Abstract
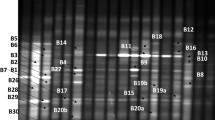
The purpose of this study was to examine the variations in the microbial community structure of soil actinomycetes in fields with continuous cropping of cotton in Xinjiang Autonomous Region, China. Soil samples were collected from four depths in fields with 7-year continuous cotton cropping. The community structure of soil actinomycetes was examined using the 16S rRNA-based polymerase chain reaction–density gradient gel electrophoresis (PCR-DGGE) techniques. The microbial diversity indices of the soil samples from different depths generally decreased along with the period of continuous cotton cropping. When the period of continuous cropping of cotton reached 5 years, the diversity indices rose again and gradually stabilized at a level slightly lower than that of soils with original ecology (i.e., 0-year cotton cropping). Cluster analysis showed that at the 1–20-cm depth, the actinomycete community structure of the soil subjected to 1-year cotton cropping was similar to that of soil subjected to 0-year cotton cropping, whereas that of soils after 3-year continuous cotton cropping showed high similarity. At the 21–40-cm depth, the actinomycete community structure showed various changes but generally recovered to its original pattern after repeated fluctuations. Principal component analysis showed that at the 1–30-cm depth, the actinomycete community structure varied similarly regardless of the period of continuous cotton cropping. In contrast, there were no clear actinomycete community structure variation trends at the 31–40-cm soil depth. Homology comparison of sequences recovered from the DGGE bands showed that the obtained sequences shared similarities >88 %. Alignment with the known homologous sequences indicated a lack of microorganisms related to soil-borne cotton diseases. Continuous cotton cropping exerted significant influences on the community structure of soil actinomycetes in Xinjiang Autonomous Region, which were largely determined by the soil depth and the period of continuous cotton cropping. The microbial diversity of soil actinomycete communities gradually recovered after 5-year continuous cropping. Thereafter, a new actinomycete community structure that was beneficial for continuous cropping of cotton was formed and stabilized each year.






Similar content being viewed by others
References
Han CL, Liu J, Zhang WF, Liu M, Huang WJ, Gao XM, Zhang HZ (2010) Biocycling of nine mineral elements of soil-cotton system in Xinjiang oasis. Acta Ecologica Sinica 30(22):6234–6241
Vargas GS, Meriles J, Conforto C, Figoni G, Basanta M, Lovera E, March GJ (2009) Field assessment of soil biological and chemical quality in response to crop management practices. World J Microb Biot 25(3):439–448
He JZ, Zheng Y, Chen CR (2010) Microbial composition and diversity of an upland red soil under long-term fertilization treatments as revealed by culture-dependent and culture-independent approaches. J Soil Sediment 8:349–358
Fu QL, Liu C, Ding NF, Lin YC, Guo B, Luo JF, Wang HL (2012) Soil microbial communities and enzyme activities in a reclaimed coastal soil chromosequence under rice-barley cropping. J Soil Sediment 12:1134–1144
Alvey S, Yang CH, Buerkert A (2003) Cereal legume rotation effects on rhizosphere bacterial community structure in west African soils. Biol Fert Soils 37:73–82
Martin I, Mun LM, Yunta F (2007) Tillage and croprotation effects on barley yield and soil nutrients on a Calciortidic Haploxeralf. Soil Till Res 92:1–9
Meriles JM, Vargas Gil S, Haro R, March GJ, Guzman CA (2008) Selected soil-borne fungi under glyphosate application and crop residues from a long-term field experiment. Biol Agric Hortic 26(2):193–205
Garbeva P, van Veen JA, van Elsas JD (2004) Microbial diversity in soil: selection of microbial populations by plant and soil type and implications for disease suppressiveness. Annu Rev Phytopathol 42:243–270
Huang JW, Yang JK, Duan YQ (2010) Bacterial diversities on unaged and aging flue-cured tobacco leaves estimated by 16S rRNA sequence analysis. Applied Microbiol Biot 88(2):553–562
Moreira D (1998) Efficient removal of PCR inhibitors using agarose-embedded DNA preparations. Nucleic Acids Res 26:3309–3310
Bruce KD, Hiorns WD, Hobman JL, Osborn AM, Strike P, Ritchie DA (1992) Amplification of DNA from native populations of soil bacteria by using the polymerase chain reaction. Appl Environ Microb 58:3413–3416
Martina K, Jan K, Tamas F, Ladislav C, Marek O, Genevieve LG, Yvan ML, Marketa SM (2008) Development of a 16S rRNA gene-based prototype microarray for the detection of selected actinomycetes genera. Anto Leeuw Int J G 94:439–453
Ovreas L, Fomey L, Daae FL (1997) Distribution of bacterioplankton in meromictic lake saelevannet, as determined by denaturing gradient gel electrophoresis of PCR. Amplified gene fragments coding for 16S rRNA. Appl Environ Microb 63:3367–3373
Graham MH, Haynes RJ (2005) Catabolic diversity of soil microbial communities under sugarcane and other land uses estimated by biolog and substrate-induced respiration methods. Appl Soil Ecol 29:155–164
Hill TCJ, Walsh KA, Harris JA (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiol Ecol 43:1–11
Daniela RDF, Raquel VF, Mario C, Teresa CDM, Marrio JP, Bruno BC, Antonio C (2012) Impact of water quality on bacterioplankton assemblage along certima river basin (central western Portugal) accessed by PCR-DGGE and multivariate analysis. Environ Monit Assess 184:471–485
Li C, Li XM, Kong WD (2010) Effect of monoculture soybean on soil microbial community in the northeast china. Plant Soil 330:423–433
Zhang XL, Li X, Zhan CG (2011) Ecological risk of long-term chlorimuron-ethyl application to soil microbial community:an in situ investigation in a continuously cropped soybean field in northeast china. Environ Sci Pollut R 18:407–415
Girvan MS, Bullimore J, Pretty JN, Osborn AM, Ball AS (2003) Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Appl Environ Microb 69:1800–1809
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40:2762–2770
Zhang Y, Du BH, Jin ZG, Li ZH, Song HN, Ding YQ (2011) Analysis of bacterial communities in rhizosphere soil of healthy and diseased cotton (Gossypium sp.) at different plant growth stages. Plant Soil 339:447–455
Robert P, Larkin (2008) Relative effects of biological amendments and crop rotations on soil microbial communities and soilborne diseases of potato. Soil Biol Biochem 40:1341–1351
Berg G, Smalla K (2009) Plant species and soil type cooperativelyshape the structure and function of microbial communities in the rhizosphere FEMS. Microbiol Ecol 68:1–13
Wei XR, Hao MD, Shao MA (2006) Changes in soil properties and the availability of soil micronutrients after 18 years of cropping and fertilization. Soil Till Res 91:120–130
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microb 59(3):695–700
Ge Y, Zhang JB, Zhang LM, Yang M, He JZ (2008) Long-term fertilization regimes and diversity of an agricultural affect bacterial community structure soil in northern china. J Soil Sediment 8:43–50
Acknowledgments
This study was supported by the National Natural Science Foundation of China (30860016) and the key disciplines of Xinjiang Normal University (Microbiology).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, W., Long, X., Huo, X. et al. 16S rRNA-Based PCR-DGGE Analysis of Actinomycete Communities in Fields with Continuous Cotton Cropping in Xinjiang, China. Microb Ecol 66, 385–393 (2013). https://doi.org/10.1007/s00248-012-0160-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-012-0160-5