Abstract
Even for processes with only a few gas species involved the detailed description of plasma-assisted conversion processes in gas mixtures requires a large amount of processes to be taken into account and a large number of neutral and charged particles must be considered. In addition, setting up the corresponding reaction kinetics model needs the knowledge of the rate coefficients and their temperature dependence for all possible reactions between those species. Reduced reaction networks offer a simplified and pragmatic way to obtain an overall reaction kinetics model, already useful for the analysis of experimental data even if not all details of chemistry can be covered. In this paper we present a derivation of a data driven reduced model for plasma-assisted conversion of methane in an helium environment. By consideration of a small number of elementary reactions, a simple model is set up. Experimental data are analyzed by a genetic algorithm that provides best-fit approximations for the open parameters of the model. In a further step non-relevant parameters of the model are identified and a further model reduction is achieved. The data driven analysis of methane conversion serves as an illustrative example of the proposed method. The parameters and reaction channels found are compared with known results from the literature. The method is described in detail. The main goal of this work is to present the potential of this data driven method for a simplified and pragmatic modeling in the increasingly important field of plasma-assisted catalytic processes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reaction kinetic models allow far-reaching predictions of complex chemical processes. The description on the basis of reaction kinetics has a tradition of decades and an enormous amount of literature exists on this topic and special applications. A starting point for the reader to get more information on this might be Refs. [1, 2] and the references therein. Still, the determination of rate coefficients is in the center of attention when creating reaction kinetic models. Although several quantum mechanical approaches allow for a number of conclusions, it is not always possible to build a complete reaction kinetic model completely on the basis of ab initio theories. Furthermore, extensive statistical analysis of experimental data is required to support or supplement theories. This is mostly complicated by large numbers of species and reactions involved. The present work discusses an approach for data driven analysis of experimental data of plasma-assisted processes, which allows to derive reduced models in a simple way. When selecting a data evaluation method it is essential to know whether a detailed knowledge of the species involved is available or whether only a few of the reactants and products involved can be observed. Furthermore, it makes a decisive difference whether a time series of concentrations of individual species is available or only stationary states can be measured. In order to cope with these difficulties, we focus on the application of genetic algorithms to analyze the experimental data. The flexibility and robustness of the method is reflected by an enormous number of publications from various scientific disciplines on the application and concrete design of genetic algorithms. Useful references on the basic ideas and practical implementation are given by Refs. [3,4,5,6,7]. Examples of applications in the field of reaction kinetics can be found in Refs. [7,8,9,10,11,12,13,14,15,16]. Especially Lapene et al. [16] give a very good overview of applications and methods in chemical physics. These references are given to name just a few and without claiming to be exhaustive. The particular details of the method depend on the problem at hand. In the present data driven analysis we face the problem of being able to track only a small number of species in the plasma-assisted conversion of methane, but with a rather good time resolution. In addition, data are available from different experiments in which individual reaction parameters were varied. The goal is, on the one hand, to find a suitable set of only a few reactions to describe the conversion process, and on the other hand, to find the appropriate rate coefficients. Both can be achieved by the genetic algorithm that is presented. In Sect. 2 the concrete experiment and the data available are described. Then, in Sect. 3 the details of the method are presented, i.e. how the reaction kinetic model is set up and how the genetic algorithm is constructed. Sect. 4 contains numerical results and Sect. 5 a discussion of the possible conclusions for further steps in model refinement. In Sect. 6 the main results are summarized.
Experiments on Plasma-based Treatment of a Methane Containing Gas Stream
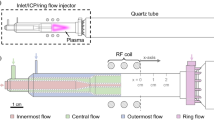
Recently, experiments were conducted to study a pretreating of hydrocarbon exhaust gas using a plasma process [17]. The intent was to study a process in which \( {\text{CH}}_{4} \) is oxidized into \(\hbox {CO}\) and \(\hbox {CO}_2\) while simultaneously consuming existing \(\hbox {O}_2\), therefore providing a kind of gas cleaning. For this purpose a radiofrequency (RF) atmospheric pressure plasma was generated in a plug flow reactor and various gas mixtures of \(\hbox {O}_2\), \(\hbox {CH}_4\) and \(\hbox {He}\) were fed into it. Helium was the dominant species, so that \(\hbox {O}_2\) and \(\hbox {CH}_4\) were only present in high dilution. Infrared spectroscopy was used to analyse the plasma conversion. This high dilution has been chosen to keep the number of relevant species small, as secondary reactions/polymerizations can be expected to be unimportant. By this means it was possible to monitor \(\hbox {CH}_4\) as well as the reaction products \(\hbox {CO}\), \(\hbox {CO}_2\) and \(\hbox {H}_2\hbox {O}\). Details on the experimental setup used and discussion of the experimental data can be found in [17] and references therein. In this paper, however, the aspect of data analysis will be in the foreground and the measured data will be taken as given and not further questioned. The special characteristics of the available data can be described as follows:
-
Complex plasma-assisted processes involving several species can be adjusted by well-defined initial conditions and power input.
-
Well equipped experimental arrangements allow the measurement of time courses of individual species.
-
Despite the wealth of information obtained by varying mixing ratios and plasma power, many species and parameter ranges are not experimentally accessible.
From this general point of view, the evaluation of existing experimental data resembles a very common situation, which is why the method discussed here may also be of importance for a number of other experiments, e. g. in plasma-assisted catalysis, where the surface species on the catalyst are usually not easy to access. The concrete data to be analyzed in this work are partial time traces of \(\hbox {CH}_4\), \(\hbox {CO}\), \(\hbox {CO}_2\) and \(\hbox {H}_2\hbox {O}\) which have been compiled in various RF discharges where the plasma power and the gas mixture have been varied. These data are supplemented by measurements of concentrations at only a single specific point in time, but obtained for several values of plasma power and initial gas composition. In total a number of 269 data points are available from the campaign reported in [17] containing scattered data of the concentrations of \(\hbox {CH}_4\), \(\hbox {CO}\), \(\hbox {CO}_2\) and \(\hbox {H}_2\hbox {O}\) at different times, plasma power and feed gas admixture.
Model Discovery Method
Selection of the Reaction Network
First, a selection of the reactants and products to be considered is made. From the combinatorial possibilities of all unimolecular and bimolecular reactions, those processes are selected that are stoichiometrically possible. In our example, a reaction network is built up for a number of \(N_s=9\) species \(X_s\), \(s=1,\ldots ,N_s\), namely \(\hbox {CH}_4\), \(\hbox {O}_2\), \(\hbox {O}\), \(\hbox {CO}_2\), \(\hbox {CO}\), \(\hbox {H}_2\hbox {O}\), \(\hbox {C}\), \(\hbox {H}_2\) and \({\hbox {e}^-}\). According to the stoichiometric selection rule a number of \(N_r=42\) reactions are possible which are listed in Table 1. The corresponding rate laws for the species densities \([X_s]\) can be written in the compact form
The stoichiometric matrix is given by the components \(\nu _{s,r}\) and the rate coefficient for a particular reaction is denoted by \(k_r\). Only a part of the densities \([X_s]\) are experimentally accessible and at the beginning of the evaluation we do not make any assumptions about the rate coefficients \(k_r\). Inspection of the reactions of Table 1 shows that the species densities obey the conservation laws
The conservation of electrons is due to the fact that it is assumed that they participate in electron impact dissociation only. It is to be noted that the combustion of methane is a well-known system [18,19,20,21] with typically very many species and reactions. Reaction pathways have also been studied extensively for plasma-assisted conversion including a large number of reactions and species [22,23,24,25,26]. In our system, however, the dilution is very high and the residence time is short. We therefore, restrict ourselves to only primary reactions. Reactions involving charged species such as ion molecule reactions or electron ion recombination are neglected due to the very low charge carrier density of \(10^{11}\,\hbox {cm}^{-3}\) in comparison to the neutral reactive species densities of the order of \(10^{17}\,\hbox {cm}^{-3}\). This model does not pretend to be complete, but it exhibits important characteristics of the observed processes. In the end, it represents a first and certainly arbitrary step towards a pragmatic model that is to be found here.
Genetic Algorithm for Extraction of Rate Coefficients
The goal of our numerical calculations is to find rate coefficients \(k_r\) that bring the results of the integrated rate equations of Eq. 1 in the best possible agreement with the experimental data. This represents a minimization problem where the difference between calculated densities and the experimental results should be minimized for all investigated time points, plasma power and starting conditions simultaneously. To solve this problem, we use a genetic algorithm which considers a population of N chromosomes which carry a number of M genes. The chromosomes are represented by vectors \(\mathbf{y }_i\), \(i=1,\ldots ,N\) with components \(y_{i,j}\), \(j=1,\ldots ,M\). Each chromosome \(\mathbf{y }_i\) is a candidate solution for the minimization problem considered and the genes represent model parameters to be optimized. In our case where rate coefficients are the model parameters, the components \(y_{i,j}\) are given by the relations
This choice ensures the positivity of the rate coefficients \(k_{i,j}\) for any \(y_{i,j}\). The number of model parameters equals the number of reaction rate coefficients, \(M=N_r\), such that \(k_{i,r}\) is the ith candidate solution for the reaction rate \(k_r\) in the rate laws of Eq. 1. The fitness \(F_i\) of the ith chromosome is defined by
where a is an appropriate normalization constant and \(f_i\) is a positive definite functional representing the deviance of our reaction kinetic model
Here, \({\hat{c}}_q\) denotes a measured density of some observable species at a particular time and for particular experimental conditions. The density \(c_{iq}\) is the corresponding numerical result obtained by integration of the rate laws of Eq. 1 using the ith candidate solution \(k_{i,r}\), \(r=1,\ldots ,N_r\) for the rate coefficients. In total a number of \(N_c\) constraints (experimental data points) is used to define the fitness functional. The factor \(w_q\) is a weight that makes it possible to assign greater importance to certain experimental points. For example, the choice of a weight \(w_q=m\), where m is an integer, would be equivalent to the m-fold repetition of an experiment, providing always the same result. An accumulated probability function \(P_i\) is introduced by
This means that if chromosome-indices i are picked randomly via the rule \(P_{i-1} < r \le P_i\), with uniform random numbers \(0 \le r \le 1\), the resulting distribution of indices reflects their fitness, i. e. more chromosomes with high fitness are selected. The practical computation then begins with a first guess for the rate coefficients \(k_r\) and the evolution to the next generation of chromosomes, i. e. a refinement of solutions, consists of the following steps indicated by roman numerals I to V. The populations \(\mathbf{y }_i\), \(i=1,\ldots ,N\) resulting from the modifications of a particular step are also labeled by (I), ..., (IV).
Step: I Hierarchy
The chromosomes \(\mathbf{y }_i\) are sorted with respect to their fitness values \(F_i\) and a resorted vector \(F^{\mathrm{(I)}}_i\) is constructed following the ordering: \(F^{\mathrm{(I)}}_i \ge F^{\mathrm{(I)}}_{i+1}\), \(i = 1,\ldots ,N-1\). This re-ordering defines the intermediate population \(\mathbf{y }^{\mathrm{(I)}}_i\).
Step: II Selection
In the selection step the lower half of the population (the solutions with small fitness) is removed and the upper half is cloned. For the resulting population of chromosomes \(\mathbf{y }^{\mathrm{(II)}}_i\) an accumulated probability function \(P_i^{\mathrm{(II)}}\) is computed according to Eq. 9 for the fitness distribution \(F^{\mathrm{(II)}}_i\). The selection can be expressed formally by writing
Step: III Crossover
First, the two best fit chromosomes are copied into the next generation of offsprings.
To obtain further \(N-2\) offsprings, pairs are picked out by taking into account the fitness probability. However, the first partner in the pairing process is always the chromosome with the best fitness \(\mathbf{y }^{\mathrm{(II)}}_1\). The second partner \(\mathbf{y }^{\mathrm{(II)}}_n\) is chosen randomly according to the probability \(P_n^{\mathrm{(II)}}\), but inbreeding, i. e. \(n=1\) is avoided. Then a crossover for a particular pair \(\mathbf{y }^{\mathrm{(II)}}_1\) and \(\mathbf{y }^{\mathrm{(II)}}_n\) takes place with a probability \(p_c\). The gene index m for crossover is chosen randomly in the range \(1,\ldots ,M\) and offspings \(\mathbf{y }^{\mathrm{(III)}}_{2j-1}\) and \(\mathbf{y }^{\mathrm{(III)}}_{2j}\), \(j=2,\ldots ,N/2\), are obtained via the interpolating crossover rule [7, 16]
The interpolation parameter is taken as \(r=1\) for \(l\ne m\), but for \(l=m\) it is sampled from a uniform distribution on the interval [0,1]. Note, that the choice \(r=0\) for \(l=m\) would result in a simple exchange crossover.
Step: IV Mutation
For each chromosome \(\mathbf{y }^{\mathrm{(III)}}_i\), \(i=2,\ldots ,N\) (again \(i=1\) is excluded to keep the best fit chromosome), a mutation takes place with a probability \(p_m\) in a single gene. The particular gene index m for mutation is chosen randomly in the range 1,...,M and the mutation is realized by an incremental change of the component \(y^{\mathrm{(III)}}_{i,m}\) [6, 16]
The values of \(H_m\) and \(L_m\) denote prescribed upper and lower bounds for the mth model parameter. A uniform random number \(-1 \le r \le 1\) and a prescribed increment \(0 < \Delta \le 1\) are used to compute the change in the gene.
Step: V Update
The resulting population \(\mathbf{y }^{\mathrm{(IV)}}_i\) is the new generation. The assignment \(\mathbf{y }^{\mathrm{(IV)}}_i \longrightarrow \mathbf{y }_i\) completes one evolutionary step and the procedure begins anew until a certain convergence criterium is fulfilled.
Parallelization Issues
It is in the nature of genetic algorithms that an increase in computational effort is often accompanied by an improvement in results or an acceleration of the computation. A large population allows a greater variance of possible solutions and this can lead to a faster and more extensive search in the solution space. Advantageously, parallel computer architectures can be used very well for this purpose, since relatively simple parallelization strategies are possible. One method which is easy to implement is the island model [27], where the algorithm sketched in Sect. 3.2 is applied not only to a single population of N individuals, but to a group of populations that are considered to live on different islands. In practice the islands are different hardware processors. In our computations we consider a number of \(N_i\) populations, i. e. a number of \(N_i\) processors, each group consisting of N individuals. For \(N_g\) generations the islands are isolated and the algorithms are used in an identical way on each island. The only difference is in the sampling of random numbers. Each island generates its own chain of random numbers which differs from all other islands. This gives diverging results for the best fit search results on the islands. After \(N_g\) evolutionary steps all individuals from all islands are evaluated together and a global ranking is established. Then the two best individuals of the global ranking replace the two best individuals of each island population. After that the computation proceeds, again with different random numbers on each island. This minimizes the communication between the processors and introduces a considerable extension of variants in the global population. This implementation is easy and has been realized by MPI routines [28].
Inclusion of Plasma Effects
The experimentally observed plasma effects are taken into account by assumed electron impact dissociation processes. Therefore, the electrons are an additional species in the model and a fundamental part of the theoretical analysis. However, the electron density is hard to determine experimentally. To find a reasonable description of the impact of electrons a linear relationship between the discharge power P and the electron density \([{\hbox {e}^-}]\) is used. This means, that the number of electrons follows a simple discharge characteristics and no saturation effects occur, where the energy of electrons might be fed into other reaction channels for higher electron energies. Such modifications would be possible by introduction of additional model parameters to describe a more complex relation between \(n_e\) and P, but this will be analysed in future studies.
Results for the time traces of the observable species \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) without plasma (top left) and with plasma power 0.55 W, 4.00 W and 7.00 W (ordered from top right to bottom right). The gas mixture at the inlet was \(\hbox {O}_2\):\(\hbox {CH}_4=2\):1. The solid lines indicate the model results obtained with the raw model with 42 reactions. The squares mark the experimental data
Results for the power dependence of the concentrations of \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) for different gas mixtures. The figures are ordered from top left to bottom right. The solid lines indicate the model results obtained with the raw model with 42 reactions. The squares mark the experimental data. For \(\hbox {H}_2\hbox {O}\) experimental data are available only for \(\hbox {O}_2\):\(\hbox {CH}_4=2\):1
Results
Numerical Setup for a Reference Case
As mentioned in Sect. 2 and Sect. 3.1, our numerical study considers a number of 269 experimental data points, representing densities of \(\hbox {CH}_4\), \(\hbox {CO}\), \(\hbox {CO}_2\) and \(\hbox {H}_2\hbox {O}\) at different times, plasma power and feed gas admixture. The numerical regression model is based on an assumed network with \(N_s=9\) species and \(N_r=42\) reactions. For the genetic algorithms a number of \(N=20\) chromosomes, each with \(M=42\) genes defines the population of a single island. The computations are performed with \(N_i=8\) processors until no significant change in the globally best fitness is detected. The number of 20 chromosomes used in the computations is just a compromise between acceptable computational effort (low N) and quality of parameter space exploration (high N). After several numerical tests it was found that \(N>20\) does not change the results anymore, i. e. the best fit quality could not be increased. During the computation the islands communicate for the global ranking every \(N_g=100\) iterations (generations). The probabilities for crossover and mutation are chosen as \(p_c=0.6\) and \(p_m=0.9\). The upper and lower bounds for the model parameters \(y_{i,j}\) are chosen as \(L_j=-15\) and \(H_j=5\) for all \(i=1,\ldots ,N\) and \(j=1,\ldots ,M\). Therefore, the rate coefficients are kept in a range \(10^{-15} \le k_r \le 10^5\). Here, dimensionless coefficients \(k_r\) result from a scaling of densities and time in the calculations. The species densities are scaled by a reference density \(n_0 = 6.0\cdot 10^{16}\) \(\hbox {cm}^{-3}\), which is equal to the nominal value of the density \([\hbox {CH}_4]\) at the inlet, and the time coordinate is scaled by the residence time \(t_0=6.92\cdot 10^{-2}\,\hbox {s}\), which is of the order of the typical residence time of the plasma in the experimental device, where \(v=0.3\,\hbox {m/s}\) is the flow speed and the plasma volume has a length of about 26 mm (see Ref. [17]). This gives values for the scaled densities and times of the order 1. As a starting guess for the rate coefficients just a small number \(k_r=10^{-5}\) has been used for all reactions \(r=1,\ldots ,N_r\). The initial condition for the integration of the rate laws of Eq. 1, is prescribed by the amount of \(\hbox {CH}_4\) and \(\hbox {O}_2\). All other densities are initially set to zero. The numerical integration of the rate laws is done with the Fortran-Routine DVODE from the package ODEPACK [29]. Prescribing all rate coefficients, the initial conditions and the time point of observation allows to obtain the values for the densities \(c_{iq}\) in the cost functional Eq. 8 and to evaluate the fitness of each chromosome in the population. The experimental data are weighted with \(w_q=1\). Then one goes through all steps of the algorithm as described in Sect. 3.2. The Fig. 1 shows the distribution of numerically found data points compared to the experimental ones. A perfect match would lie on the straight line, but due to uncertainties in the measurements actually this would mean overfitting. Therefore, the regression gives a scattered distribution close to the perfect match. To quantify the quality of the regression the following parameters are introduced
where
Actually they all consider just the sum of squared residuals \((c_q - {\hat{c}})^2\), but in each case a different normalization factor is used to get dimensionless measures. Note that \(1-SSR_3\) is identical with the so-called coefficient of determination \(R^2\). The plot in Fig. 1 illustrates the final distribution of numerical results vs experimental data, therefore the convergence of the algorithm. The results are shown and analysed in detail in Figs. 2 and 3. The corresponding measures for the residuals are \(SSR_1 = 0.16\), \(SSR_2 = 0.03\) and \(SSR_3 = 0.06\). All of these measures demonstrate a fairly good quality of the regression to fit the data points.
Results for the time traces of the non-observable species \(\hbox {O}_2\), \(\hbox {O}\), \(\hbox {H}_2\) and \(\hbox {C}\) without plasma (top left) and with plasma power 0.55 W, 4.00 W and 7.00 W (ordered from top right to bottom right). The solid lines indicate the model results obtained with the full model with 42 reactions. The squares mark the experimental data evaluated using the assumed conservation laws Eqs. 3, 4 and 5
Results for the time traces of the observable species \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) with plasma power 4.00 W for four model variants. Top left: constrained model, top right: \(\alpha =0\)-model, bottom left: \(\alpha =1\)-model, bottom right: minimal model. The solid lines indicate the model results and the squares mark the experimental data. The gas mixture at the inlet was \(\hbox {O}_2\):\(\hbox {CH}_4=2\):1
Distribution of numerically obtained points vs experimental data points for four different model variants. Top left: constrained model, top right: \(\alpha =0\)-model, bottom left: \(\alpha =1\)-model, bottom right: minimal model. Refering to the same order, the corresponding values for the regression measures are \(SSR_1 = 0.14,\,0.27,\,0.18,\,0.11\), \(SSR_2 = 0.04,\,0.04,\,0.03,\,0.04\) and \(SSR_3 = 0.06,\,0.08,\,0.06,\,0.08\), respectively
Results for the time traces of the non-observable species \(\hbox {O}_2\), \(\hbox {O}\), \(\hbox {H}_2\) and \(\hbox {C}\) with plasma power 4.00 W for four model variants. Top left: constrained model, top right: \(\alpha =0\)-model, bottom left: \(\alpha =1\)-model, bottom right: minimal model. The solid lines indicate the model results and the squares mark the experimental data evaluated using the assumed conservation laws Eqs. 3, 4 and 5
Regression of Experimental Data
The results of the regression with the numerical setup of the reference case described in Sect. 4.1 and shown in Fig. 1 are plotted again in figures Figs. 2 and 3. This time however, as density vs time and density vs plasma power, respectively, together with the corresponding experimental results. It is clearly visible that the regression curves reflect the experimentally observed trends for the temporal evolution and the power dependence relatively well. In Fig. 4 the numerically found temporal variations of the densities of \(\hbox {O}_2\), \(\hbox {O}\), \(\hbox {H}_2\) and \(\hbox {C}\) are shown. Even though these densities were not accessible experimentally, derived data points for the densities of \(\hbox {H}_2\), \(\hbox {C}\) and \(\hbox {O}_2+\hbox {O}\) have been added which have been calculated from the measured data using the assumed conservation laws Eqs. 3, 4 and 5. It can be seen that also these indirectly found data points are relatively well recovered by the regression model. Of course, the results for the densities of \(\hbox {O}_2\) and \(\hbox {O}\) are a model extrapolation and at this point we would like to emphasize that these results must be considered with caution as long as no further measurement points for \(\hbox {O}_2\) and \(\hbox {O}\) can be added. In fact, experience has shown that other sets of rate coefficients and reaction channels provide similarly good agreement with the experimental observations of \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\), but quite different results for the densities of \(\hbox {O}_2\), \(\hbox {O}\), \(\hbox {H}_2\) and \(\hbox {C}\). This is illustrated in Figs. 5, 6 and 7. The results shown there are obtained with model assumptions different from the approach employed above. The model underlying the results of Figs. 1, 2, 3 and 4 will be called the “raw model”, where all 42 reactions of Table 1 are used without further weighting (all \(w_q=1\)) or other constraints. Obviously, the raw model works reasonably well to fit the data. However, it has been observed, that for some different starting guesses \(k_{i,r}\) some slight conversion can take place in the absence of plasma (top left figure in Fig. 2). It is assumed that this is not really reflecting the experiments, even though a detailed proof is missing. Therefore, an additional information is taken into account: The model should not show any \(\hbox {CH}_4\)-conversion if no plasma is present. As a consequence, the reactions \(R_1\), \(R_8\), \(R_9\) and \(R_{10}\) of Table 1 should be discarded, i. e. \(k_1=k_8=k_9=k_{10}=0\). According to the experimental trends it is also assumed that even for high plasma powers a finite amount of \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) still remains. This requires that the processes \(R_{36}\), \(R_{37}\), \(R_{38}\), \(R_{39}\) and \(R_{41}\) of Table 1 are not involved, i. e. \(k_{36}=k_{37}=k_{38}=k_{39}=k_{41}=0\). This reduces the number of possible reactions in the model to \(N_r = 32\). We will call this set of reactions the “constrained model”, even though it is just an assumption of expected trends. At this point we would like to note that the numerical algorithm also yields negligible rate coefficients for the corresponding processes, if the weights for the data points corresponding to plasma power \(P=0\) are increased to \(w=20\) and additional (fictitious) points are inserted for the densities of \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) at a power \(P > 8\,\text{ W }\). This means that the algorithm was able to identify these insignificant reactions as such, if an appropriate constraint is taken into account. In addition to this and to demonstrate how additional data points which remove some uncertainties in the densities of \(\hbox {O}_2\) and \(\hbox {O}\) comply with the observed data we combine the constrained model with a specification of fictitious data points, defining a mixture \(\hbox {O}_2\):\(\hbox {O}\) for a time \(t=t_*\gg t_0\). The parameter \(t_*\) is chosen to represent a point in time, when a stationary equilibrium is expected according to previous observations. The parameter \(\alpha \) is introduced to prescibe the densities according to the following relations which are a consequence of the conservation law Eq. 5
where \(0\le \alpha \le 1\) and
The indices 0 and \(*\) label the densities at time \(t=0\) (initial conditions in the experiment) and time \(t=t_*\), respectively. The parameter \(\alpha \) is set to either 0 or 1. These values represent the two limiting cases, in which either \([\hbox {O}]_*\) disappears and \([\hbox {O}_2]_*\) fulfills the mass balance (\(\alpha =0\)), or \([\hbox {O}_2]_*\) disappears and \([\hbox {O}]_*\) is constrained by the mass balance (\(\alpha =1\)). We call these models the \(\alpha =0\)-model and the \(\alpha =1\)-model, respectively. Finally, a fourth model is considered which consists of only 6 reactions, which are labeled by the symbol \(\bigstar \) in Table 1. This model was not the result of a systematic consideration, but arose from the attempt to use only reactions whose reaction rates are known from the literature. We call this the “minimal model”. The plots in Fig. 5 show exemplary results for the temporal evolution of the observable densities for a power of \(P=4.00\,\text{ W }\). For all the four alternative models the fit curves match fairly well the data points. This is confirmed by Fig. 6, where the entire set of fit results are compared to the experimental data. The corresponding error quality measures \(SSR_1\), \(SSR_2\) and \(SSR_3\) are given in the caption and show similar values as for the raw model regression. Although differences in regression quality can be seen in individual species, the overall quality remains similar. The same quality is found for plots similar to those shown in Figs. 2, 3 and 4 for the other time series and plasma power scans. However, considering the extrapolated results for the non-observable quantities shown in the plots of Fig. 7, large differences between the models become visible. One can conclude, that, depending on the concrete structure of the assumed reaction kinetic model a variety of different solutions can result, which fulfill the requirement of fitting \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\) similarly good. In the same way the sum of densities of \(\hbox {O}_2\) and \(\hbox {O}\) shows only small variations On the other hand, the ratio \(\hbox {O}_2\):\(\hbox {O}\) depends very sensitively on the model and on numerical details and also the concrete values for the rate coefficients can differ by orders of magnitude depending on the specifics of the calculation. As far as possible a comparison of rate coefficients from our numerical calculations and literature values is undertaken in Table 2 and in the discussion in Sect. 4.4. In fact, when applying genetic algorithms, one generally has the problem of determining the optimum of possible solutions and their quality. In the case considered here, the problem seems most likely to be in the fact that the solution space is only insufficiently constrained. In the end, the useful results only reflect that the proposed model networks fulfill conservation laws Eqs. 3, 4 and 5 which fit the experimental data. The conservation laws introduce a constraint on the densities of \(\hbox {H}_2\) (Eq. 3), on \(\hbox {C}\) (Eq. 4) and on the sum \(2\hbox {O}_2+\hbox {O}\) (Eq. 4), but not on \(\hbox {O}_2\) and \(\hbox {O}\) separately. According to this general argumentation, also the differences in the models can be explained for points in time far outside the experimentally observable time series and for very fast processes, which cannot be resolved by experimental data. The findings up to now can be summarized as follows:
-
Satisfactory approximation of the experimental trends was found using a raw model with 9 species and 42 reactions and no additional information or assumption used as constraint.
-
The assumption that the plasma effects are described by a linear relation of electron density and plasma power and reactions including electron impact dissociation works very well to take into account the dependence of species densities on plasma power.
-
The method provides good results for an interpolation in the experimentally investigated parameter range , i.e. for times \(t < t_0\), for a plasma power \(0\,\text{ W } \le P < 8 \,\text{ W }\), initial admixtures \(\hbox {CH}_4\):\(\hbox {O}_2\) of 1:2, 1:1, 2:1 and 4:1 and for the species \(\hbox {CH}_4\), \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\).
-
The good agreement of the experimental data with the regression model suggests that the conservation laws the model implies are valid, i. e. Eqs. 2–5 describe the experimental situation well. This result is probably due to the fact that a high helium dilution was used in the experiments and thus the number of species remained small.
-
Such good interpolation results are also compatible with certain constraints imposed on species that cannot be observed or for data points beyond the experimental limit. This is demonstrated by numerical analysis of four alternative models (the constrained model, the \(\alpha =0\)- and \(\alpha =1\)-model and the minimal model), which gives good regression for the experimental data too.
-
One can notice a sensitive dependence on numerical details when trying to determine the time dependence of the species \([\hbox {O}_2]\) and \([\hbox {O}]\), which cannot be observed. This can be explained by the conservation law Eq. 5, because the model provides a condition for the sum \(2[\hbox {O}_2]+[\hbox {O}]\), but not for its individual parts.
-
The lack of suitable constraints for improving the unambiguity of the model results is also reflected in the found number of relevant reaction mechanisms and their rate coefficients. A wide range of reaction networks and associated rate coefficients are compatible with a good regression of the experimental data.
-
However, the genetic algorithm offers the possibility to consider additional experimental data and constraints in a very simple and flexible way.
In order to advance the modelling and to derive reliable rate coefficients, further a priori information should be included. In the ideal case detailed time traces of all species of a proposed model can be observed. However, before we get there, we want to emphasize the successful application of the presented algorithm and discuss the results of the model discovery in more detail.
Lower Limit of \(\hbox {O}_2\)-Depletion
Even though the extrapolation to densities of \(\hbox {O}_2\) and \(\hbox {O}\) separately is uncertain, the good interpolation of the density of the sum \(2\hbox {O}_2+\hbox {O}\) can be used to derive a lower limit for the conversion of \(\hbox {O}_2\). For this purpose coefficients for \(\hbox {O}_2\)-depletion are introduced as
The coefficient \(\Gamma _{true}\) is actually the quantitiy of interest, which tells us the amount of molecular oxygen, reduced by the conversion process. For complete depletion one finds \(\Gamma _{true} = 1\), whereas \(\Gamma _{true} = 0\) if no \(\hbox {O}_2\) has been lost by production of \(\hbox {CO}_2\), \(\hbox {CO}\) and \(\hbox {H}_2\hbox {O}\). Due to the lack of knowledge of \([\hbox {O}_2]\) the true depletion is not known. But one can give a lower bound for the depletion efficiency by \(\Gamma _{min}\), because \(\Gamma _{true}\) is always larger than \(\Gamma _{min}\). This coefficient is plotted in Fig. 8 as a function of plasma power and initial admixture ration \([\hbox {O}_2]_0/[\hbox {CH}_4]_0\). It can be seen that a complete depletion can be expected for a plasma power almost proportional to the square of mixing ratio, i. e. , the higher the initial content of \(\hbox {O}_2\), the more plasma power is needed to ensure reliable reduction.
Characteristics of the Minimal Model
The very surprising success of the minimal model gives reason to have a closer look on the implications of this good regression result. First we want to compare the coefficients with values from the literature. The table Table 2 lists those reactions for which numerical values could be found in the NIST database [30]. They are compared with the results of the regression with the raw model and the minimal model. One recognizes that the raw model reflects the negligibility of some reactions quite well and recognizes some essential processes as such. The minimal model, however, does not show this. The numerical values are far away from the literature values. From this it can be concluded that the raw model is quite capable of identifying important reaction channels, but that the minimal model should only be understood as a fit model with effective rate coefficients. The reaction channels in this very simple network should be understood as effective overall reactions. On the other hand, the simplicity of the minimal model allows an analytical treatment of the steady state. Inspection of the corresponding rate laws gives two possibilities for stationary densities:
The first solution is given
and a second solution is
If one assumes that only \(\hbox {CH}_4\) and \(\hbox {O}_2\) are present at \(t=0\) – like in the experiment – the constants \(\hbox {Z}_1\), \(\hbox {Z}_2\) and \(\hbox {Z}_3\) are
Both solutions show a complete conversion of \(\hbox {CH}_4\) and \(\hbox {O}_2\). In the first case, the entire initial content of carbon has been transferred to \(\hbox {C}\) and the densities of \(\hbox {CO}_2\) and \(\hbox {CO}\) are zero. In the second case the possibility of finite \(\hbox {CO}_2\) and \(\hbox {CO}\) densities remains, but \(\hbox {H}_2\hbox {O}\) disappears completely. However, the first solution can be excluded, because if \(\hbox {CH}_4\) and \(\hbox {O}_2\) have already disappeared, there is no channel to transfer carbon from \(\hbox {CO}_2\) and \(\hbox {CO}\) to \(\hbox {C}\). Thus, any carbon present when \(\hbox {CH}_4\) approaches zero remains in the species \(\hbox {CO}_2\) and \(\hbox {CO}\) and there is no process, which can decompose \(\hbox {CO}_2\) and \(\hbox {CO}\) together and thus pushing both densities to zero. The composition of the gas in the stationary state depends only on the initial values \([\hbox {CH}_4]_0\) and \([\hbox {O}_2]_0\) and the ratio \(k_{4}/k_{21}\), which might be expressed by measurable quantities of the final state
Conclusion and Next Steps
What was found in the previous sections? Considering the aspect of finding a suitable regression model able to interpolate the experimental data, it was shown that the genetic algorithm presented is capable of identifying simple and small models. For different assumptions about reaction channels, reactants and products, rate coefficients could be determined that allow to describe the experimental data very well. Moreover, irrelevant reaction mechanisms could be identified, thus providing an indication for a further reduction of the models.
However, if one considers the concrete physical processes underlying the model assumptions, it becomes apparent that the numerical method is not able to identify whether a reaction is physically meaningful. This has led to calculated rate coefficients sometimes differing considerably from known literature values (see Table 2, where a few rate coefficients match quite well and others deviate strongly). Also, various well-functioning models sometimes contradicted each other considerably. An extreme example of this is the minimal model, which gets by with a very small number of reactions and neglects mechanisms that are relevant in other useful regression models. It should be noted here that Ref. [17] also discusses a useful regression model that differs significantly from those presented here. However, in our data driven approach, the choice of the rate coefficients and their absolute values is in the first order unimportant as along as the model reproduces the data and allows some extrapolation. The comparison of individual rate coefficients with literature values may then serve as an indicator for open points in the model, whether further reaction mechanisms should be invoked in the future. At the moment the data do not enforce us to that.
Nevertheless, the results for the \(\hbox {H}_2\) densities in Figs.4 and 7, for example, are not very satisfactory. The accumulation of \(\hbox {H}_2\) is not very likely in an oxidizing environment. A way out of this inadequacy, for example, would be to consider hydroxyl \(\hbox {OH}\) as another reactive species. Various studies have already shown that there are many indications that \(\hbox {OH}\) plays an important role in the conversion of \(\hbox {CH}_4\) [23,24,25] and must also be considered as an important channel for the production of \(\hbox {CO}_2\) via the reaction pathway \({\hbox {OH} + \hbox {CO} \longrightarrow \hbox {H} + \hbox {CO}_2}\) [36]. Indeed, it would have been easy to include other species like OH in the models and look at other reaction pathways. But this route would only have introduced more unknowns into the models that would have to be clarified by experimental data, i.e., the problem of extrapolation to non-measurable quantities discussed in the previous sections would have become even greater.
At this point, it is important to remember, that the fundamental difference between a detailed forward calculation and the model discovery algorithm is that in the former case the rate coefficients and the reaction channels are specified, while in the latter only the reaction channels are provided, and the algorithm looks for an optimal solution of how these channels could be used by the reaction network to reproduce the experimental data. Physical laws can help by constraining the rate coefficients. But an educated guess about the basic reaction mechanisms is essential if one wants to build a realistic model that goes beyond the regression of a few measured data.
Now what is the best way to use such a method? It could be used to extend a model step by step as new experimental data become known. The flexibility of the numerical method makes it quick and easy to “try out” additional effects that could serve as explanations without having to calculate or guess rate coefficients. In addition, competing effects could be added to existing models to investigate which reaction dominates a process, or under what conditions, a reaction becomes irrelevant. An important application is also in the identification of the relevant experimental domains to compare different models. For the \(\hbox {CH}_4\) conversion discussed here, a few measurements of atomic or molecular oxygen would already be sufficient to exclude some of the presented models as unrealistic. Likewise, the measurement of \(\hbox {H}_2\) would provide information on whether, for example, a species such as the hydroxyl \(\hbox {OH}\) is necessary to explain the experiments. In addition, a refined evaluation of the results from the genetic algorithm opens up the possibility of learning more about the significance and correlation of individual reactions. However, these refined methods are being developed at the moment and will be presented in a later paper. In this work, we have limited ourselves to presenting the basic framework for this type of study, which will be used further on for the analysis of plasma-assisted processes.
Summary
In this work a reaction kinetic analysis of experiments on the plasma conversion of \(\hbox {CH}_4\):\(\hbox {O}_2\):\(\hbox {He}\) gas mixtures in RF discharges was carried out. The aim was to formulate a model as simple as possible with few species and reaction channels, which allows the representation of experimental findings with only a few model parameters. Furthermore, the model should be flexible enough to consider future experimental findings and other information (or assumptions) about the underlying processes. To cope with both purposes, genetic algorithms prove to be useful tools that allow not only the calculation of model parameters but also the classification and comparison of different models. For the particular problem in the context of the plasma-assisted processes in RF plasma discharges considered here, a genetic algorithm has been developed and implemented. The details of the numerical structure and practical application were presented in detail. By considering a fairly extensive model based on 42 reactions and including the simplifying assumption for the plasma effects, that only a few electron impact dissociation processes are relevant, a fairly good match with the available experimental data has been obtained. This can serve as a basic framework for an analysis of further models which can be derived from it. Some examples of model reductions have been presented in this paper and have been examined and evaluated using the genetic algorithm. All these discussed models are suitable to reconstruct the experimental data. However, the predictions for species that cannot be observed and for parameter ranges that are not supported by experimental data can differ considerably from model to model. But this is not a fundamental problem, it only reflects the gaps in the currently available data. As soon as new data are added, it will be possible to use the contradictory results to exclude certain models. Nevertheless, the discussed examples show that our approach allows a convenient numerical investigation of different model assumptions within minutes or a few hours. In the sense of the objective of reduced reaction-kinetic models, our method allows to test different consequences of the model assumptions in a clearly arranged way in order to verify the predictions later in experiments. Among the model variants for the \(\hbox {CH}_4\)-\(\hbox {O}_2\)-conversion presented here, the model with only 6 reactions is particularly noteworthy, which partly allows analytical results. This allows a very convenient verification of the hypotheses and gives a direction for further investigations. In summary, the genetic algorithm presented here is a very flexible and simple method to systematically develop a reaction network for a variety of different plasma-assisted processes. The Fortran routines used for the simulations presented here can be obtained from the authors upon request.
References
Vallance C, An Introduction to chemical kinetics. IOP Publishing, 2017, ISBN 978-1-6817-4664-7 (ebook)
Turányi T, Tomlin AS (2014) Analysis of kinetic reaction mechanisms. Springer. ISBN 978-3-662-44562-4 (eBook)
Michalewicz Z, Hinterding R, Michalewicz M (1997) Evolutionary algorithms. In: Fuzzy evolutionary computation, edited by W. Kluwer Academic, Pedrycz
Olinsky AD, Quinn JT, Mangiameli PM, Chen SK (2004) A genetic algorithm approach to nonlinear least squares estimation. Int J Math Educ Sci Technol 35(2):207–217
Chambers L (ed) (1998) Practical handbook of genetic algorithms. CRC Press, Boca Raton
Michalewicz Z, Janikow CZ, Krawczyk JB (1992) A modified genetic algorithm for optimal control problems. Comput Math Applic 23(12):83–94
Elliot L, Ingham DB, Kyne AG, Mera NS, Pourkashanian M, Wilson CW (2004) Genetic algorithms for optimisation of chemical kinetics reaction mechanisms. Prog Energy Combust Sci 30:297–328
Cailler M, Darabiha N, Fiorina B (2020) Development of a virtual optimized chemistry method. application to hydrocarbon/air combustion. Combust Flame 211:281–302
Searson DP, Willis MJ, Wright A (2012) Reverse engineering chemical reaction networks from time series data. In: Varmuzza MK, Bonchev D (eds) Statistical modelling of molecular descriptors in QSAR/QSPR. Wiley-VCH, Dehmer
Tsuchiya M, Ross J (2001) Application of genetic algorithm to chemical kinetics: systematic determination of reaction mechanism and rate coefficients for a complex reaction network. J Phys Chem A 105:4052–4058
Edwards K, Edgar TF, Manousiouthakis VI (1998) Kinetic model reduction using genetic algorithms. Comput chem Engng 22:239–246
Fatemi S, Masoori M, Bozorgmehry Boozarjomehry R (2005) Application of genetic algorithm in kinetic modeling and reaction mechanism studies. Iran J Chem Chem Eng 24:37
Kadivar A, Sadeghi MT, Sotudeh-Gharebagh R, Mahmudi M (2009) Estimation of kinetic parameters for hydrogenation reactions using a genetic algorithm. Chem Eng Technol 32(10):1588–1594
Tognazzi S, Tristo M, Tschaikowski M, Vandin A (2017) EGAC: a genetic algorithm to compare chemical reaction networks. GECCO ’17: Proceedings of the Genetic and Evolutionary Computation Conference, pp 833–840
Niu B, Jia M, Xu G, Chang Y, Xie M (2018) Efficient approach for the optimization of skeletal chemical mechanisms with multiobjective genetic algorithm. Energy Fuels 32(6):7086–7102
Lapene A, Debenest G, Quintard M, Castanier LM, Gerritsen MG, Kovscek AR (2014) Kinetics oxidation of heavy oil. 2. application of genetic algorithm for evaluation of kinetic parameters. Energy Fuels 2015 29(2):1119–1129
Urbanietz T, Stewig C, Böke M, von Keudell A (2021) Oxygen removal from a hydrocarbon containing gas stream by plasma catalysis. Plasma Chem Plasma Process 41:619–642
http://combustion.berkeley.edu/gri-mech/; Smith Gregory P, Golden David M, Frenklach Michael, Moriarty Nigel W, Eiteneer Boris, Goldenberg Mikhail, Thomas Bowman C, Hanson Ronald K, Song Soonho, Gardiner Jr William C, Lissianski Vitali V, Qin Zhiwei. http://www.me.berkeley.edu/gri_mech/
Vardanyan IA, Nalbandyan AB (1985) On the mechanism of thermal oxidation of methane. Int. J. Chem Kinet. 17:901–924
Hughes KJ, Turányi T, Clague AR, Pilling MJ (2001) Development and testing of a comprehensive chemical mechanism for the oxidation of methane. Int J Chem Kinet 33(9):513–538
Mantashyan AA, Makaryan EM, Charchyan AV (2019) Kinetic analysis of the oxidative conversion of methane in slow combustion. i. key steps of the chemical mechanism. Combust Explos Shock Waves 55:513–525
Qian M, Zhong W, Kang J, Liu S, Ren C, Zhang J, Wang D (2020) Global modeling on partial oxidation of methane to oxygenates and syngas in non-equilibrium plasma. Japanese J Appl Phys 59:066003
Bogaerts A, De Bie Ch, Snoeckx R, Kozák T (2017) Plasma based CO2 and CH4 conversion: a modeling perspective. Plasma Process Polym 14:e1600070
De Bie Ch, van Dijk J, Bogaerts A (2015) The dominant pathways for the conversion of methane into oxygenates and syngas in an atmospheric pressure dielectric barrier discharge. J Phys Chem C 119:22331–22350
Zhou LM, Xue B, Kogelschatz U, Eliasson B (1998) Partial oxidation of methane to methanol with oxygen or air in a nonequilibrium discharge plasma. Plasma Chem Plasma Process 18(3)
Zhou J, Xu Y, Zhou X, Gong J, Yin Y, Zheng H, Guo H (2011) Direct oxidation of methane to hydrogen peroxide and organic oxygenates in a double dielectric plasma reactor. Chem Sus Chem 4:1095–1098
Corcoran AL, Wainwright RL (1994) A parallel island model genetic algorithm for the multiprocessor scheduling problem. In: Proceedings of the 1994 ACM/SIGAPP Symposium on Applied Computing March 6–8. ACM Press, pp 483–487
The MPI Forum webpage. http://mpi-forum.org
Manion JA, Huie RE, Levin RD, Burgess DR Jr, Orkin VL, Tsang W, McGivern WS, Hudgens JW, Knyazev VD, Atkinson DB, Chai E, Tereza AM, Lin C-Y, Allison TC, Mallard WG, Westley F, Herron JT, Hampson RF, Frizzell DHNIST Chemical Kinetics Database, NIST Standard Reference Database 17, Version 7.0 (Web Version), Release 1.6.8, Data version 2015.09, National Institute of Standards and Technology, Gaithersburg, Maryland, 20899-8320. Web address: https://kinetics.nist.gov/
Fujii N, Sagawai S, Sato T, Nosaka Y, Miyama H (1989) Study of the thermal dissociation of N2O and CO2 using O(3P) atomic resonance absorption spectroscopy. J Phys Chem 93:5474–5478
Mick H-J, Burmeister M, Roth P (1993) Atomic resonance absorption spectroscopy measurements on high-temperature CO dissociation kinetics. AIAA J. 31:671–676
Tsang W, Hampson RF (1986) Chemical kinetic data base for combustion chemistry. Part I. Methane and related compounds. J Phys Chem Ref Data 15
Dean AJ, Davidson DF, Hanson RK (1991) A shock tube study of reactions of C atoms with H2 and O2 using excimer photolysis of C3O2 and C atom atomic resonance absorption spectroscopy. J Phys Chem 95:183–191
Karach SP, Osherov VI (1999) Ab initio analysis of the transition states on the lowest triplet h2o2 potential surface. J Chem Phys 110:11918–11927
Barker JR, Stanton JF, Lam Nguyen T (2020) Semiclassical transition state theory/master equation “kinetics of HO + CO: Performance evaluation’’. Int J Chem Kinet 52:1022–1045
Acknowledgements
This project is supported by the BMBF in the framework of the L3 subproject of the Carbon2Chem consortium and by the DFG (German Science Foundation) within the framework of the Coordinated Research Centre SFB 1316 at Ruhr-University Bochum.
Funding
Open Access funding enabled and organized by Projekt DEAL..
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reiser, D., von Keudell, A. & Urbanietz, T. Determining Chemical Reaction Systems in Plasma-Assisted Conversion of Methane Using Genetic Algorithms. Plasma Chem Plasma Process 41, 793–813 (2021). https://doi.org/10.1007/s11090-021-10159-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11090-021-10159-6