 28:2223–2230
Imachi H, Nobu MK, Nakahara N, Morono Y, Takai KJN (2020) Isolation of an archaeon at the prokaryote–eukaryote interface. Nature 577:519–525
Jin M, Guo X, Zhang R, Qu W, Gao B, Zeng R (2019) Diversities and potential biogeochemical impacts of mangrove soil viruses. Microbiome 7:58
Kuzyakov Y, Mason-Jones K (2018) Viruses in soil: nano-scale undead drivers of microbial life, biogeochemical turnover and ecosystem functions. Soil Biol Biochem 127:305–317
Li D, Liu CM, Luo R, Sadakane K, Lam TW (2015) MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31:1674–1676
Li H (2013) Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:1303.3997
Liu L, Gong T, Tao W, Lin B, Li C, Zheng X, Zhu S, Jiang W, Zhou R (2019) Commensal viruses maintain intestinal intraepithelial lymphocytes via noncanonical RIG-I signaling. Nat Immunol 20:1681–1691
Liu S, Wang Z, Niu J, Dang K, Zhang S, Wang S, Wang Z (2021) Changes in physicochemical properties, enzymatic activities, and the microbial community of soil significantly influence the continuous cropping of Panax quinquefolius L. (American ginseng). Plant Soil 463:427–446
Malki K, Rosario K, Sawaya NA, Székely AJ, Tisza MJ, Breitbart M (2020) Prokaryotic and viral community composition of freshwater springs in Florida, USA. Mbio 11:e00436-e420
Min J, Xun G, Rui Z, Wu Q, Boliang G, Runying Z (2019) Diversities and potential biogeochemical impacts of mangrove soil viruses. Microbiome 7:58
Nicola S, Jacques I, Levi W, Dirk G, Larisa M, Garrett WS, Curtis H (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60
O’Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B, Smith-White B, Ako-Adjei D (2016) Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res 44:D733–D745
Paez-Espino D, Eloe-Fadrosh EA, Pavlopoulos GA, Thomas AD, Huntemann M, Mikhailova N, Rubin E, Ivanova NN, Kyrpides NC (2016) Uncovering earth’s virome. Nature 536:425–430
Pérez-Pantoja D, Donoso R, Agulló L, Córdova M, Seeger M, Pieper DH, González B (2012) Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environ Microbiol 14:1091–1117
Pervaiz ZH, Iqbal J, Zhang Q, Chen D, Wei H, Saleem M (2020) Continuous cropping alters multiple biotic and abiotic indicators of soil health. Soil Syst 4:59
Pratama AA, Elsas JDv (2018) The ‘neglected’ soil virome – potential role and impact. Trends Microbiol 26:649–662
Tisza MJ, Pastrana DV, Welch NL, Stewart B, Peretti A, Starrett GJ, Pang YYS, Krishnamurthy SR, Pesavento PA, McDermott DH (2020) Discovery of several thousand highly diverse circular DNA viruses. Elife 9:e51971
Wagner GP, Kin K, Lynch VJ (2012) Measurement of mRNA abundance using RNA-seq data: RPKM measure is inconsistent among samples. Theory Biosci 131:281–285
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang B, Liu GB, Xue S, Zhu BB (2011) Changes in soil physico-chemical and microbiological properties during natural succession on abandoned farmland in the Loess Plateau. Environmental Earth Sciences 62:915–925
Wang W, Ren J, Tang K, Dart E, Ignacio-Espinoza JC, Fuhrman JA, Braun J, Sun F, Ahlgren NA (2020) A network-based integrated framework for predicting virus–prokaryote interactions. NAR Genom Bioinform 2:lqaa044
Warwick-Dugdale J, Buchholz HH, Allen MJ, Temperton B (2019) Host-hijacking and planktonic piracy: How phages command the microbial high seas. Virol J 16:15
Zhang W, Long XQ, Huo XD, Chen YF, Lou K (2013) 16S rRNA-Based PCR-DGGE analysis of actinomycete communities in fields with continuous cotton cropping in Xinjiang, China. Microb Ecol 66:385–393
Zhang Y, Gu Y, Ren H, Wang S, Zhong H, Zhao X, Ma J, Gu X, Xue Y, Huang S, Yang J, Chen L, Chen G, Qu S, Liang J, Qin L, Huang Q, Peng Y, Li Q, Wang X, Kong P, Hou G, Gao M, Shi Z, Li X, Qiu Y, Zou Y, Yang H, Wang J, Xu G, Lai S, Li J, Ning G, Wang W (2020) Gut microbiome-related effects of berberine and probiotics on type 2 diabetes (the PREMOTE study). Nat Commun 11:5015
Zimmerman AE, Howard-Varona C, Needham DM, John SG, Worden AZ, Sullivan MB, Waldbauer JR, Coleman ML (2020) Nat Rev Microbiol 18:21–34
Acknowledgements
This work is supported by the National Key Research and Development Program of China for Traditional Chinese Medicine Modernization (2017YFC1702600), the National Natural Science Foundation (31600148), and the Natural Science Foundation of Shandong Province (ZR2021MC018). We would like to thank James Voordeckers for English language editing.
Author information
Authors and Affiliations
Contributions
Xiangyu Fan and Changhua Hu designed the research; Xiangyu Fan, Mengzhi Ji, Muyuan Li, Kaili Sun, Zhen Tian, Rongfeng Gao and Yang Liu performed research and analyzed the data; Xiangyu Fan wrote the paper. Guojian Liao helped to modify the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper. The authors declare no conflict of interest.
Additional information
Responsible Editor: Stéphane Compant.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Figure S1.
The picture of experimental settings. (a) Original soil. (b) Continuous cropping soil. (PNG 4987 kb)

Figure S2.
The refraction curve with maximum read depth for the 16S rRNA gene sequences. (PNG 158 kb)

Figure S3.
Virus-host interaction network in genus level. Hexagons represent viruses and circles represent hosts. Blue hexagons represent viruses which lie in both soil types. Cyan hexagons represent viruses which lie in Original_soil sample. Purple hexagons represent viruses which lie in CC_soil sample. Line represents interaction between virus and its host(s). (PNG 124 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fan, X., Ji, M., Li, M. et al. Composition of prokaryotic and viral community in continuously cropped field of Coptis chinensis Franch. Plant Soil 481, 97–109 (2022). https://doi.org/10.1007/s11104-022-05620-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05620-0